Abstract—
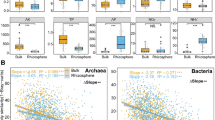
Microbiomes of strongly skeletal residual calcareous pelozems (Skeletal Leptosols (Loamic)), carbopetrozems (Calcaric Leptosols (Protic)), petrozems (Skeletal Leptosols (Protic)), and cryozems (Oxyaquic Cryosols (Loamic)) in the north of Novaya Zemlya were studied by the methods of molecular biology. The number of copies of 16S rRNA genes was small and ranged from 2.30 × 107 to 1.63 × 109 gene copies/g soil for archaea and from 3.47 × 108 to 2.26 × 1011 gene copies/g soil for bacteria; the number of copies of ribosomal genes ITS rRNA of fungi varied from 8.87 × 106 to 7.56 × 109 gene copies/g soil. The content of copies of ribosomal genes of all groups of microorganisms sharply decreased down the soil profiles. Bacteria predominated among prokaryotes (up to 90%). The greatest abundance (20%) was manifested by the phyla Proteobacteria, Actinobacteria, and Acidobacteria; the abundances of Bacteroidetes, Firmicutes, Verrucomicrobia, Gemmatimonadetes, and Chloroflexi were about 1–10%. The Archaea domain represented mainly by the Ferroplasma genus (phylum Euryarchaeota), accounted for ≤4% of prokaryotes. The taxonomic diversity of prokaryotes increased down the soil profiles and reached maximum values in the suprapermafrost horizons, where the number of candidate phyla typical of marine ecosystems—Latescibacteria, Tectomicrobia, Parcubacteria, Saccaribacteria, Hydrogenedentes, Peregrinibacteria, Ignavibacteria, and Gracilibacteria—was high.





Similar content being viewed by others
REFERENCES
E. V. Blagodatskaya, M. V. Semenov, and A. V. Yakushev, Activity and Biomass of Soil Microorganisms in a Changing Environment (KMK Press, Moscow, 2016) [in Russian].
M. V. Korneykova, D. A. Nikitin, A. V. Dolgikh, and A. S. Soshina, “Soil mycobiota of the Apatity city, Murmansk region,” Mikologiya Fitopatol. 54 (4), 264–277 (2020). https://doi.org/10.31857/S0026364820040078
M. V. Korneykova and D. A. Nikitin, “Qualitative and quantitative characteristics of the soil microbiome in the impact zone of the Kandalaksha aluminum smelter,” Eurasian Soil Sci. 54 (6), 897–906 (2021). https://doi.org/10.1134/S1064229321060089
L. V. Lysak, I. A. Maksimova, D. A. Nikitin, A. E. Ivanova, A. G. Kudinova, V. S. Soina, and O. E. Marfenina, “Microbial communities of soils in East Antarctica,” Moscow Univ. Bull. Ser. 16: Biol. 73 (3), 132–140 (2018).
O. E. Marfenina, D. A. Nikitin, and A. E. Ivanova, “The structure of fungal biomass and diversity of cultivated micromycetes in Antarctic soils (Progress and Russkaya stations),” Eurasian Soil Sci. 49 (8), 934–941. https://doi.org/10.1134/S106422931608007X
D. A. Nikitin, O. E. Marfenina, and I. A. Maksimova, “The use of the succession approach in studying the species composition of microscopic fungi and the content of fungal biomass in Antarctic soils,” Mikologiya Fitopatol. 51 (4), 211–219 (2017).
