Abstract
Background
De novo genome assembly of next-generation sequencing data is one of the most important current problems in bioinformatics, essential in many biological applications. In spite of significant amount of work in this area, better solutions are still very much needed.
Results
We present a new program, SAGE, for de novo genome assembly. As opposed to most assemblers, which are de Bruijn graph based, SAGE uses the string-overlap graph. SAGE builds upon great existing work on string-overlap graph and maximum likelihood assembly, bringing an important number of new ideas, such as the efficient computation of the transitive reduction of the string overlap graph, the use of (generalized) edge multiplicity statistics for more accurate estimation of read copy counts, and the improved use of mate pairs and min-cost flow for supporting edge merging. The assemblies produced by SAGE for several short and medium-size genomes compared favourably with those of existing leading assemblers.
Conclusions
SAGE benefits from innovations in almost every aspect of the assembly process: error correction of input reads, string-overlap graph construction, read copy counts estimation, overlap graph analysis and reduction, contig extraction, and scaffolding. We hope that these new ideas will help advance the current state-of-the-art in an essential area of research in genomics.
Similar content being viewed by others
Background
Next-generation sequencing (NGS) technologies are causing an unprecedented revolution in biological sciences. The ability to obtain the genome sequence of a species quickly and at a relatively low cost has tremendous biological applications to cancer research, genetic disorders, disease control, neurological research, personalized medicine, etc. NGS technologies such as Illumina, 454, APG, Helicos, Pacific Biosciences, and Ion Torrent [1] produce huge outputs at ever decreasing costs, enabling ambitious projects such as the Genome 10 K Project, [2] (http://www.genome10k.org) whose goal is to obtain the genomes of 10,000 vertebrate species, the 1000 Genomes Project, [3] (http://www.1000genomes.org) that proposes to obtain the genomes of 1000 genetically varying humans, or the Human Microbiome project, [4] (commonfund.nih.gov/Hmp) whose aim is to characterize the microbial communities found at several different sites on the human body.
NGS machines produce short pieces of DNA, called reads, that often need to be assembled together into longer sequences. The ability to de novo assemble genomic data is crucial for the success of many applications including gene expression analysis, structural variation detection, and metagenomics, [5]. There is increased demand in both computing power and algorithmic ideas in order to cope with the increasingly popular NGS data. Great work has been done on creating improved assembly programs, e.g., [6–15], as well as surveying various techniques, [5], or critically and thoroughly evaluating the existing assemblers [16, 17]; we refer the reader to the latter three papers for references to other genome assembly programs or related work. In spite of considerable advances, much improvement is still needed in the current state-of-the-art technology for de novo genome assembly [18].
Reads that overlap significantly offer a good indication that they may come from the same region of the genome. Two main approaches are used in building assemblies. Both use overlaps between the reads but in different ways. In the string-overlap graph approach [19, 20] sufficiently long overlaps between reads are used as edges to connect vertices that represent reads. In the de Bruijn graph approach [21, 22] reads are broken into k-mers that are used as vertices connected by edges representing overlaps of length k−1. However, regardless of the model, the problems underlying both approaches can be shown to be NP-hard [23]. The de Bruijn graph approach seems counterintuitive, as it breaks the reads into shorter k-mers, which appears contrary to the technological efforts to produce longer reads. Nevertheless, most of the top assemblers to date use the de Bruijn graph approach.
We propose a new assembler, SAGE (String-overlap Assembly of GEnomes), that is string-overlap graph based. SAGE includes innovations in almost every aspect of the assembly process: error correction of input reads, string-overlap graph construction, read copy counts estimation, overlap graph analysis and reduction, contig extraction, and scaffolding. We have tested SAGE on short and medium-size genomes against several of the very best assemblers, ABySS [9], SGA [12], SOAPdenovo2 [11], and SPAdes [14] and showed that it performs very well.
Implementation
The algorithms used in SAGE are described here. Some of the existing ideas are included as well in order to give a complete description.
Error correction
All NGS datasets contain errors that make any usage of such data, and genome assembly in particular, very difficult. We have used a new program, RACER [24], that consistently exceeds the error correcting performance of existing programs. All datasets have been corrected with RACER before being assembled with SAGE.
