Abstract
Background
Osteosarcoma (OS) is a common malignant tumor that predominantly occurs in adolescents. Its most common metastasis is to the lungs. As shown in our earlier study, lysosome-associated membrane glycoprotein 3 (LAMP3) is highly upregulated in metastatic OS. However, its role in the regulation of OS cell viability and apoptosis remains unknown.
Methods
We knocked down and overexpressed LAMP3 in OS cells and assessed the cell viability and apoptosis. Then, we investigated the expression of apoptosis-associated genes to identify the downstream gene(s) of LAMP3.
Results
Knockdown of LAMP3 significantly inhibited OS cell viability and promoted apoptosis. TP53, which is involved in the apoptosis pathway, was found to be highly upregulated after knockdown of LAMP3. Overexpression of LAMP3 significantly increased cell viability and abrogated apoptosis. Importantly, subsequent knockdown of TP53 partially suppressed the increased OS cell apoptosis induced by the inhibition of LAMP3, suggesting that TP53 is a key functional downstream gene of LAMP3.
Conclusions
Our findings suggest that LAMP3 promotes OS cell viability and survival by regulating TP53 expression.
Similar content being viewed by others
Introduction
Originating from mesenchymal cells, osteosarcoma (OS) is a very common tumor occurring in the long bones. It presents with spindle cells and aberrant osteoid formation [1]. Exposure to radiation is a known cause, and it coexists with other disorders, such as Li-Fraumeni disease and hereditary retinoblastoma [2]. The first peak of OS incidence is in 10- to 16-year old adolescents, while the second peak occurs in people over 50 years of age [2, 3]. The overall world incidence is 1 to 3 cases per million people per year [4].
The survival rate of OS patients has been improved to a certain degree with conventional therapies, including preoperative chemotherapy, surgical resection and postoperative chemotherapy. This improvement is also thanks to identifications of potential prognostic factors and advances in clinical therapy and OS research. However, patients with OS recurrence or an advanced stage of the disease always show poor responses to these conventional chemotherapies, with most relapsing [4, 5].
In addition, OS often metastasizes via the hematogenous route to other organs, such as the lungs (95%), kidneys (12%) and other bones (50%) [6, 7]. Advances in clinical technology have significantly increased the survival rates of patients with localized OS (by more than 50%). However, most patients had less than a 20% survival chance when metastases occurred [2, 8]. For example, the 5-year survival rate for OS patients with metastasis to the lungs was just 11% when they were treated with high-dose methotrexate, doxorubicin, bleomycin, cyclophosphamide and dactinomycin, together with surgery. This only increased to 11.8% when combined with treatment with carboplatin and ifosfamide [9]. The mechanism of OS metastasis clearly needs more study.
Gene therapy and targeted treatment have gained the attention of the medical community in recent years due to their potential to elucidate disease mechanisms and affect change at the molecular level. Identification of specific genes is critical for their success.
Some molecular targets involved in OS metastasis have been found. APEX1 and HER2/neu (ErbB2) were found to correlate well with recurrence and metastasis in OS patients [10]. NFIB was confirmed to be associated with metastasis in OS patients and the TP53 pathway was not only determined to be involved in primary OS development but is also a key factor for metastasis [11]. Several biopathways, including the HIF-1α & AP-1 and ERK & PI3K/AKT pathways, were also confirmed to be highly relevant for OS metastasis [11].
LAMP3 is a member of the lysosome-associated membrane glycoprotein (LAMP) family. It codes a 416-amino acid protein [12]. It is primarily reported in lung tissues, and it is found to be overexpressed in several primary cancers, such as breast, lung and liver cancer [13]. Furthermore, LAMP3 is correlated to the hypoxia regulation progress, which makes it a good biomarker for breast cancer [14], and epithelial LAMP3 expression is reported to be a prognostic biomarker for esophageal squamous cell carcinoma [15]. Importantly, LAMP3 has been reported to be associated with many tumor metastases, such as cervical cancer and osteosarcoma [12, 16].
In our previous study, LAMP3 was found to be one of the top upregulated genes in OS lung metastasis tissue compared to conventional OS tissue [16]. However, its role in the regulation of OS cell viability and apoptosis is still unclear.