D. A. Nikitin, M. V. Semenov, A. A. Semikolennykh, I. A. Maksimova, A. V. Kachalkin, and A. E. Ivanova, “Fungal biomass and species diversity of the cultivated mycobiota of soils and substrates of Northbrook, Franz Josef Land,” Mikologiya Fitopatol. 53 (4), 210–222 (2019). https://doi.org/10.1134/S002636481904010X
D. A. Nikitin, E. A. Ivanova, A. D. Zhelezova, M. V. Semenov, R. G. Gadzhiumarov, A. K. Tkhakakhova, T. I. Chernov, N. A. Ksenofontova, and O. V. Kutovaya, “Assessment of the impact of no-till technology and plowing on the microbiome of southern agrochernozems,” Eurasian Soil Sci. 53 (12), 1782–1793 (2020). https://doi.org/10.1134/S106422932012008X
D. A. Nikitin, L. V. Lysak, D. V. Badmadashiev, S. S. Kholod, N. S. Mergelov, A. V. Dolgikh, and S. V. Goryachkin, “Biological activity of soils in the north of the Novaya Zemlya Archipelago: effect of the largest glacial sheet in Russia,” Eurasian Soil Sci. 54 (10), 1496–1516. https://doi.org/10.1134/S1064229321100082
D. A. Nikitin, L. V. Lysak, O. V. Kutovaya, and T. A. Gracheva, “Ecological-trophic structure and taxonomic characteristics of the communities of soil microorganisms in the northern part of the Novaya Zemlya Archipelago,” Eurasian Soil Sci. 54 (11), 1689-1704 (2021). https://doi.org/10.1134/S1064229321110107
D. A. Nikitin and M. V. Semenov, “Characterization of Franz Josef Land Soil Mycobiota by Microbiological Plating and Real-time PCR,” Microbiology (Moscow) 91 (1), 56–66 (2022). https://doi.org/10.1134/S002626172201009X
M. V. Semenov, “Metabarcoding and metagenomics in soil ecology research,” Zhurn. Obshch. Biol. 80 (6), 403–417 (2019). https://doi.org/10.1134/S004445961906006X
M. V. Semenov, N. A. Manucharova, and A. L. Stepanov, “Biomass and taxonomic structure of microbial communities in soils of the right-bank basin of the Oka River,” Eurasian Soil Sci. 52 (8), 971–981. https://doi.org/10.1134/S106422931908012X
P. Baldrian, “The known and the unknown in soil microbial ecology,” FEMS Microbiol. Ecol. 95 (2), fiz005 (2019). https://doi.org/10.1093/femsec/fiz005
A. A. Belov, V. S. Cheptsov, N. A. Manucharova, and Z. S. Ezhelev, “Bacterial communities of Novaya Zemlya Archipelago ice and permafrost,” Geosciences 10 (2), 67 (2020).https://doi.org/10.3390/geosciences10020067
J. E. Box, W. T. Colgan, T. R. Christensen, N. M. Schmidt, M. Lund, F. J. W. Parmentier, R. Brown, et al., “Key indicators of Arctic climate change: 1971–2017,” Environ. Res. Lett. 4 (4), 045010 (2019). https://doi.org/10.1088/1748-9326/aafc1b
J. G. Caporaso, J. Kuczynski, J. Stombaugh, K. Bittinger, F. D. Bushman, E. K. Costello, N. Fierer, et al., “QIIME allows analysis of high-throughput community sequencing data,” Nat. Methods 7 (5), 335–336 (2010). https://doi.org/10.1038/nmeth.f.303
M. Dopson, C. Baker-Austin, A. Hind, J. P. Bowman, P. L. Bond, “Characterization of Ferroplasma isolates and Ferroplasma acidarmanus sp. nov., extreme acidophiles from acid mine drainage and industrial bioleaching environments,” Appl. Environ. Microbiol. 70 (4), 2079–2088 (2004).
P. A. Figueroa-Gonzalez, T. L. Bornemann, P. S. Adam, J. Plewka, F. Revesz, C. Hagen, A. Tancsics, J. Probst, “Saccharibacteria as organic carbon sinks in hydrocarbon-fueled communities,” Front. Microbiol. (2020). https://doi.org/10.3389/fmicb.2020.587782
C. G. Flocco, W. P. Mac Cormack, and K. Smalla, “Antarctic soil microbial communities in a changing environment: their contributions to the sustainability of Antarctic ecosystems and the bioremediation of anthropogenic pollution,” in The Ecological Role of Microorganisms in the Antarctic Environment (Springer, Cham., 2019), pp. 133–161.https://doi.org/10.1007/978-3-030-02786-5_7