Bidirected graph
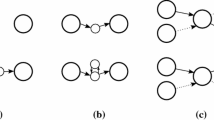
Assume a dataset of n input reads of length ℓ each, sequenced from a genome of length L. The string-overlap graph [20] has the reads as vertices. There is an edge between two vertices if there is an overlap between the sequences of the reads in the vertices (or their reverse complements) of length higher than a given threshold, M, the minimum overlap size. In order to avoid the complication due to double strandedness of DNA, [25] introduced the bidirected overlap graph, where a read and its reverse complement are represented by the same vertex and an edge has an orientation at each end point, depending on whether the read or its reverse complement is used in producing the overlap defining the edge. Three possible types of edges are thus obtained. Each edge has a string associated with it, obtained from the strings of the reads according to their overlaps. For instance, two strings xy and yz (y is the overlap) produce the string xyz for the edge. Assuming no errors in the reads, a consistent path through the graph (for each vertex, the orientation of the ingoing edge must match the orientation of the outgoing edge) spells a substring of the genome. That is also the way of associating a string with a path in the graph.
String-overlap graph construction
In order to efficiently find all overlaps of length at least M between reads, we make the following observation. Whenever two reads share an overlap of length M or more bases, there exists a prefix or suffix of length at least M in one of the reads that occurs as a substring of the other read. Therefore, we build a hash table with all prefixes and suffixes of all reads (and reverse complements) of length min{64,M}. A fast computation of these is enabled by a 2-bit representation of the DNA bases and computation of the prefixes and suffixes as 64-bit integers by fast bit operations.
After the hash table is built, a search is performed for all substrings of length min{64,M} of all reads. This is done in one pass through all reads with expected constant time search per substring and fast computation of the next substring, again using efficient bit operations. Each successful search is followed by a fast check for a valid overlap. Whenever an overlap is found, the corresponding edge is inserted in the graph.
Space-efficient transitive reduction
The string-overlap graph can be very large and a transitive reduction is performed to significantly decrease its size. An edge e=(r1,r2) is transitive if there is a read r3 and edges e1=(r1,r3),e2=(r3,r2) such that the string of the edge e is the same as the one of the path (e1,e2). Noticing that the overlaps producing the edges e1 and e2 are longer, and thus more reliable, than the one producing e, the transitive edge e can be eliminated.
Myers [20] gave a linear expected time algorithm for transitive reduction. While Myers’ algorithm is very efficient, the graph has to be built before being reduced, thus creating a space bottleneck. We have modified Myers’ algorithm to reduce the graph as it is being built. Myers’ essential observation is that the edges adjacent to a vertex have to be considered in increasing order of their lengths. We maintain this order and in addition attempt to reduce the graph as locally as possible. That is, we build only the part of the graph necessary to determine the transitively reducible edges for a given vertex v. These edges are marked for elimination but not yet removed. Once all vertices whose transitive reduction can be influenced by the edges incident with v have been investigated, the edges of v marked for elimination can be removed, thus reducing the space during construction. The running time for building the transitively reduced graph remains the same as if the complete string-overlap graph is first constructed and then transitive edges are removed, however the space decreases very much. This is essential for the entire SAGE algorithm since graph construction is the most space consuming step.
Graph simplification
The next step is further simplifying the graph. First, any path consisting exclusively of vertices of indegree and outdegree one is compressed to a single edge, subsequently called composite edge; edges that are not composite are called simple. The string spelled by the composite path is stored with the new edge along with the information concerning the reads corresponding to the collapsed vertices.
Our error correction procedure is very effective, however, errors remain in the corrected dataset. The correcting step does not remove any reads in order to avoid breaks in coverage. Overlaps between a correct read and one containing errors most likely result in short “dead-end” paths in the graph. Composite path compression might produce dead-end paths consisting of single edges. Whenever the number of reads in one such edge is lower than an experimentally determined threshold, the edge is removed.
Sometimes such erroneous paths can connect back into the graph, resulting in “bubbles”. A bubble is the event of two disjoint single paths between two vertices such that their strings are highly similar but their number of reads is very different. In such a case, if one of the two paths has much lower coverage, then it is removed.
