Abstract—
The interaction of B7-1/B7-2 ligands with CD28/CTLA-4 receptors plays a key role in the regulation of the immune response. The aim of this study was to find and study peptides that interact with the human B7-2 molecule. In the course of the work, three rounds of affinity selection were carried out and individual phage clones were selected, which include peptides with varying degrees of interaction with the coregulatory target B7-2. As a result of DNA sequencing of selected phages, nucleotide sequences encoding peptides that specifically bind to B7-2 were obtained. The identified peptides can be used as a basis for the development of immunotherapeutic drugs for regulating the immune response in the treatment of oncological diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
B7 is an important coregulatory molecule that is expressed on the surface of human antigen-presenting cells (APCs) in two forms: B7-1 and B7-2. B7 ligand receptors on the surface of T cells are CD28/CTLA-4 molecules. Interaction of B7-1 or B7-2 with CD28 promotes T cell activation, proliferation and cytokine secretion. Interaction of B7-1 or B7-2 with CTLA-4 reduces the activation and proliferation of T-lymphocytes [1]. The blockade of the costimulatory pathways for the regulation of the immune response can provide an effective therapy for autoimmune diseases and prevent transplant rejection. The role of blockers can be played by peptides characterized by high specificity of interaction with the target and low molecular weight, which can reduce the number of side effects [2].
Phage peptide libraries are one of the sources for the search for peptides capable of selectively interacting with proteins, lipids, and carbohydrates [3]. To obtain peptides that specifically interact with a molecular target, several cycles of affinity selection of bacteriophages from phage peptide libraries are carried out, followed by sequencing of a DNA region encoding a foreign peptide.
The aim of this work is to search for and study peptides that specifically interact with the coregulatory molecule B7-2 using phage peptide libraries.
RESULTS AND DISCUSSION
Selection of bacteriophages interacting with the coregulatory molecule B7-2. Affinity selection implies selection from a phage peptide library of bacteriophages carrying on their surface foreign peptides that specifically interact with the target—the coregulatory molecule B7-2 containing the Fc region of human IgG1. Selection is carried out due to the formation of a complex of a bacteriophage, protein B7-2 and a magnetic particle, on the surface of which there is a protein G, which specifically interacts with the Fc region of human IgG1. The eluted heterogeneous mixture of bacteriophages is amplified in bacterial culture and used for the next round of selection.
We used the GerLab phage peptide library based on the filamentous bacteriophage fd. To select bacteriophages interacting with recombinant human B7-2 protein (SinoBiological, China), three cycles of affinity selection were carried out. To track the enrichment of the phage population, after each cycle, the biological titer of bacteriophages eluted from magnetic particles was determined and after their amplification in a bacterial culture by the Grazia method [4]. After the second cycle, an increase in the titer of bacteriophages in the eluate was observed. After the third cycle, the titer did not increase, which, possibly, indicates that the saturation limit of the phage population was reached. The change in the titer of bacteriophages during three cycles of affinity selection, as a result of which bacteriophages with the highest affinity for the B7-2 target were obtained, are shown in Table 1.
Nine clones were selected from Petri dishes containing single plaques of the third cycle phage eluate for DNA isolation and subsequent sequencing of the DNA region encoding the foreign peptide.
Identification of amino acid sequences of peptides. Sequenced DNA regions encoding foreign peptides of nine clones were analyzed using the BioEdit 7.2 program and then converted into amino acid sequences. The amino acid sequence of CLAACLGAC is presented in 66.7% of the analyzed phage clones, while the sequences CPSASSGLTC, QMPALMQQ, and AHIGVVSP in 11.1% of clones (Table 2).
In further work, we used clones of phages nos. 1, 2, 6, and 8, which carry on their surface four unique foreign peptides interacting with the B7-2 protein.
Immunochemical properties of the selected peptides. The immunochemical properties of peptides from four phage clones were analyzed by immunoblotting and enzyme-linked immunosorbent assay.
The results of immunoblotting are shown in Fig. 1. The highest optical density, equal to 50, was observed in clone no. 1 (peptide CLARCLGRC). In clones #6 (peptide AHIEVVSP) and #8 (QMPALMQQ peptide) the optical density was lower and amounted to 44 and 23, respectively, which indicates a lower affinity of these peptides for the recombinant protein B7-2. The optical density of clone 2 (peptide CPSASSQLTC) was –10; therefore, this peptide does not bind to the recombinant protein B7-2. The absorbance values for the positive and negative controls were 173 and 0, respectively.
Immunoblot hybridization of the studied phage clones no. 1, 2, 6 and 8 with the recombinant protein B7-2. (a) Dot blot membrane; (b) histogram of optical density values. The positive control (C+) is the recombinant protein B7-2, the negative control (C–) is the fd bacteriophage that does not contain a foreign insert.
The ELISA results are shown in Fig. 2. The highest values of optical density were observed at clones #1 (peptide CLARCLGRC) and #6 (AHIEVVSP peptide): 0.83 ± 0.02 and 0.86 ± 0.03, respectively, which confirms the high affinity of these peptides for the recombinant protein B7-2. ELISA with clones 2 and 4 showed a low specific interaction with the B7-2 target (0.06 ± 0.00 and 0.09 ± 0.01, respectively).
