Abstract—
Two variants of circularly permuted BrUSLEE, a green fluorescent protein with a short fluorescence lifetime, have been engineered. We characterized the pH-dependence of fluorescence decay kinetics of these fluorophores. It was shown that both permutants (cpBrUS and cpBrUS-145) exhibit three-component fluorescence decay kinetics, with the lifetime of the one component varying within the ~3000–300 ps range upon pH shift from 5.5 to 9.0. At the same time, the original BrUSLEE does not show a significant change in the fluorescence decay kinetics within the physiologically relevant pH-range of 6.0–8.5. The described pH-dependence allows considering the BrUSLEE permutants as pH indicators with the fluorescence lifetime readout.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
One of the factors for maintaining homeostasis and controlling the normal physiological functions of cells is the specific acidity characteristic of different subcellular compartments [1, 2]. Monitoring changes in intracellular pH in situ can provide valuable information about cellular metabolism and a deeper understanding of physiological and pathological processes [3, 4]. Most of the conventional fluorescent indicators, being a powerful tool for real-time bioimaging, are capable of only a relative assessment of changes in the studied parameter in the cell [5–7].
The aim of this work is to engineer genetically encoded indicators for quantitative measurement of pH based on the fluorescent protein BrUSLEE [8], which has previously demonstrated its potential as a label for FLIM (fluorescence lifetime imaging microscopy) with a short fluorescence lifetime.
RESULTS AND DISCUSSION
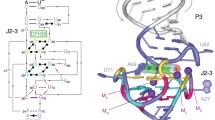
To increase the sensitivity of the fluorescent protein chromophore to the chemical environment, we performed circular permutation of BrUSLEE (Fig. 1). The principle of the permutation method consists in the rearrangement of the elements of the primary structure of the protein with the general preservation of its three-dimensional packing and the displacement of the ends into the region of the chromophore environment. The site containing 144–149 a.a. was chosen as the permutation point (according to the a.a. numbering of avGFP, Aequorea victoria Green Fluorescent Protein [9]), by analogy with the existing permutants based on GFP (Green Fluorescent Protein), which have proven themselves in the design of fluorescent indicators (in particular, GCaMP6s [10]). Two alternative variants of the primary structure of permutants were proposed: with the removal (cpBrUS-145) and with the preservation (cpBrUS) of the residue at position 145 (the Met145 residue is an important determinant of the fluorescent properties of BrUSLEE [11]).
Scheme for the engineering of permuted variants of the fluorescent protein BrUSLEE. The regions encoding the fluorescent protein are shown as green boxes. Asterisks mark the permutation points. Numbers indicate the corresponding amino acid residues according to the avGFP numbering [9].
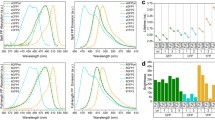
Next, we studied the pH dependence of the fluorescence lifetime of the original BrUSLEE protein and the engineered circular permutants (cpBrUS-145 and cpBrUS) in vitro, on the aqueous samples of isolated proteins using time-resolved spectroscopy with single-photon excitation by a picosecond laser with a central emission wavelength of ~450 nm (Table 1, Fig. 2).
The fluorescence decay kinetics of the fluorophores of both permutants demonstrate a pronounced dependence on the samples’ pH (Fig. 2a); they are adequately fitted by a three-exponential model, with the amplitude-normalized lifetime of the third component (τ3 × A3) showing a tenfold change (~3000–300 ps) in the pH range of 5.5–9.0 (Fig. 2b, Table 1). The shape of the graph describing dependence of τ3 × A3 on pH is close to linear, which potentially facilitates the calibration of the fluorescent signal for experimental measurement of the pH value. It should be noted that the cpBrUS-145 permutant demonstrates a greater dynamic range of its fluorescence response than cpBrUS (sixfold versus twofold change in the mean fluorescence lifetime, respectively), which indicates the ability of the Met145 residue to limit the conformational mobility of the chromophore. On the contrary, the original BrUSLEE protein shows only an insignificant sensitivity in a narrow range of pH values 4.5–5.5 (Fig. 2a), and while fitted with multicomponent exponential model, none of the fluorescence decay's components of the BrUSLEE shows significant changes in the pH range of 6.0–8.5, which is of relevance from a physiological point of view (Table 1).
EXPERIMENTAL
Engineering the circular permutants of BrUSLEE. We constructed DNA vectors carrying cpBrUSLEE coding sequences. For this, a tandem repeat of the BrUSLEE coding sequences, connected by an oligonucleotide linker encoding the amino acid sequence of (Ser-Gly-Thr)2 composition, was first obtained by PCR amplification [8, 12]; the following primers were used: 5'-ATGCGGATCCATGGTGAGCAAGGGCGAG-3', 5'-ATGCGTCGACCTTGTACAGCTCGTCCATGC-3', 5'-ATGCGTCGACAGCGGGACTAGCGGGACTATGGTGAGCAAGGGCGAG-3', 5'-ATGCAAGCTTTTACTTGTACAGCTCGTCCATGC-3'; PCR program: 95°C, 5 min, (95°C, 30 s; 58°C, 30 s; 72°C, 30 s) × 18, 72°C, 7 min. This intermediate construct served as a template for PCR amplification of DNA sequences encoding cpBrUSLEE variants using a forward (5'-end) primer corresponding to the sequence encoding the start C-terminal part of the first copy of cpBrUSLEE (starting from 145 or 146 a.a.) (5'-ATGCGGATCCATGAACAGCCACAACGTC-3' or 5'‑ATGCGGATCCAACAGCCACAACGTCTAT-3'), and a reverse (3'-terminal) primer complementary to the DNA sequence encoding the end N-terminal part of the second copy of cpBrUSLEE (1–144 a.a.) (5'‑AAGCTTTTAATGCGTTGTACTCCAGCTTGTGCC-3'), with the following PCR program: 95°C, 5 min, (95°C, 30 s; 55°C, 30 s; 72°C, 30 s) × 18, 72°C, 7 min.