Egde multiplicity
Assuming complete coverage and no errors, the genome would be represented as the string corresponding to a path in the graph. The number of times an edge is traversed by this path equals the number of times the string associated with the edge occurs in the genome. Myers [20] introduced the A-statistics to identify unique, or single-copy, edges. We generalize these statistics in order to be able to obtain accurate estimates of the number of occurrences of the string associated with each edge in the genome.
Given an edge e containing k reads such and whose associated string has length d, assuming the reads are sampled uniformly from the genome the probability that e has multiplicity m, that is, its string occurs m times in the genome, is given in (1).
In order to estimate the actual number of copies e has in the genome, we define the logarithm of the ratio between the probability of e having m or m+1 copies; see (2).
For m=1, this log-odds ratio gives the A-statistics of Myers [20].
Genome and insert size estimation
When defining the edge multiplicity probability and log-odds ratios, the genome length L is necessary but unknown. The values R(e,1) are used to estimate L using the bootstrap algorithm of Myers [20]. The arrival rate for a unique edge of length d and k reads is expected to be close to that of the entire genome, which gives the estimate: . Edges of length 1000 or more are initially assumed unique. Subsequent estimates of L are used for computing the R(e,1) ratios and only those edges with R(e,1)≥20 are kept as unique in future iterations. This bootstrap procedure is repeated until the set of unique edges does not change. In practice this happened in five iterations or less and a good estimate for L was obtained; see Table 1.
Often the dataset does not include information concerning the insert size, that is, the distance between the two reads of the same mate pair. The mean, μ, and standard deviation, σ, of the insert size distribution are estimated by considering only those mate pairs that belong to the same edge in the string-overlap graph where the distance between the reads in the edge is known.
Maximum likelihood assembly
Assembling a genome as the shortest string that contains the given reads suffers from an important drawback, that of overcompressing the genome or overcollapsing the repeats. This was already noticed by Myers [19], where the maximum likelihood reconstruction of the genome was proposed, that is, instead of the shortest genome, the one that is most likely to have produced the genome is searched for. Medvedev and Brudno [26] considered a very interesting approach to maximum likelihood assembly that is suitable for our purpose.
Our goal is to produce good estimates for the read copy counts, that is, for each read, the number of times its sequence appears in the genome. For a read r
i
, assume its copy count is c
i
. The c
i
values are unknown. What is known are their observed values, say x
i
, and the likelihood,  , that needs to be maximized, shown in (3).
, that needs to be maximized, shown in (3).
Maximizing  is the same as minimizing its negative logarithm, see (4).
is the same as minimizing its negative logarithm, see (4).
where C is a constant independent of the c i ’s.
For each i, the path in the graph spelling the genome sequence traverses the edge containing the read r i exactly c i times. Since is separable convex, maximizing the likelihood is reduced to a convex min-cost bidirected flow problem in a network build on the string-overlap graph. First, each convex cost function is given a three-piece linear approximation. Then the bidirected flow problem is reduced to a directed flow problem that is solved using the CS2 algorithm [27] downloaded from http://www.igsystems.com/cs2. We refer to [26] for details.
Read copy counts estimation
As explained in the previous section, the solution to the bidirected flow problem gives an estimation of the copy counts c i ’s of the reads. Crucial are the bounds we set on the capacities of the edges. For vertices, we must set a lower bound of 1 since the reads in vertices must be included in the assembly. For edges, we use the R(e,m) statistics that we introduced previously. We consider only long edges (1000 bp or more) for which the statistics should work well. For one such edge e, if R(e,1)≥T, for some threshold T (T=3 works well in practice), then we set the lower and upper bounds on the capacity of the edge as l(e)=u(e)=1. If R(e,1)<3, then we find the smallest m such that R(e,m−1)≤−T and R(e,m)≥T and set the lower bound l(e)=m and the upper bound to some large value, u(e)=∞. In case the above procedure fails to assign lower bounds, we set l(e)=1 and u(e)=∞. For the composite edges shorter than 1000 bp but containing at least 30 reads, we assign l(e)=1 and u(e)=∞. The remaining edges receive the trivial l(e)=0, u(e)=∞.
The estimation of the copy counts obtained using the above procedure is very good. Table 2 gives the comparison between our procedure and the one of [26]. The latter one works well for synthetic datasets, as reported in [26], but not for real data.