TP53 is widely accepted as one of the most important tumor suppressor genes. It acts as a central regulator of multiple biological processes, including cell proliferation and apoptosis [10, 17, 18]. Notably, TP53 was proved to be an effective prognostic biomarker for OS patients [19]. Whether TP53 is a crucial regulator of the LAMP3 pathway in human OS remains to be investigated. Comprehensive understanding of its functional network will significantly benefit clinical treatment.
In this study, we investigated the impact of LAMP3 on OS cell viability and apoptosis, and then identified the functional downstream gene of LAMP3. Our findings revealed that LAMP3 increased OS cell viability and survival through the regulation of TP53 expression.
Materials and methods
Cell culture and RNA interference
U2OS and OS-732 cell lines were respectively obtained from ATCC (American Type Culture Collection) and the Chinese Academy of Sciences. Cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum (FBS), 2 mM glutamine, 100 μg/ml streptomycin and 100 U/ml penicillin at 37 °C in a humidified atmosphere containing 5% CO2. To knockdown LAMP3 and TP53, commercially validated siLAMP3 and/or siTP53 were synthesized by GenePharma and transfected to cells using Lipofectamine RNAi MAX (Life Technologies) according to the manufacturer’s instructions. Overexpression of LAMP3 was accomplished through transfection to cells with expression plasmids from GeneCopoeia company (cat. no. EX-A6482-M02) using the Lipofectamine 3000 system (Thermo Fisher Scientific) according to the manufacturer’s protocol. Two days later, the medium was replaced with fresh full culture medium. After transfection for about 72 h, the cells were collected for western blot or quantitative RT-PCR.
MTT assay
OS-732 and U2OS cells were seeded in 96-well cell culture plates with 2500 cells/well, and cultured for 24 h. Then, the cells were incubated with LAMP3/TP53 siRNA for 48 h. After that, the cells were treated with 100 μl 5 g/l MTT for 4 h. 100 μl DMSO was added to each well 15 min before analysis. OD570 nm was measured. Each assay was repeated three times.
Western blotting
The expression levels of LAMP3, E-cadherin, and β-actin proteins were determined via western blotting. Cells were lysed with RIPA buffer (Beyotime Institute of Biotechnology) and protease inhibitor cocktail (Sigma-Aldrich). Then, proteins were added to each well of an SDS-PAGE setup. After protein concentration and separation, the proteins were carefully transferred to polyvinylidene fluoride (PVDF) membrane and treated with 5% non-fat dry milk in TBST buffer for blocking. After three washes, the membranes were incubated with primary antibodies at 4 °C overnight with shaking. The next day, the washed membranes were protected from light and exposed to HRP-conjugated secondary antibodies at room temperature for 1 h.
The protein levels were determined using a chemiluminescence substrate. In this section, the antibodies were: anti-LAMP3 antibody (Abcam, ab83659), cleaved caspase-3 (Asp175) antibody (Cell Signaling, 9661), mouse anti-β-actin (Abcam), anti-mouse HRP (Sigma), and anti-rabbit HRP (Sigma).