Protein expression and purification. The sequences encoding the cpBrUS and cpBrUS-145 permutants were cloned into the pQE30 vector (Qiagen, United States) with the 6His tag at N-terminus, expressed in strain E. coli XL1 Blue (Invitrogen, United States) and purified using TALON metal affinity resin (Clontech, United States) [13].
Fluorescence lifetime spectroscopy of purified proteins. Measurements were made using a time-resolved miniTau fluorescence spectrometer (Edinburgh Instruments, UK) in a 50-ns window divided into 2048 time channels. Fluorescence was excited with an EPL-450 picosecond laser (Edinburgh Instruments) with a central emission wavelength of 445.6 nm and a repetition rate of 20 MHz; photons were counted in the spectral range 475–525 nm. Data processing and visualization, determination of χ2 (Pearson’s test) was carried out using the Fluoracle 2.5.1 software (Edinburgh Instruments).
CONCLUSIONS
Two circular permutants of BrUSLEE, with both preserved (cpBrUS) and removed (cpBrUS-145) Met residue at position 145, have been engineered. In contrast to the original BrUSLEE, both permutants demonstrated a pronounced dependence of the fluorescence decay kinetics on pH in the physiological range of 6.0–8.5. Thus, circular permutation significantly increases the sensitivity of the BrUSLEE chromophore to pH changes, and both BrUSLEE circular permutants can be considered as promising probes for quantitative pH measurements.
REFERENCES
Casey, J.R., Grinstein, S., and Orlowski, J., Nat. Rev. Mol. Cell Biol., 2010, vol. 11, pp. 50–61. https://doi.org/10.1038/nrm2820
Demaurex, N., Physiology, 2002, vol. 17, pp. 1–5. https://doi.org/10.1152/physiologyonline.2002.17.1.1
Boron, W.F., Adv. Physiol. Educ., 2004, vol. 28, pp. 160–179. https://doi.org/10.1152/advan.00045.2004
Lagadic-Gossmann, D., Huc, L., and Lecureur, V., Cell Death Differ., 2004, vol. 11, pp. 953–961. https://doi.org/10.1038/sj.cdd.4401466
Zhu, H., Fan, J., Xu, Q., Li, H., Wang, J., Gao, P., and Peng, X., Chem. Commun., 2012, vol. 48, p. 11766. https://doi.org/10.1039/c2cc36785h
Ermakova, Y.G., Pak, V.V., Bogdanova, Y.A., Kotlobay, A.A., Yampolsky, I.V., Shokhina, A.G., Panova, A.S., Marygin, R.A., Staroverov, D.B., Bilan, D.S., Sies, H., and Belousov, V.V., Chem. Commun., 2018, vol. 54, pp. 2898–2901. https://doi.org/10.1039/C7CC08740C
Martynov, V.I., Pakhomov, A.A., Deyev, I.E., and Petrenko, A.G., Biochim. Biophys. Acta Gen. Subj., 2018, vol. 1862, pp. 2924–2939. https://doi.org/10.1016/j.bbagen.2018.09.013
Mamontova, A.V., Solovyev, I.D., Savitsky, A.P., Shakhov, A.M., Lukyanov, K.A., and Bogdanov, A.M., Sci. Rep., 2018, vol. 8, p. 13224. https://doi.org/10.1038/s41598-018-31687-w
www.fpbase.org/protein/avgfp/
Chen, T.-W., Wardill, T.J., Sun, Y., Pulver, S.R., Renninger, S.L., Baohan, A., Schreiter, E.R., Kerr, R.A., Orger, M.B., Jayaraman, V., Looger, L.L., Svoboda, K., and Kim, D.S., Nature, 2013, vol. 499, pp. 295–300.
Mamontova, A.V., Shakhov, A.M., Lukyanov, K.A., and Bogdanov, A.M., Biomolecules, 2020, vol. 10, p. 1547.
FPbase: BrUSLEE. www.fpbase.org/protein/bruslee/.
TALON® Metal Affinity, Resins. and User, Manual., http://wolfson.huji.ac.il/purification/PDF/Tag_Protein_Purification/Ni-NTA/TALON_UserManual.pdf.
Funding
This work was supported by the Russian Foundation for Basic Research (project no. 19-34-60019, supervisor A.V. Mamontova).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
This article does not contain any research involving humans and animals as research objects.
Conflict of Interests
The authors declare they have no conflicts of interest.
Additional information
Abbreviations: BrUSLEE. bright ultimately short lifetime enhanced emitter; cp, circular permuted; avGFP, Aequorea victoria green fluorescent protein; GFP, green fluorescent protein.
Corresponding autor: phone: +7 (903) 746-08-49
Rights and permissions
Open Access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mamontova, A.V., Simonyan, T.R., Lukyanov, K.A. et al. Circular Permutants of BrUSLEE Protein as Fluorescent pH Indicators. Russ J Bioorg Chem 48, 850–853 (2022). https://doi.org/10.1134/S106816202204015X
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S106816202204015X