Further graph simplification
Based on the flow computed above, several further simplifications are performed to the graph. For a vertex, r, with only one incoming edge, (s,r) and outgoing edges (r,r i ),1≤i≤k, we remove the vertex r and its adjacent edges and add edges (s,r i ), 1≤i≤k. The flow on the edge (s,r i ) is the same as it was on the edge (r,r i ). A similar modification is done for vertices with only one outgoing edge.
For a vertex r that has a self loop (r,r) and only two adjacent edges, (s,r) and (r,t), we remove the vertex r and its edges and replace them with (s,t). Note that the flow on (s,t) is the same as it was on either (s,r) or (r,t).
Mate pair support
A vertex having more than one incoming edge and more than one outgoing edge is “ambiguous”. Mate pairs are used in connection with the flow to solve some of these ambiguities. For a vertex r, an incoming edge (s,r) and an outgoing edge (r,t), we say that a mate pair (r1,r2) supports the path (s,r,t) through r if all paths of length within the range μ±3σ from r1 to r2 include the path (s,r,t). Note that this is significantly more general than simply having r1 on the edge (s,r) and r2 on the edge (r,t). There may be many paths of length μ±3σ between r1 and r2.
The support required in SAGE for merging a pair of adjacent edges is at least 5. When edges (s,r) and (r,t) are merged, an edge (s,t) is added with flow equal to the minimum flow on (s,r) and (r,t). The edge with lower flow out of (s,r) and (r,t) is then deleted and the other has its flow decreased by the minimum of the two. If both have the same flow, then both are deleted. Often, r has only four adjacent edges and so it is completely resolved by this procedure.
In order to save space, we compute the paths of length up to μ+3σ from each node and then consider the reads on all edges incident to that node.
Due to lack of coverage in some regions or errors in the reads, some mate pairs may have no path to connect them in the graph. We can still use their support in such cases as follows. Consider a mate pair (r1,r2) such that r1 belongs to the edge e1 and r2 belongs to the edge e2. If the sum of the distances from r1 to the end of the edge e1 and from the corresponding end of e2 to r2 is less than μ+3σ, then (r1,r2) supports the edges (e1,e2) merging. This type of support from mate pairs is less reliable and we strengthen it by considering only edges with non-zero flow or having sufficiently long associated paths (at least 100 bp).
Assuming sufficient support is accumulated to merge the edges e1 and e2, the distance between their ends is estimated based on μ. If they overlap, there will be no gap, otherwise the gap is filled with N’s.
Finally, the set of output contigs consists of the strings of minimum required length (default is 100) that are associated with edges of non-zero flow.
SAGE overview
We present here a brief overview of the main stages of SAGE using all the procedures presented above.

Results and discussion
Assemblers
We have chosen for comparison several leading de novo assembly programs, according to their ranking by the Assemblathon 1 competition [16] and the GAGE survey [17] which rank ALLPATHS [8] and SOAPdenovo [10], both de Bruijn graph based, at the top. We have tested against the improved SOAPdenovo2 [11], but we could not use ALLPATHS since it is unable to assemble single-library real datasets. We have also included ABySS [9] as one of the most widely used and frequently updated programs and SPAdes [14] as one of the best new assemblers for bacterial genomes [28], both de Bruijn graph based. Our comparison is by no means exhaustive. The main point is to show that SAGE is competitive.
The most notable exception from the domination of the de Bruijn graph-based assemblers is the recent SGA program [12], which is string-overlap graph based. SGA uses compressed data structures to keep the memory requirements low, with the main aim of being able to run on low-end computing clusters. Nevertheless, SGA is a successful assembler, ranked third in the Assemblathon 1 competition [16]. Our efforts are complementary to those of [12] in the attempt of producing better assemblers that are string-overlap graph based.
Datasets
We compare the assemblers on real datasets only. We have downloaded a number of datasets from the NCBI web site (http://www.ncbi.nlm.nih.gov), with varied read length and genome size, together with a C.elegans dataset, from http://www.wormbase.org, that has been previously used by [12]. As noticed by [12], the genome of C.elegans is an example of an excellent test case for assembly algorithms due to its long (100 MB) and accurate reference sequence, free of SNPs and structural variants. The accession numbers for all datasets and their reference genomes are given in Table 3, together with all their parameters.
