Abstract
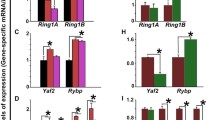
It has been well established that a previous pregnancy exhibits a beneficial effect on the subsequent pregnancy. However, the underlying mechanisms have not been defined. We hypothesized that multiparity may affect decidualization process during early pregnancy. To test this hypothesis, we analyzed global gene changes associated with multiparity in the mouse uterus using RNA-sequencing (RNA-seq). We identified a total of 131 differentially expressed genes (fold change > 2 and false discovery rate < 0.05), of which 58 were downregulated and 73 genes were upregulated in the second pregnancy (SP) compared to the first pregnancy. Functional clustering analysis showed that genes involved in stress response were significantly enriched. Most importantly, a significant portion of differentially expressed genes, 14 genes or 10.7%, overlapped with the gene list associated with decidualization. Quantitative reverse transcription (RT) polymerase chain reaction (qRT-PCR) analysis confirmed a decreased expression of 4 genes (Klk1, kallikrein 1; H2-Eb1, histocompatibility 2 class II antigen E beta; Mmp7, matrix metallopeptidase 7; Pdpn, podoplanin) and an increase in expression of 2 genes (Thy1, thymus cell antigen 1; Ptgs2, prostaglandin-endoperoxide synthase 2) in SP. Beyond protein-coding genes, we also identified a differentially expressed long noncoding RNA AI506816. Our data provide new insights into the molecular mechanisms underlying the beneficial effect of multiparity.
Similar content being viewed by others
References
Trupin LS, Simon LP, Eskenazi B. Change in paternity: a risk factor for preeclampsia in multiparas. Epidemiology. 1996;7(3): 240–244.
Roberts JM, Cooper DW. Pathogenesis and genetics of preeclampsia. Lancet. 2001;357(9249):53–56.
Duckitt K, Harrington D. Risk factors for pre-eclampsia at antenatal booking: systematic review of controlled studies. BMJ. 2005; 330(7491):565.
Eskenazi B, Fenster L, Sidney S. A multivariate analysis of risk factors for preeclampsia. JAMA. 1991;266(2):237–241.
Lie RT, Rasmussen S, Brunborg H, et al. Fetal and maternal contributions to risk of pre-eclampsia: population based study. BMJ. 1998;316(7141):1343–1347.
Macgillivray I. Some observations on the incidence of preeclampsia. J Obstet Gynaecol Br Emp. 1958;65(4):536–539.
Eras JL, Saftlas AF, Triche E, et al. Abortion and its effect on risk of preeclampsia and transient hypertension. Epidemiology. 2000; 11(1):36–43.
Zhang J, Zeisler J, Hatch MC, Berkowitz G. Epidemiology of pregnancy-induced hypertension. Epidemiol Rev. 1997; 19(2): 218–232.
Esplin MS, Fausett MB, Fraser A, et al. Paternal and maternal components of the predisposition to preeclampsia. N Engl J Med. 2001;344(12):867–872.
Harlap S, Paltiel O, Deutsch L, et al. Paternal age and preeclampsia. Epidemiology. 2002;13(6):660–667.
Tan CY, Ho JF, Chong YS, et al. Paternal contribution of HLAG*0106 significantly increases risk for pre-eclampsia in multigravid pregnancies. Mol Hum Reprod. 2008;14(5):317–324.
Skjaerven R, Wilcox AJ, Lie RT. The interval between pregnancies and the risk of preeclampsia. N Engl J Med. 2002;346(1):33–38.
Beaty TH, Skjaerven R, Breazeale DR, Liang KY. Analyzing sibship correlations in birth weight using large sibships from Norway. Genet Epidemiol. 1997;14(4):423–433.
Hinkle SN, Albert PS, Mendola P, et al. The association between parity and birthweight in a longitudinal consecutive pregnancy cohort. Paediatr Perinat Epidemiol. 2014;28(2):106–115.
Shah PS. Parity and low birth weight and preterm birth: a systematic review and meta-analyses. Acta Obstet Gynecol Scand. 2010;89(7):862–875.
Chaouat G, Kolb JP, Kiger N, Stanislawski M, Wegmann TG. Immunologic consequences of vaccination against abortion in mice. J Immunol. 1985;134(3):1594–1598.
Kiger N, Chaouat G, Kolb JP, Wegmann TG, Guenet JL. Immunogenetic studies of spontaneous abortion in mice. Preimmunization of females with allogeneic cells. J Immunol. 1985;134(5): 2966–2970.
Brown LY, Bonney EA, Raj RS, Nielsen B, Brown S. Generalized disturbance of DNA methylation in the uterine decidua in the CBA/J x DBA/2 mouse model of pregnancy failure. Biol Reprod. 2013;89(5):120.
Ghulmiyyah LM, Tamayo E, Clark SM, et al. Effect of a previous pregnancy on vascular function in endothelial nitric oxide synthase 3 knockout mice. Am J Obstet Gynecol. 2007;197(3): 279 e271–e275.
Zhang S, Lin H, Kong S, et al. Physiological and molecular determinants of embryo implantation. Mol Aspects Med. 2013;34(5): 939–980.
Dey SK, Lim H, Das SK, et al. Molecular cues to implantation. Endocr Rev. 2004;25(3):341–373.
Wang H, Dey SK. Roadmap to embryo implantation: clues from mouse models. Nat Rev Genet. 2006;7(3):185–199.
Blois SM, Klapp BF, Barrientos G. Decidualization and angiogenesis in early pregnancy: unravelling the functions of DC and NK cells. J Reprod Immunol. 2011;88(2):86–92.