Quantitative RT-PCR
To obtain total RNA, cells were lysed with TRIzol reagent (Invitrogen). The SuperScript III First-Strand Synthesis Kit (Invitrogen) was used to synthesize cDNA. Then quantitative RT-RCR was performed using a SYBR green master mix system (7900HT, Applied Biosystems). The 2-ΔΔCt method was used to calculate the mRNA levels. The GAPDH gene was used as the internal control. The primer sequences were:
Gene | Forward primer | Reverse primer |
LAMP3 | AGCAAGCACCTCACCAAACTT | TGTAGTCGCTGGGGTAGTTGT |
BIRC2 | AGCACGATCTTGTCAGATTGG | GGCGGGGAAAGTTGAATATGTA |
BIRC3 | TTTCCGTGGCTCTTATTCAAACT | GCACAGTGGTAGGAACTTCTCAT |
BNIP2 | TCCTAGTGATGGCTCTGTATTGT | ACTATTCTCTGACGGTGTGTCT |
BNIP3 | CAGGGCTCCTGGGTAGAACT | CTACTCCGTCCAGACTCATGC |
BNIP3L | TTGGATGCACAACATGAATCAGG | TCTTCTGACTGAGAGCTATGGTC |
CASP3 | AGAGGGGATCGTTGTAGAAGTC | ACAGTCCAGTTCTGTACCACG |
CD27 | CAGAGAGGCACTACTGGGCT | CGGTATGCAAGGATCACACTG |
CD40LG | ACATACAACCAAACTTCTCCCCG | GCAAAAAGTGCTGACCCAATCA |
BIRC6 | TAGTGTATGCCTCGTTTGTTGG | TTCTGTGTGTGCTCACCTTTC |
BRAF | CCCCAAGTCACCACAAAAACC | CGGACTGTAACTCCACACCTT |
TP53 | CAGCACATGACGGAGGTTGT | TCATCCAAATACTCCACACGC |
CASP1 | TTTCCGCAAGGTTCGATTTTCA | GGCATCTGCGCTCTACCATC |
ABL1 | TGAAAAGCTCCGGGTCTTAGG | TTGACTGGCGTGATGTAGTTG |
AKT1 | AGCGACGTGGCTATTGTGAAG | GCCATCATTCTTGAGGAGGAAGT |
BAD | CCCAGAGTTTGAGCCGAGTG | CCCATCCCTTCGTCGTCCT |
BAK1 | GTTTTCCGCAGCTACGTTTTT | GCAGAGGTAAGGTGACCATCTC |
BAX | CCCGAGAGGTCTTTTTCCGAG | CCAGCCCATGATGGTTCTGAT |
BID | ATGGACCGTAGCATCCCTCC | GTAGGTGCGTAGGTTCTGGT |
BIK | GACCTGGACCCTATGGAGGAC | CCTCAGTCTGGTCGTAGATGA |
CASP10 | AGAAACCTGCTCTACGAACTGT | GGGAAGCGAGTCTTTCAGAAG |
CASP14 | CGCCTGGCCCTAATACTGTG | GGGTCTCTTTTCATGGTGCTTTC |
GAPDH | GGAGCGAGATCCCTCCAAAAT | GGCTGTTGTCATACTTCTCATGG |
Apoptosis assay
An Annexin V-FITC/PI apoptosis Detection Kit (Abcam) was used to measure cell apoptosis. Cells were washed using PBS and then re-suspended in 200 μl binding buffer. After that, 5 μl Annexin V-FITC solution and 5 μl propidium iodide (PI) were incubated with the cells at 4 °C in the darkroom for 15 min. Samples were then analyzed via flow cytometry with Annexin V-(FL1-H) and PI-(FL2-H) on a BD FACSCalibur platform.
Data acquisition
A tag cloud showing genes frequently mentioned to be associated with OS was acquired from the Osteosarcoma Database (http://osteosarcoma-db.uni-muenster.de/index.php). This database, which contains 911 protein-coding genes and 81 microRNAs associated with osteosarcoma (derived from 1331 abstracts), provides a structured view of the state of knowledge on osteosarcoma, relying on literature mining and manual annotation of PubMed abstracts [20]. The top genes related with OS can be searched and easily shown on the homepage of the website.
Statistical analysis
All experiments were performed in at least three replicates. Data are presented as means ± SEM. Student’s t-test was analyzed to calculate the significance of means between two groups. Repeated analysis of variance (ANOVA) was performed to evaluate the difference in cell growth between the control and LAMP3 or TP53 siRNA groups. p < 0.05 was considered significant.
Results
LAMP3 promotes OS cell viability
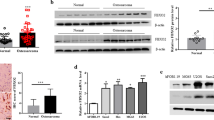
To investigate the roles of LAMP3 in OS cells, we transfected siLAMP3 into OS cells and then examined cell viability. The mRNA and protein expression levels of LAMP3 were significantly decreased after knockdown of LAMP3 in OS-732 and U2OS cells (Fig. 1 and Additional file 1: Figure S1). Although cells in the siLAMP3 group showed a minor growth increase, which may due to the gradual decrease in gene expression due to transient transfection over time, the cell viability was significantly inhibited. Overexpression of LAMP3 led to evident increased cell proliferation, indicating that LAMP3 is required for cell viability and promotes OS tumor cell growth.