Evaluation method
The most recent version of each assembler has been used and each program was run for all possible k-mers or minimum overlap sizes on each dataset. The k-mer or minimum overlap size producing the highest scaffold N50 value is reported. N50 is representative for the quality of the assembly, however producing high values of N50 is not sufficient by itself, as it says nothing of the quality of the contigs. In particular, misassembled contigs can artificially increase N50. Therefore, we have used the QUAST comprehensive evaluation tool [29] to compare the quality of the assemblies.
The most relevant parameter is NGA50, which is computed by aligning the contigs to the reference genome, splitting them at misassembly breakpoints, eliminating unaligned parts, and then computing the N50 of the obtained contigs with respect to the length of the reference genome.
We present also the NGA75, the length of the largest alignment, the fraction of genome covered, the number of unaligned contigs, the average number of indels and mismatches for each 100 kbp, and the number of (local) misassemblies as the most important parameters computed by QUAST. All the information given by QUAST is included in the Additional file 1. Details on how each program was run and its output was evaluated by QUAST, including the precise commands used, are also given in the Additional file 1.
Comparison
The best assemblies produced by all the assemblers considered are compared and presented in Tables 4, 5, 6, 7, 8, 9, 10, 11. Whenever meaningful, we present also the average of the results. SAGE has the best NGA50, NGA75, and length of the longest aligned contig for most datasets. For NGA50, the average of SAGE is 50% better than the second one, from SPAdes. For genome coverage, SAGE and ABySS are tied for the first place. ABySS has the lowest number of unaligned contigs for the bacterial datasets but the highest for C.elegans for which SAGE performed the best. SGA produces the fewest misassemblies, followed closely by SOAPdenovo2. For local misassemblies, SGA is the best. Overall, SGA produces the fewest errors but also the lowest NGA50, NGA75, and longest contig values. Concerning the average number of indels and mismatches per 100 kbp, significant differences are seen between datasets, with 2, 3, 4 having many more errors than the other ones. All assemblers produce similar values with ABySS, SGA and SAGE at the top, with almost the same number of errors.
The time and space comparison is presented in Table 12. In order to facilitate comparison we present the time (seconds) and space (megabytes) per input mega base pairs. This way we can also compute averages. ABySS uses the least amount of space and SOAPdenovo2 is the fastest, with SAGE coming closely in second place for both time and space. Actual time and space values are presented in the Additional file 1.
Conclusions
Myers [20] suggests that string-overlap graph based assemblers should perform better than those based on the de Bruijn graph and our work aims at supporting his prediction.
SAGE builds upon great existing work and brings several new ideas, such as the efficient computation of the transitive reduction of the string overlap graph, the use of (generalized) edge multiplicity statistics for improved estimation of copy counts, and the improved use of mate pairs and flow for supporting edge merging.
We believe that our work shows that the potential of string-overlap graph-based assemblers is higher than previously thought. SAGE is currently able to successfully handle short and medium-size genomes but future versions will handle mammalian genomes as well. Also, we plan to work on reducing the number of misassemblies produced by SAGE.
We hope that some of the ideas presented will be used also by others in order to boost the development of this type of assemblers and further improve the current state-of-the-art. As read length is going to grow, we expect that string-overlap graph-based assemblers will have a better chance to improve.
Data access
All datasets are available from the NCBI, except C.elegans which is from http://www.wormbase.org. The genome assemblers are available as follows: ABySS at http://www.bcgsc.ca/platform/bioinfo/software/abyss, SGA at github.com/jts/ sga, SOAPdenovo2 at soapdenovo2.sourceforge.net/, and SPAdes at bioinf.spbau.ru/spades. The CS2 program is available at http://www.igsystems.com/cs2.
Availability and requirements
Project name: SAGEProject home page:http://www.csd.uwo.ca/~ilie/SAGE/
Operating system(s): Platform independent
Programming language: C++
Other requirements: none
License: GNU
Any restrictions to use by non-academics: none
References
Metzker ML: Sequencing technologies - the next generation. Nat Rev Genet. 2010, 11 (1): 31-46. 10.1038/nrg2626.