EXPERIMENTAL
Recombinant protein B7-2. We used a commercial recombinant protein consisting of the extracellular domain (Met1-His239) of the human B7-2 protein and Fc region IgG1 person (Pro100-Lys330) with 6His-tag on the C-end (Sino Biological, PRC).
Phage peptide library. The phage peptide library GerLab based on the filamentous bacteriophage fd used in this work was obtained from the laboratory of Professor J.M. Gershoni [5]. It consisted of a mixture of eight phage peptide libraries made on the basis of a phagemid vector of the p88 type and displaying foreign randomized peptides 6, 8, 10, 12 aa in length, as well as peptides closed in a loop, in the composition of the main surface protein pVIII.
Affinity selection of peptides from a phage peptide library. Affinity selection of the phage library against the recombinant protein B7-2, including three cycles, was carried out using magnetic particles (MP) DynabeadsTM Protein G (Invitrogen, Thermo Fisher Scientific, United States), which are homogeneous superparamagnetic beads 2.8 μm in size.
The selection began with washing the MPs with a washing solution (0.1% polysorbate 20 in Tris buffer solution). Then, 1 mL of a blocking solution (5% bovine serum albumin (BSA) in Tris-buffered solution) was added and left for 1 h at 4°C. At this time, the phage library and the recombinant B7-2 protein were mixed in a wash solution and incubated for 20 min at room temperature. The resulting mixture was added to the washed MP and incubated for 15 min at room temperature. The formed MP—phage—target complex was washed from unbound bacteriophages, and the bound phages were eluted. Then the MP was removed and a neutralizing solution (1 M Tris-HCl) was added. The resulting eluate (50 μL) was inoculated into 5 mL of culture, the optical density was measured on a NanoPhotometer N50 spectrophotometer (Implen, Germany) at a wavelength of 600 nm (it was 0.6) and incubated overnight at 37°C.
Titration of bacteriophages, amplification of the eluate, and production of individual phage clones were performed in accordance with the manufacturer’s manual (Ph.D.™ Phage Display Libraries Instruction Manual; NEB, United States) [6] using the strain E. coli DH5αF'+ (NEB, United States).
Isolation of single-stranded DNA of bacteriophages of the library. In 500 μL of the bacteriophage suspension, 100 μL of iodide buffer solution and 250 μL of 96% ethanol were added, the sediment was resuspended, and incubated for 10 min at room temperature. Then ssDNA was precipitated by centrifugation for 10 min at 13 000 g and 4°C. The supernatant was removed, the precipitate was washed with 500 μL of 70% ethyl alcohol. Again, ssDNA was precipitated by centrifugation for 10 min at 13 000 g and 4°C. Then the precipitate of ssDNA was dried in a vacuum and dissolved in 30 µL of distilled water [7].
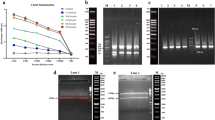
Control of ssDNA isolation was performed by separating nucleic acids in 1% agarose gel.
Sequencing. Sequencing of nucleotide sequences encoding foreign peptides that specifically interact with B7-2 was carried out at the Genomika Center for Collective Use (Novosibirsk Akademgorodok) on an ABI 3130XL capillary sequencer (Genetic Analyzer, Applied Biosystems, United States) using a sequencing primer -96gIII(5'-HO -CCCTCATAGTTAGCGTAACG-3') at a concentration of 1 pM.
Immunoblotting (dot blot). Phage clones (1 μL, 108 virions) were applied to a nitrocellulose membrane once and dried in air. As a negative control (C–) we used bacteriophage fd (1 μL, 108 virions), not containing a foreign insert; as a positive control (C+), recombinant protein B7-2 (Sino Biological, China; 1 μL, 2.5 μg/mL). Cellulose was incubated with a blocking Tris buffer solution (15 mL) containing 1% BSA and 0.1% polysorbate 20 to prevent nonspecific interaction [8].
After three times washing with Tris buffer solution with 0.1% polysorbate 20, 1.5 mL of recombinant B7-2 protein (2.5 μg/ml) in a blocking solution was applied and incubated at room temperature for 15 min.
After washing three times with a washing solution, 15 mL of a solution of goat antibodies against the Fc region of human IgG conjugated with alkaline phosphatase (Invitrogen, United States) was applied at a working dilution of 1 : 5000 in blocking solution and incubated at room temperature for 15 min.
After washing three times with Tris buffer solution with 0.1% polysorbate 20, a mixture of 5-bromo-4-chloro-3-indolyl phosphate (BCIP, 0.21 mg/mL) and nitro blue tetrazolium (NBT, 0.42 mg/mL in water) with a volume of 15 mL, incubated at room temperature for 15 min in the dark.
After washing three times with water, the nitrocellulose membrane was dried in air in the dark. The signals on the membrane were converted into a computer image using the ImageJ application (https://imagej.nih.gov/ij/).