LAMP3 regulated cell viability in osteosarcoma (OS) cells. a, b, d and e – Western blots showing the efficiency of LAMP3 knockdown in OS-732 (a, b) or U2OS (d, e) cells at the mRNA (a, d) and protein (b, e) levels. β-actin served as the loading control. g and h – Western blots showing the efficiency of LAMP3 overexpression. c, f and i – MTT assay results showing that OS cell viability decreased after the inhibition of LAMP3 (C) and increased following overexpression of LAMP3 (f) compared with the control (i). *p < 0.05, **p < 0.01, ***p < 0.001, n = 3
LAMP3 inhibits OS cell apoptosis
To investigate how LAMP3 affects cell viability, we assessed apoptosis after knockdown of LAMP3. As shown in Fig. 2, inhibition of LAMP3 increased the expression level of cleaved caspase-3, a protein marker for the apoptosis in OS-732 and U2OS cells, suggesting that LAMP3 might positively regulate OS cell viability by inhibiting cell apoptosis. Consistent with the western blot results for cleaved caspase-3, the percentage of late apoptotic cells increased from 3.82 to 5.14% in OS-732 cells and that of early apoptotic cells increased from 0.8 to 0.94%. Similar results were observed for siLAMP3 U2OS cells. When we upregulated LAMP3 expression in OS-732 cells, we found opposite effect. Overexpression of LAMP3 inhibited the expression of cleaved caspase-3, and the apoptotic percentage decreased from 1.01 to 0.4% (early apoptosis) and 5.24 to 4.18% (late apoptosis) of cells. These results showed LAMP3 could inhibit cell apoptosis.
LAMP3 regulated cell apoptosis in osteosarcoma (OS) cells. a and c – Apoptosis assay results showing that the percentage of apoptotic cells increased after inhibition of LAMP3 compared with the control. E – The apoptotic rate significantly decreased when LAMP3 was overexpressed compared with the control cells. b and d – Western blots showing the upregulated expression of cleaved-caspase-3 (b) and TP53 (d) after knockdown of LAMP3 in either OS-732 or U2OS cells. f – Representative example of the downregulated expression of cleaved-caspase-3 and TP53 at the protein level with exogenous LAMP3 expression. *p < 0.05, **p < 0.01, ***p < 0.001, n = 3
TP53 is the top upregulated gene after knockdown of LAMP3 in OS cells
To understand how LAMP3 regulates OS cell apoptosis, we examined the apoptotic gene expression profiles by inhibiting LAMP3 in OS cells. We observed that the mRNA levels of TP53, CASP1 and ABL1 significantly increased, while BIRC6 and BRAF mRNA expressions significantly decreased (Fig. 3a). Those results suggest that these genes might play important roles in the LAMP3-mediated regulation of OS cell apoptosis. Notably, TP53 was the top upregulated gene, indicating that TP53 is the major response downstream gene of LAMP3 in the regulation of OS cell apoptosis.
TP53 was upregulated and partially reversed the effect of LAMP3 on osteosarcoma (OS) cell viability and apoptosis. a – Quantitative RT-PCR results showing the mRNA levels of apoptotic genes after knockdown of LAMP3. b – Western blot confirmed the knockdown efficiency of TP53 with β-actin as the loading control. c – Apoptosis assay results showing that the percentage of apoptotic cells was decreased after inhibition of TP53 in siLAMP3 cells. d – MTT assay results showing that OS cell viability was increased after the inhibition of TP53 in siLAMP3 cells. *p < 0.05, **p < 0.01, ***p < 0.001, n = 3
Further knockdown of TP53 partially reverses the effect of LAMP3 on OS cell apoptosis and viability
To investigate the role of TP53 in OS cell apoptosis promoted by inhibition of LAMP3, we further knocked down TP53 in siLAMP3 cells (Fig. 3b). We observed a similar increased percentage of late apoptotic cells, from 2.97 to 5.64% after knockdown of LAMP3 in OS-732 cells, but this increase could be partially inhibited by further knockdown of TP53, whose ratio was 4.42% (Fig. 3c). Similar inhibition was also observed in early apoptotic cells (Fig. 3c). Consistently with the inhibition of cell apoptosis, the decreased cell viability in the siLAMP3 group compared to the siControl group was also partially inhibited by the further knockdown of TP53 (Fig. 3d). These results show that TP53 is the key downstream gene of LAMP3 in regulation of OS cell apoptosis and viability.