Haussler D, O’Brien SJ, Ryder OA, Barker FK, Clamp M, Crawford AJ, Hanner R, Hanotte O, Johnson WE, McGuire JA, Miller W, Murphy RW, Murphy WJ, Sheldon FH, Sinervo B, Venkatesh B, Wiley EO, Allendorf FW, Amato G, Baker CS, Bauer A, Beja-Pereira A, Bernardi G, Bonvicino CR, Burke T, Cracraft J, Diekhans M, Edwards S, Bermingham E, et al: Genome 10K: a proposal to obtain whole-genome sequence for 10,000 vertebrate species. J Hered. 2009, 100 (6): 659-674.
Siva N: 1000 Genomes project. Nat Biotech. 2008, 26 (3): 256-
Turnbaugh PJ, Ley RE, Hamady M, Fraser-Liggett CM, Knight R, Gordon JI: The human microbiome project. Nature. 2007, 449 (7164): 804-810. 10.1038/nature06244.
Nagarajan N, Pop M: Sequence assembly demystified. Nat Rev Genet. 2013, 14 (3): 157-167. 10.1038/nrg3367.
Dohm JC, Lottaz C, Borodina T, Himmelbauer H: SHARCGS, a fast and highly accurate short-read assembly algorithm for de novo genomic sequencing. Genome Res. 2007, 17 (11): 1697-1706. 10.1101/gr.6435207.
Zerbino DR, Birney E: Velvet: algorithms for de novo short read assembly using de bruijn graphs. Genome Res. 2008, 18 (5): 821-829. 10.1101/gr.074492.107.
Butler J, MacCallum I, Kleber M, Shlyakhter IA, Belmonte MK, Lander ES, Nusbaum C, Jaffe DB: ALLPATHS: De novo assembly of whole-genome shotgun microreads. Genome Res. 2008, 18 (5): 810-820. 10.1101/gr.7337908.
Simpson JT, Wong K, Jackman S D Schein, Jones SJ, Birol İ: ABySS: a parallel assembler for short read sequence data. Genome Res. 2009, 19 (6): 1117-1123. 10.1101/gr.089532.108.
Li R, Zhu H, Ruan J, Qian W, Fang X, Shi Z, Li Y, Li S, Shan G, Kristiansen K, Li S, Yang H, Wang J, Wang J: De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 2010, 20: 265-272. 10.1101/gr.097261.109.
Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J, He G, Chen Y, Pan Q, Liu Y, Tang J, Wu G, Zhang H, Shi Y, Liu Y, Yu C, Wang B, Lu Y, Han C, Cheung DW, Yiu S-M, Peng S, Xiaoqian Z, Liu G, Liao X, Li Y, Yang H, Wang J, Lam T-W, Wang J: SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience. 2012, 1 (1): 18-10.1186/2047-217X-1-18.
Simpson JT, Durbin R: Efficient de novo assembly of large genomes using compressed data structures. Genome Res. 2012, 22 (3): 549-556. 10.1101/gr.126953.111.
Li H: Exploring single-sample SNP and INDEL calling with whole-genome de novo assembly. Bioinformatics. 2012, 28 (14): 1838-1844. 10.1093/bioinformatics/bts280.
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA: SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol. 2012, 19 (5): 455-477. 10.1089/cmb.2012.0021.
Zimin AV, Marçais G, Puiu D, Roberts M, Yorke JA, Salzberg S L: The MaSuRCA genome assembler. Bioinformatics. 2013, 29 (21): 2669-2677. 10.1093/bioinformatics/btt476.
Earl D, Bradnam K, John JS, Darling A, Lin D, Fass J, Yu HOK, Buffalo V, Zerbino DR, Diekhans M, Ariyaratne PN, Sung W-K, Ning Z, Haimel M, Simpson JT, Fonseca NA, Birol I, Docking TR, Ho IY, Rokhsar DS, Chikhi R, Lavenier D, Chapuis G, Naquin D, Maillet N, Schatz MC, Kelley DR, Phillippy AM, Koren S, NguyenN, et al: Assemblathon 1: a competitive assessment of de novo short read assembly methods. Genome Res. 2011, 21 (12): 2224-2241. 10.1101/gr.126599.111.
Salzberg SL, Phillippy AM, Zimin A, Puiu D, Magoc T, Koren S, Treangen TJ, Schatz MC, Delcher AL, Roberts M, Marçais G, Pop M, Yorke JA: GAGE: a critical evaluation of genome assemblies and assembly algorithms. Genome Res. 2012, 22 (3): 557-567. 10.1101/gr.131383.111.