Enzyme-linked immunosorbent assay (ELISA). For ELISA, medium sorption 96-well plates (Jet Biofil, China) were used. The wells were filled with 100 μL of a solution of recombinant protein B7-2 (Sino Biological, China) at a concentration of 2.5 μg/mL in Tris buffer solution (pH 8.6) and sorbed at 4°C overnight. Then, nonspecific binding sites were blocked by adding 200 μL of Tris buffer solution (pH 7.0) containing 5% BSA, and incubation in a PST-60HL thermo-shaker for plates (BioSan, Latvia) at 37°C and 200 rpm for 2 h. After removing the blocking solution, bacteriophages were added to the wells (100 μL in blocking solution, 108 virions) and incubated for 1 h at 37°C. As a negative control (C–), we used bacteriophage fd (100 μL in a blocking solution, 108 virions), not containing a foreign insert; as a positive control (C+), recombinant protein B7-2 (Sino Biological, China; 100 μL, 2.5 μg/mL). After washing three times with a washing solution containing Tris-buffer solution with 0.5% polysorbate 20, 100 μL of conjugate of monoclonal antibody against filamentous bacteriophage M13 (GE Healthcare Life Sciences, United States), labeled with horseradish peroxidase, was added in the selected working dilution 1 : 5000 and incubated for 1 h at 37°C. After washing six times with washing solution, a substrate based on 3,3', 5,5'-tetramethylbenzidine (TMB; Sigma-Aldrich, United States) was added to the wells. The reaction was stopped by adding 50 μL of 1 M sulfuric acid to each well.
The results were detected using an iMark plate photometer (Bio-Rad, United States) at a wavelength of 450 nm. The mean values of optical density and errors of the mean values (M ± SE) were calculated, according to the data obtained; diagrams were plotted indicating the values of the standard deviation [nine].
CONCLUSIONS
As a result of three cycles of affinity selection using the GerLab phage peptide library and the commercial recombinant B7-2 protein, four unique peptide sequences interacting with the human coregulatory protein B7-2 were selected. Using immunoblotting and ELISA methods, it was shown that phage clones no. 1 (peptide CLARCLGRC) and no. 6 (peptide AHIEVVSP) have the highest affinity for the recombinant protein B7-2, while phage clones no. 2 (peptide CPSASSQLTC) and no. 8 (peptide QMPALMQQ) show no specific interaction with the target.
The identified peptides with high affinity for the human coregulatory protein B7-2 can serve as a basis for calculating and designing artificial immunogens for use in the development of immunotherapeutic agents for the treatment of cancer.
REFERENCES
Chapoval, A.I., Chapoval, S.P., Shcherbakova, N.S., and Shcherbakov, D.N., Russ. J. Bioorg. Chem., 2019, vol. 45, pp. 225–240. https://doi.org/10.1134/S0132342319040110
Titov, M.I., Vestn. St.-Peterb. Univ., 2013, no. 4, pp. 86–102.
Urban, I.G., Mosmaier, M.A., Bosma, T., and Prassler, I., RF Patent no. RU2702087S2, 2019.
Borgoyakova, M.B. and Il’ichev, A.A., Bakteriofagi. Praktikum po molekulyarnoi virusologii. Ucheb.-metod. posobie (Bacteriophages. Workshop on Molecular Virology: A Study Guide), Novosibirsk: Novosib. Gos. Univ., 2013.
Ryvkin, A., Ashkenazy, H., Weiss-Ottolenghi, Y., Piller, C., Pupko, T., and Gershoni, J.M., Nucleic Acids Res., 2018, vol. 46, p. 52. https://doi.org/10.1093/nar/gky077
Ph.D™ Phage Display Libraries Instructional Manual, New England BioLabs Inc.
Wilson, R.K., BioTechniques, 1993, vol. 15, pp. 414–416, 418–420, 422.
Reitinger, S., Petriv, I., Mehr, K., Hansen, C.L., and Withers, S.G., J. Virol. Methods, 2012, vol. 185, pp. 171–174. https://doi.org/10.1016/j.jviromet.2012.06.021
Shcherbakova, N.S., Chikaev, A.N., and Karpenko, L.I., Mol. Genet., Mikrobiol., Virusol., 2012, no. 1, pp. 20–25.
Funding
This work was supported by the Russian Foundation for Basic Research (grants no. 19-44-220008, 17-54-33003) and the state assignment of the Ministry of Education and Science of Russia (no. FZMW-2020-0007).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
This article does not describe any research involving humans and the use of animals as research subjects.
Conflict of Interests
The authors declare that they have no conflicts of interest.
Additional information
Corresponding author: phone: +7 (3852) 298-142.
Rights and permissions
Open Access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kolosova, E.A., Viktorina, O.E., Chapoval, A.I. et al. Search for Peptides Specifically Binding with the B7-2 Costimulatory Molecule. Russ J Bioorg Chem 47, 1220–1224 (2021). https://doi.org/10.1134/S106816202106011X
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S106816202106011X