TP53 is top dysregulated gene in OS patients
To explore the clinical relevance of TP53, we examined the expression of TP53 gene in OS patients in the online database. We found that TP53 was a top dysregulated gene in OS patients (Fig. 4), representing genes frequently mentioned to be associated with OS. Those data suggest the strong clinical relevance of LAMP3’s key downstream gene, TP53, in OS patients. Therefore, activation of the TP53 pathway may be a promising way to treat OS in such patients.
TP53 is one of the top dysregulated genes in osteosarcoma (OS) patients according to the previous study (doi: https://doi.org/10.1093/database/bau042)S
Discussion
Our previous study showed that LAMP3 was significantly upregulated in osteosarcoma (OS) lung metastasis tissue compared to its expression in conventional OS tissue. Here, we showed that inhibition of LAMP3 in OS cells reduced viability and increased apoptosis through the regulation of TP53 expression. Therefore, activating TP53 pathway may be an alternative way to treat OS patients with higher levels of LAMP3.
It is reported that TP53 mutations result in functional defects of tumor suppression, such as resistance to DNA damage and extensive proliferation [10], and enhanced proliferation, invasion and drug resistance in vitro [18]. Although the mechanisms of OS are complicated and include multiple genetic alterations, most OS progressions were caused by a preexisting genomic instability that is independent of TP53 deficiency. Such instability could involve TP53 and its regulated genes: RB1, ATRX, DLG2 and the PI3K/AKT/mTOR pathway genes [11]. Importantly, polymorphisms of the TP53 gene are associated with higher risk or survival for OS in the Chinese population [21].
In this study, we found that inhibition of LAMP3 greatly increased the expression of TP53 in OS cells. Silencing of TP53 combined with siLAMP3 did not fully restore the viability of OS cells, suggesting that other factors that could interfere with cell proliferation might occur following the double knockdown in OS cells. As a key tumor-associated gene, TP53 might be a molecular target in clinical treatment of certain OS patients, e.g., those with LAMP3 overexpression. However, Bisio demonstrated that LAMP3 itself is bound and transcriptionally activated by TP53 protein in the context of doxorubicin plus TNF-alpha induction [22], adding a layer of complexity to this system. Further investigation is needed.
We also detected other upregulated genes in siLAMP3 OS cells, including CASP1 and ABL1. Caspase-1 is an inflammatory caspase. Once activated, its downstream executioner caspases can be induced, which is pivotal to induce apoptosis [23]. ABL1 is a proto-oncogene encoding tyrosine kinase. Its expression level is associated with the formation of hematopoietic malignancy and the regulation of apoptosis in T-lymphocytes [24]. Extensive studies have demonstrated that ABL1 can work as a co-activator and positive cofactor of TP53 in activating genes involved in cell cycle arrest and apoptosis in cases of DNA damage [25,26,27].
We also found that BIRC6 and BRAF were downregulated after knockdown of LAMP3. BIRC6 is a member of the inhibition of apoptotic proteins (IAPs) family, and it regulates the degradation of caspase-9. It negatively regulates apoptosis by facilitating the degradation of TP53 [28]. BRAF usually serves as a biomarker for many tumors, including colorectal cancer, urachal carcinoma and melanoma cells [29]. This is consistent with its expression pattern in the inhibition of LAMP3 in OS cells. Therefore, these less changed genes, might also contribute to the regulation of LAMP3 in OS cell apoptosis and viability. Further studies are needed to classify it.
Future research should also be considered to reveal the details of: LAMP3 regulation of TP53 expression in OS cells; the in vivo phenotypes; the interplay between those downstream genes and LAMP3; and the other mechanisms of LAMP3 in the regulation of OS cell viability.
