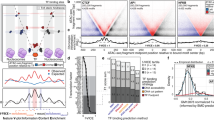
Abstract.
This commentary on the inspiring works and ideas by Langowski, Mangeol et al., Lee et al., Bundschuh and Gerland, Schiessel, Vaillant et al., Lesne and Victor, Claudet and Bednar, Fuks, Allemand et al., and Blossey, all appearing in this issue (Eur. Phys. J. E 19 (2006)), expresses our felt need of novel approaches to chromatin modeling.
Similar content being viewed by others
References
J.D. Watson, F.H. Crick, Nature 171, 737 (1953).
H.E. Avery, C.M. MacLeod, M. McCarty, J. Exp. Med. 79, 137 (1944).
U. Bockelmann, Curr. Opin. Struct. Biol. 14, 368 (2004).
P. Mangeol, Probing DNA and RNA single molecules with a double optical tweezer, this issue, p. 311.
C.H. Lee, Comparison of the measured phase diagrams in the force-temperature plane for the unzipping of two different natural DNA sequences, this issue, p. 339.
C. Anselmi, Biophys. Chem. 95, 23 (2002).
T.E. Cloutier, J. Widom, Proc. Natl. Acad. Sci. U.S.A. 102, 3645 (2005).
J. Yan, J.F. Marko, Phys. Rev. Lett. 93, 108108 (2004).
M.T. Rivero, Exp. Cell Res. 295, 161 (2004).
A.E. Vinogradov, Nucleic Acids Res. 31, 1838 (2003).
R. Bundschuh, U. Gerland, Dynamics of intramolecular recognition: Base-pairing in DNA/RNA near and far from equilibrium, this issue, p. 319.
S.E. Halford, J.F. Marko, Nucleic Acids Res. 32, 3040 (2004).
R.V. Polozov, J. Biomol. Struct. Dyn. 16, 1135 (1999).
A. Bird, Genes Dev. 16, 6 (2002).
K.D. Robertson, Nat. Rev. Genet. 6, 597 (2005).
T.M. Geiman, K.D. Robertson, J. Cell. Biochem. 87, 117 (2002).
R.D. Kornberg, Science 184, 868 (1974).
M. Noll, Nature 251, 249 (1974).
R.D. Kornberg, Y. Lorch, Cell 98, 285 (1999).
R.T. Kamakaka, S. Biggins, Genes Dev. 19, 295 (2005).
C.L. Peterson, M.A. Laniel, Curr. Biol. 14, R546 (2004).
A. Sivolob, C. Lavelle, A. Prunell, J. Mol. Biol. 326, 49 (2003).
H. Schiessel, The nucleosome: A transparent, slippery, sticky and yet stable DNA-protein complex, this issue, p. 251.
A. Sivolob, A. Prunell, Philos. Trans. R. Soc. London, Ser. A 362, 1519 (2004).
F. De Lucia, J. Mol. Biol. 285, 1101 (1999).
A. Bancaud, Structural dynamics of single chromatin fibres revealed by torsional manipulation, submitted.
C. Lavelle, A. Sivolob, A. Prunell, in preparation.
C.P. Prior, Cell 34, 1033 (1983).
J. Mozziconacci, J.M. Victor, J. Struct. Biol. 143, 72 (2003).
M. Alilat, J. Mol. Biol. 291, 815 (1999).
A. Hamiche, Proc. Natl. Acad. Sci. U.S.A. 93, 7588 (1996).
P.B. Becker, EMBO J. 21, 4749 (2002).
A. Flaus, T. Owen-Hughes, Curr. Opin. Genet. Dev. 14, 165 (2004).
C.J. Brandl, K. Struhl, Mol. Cell. Biol. 10, 4256 (1990).
M. Kanduri, Mol. Cell. Biol. 22, 3339 (2002).
N. Gilbert, J. Allan, Proc. Natl. Acad. Sci. U.S.A. 98, 11949 (2001).
C. Vaillant, Formation and positioning of nucleosomes: Effect of sequence-dependent long-range correlated structural disorder, this issue, p. 263.
Y.Y. Hsu, Y.H. Wang, J. Biol. Chem. 277, 17315 (2002).
D.T. Kirkpatrick, Mol. Cell. Biol. 19, 7661 (1999).
C. Anselmi, Biophys. J. 79, 601 (2000).
V.G. Levitsky, Nucleic Acids Res. 32 (Web Server issue), W346-9 (2004).
A. Lesne, J.-M. Victor, Chromatin fiber functional organization: Some plausible models, this issue, p. 279.
J. Zlatanova, S.H. Leuba, J. Mol. Biol. 331, 1 (2003).
C. Claudet, J. Bednar, Pulling the chromatin, this issue, p. 331.
T.A. Blank, P.B. Becker, J. Mol. Biol. 252, 305 (1995).
F. Fuks, The view from the biochemist, this issue, p. 367.
A. Benecke, Chromatin code, local non-equilibrium dynamics, and the emergence of transcription regulatory programs, this issue, p. 353.
Y. Huyen, Nature 432, 406 (2004).
S. Henikoff, Y. Dalal, Curr. Opin. Genet. Dev. 15, 177 (2005).
E.R. Foster, J.A. Downs, FEBS J. 272, 3231 (2005).
A.S. Belmont, Curr. Opin. Cell. Biol. 11, 307 (1999).
A.S. Belmont, K. Bruce, J. Cell. Biol. 127, 287 (1994).
K. Byrd, V.G. Corces, J. Cell. Biol. 162, 565 (2003).
D. Sproul, N. Gilbert, W.A. Bickmore, Nat. Rev. Genet. 6, 775 (2005).
D. Carter, Nat. Genet. 32, 623 (2002).
P.R. Cook, J. Cell. Sci. 116, 4483 (2003).
J.R. Swedlow, T. Hirano, Mol. Cell 11, 557 (2003).
B.D. Lavoie, E. Hogan, D. Koshland, J. Cell. Biol. 156, 805 (2002).
S. Almagro, J. Biol. Chem. 279, 5118 (2004).
J.F. Marko, M.G. Poirier, Biochem. Cell. Biol. 81, 209 (2003).
J. Mozziconacci, FEBS Lett. 580, 368 (2006).
T. Cremer, C. Cremer, Nat. Rev. Genet. 2, 292 (2001).
T. Cremer, Biol. Cell 96, 555 (2004).
T. Misteli, Bioessays 27, 477 (2005).
S. Kozubek, Chromosoma 108, 426 (1999).
T. Misteli, Cell 119, 153 (2004).
L. Parada, T. Misteli, Trends Cell. Biol. 12, 425 (2002).
L.A. Parada, P.G. McQueen, T. Misteli, Genome Biol. 5, R44 (2004).
D. Gerlich, Cell 112, 751 (2003).
E.M. Manders, Chromosome Res. 11, 537 (2003).
J. Langowski, Polymer chain models of DNA and chromatin, this issue, p. 241.
G. Li, Nat. Struct. Mol. Biol. 12, 46 (2005).
S. Henikoff, T. Furuyama, K. Ahmad, Trends Genet. 20, 320 (2004).
R. Blossey, Regulating chromatin: On code and dynamic models, this issue, p. 371.
J.-F. Allemand, Loops in DNA: An overview of experimental and theoretical approaches, this issue, p. 293.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lavelle, C., Benecke, A. Chromatin physics: Replacing multiple, representation-centered descriptions at discrete scales by a continuous, function-dependent self-scaled model. Eur. Phys. J. E 19, 379–384 (2006). https://doi.org/10.1140/epje/i2005-10059-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1140/epje/i2005-10059-9