Abstract
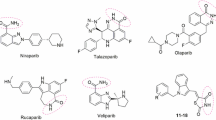
The nuclear protein poly (ADP-ribose) polymerase-1 (PARP-1) plays an important role in the signaling and repair of DNA. PARP-1 catalyses the covalent binding of poly (ADP-ribose) polymers to its subunit, as well as to other acceptor proteins, using NAD+ as a donor of ADP-ribose. Inhibitors of poly (ADP-ribose) polymerase have been shown to be effective in improving radiation therapy and chemotherapy of cancer in clinical testing. Development of new poly (ADP-ribose) polymerase-1 inhibitors based on derivatives of natural compounds such as NAD+ represents a novel and promising strategy. The structure of the complex of human poly (ADP-ribose) polymerase-1 with NAD+ can be a starting point for the rational design of small molecule inhibitors based on NAD+ derivatives. Moreover, there is no crystal structure of the complex of poly (ADP-ribose) polymerase-1 with nicotinamide adenine dinucleotide (NAD+) available yet. In this work, using molecular modeling approaches, we have predicted NAD+ binding modes to PARP-1 at the donor binding site of the catalytic domain. Using structures of PARP-1 homologs in a complex with NAD+, we predict the pharmacophore restraints of NAD+ binding to PARP-1. Based on the clustering of PARP-1 conformations in a complex with cocrystallized inhibitors and the predicted pharmacophore restraints, we propose several possible models of NAD+ binding to PARP-1 at the donor binding site of the catalytic domain. According to the predicted models, two conformations for the pyrophosphate group of NAD+ in complex with PARP-1 at the donor binding site are possible. The proposed models of NAD+ binding to PARP-1 can be validated by the quantitative structure–activity analysis of NAD+ derivatives. We designed two NAD+ derivatives, which can be used to validate the predicted NAD+ binding models.
Similar content being viewed by others
References
Barkauskaite, E., Jankevicius, G., and Ahel, I., Structures and mechanisms of enzymes employed in the synthesis and degradation of PARP-dependent protein ADP-ribosylation, Mol. Cell, 2015, vol. 58, no. 6, pp. 935–946. doi 10.1016/j.molcel.2015.05.007
Basu, B., Sandhu, S.K., and de Bono, J.S., PARP inhibitors, Drugs, 2012, vol. 72, no. 12, pp. 1579–1590.
Domagala, P., Huzarski, T., Lubinski, J., Gugala, K., and Domagala, W., PARP-1 expression in breast cancer including BRCA1-associated, triple negative and basal-like tumors: Possible implications for PARP-1 inhibitor therapy, Breast Cancer Res. Treat., 2011, vol. 127, no. 3, pp. 861–869. doi 10.1007/s10549-011-1441-2
Ferraris, D., Evolution of poly (ADP-ribose) polymerase-1 (PARP-1) inhibitors. From concept to clinic, J. Med. Chem., 2010, vol. 53, no. 12, pp. 4561–4584. doi 10.1021/jm100012m
Harder, E., Damm, W., Maple, J., Wu, C., Reboul, M., Xiang, J.Y., Wang, L., Lupyan, D., Dahlgren, M.K., Knight, J.L., Kaus, J.W., Cerutti, D.S., Krilov, G., Jorgensen, W.L., Abel, R., and Friesner, R.A., OPLS3: A force field providing broad coverage of drug-like small molecules and proteins, J. Chem. Theory Comput., 2015, vol. 12, no. 1, pp. 281–296. doi 10.1021/acs.jctc.5b00864
Hay, T., Jenkins, H., Sansom, O.J., Martin, N.M.B., Smith, G.C.M., and Clarke, A.R., Efficient deletion of normal Brca2-deficient intestinal epithelium by poly(ADPribose) polymerase inhibition models potential prophylactic therapy, Cancer Res., 2005, vol. 65, pp. 10145–10148. doi 10.1158/0008-5472