Conclusions
Our study showed that LAMP3 promotes OS cell viability and inhibits apoptosis. TP53 was found to be the top upregulated gene upon LAMP3 knockdown. Further knockdown of TP53 partially reverse the effect of LAMP3 on OS cell viability and apoptosis. Human clinical data showed TP53 was the top dysregulated gene in OS patients. Thus, activating TP53 may be a good clinical approach to treat OS patients with higher expression of LAMP3.
Abbreviations
- IAPs:
-
Inhibition of apoptotic proteins
- LAMP3:
-
Lysosome-associated membrane glycoprotein 3
- OS:
-
Osteosarcoma
- PVDF:
-
Polyvinylidene fluoride
References
Gill J, Ahluwalia MK, Geller D, Gorlick R. New targets and approaches in osteosarcoma. Pharmacol Ther. 2013;137(1):89–99.
Abarrategi A, Tornin J, Martinez-Cruzado L, Hamilton A, Martinez-Campos E, Rodrigo JP, et al. Osteosarcoma: cells-of-origin, Cancer stem cells, and targeted therapies. Stem Cells Int. 2016;2016:3631764.
Osborne TS, Khanna C. A review of the association between osteosarcoma metastasis and protein translation. J Comp Pathol. 2012;146(2–3):132–42.
Jackson TM, Bittman M, Granowetter L. Pediatric malignant bone tumors: a review and update on current challenges, and emerging drug targets. Curr Probl Pediatr Adolesc Health Care. 2016;46(7):213–28.
Wang Z, Li B, Ren Y, Ye Z. T-cell-based immunotherapy for osteosarcoma: challenges and opportunities. Front Immunol. 2016;7:353.
Saeter G, Hoie J, Stenwig AE, Johansson AK, Hannisdal E, Solheim OP. Systemic relapse of patients with osteogenic sarcoma. Prognostic factors for long term survival. Cancer. 1995;75(5):1084–93.
De AHC, Busam K, Salob S. Cutaneous metastasis of osteosarcoma. J Am Acad Dermatol. 2003;49:457–60.
Gok Durnali A, Paksoy Turkoz F, Ardic Yukruk F, Tokluoglu S, Yazici OK, Demirci A, et al. Outcomes of adolescent and adult patients with lung metastatic osteosarcoma and comparison of synchronous and Metachronous lung metastatic groups. PLoS One. 2016;11(5):e0152621.
Meazza C, Scanagatta P. Metastatic osteosarcoma: a challenging multidisciplinary treatment. Expert Rev Anticancer Ther. 2016;16(5):543–56.
Shaikh AB, Li F, Li M, He B, He X, Chen G, et al. Present advances and future perspectives of molecular targeted therapy for osteosarcoma. Int J Mol Sci. 2016;17(4):506.
Maximov VV, Aqeilan RI. Genetic factors conferring metastasis in osteosarcoma. Future Oncol. 2016;12(13):1623–44.
Kanao H, Enomoto T, Kimura T, Fujita M, Nakashima R, Ueda Y, et al. Overexpression of LAMP3/TSC403/DC-LAMP promotes metastasis in uterine cervical cancer. Cancer Res. 2005;65(19):8640–5.
Nagelkerke A, Bussink J, Mujcic H, Wouters BG, Lehmann S, Sweep FC, et al. Hypoxia stimulates migration of breast cancer cells via the PERK/ATF4/LAMP3-arm of the unfolded protein response. Breast Cancer Res. 2013;15(1):R2.
Nagelkerke A, Mujcic H, Bussink J, Wouters BG, van Laarhoven HW, Sweep FC, et al. Hypoxic regulation and prognostic value of LAMP3 expression in breast cancer. Cancer. 2011;117(16):3670–81.
Liao X, Chen Y, Liu D, Li F, Li X, Jia W. High expression of LAMP3 is a novel biomarker of poor prognosis in patients with esophageal squamous cell carcinoma. Int J Mol Sci. 2015;16(8):17655–67.
Liu K, He Q, Liao G, Han J. Identification of critical genes and gene interaction networks that mediate osteosarcoma metastasis to the lungs. Exp Ther Med. 2015;10(5):1796–806.
Papiol S, Arias B, Barrantes-Vidal N, Guitart M, Salgado P, Catalan R, et al. Analysis of polymorphisms at the tumor suppressor gene p53 (TP53) in contributing to the risk for schizophrenia and its associated neurocognitive deficits. Neurosci Lett. 2004;363(1):78–80.