Bradnam KR, Fass JN, Alexandrov A, Baranay P, Bechner M, Birol I, Boisvert S, Chapman JA, Chapuis G, Chikhi R, Chitsaz H, Chou W-C, Corbeil J, Fabbro CD, Docking TR, Durbin R, Earl D, Emrich S, Fedotov P, Fonseca NA, Ganapathy G, Gibbs RA, Gnerre S, Godzaridis E, Goldstein S, Haimel M, Hall G, Haussler D, Hiatt JB, Ho IY, et al: Assemblathon 2: evaluating de novo methods of genome assembly in three vertebrate species. GigaScience. 2013, 2 (1): 1-31. 10.1186/2047-217X-2-1.
Myers EW: Toward simplifying and accurately formulating fragment assembly. J Comput Biol. 1995, 2 (2): 275-290. 10.1089/cmb.1995.2.275.
Myers EW: The fragment assembly string graph. Bioinformatics. 2005, 21 (suppl 2): 79-85.
Idury RM, Waterman MS: A new algorithm for DNA sequence assembly. J Comput Biol. 1995, 2 (2): 291-306. 10.1089/cmb.1995.2.291.
Pevzner PA, Tang H, Waterman MS: An Eulerian path approach to dna fragment assembly. Proc Nat Acad Sci. 2001, 98 (17): 9748-9753. 10.1073/pnas.171285098.
Medvedev P, Georgiou K, Myers G, Brudno M:Computability of models for sequence assembly. Algorithms in Bioinformatics, Lecture Notes in Bioinformatics 4645. 2007, Springer, 289-301.
Ilie L, Molnar M: RACER: Rapid and accurate correction of errors in reads. Bioinformatics. 2013, 29 (19): 2490-2493. 10.1093/bioinformatics/btt407.
Kececioglu JD: Exact and approximation algorithms for DNA sequence reconstruction. PhD thesis. The University of Arizona; 1991,
Medvedev P, Brudno M: Maximum likelihood genome assembly. J Comput Biol. 2009, 16 (8): 1101-1116. 10.1089/cmb.2009.0047.
Goldberg AV: An efficient implementation of a scaling minimum-cost flow algorithm. J Algorithms. 1997, 22 (1): 1-29. 10.1006/jagm.1995.0805.
Magoc T, Pabinger S, Canzar S, Liu X, Su Q, Puiu D, Tallon LJ, Salzberg SL: Gage-b: an evaluation of genome assemblers for bacterial organisms. Bioinformatics. 2013, 29 (14): 1718-1725. 10.1093/bioinformatics/btt273.
Gurevich A, Saveliev V, Vyahhi N, Tesler G: QUAST: quality assessment tool for genome assemblies. Bioinformatics. 2013, 29 (8): 1072-1075. 10.1093/bioinformatics/btt086.
Acknowledgements
We would like to thank Jared Simpson for helping installing SGA and Paul Medvedev for sharing his code for copy count estimates. Performance evaluation has been performed using the facilities of the Shared Hierarchical Academic Research Computing Network (SHARCNET: http://www.sharcnet.ca) and Compute/Calcul Canada.
Funding
LI and RS-O have been partially supported by grants from the Natural Sciences and Engineering Research Council of Canada (NSERC).
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
LI proposed the string-overlap graph approach, the generalized edge multiplicity statistics, and the efficient transitive reduction, BH implemented the SAGE algorithm, and RS-O proposed various algorithmic improvements. LI, BH, and RS-O met regularly and discussed various ideas that were implemented and tested by BH. MM installed the competing programs and the evaluation software, performed all final tests and comparisons, and wrote the SAGE manual. LI wrote the manuscript that was read and approved by all authors.
Electronic supplementary material
12859_2014_6583_MOESM1_ESM.pdf
Additional file 1: The supplementary material includes all the details on how each program was run and its output was evaluated by QUAST, including the precise commands used. All details for evaluating the assemblies, as given by QUAST, are also included, as well as the actual time and space values. (PDF 306 KB)
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Ilie, L., Haider, B., Molnar, M. et al. SAGE: String-overlap Assembly of GEnomes. BMC Bioinformatics 15, 302 (2014). https://doi.org/10.1186/1471-2105-15-302
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2105-15-302