Barrientos G, Freitag N, Tirado-Gonzalez I, et al. Involvement of galectin-1 in reproduction: past, present and future. Hum Reprod Update. 2014;20(2):175–193.
Gellersen B, Brosens JJ. Cyclic decidualization of the human endometrium in reproductive health and failure. Endocr Rev. 2014;35(6):851–905.
Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25(9): 1105–1111.
Trapnell C, Williams BA, Pertea G, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28(5):511–515.
Zhu Y, King BL, Parvizi B, et al. Integrating computationally assembled mouse transcript sequences with the Mouse Genome Informatics (MGI) database. Genome Biol. 2003;4(2):R16.
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods. 2008;5(7):621–628.
Brown GR, Hem V, Katz KS, et al. Gene: a gene-centered information resource at NCBI. Nucleic Acids Res. 2015;43(database issue):D36–D42.
Wang Z, Gerstein M, Snyder M. RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet. 2009;10(1):57–63.
Ruocco MG, Chaouat G, Florez L, Bensussan A, Klatzmann D. Regulatory T-cells in pregnancy: historical perspective, state of the art, and burning questions. Front Immunol. 2014;5:389.
Chen T, Darrasse-Jeze G, Bergot AS, et al. Self-specific memory regulatory T cells protect embryos at implantation in mice. J Immunol. 2013;191(5):2273–2281.
Rowe JH, Ertelt JM, Xin L, Way SS. Pregnancy imprints regulatory memory that sustains anergy to fetal antigen. Nature. 2012; 490(7418):102–106.
Dos Santos CO, Dolzhenko E, Hodges E, Smith AD, Hannon GJ. An epigenetic memory of pregnancy in the mouse mammary gland. Cell Rep. 2015;11(7): 1102–1109.
Bernstein L, Depue RH, Ross RK, et al. Higher maternal levels of free estradiol in first compared to second pregnancy: early gestational differences. J Natl Cancer Inst. 1986;76(6): 1035–1039.
Bridges RS, Byrnes EM. Reproductive experience reduces circulating 17beta-estradiol and prolactin levels during proestrus and alters estrogen sensitivity in female rats. Endocrinology. 2006; 147(5):2575–2582.
Liu JL, Wang TS. Systematic analysis of the molecular mechanism underlying decidualization using a text mining approach. PLoS One. 2015;10(7):e0134585.
Chan CS, Harvey MB, Clements JA. Temporal and tissue-specific expression of kallikrein (Klk) genes and identification of a novel Klk messenger ribonucleic acid transcript during early development in the mouse. Biol Reprod. 1999;61(3):621–628.
Rajapakse S, Yamano N, Ogiwara K, et al. Estrogen-dependent expression of the tissue kallikrein gene (Klk1) in the mouse uterus and its implications for endometrial tissue growth. Mol Reprod Dev. 2007;74(8):1053–1063.
Weiss A, Goldman S, Shalev E. The matrix metalloproteinases (MMPS) in the decidua and fetal membranes. Front Biosci. 2007;12:649–659.
Rechtman MP, Zhang J, Salamonsen LA. Effect of inhibition of matrix metalloproteinases on endometrial decidualization and implantation in mated rats. J Reprod Fertil. 1999;117(1): 169–177.
Meuleman T, Lashley LE, Dekkers OM, et al. HLA associations and HLA sharing in recurrent miscarriage: A systematic review and meta-analysis. Hum Immunol. 2015;76(5):362–373.
Volchek M, Girling JE, Lash GE, et al. Lymphatics in the human endometrium disappear during decidualization. Hum Reprod. 2010;25(10):2455–2464.
Schuring AN, Schulte N, Kelsch R, et al. Characterization of endometrial mesenchymal stem-like cells obtained by endometrial biopsy during routine diagnostics. Fertil Steril. 2011;95(1): 423–426.
Gargett CE, Schwab KE, Zillwood RM, Nguyen HP, Wu D. Isolation and culture of epithelial progenitors and mesenchymal stem cells from human endometrium. Biol Reprod. 2009;80(6): 1136–1145.
Chakraborty I, Das SK, Wang J, Dey SK. Developmental expression of the cyclo-oxygenase-1 and cyclo-oxygenase-2 genes in the peri-implantation mouse uterus and their differential regulation by the blastocyst and ovarian steroids. J Mol Endocrinol. 1996; 16(2):107–122.
Lim H, Paria BC, Das SK, et al. Multiple female reproductive failures in cyclooxygenase 2–deficient mice. Cell. 1997;91(2): 197–208.
Wang H, Ma WG, Tejada L, et al. Rescue of female infertility from the loss of cyclooxygenase-2 by compensatory upregulation of cyclooxygenase-1 is a function of genetic makeup. J Biol Chem. 2004;279(11): 10649–10658.
Wu L, Candille SI, Choi Y, et al. Variation and genetic control of protein abundance in humans. Nature. 2013;499(7456):79–82.
Fatica A, Bozzoni I. Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet. 2014;15(1):7–21.
Wang Y, Xu Z, Jiang J, et al. Endogenous miRNA sponge lincRNA-RoR regulates Oct4, Nanog, and Sox2 in human embryonic stem cell self-renewal. Dev Cell. 2013;25(1):69–80.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, JL., Zuo, RJ., Peng, Y. et al. The Impact of Multiparity on Uterine Gene Expression and Decidualization in Mice. Reprod. Sci. 23, 687–694 (2016). https://doi.org/10.1177/1933719115612131
Published:
Issue Date:
DOI: https://doi.org/10.1177/1933719115612131