Li J, Yang L, Gaur S, Zhang K, Wu X, Yuan YC, et al. Mutants TP53 p.R273H and p.R273C but not p.R273G enhance Cancer cell malignancy. Hum Mutat. 2014;35(5):575–84.
Chen Z, Guo J, Zhang K, Guo Y. TP53 mutations and survival in osteosarcoma patients: a meta-analysis of published data. Dis Markers. 2016;2016:4639575.
Poos K, Smida J, Nathrath M, Maugg D, Baumhoer D, Neumann A, et al. Structuring osteosarcoma knowledge: an osteosarcoma-gene association database based on literature mining and manual annotation. Database (Oxford). 2014(12):2159–60.
Ru J-Y, Cong Y, Kang W-B, Yu L, Tin Guo ZJ-N. Polymorphisms in TP53 are associated with risk and survival of osteosarcoma in a Chinese population. Int J Clin Exp Pathol. 2015;8(3):3198–203.
Bisio A, Zamborszky J, Zaccara S, Lion M, Tebaldi T, Sharma V, et al. Cooperative interactions between p53 and NFkappaB enhance cell plasticity. Oncotarget. 2014;5(23):12111–25.
Ke Y, Tang H, Ye C, Lei CT, Wang YM, Su H, et al. Role and Association of Inflammatory and Apoptotic Caspases in renal Tubulointerstitial fibrosis. Kidney Blood Press Res. 2016;41(5):643–53.
Bueno MJ, Perez de Castro I, Gomez de Cedron M, Santos J, Calin GA, Cigudosa JC, et al. Genetic and epigenetic silencing of microRNA-203 enhances ABL1 and BCR-ABL1 oncogene expression. Cancer Cell. 2008;13(6):496–506.
Dias SS, Milne DM, Meek DW. C-Abl phosphorylates Hdm2 at tyrosine 276 in response to DNA damage and regulates interaction with ARF. Oncogene. 2006;25(50):6666–71.
Gong JG, Costanzo A, Yang HQ, Melino G, Kaelin WG Jr, Levrero M, et al. The tyrosine kinase c-Abl regulates p73 in apoptotic response to cisplatin-induced DNA damage. Nature. 1999;399(6738):806–9.
Sionov RV, Coen S, Goldberg Z, Berger M, Bercovich B, Ben-Neriah Y, et al. c-Abl regulates p53 levels under normal and stress conditions by preventing its nuclear export and ubiquitination. Mol Cell Biol. 2001;21(17):5869–78.
Tang W, Xue R, Weng S, Wu J, Fang Y, Wang Y, et al. BIRC6 promotes hepatocellular carcinogenesis: interaction of BIRC6 with p53 facilitating p53 degradation. Int J Cancer. 2015;136(6):E475–87.
Yang J, Li S, Wang B, Wu Y, Chen Z, Lv M, et al. Potential biomarkers for anti-EGFR therapy in metastatic colorectal cancer. Tumour Biol. 2016;37(9):11645–55.
Availability of data and materials
The datasets used and/or analyzed during this study are available from the corresponding author on reasonable request.
Author information
Authors and Affiliations
Contributions
Weidong Zhang and Shaoxian Liu designed the research. Shaoxian Liu and Junyi Yue contributed to the collection and assembly of the data. Wei Du and Jian Han contributed to data analysis and interpretation. All the authors contributed to writing the paper and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional file
Additional file 1:
Figure S1. A – The full image of the western blot of LAMP3 detection with anti-LAMP3 from Abcam (#ab83659) in osteosarcoma (OS) cells. The rectangular box outlines the LAMP3 protein bands shown in Fig. 1a and d. B – Western blot analysis showed that LAMP3 protein expression was sharply inhibited by tunicamycin treatment in U2OS cells. (TIF 547 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Liu, S., Yue, J., Du, W. et al. LAMP3 plays an oncogenic role in osteosarcoma cells partially by inhibiting TP53. Cell Mol Biol Lett 23, 33 (2018). https://doi.org/10.1186/s11658-018-0099-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s11658-018-0099-8