Jacobson, M.P., Friesner, R.A., Xiang, Z., and Honig, B., On the role of crystal packing forces in determining protein side-chain conformations, J. Mol. Biol., 2002, vol. 320, pp. 597–608. doi 10.1016/S0022-2836(02)00470-9
Jacobson, M.P., Pincus, D.L., Rapp, C.S., Day, T.J.F., Honig, B., Shaw, D.E., and Friesner, R.A., A hierarchical approach to all-atom protein loop prediction, Proteins: Struct., Funct. Bioinf., 2004, vol. 55, pp. 351–367. doi 10.1002/prot.10613
Jørgensen, R., Wang, Y., Visschedyk, D., and Merrill, A.R., The nature and character of the transition state for the ADP-ribosyltransferase reaction, EMBO Rep., 2008, vol. 9, no. 8, pp. 802–809. doi 10.1038/embor.2008.90
Kinoshita, T., Nakanishi, I., Warizaya, M., Iwashita, A., Kido, Y., Hattori, K., and Fujia, T., Inhibitor-induced structural change of the active site of human poly(ADP-ribose) polymerase, FEBS Lett., 2004, vol. 556, nos. 1–3, pp. 43–46. doi 10.1016/S0014-5793(03)01362-0
Lee, Y.M., Babu, C.S., Chen, Y.C., Milcic, M., Qu, Y., and Lim, C., Conserved structural motif for recognizing nicotinamide adenine dinucleotide in poly(ADP-ribose) polymerases and ADP-ribosylating toxins: Implications for structure-based drug design, J. Med. Chem., 2010, vol. 53, no. 10, pp. 4038–4049. doi 10.1021/jm1001106
Lin, H., Nicotinamide adenine dinucleotide: Beyond a redox coenzyme, Org. Biomol. Chem., 2007, vol. 5, no. 16, pp. 2541–2554. doi 10.1039/B706887E
Luo, X. and Kraus, W.L., On PAR with PARP: Cellular stress signaling through poly(ADP-ribose) and PARP-1, Gen. Dev., 2012, vol. 26, no. 5, pp. 417–432. doi 10.1101/gad.183509.111
Martin, D.S., Bertino, J.R., and Koutcher, J.A., ATP depletion+ pyrimidine depletion can markedly enhance cancer therapy: Fresh insight for a new approach, Cancer Res., 2000, vol. 60, no. 24, pp. 6776–6783.
Peralta-Leal, A., Rodriguez, M.I., and Oliver, F.J., Poly(ADP-ribose) polymerase-1 (PARP-1) in carcinogenesis: Potential role of PARP inhibitors in cancer treatment, Clin. Transl. Oncol., 2008, vol. 10, no. 6, pp. 318–323.
Ruf, A., de Murcia, J.M., de Murcia, G., and Schulz, G.E., Structure of the catalytic fragment of poly(AD-ribose) polymerase from chicken, Proc. Natl. Acad. Sci. U.S.A., 1996, vol. 93, no. 15, pp. 7481–7485.
Sherstyuk, Y.V., Zakharenko, A.L., Kutuzov, M.M., Chalova, P.V., Sukhanova, M.V., Lavrik, O.I., Silnikov, V.N., and Abramova, T.V., A versatile strategy for the design and synthesis of novel ADP conjugates and their evaluation as potential poly(ADP-ribose) polymerase 1 inhibitors, Mol. Diversity, 2016, vol. 1–13. doi 10.1007/s11030-016-9703-x
Zhu, Q., Wang, X., Chu, Z., He, G., Dong, G., and Xu, Y., Design, synthesis and biological evaluation of novel imidazo[4,5-c]pyridinecarboxamide derivatives as PARP-1 inhibitors, Bioorg. Med. Chem. Lett., 2013, vol. 23, no. 7, pp. 1993–1996. doi 10.1016/j.bmcl.2013.02.032
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © N.V. Ivanisenko, D.A. Zhechev, V.A. Ivanisenko, 2016, published in Vavilovskii Zhurnal Genetiki i Selektsii, 2016, Vol. 20, No. 6, pp. 857–862.
Rights and permissions
About this article
Cite this article
Ivanisenko, N.V., Zhechev, D.A. & Ivanisenko, V.A. Structural modeling of NAD+ binding modes to PARP-1. Russ J Genet Appl Res 7, 574–579 (2017). https://doi.org/10.1134/S2079059717050070
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S2079059717050070