R. A. Garrett and H. P. Klenk, Archaea: Evolution, Physiology, and Molecular Biology (John Wiley & Sons, 2008).
F. O. Glöckner, P. Yilmaz, C. Quast, J. Gerken, A. Beccati, A. Ciuprina, G. Brunsa, P. Yarzac, J. Pepliesc, R. Westram, and W. Ludwig, “25 Years of serving the community with ribosomal RNA gene reference databases and tools,” J. Biotechnol. 261, 169–176 (2017). https://doi.org/10.1007/978-3-030-02786-5_7
C. A. Guerra, A. Heintz-Buschart, J. Sikorski, A. Chatzinotas, N. Guerrero-Ramirez, S. Cesarz, L. Beaumelle, et al., “Blind spots in global soil biodiversity and ecosystem function research,” Nature communic. 11 (1), 1–13 (2020). https://doi.org/10.1038/s41467-020-17688-2
C. M. Hansel, S. Fendorf, P. M. Jardine, and C. A. Francis, “Changes in bacterial and archaeal community structure and functional diversity along a geochemically variable soil profile,” Appl. Environ. Microbiol. 74, 1620–1633 (2008). https://doi.org/10.1128/AEM.01787-07
R. Jacoby, M. Peukert, A. Succurro, A. Koprivova, and S. Kopriva, “The role of soil microorganisms in plant mineral nutrition-current knowledge and future directions,” Frontiers Plant Sci. 8, 1617 (2017). https://doi.org/10.3389/fpls.2017.01617
P. H. Janssen, “Identifying the dominant soil bacterial taxa in libraries of 16S RRNA and 16S RRNA genes,” Appl. Environ. Microbiol. 72 (3), 1719–1728 (2006). https://doi.org/10.1128/AEM.72.3.1719-1728.2006
H. M. Kim, J. Y. Jung, E. Yergeau, C. Y. Hwang, L. Hinzman, S. Nam, S. G. Hong, O. Kim, J. Chun, and Y. K. Lee, “Bacterial community structure and soil properties of a subarctic tundra soil in council, Alaska,” FEMS Microbiol. Ecol. 89 (2), 465–475 (2014). https://doi.org/10.1111/1574-6941.12362
K. D. Kits, C. J. Sedlacek, E. V. Lebedeva, P. Han, A. Bulaev, P. Pjevac, A. Daebeler, et al., “Kinetic analysis of a complete nitrifier reveals an oligotrophic lifestyle,” Nature 549, 269–272 (2017). https://doi.org/10.1038/nature23679
L. A. Malard and D. A. Pearce, “Microbial diversity and biogeography in Arctic Soils,” Environ. Microbiol. Rep 10 (6), 611–625 (2018). https://doi.org/10.1111/1758-2229.12680
M. Podar, C. B. Abulencia, M. Walcher, D. Hutchison, K. Zengler, J. A. Garcia, T. Holland, D. Cotton, L. Hauser, and M. Keller, “Targeted access to the genomes of low-abundance organisms in complex microbial communities,” Appl. Environ. Microbiol. 10 (73), 3205–3214 (2007). https://doi.org/10.1128/AEM.02985-06
G. Pold, J. P. Schimel, and S. A. Sistla, “Soil bacterial communities vary more by season than with over two decades of experimental warming in Arctic tussock tundra,” Elementa: Science of the Anthropocene 9 (1) (2021). https://doi.org/10.1525/elementa.2021.00116
E. Post, R. B. Alley, T. R. Christensen, M. Macias-Fauria, B. C. Forbes, M. N. Gooseff et al., “Virginia and Muyin Wang. The polar regions in a 2°C warmer world,” Sci. Adv. 5 (12), 12 (2019). https://doi.org/10.1126/sciadv.aaw9883
E. Pruesse, C. Quast, K. Knittel, B. M. Fuchs, W. Ludwig, J. Peplies, and F. O. Glöckner, “SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB,” Nucleic Acid Res. 35 (21), 7188–7196 (2007). https://doi.org/10.1093/nar/gkm864
C. Rinke, P. Schwientek, A. Sczyrba, N. N. Ivanova, I. J. Anderson, J.-F. Cheng, A. Darling, et al., “Insights into the phylogeny and coding potential of microbial dark matter,” Nature 499 (7459), 431–437 (2013). https://doi.org/10.1038/nature12352
C. M. Sieber, A. J. Probst, A. Sharrar, B. C. Thomas, M. Hess, S. G. Tringe, and J. F. Banfield, “Recovery of genomes from metagenomes via a dereplication, aggregation and scoring strategy,” Nature Microbiol. 3 (7), 836 (2018).
J. S. Singh and V. K. Gupta, “Soil microbial biomass: a key soil driver in management of ecosystem functioning,” Sci. Total Environ. 634, 497–500 (2018).
A. Spang, R. Hatzenpichler, C. Brochier-Armanet, and T. Rattei, “Distinct Gene Set in Two Different Lineages of Ammonia-Oxidizing Archaea Supports the Phylum Thaumarchaeota,” Trends in Microbiology 18 (8), 331–340 (2010). https://doi.org/10.1016/j.tim.2010.06.003
A. D. Steen, A. Crits-Christoph, P. Carini, K. M. DeAngelis, N. Fierer, K. G. Lloyd, and J. C. Thrash, “High proportions of bacteria and archaea across most biomes remain uncultured,” The ISME J. (2019). https://doi.org/10.1038/s41396-019-0484-y
B. M. Tripathi, H. M. Kim, J. Y. Jung, S. Nam, H. T. Ju, M. Kim, and Y. K. Lee, “Distinct taxonomic and functional profiles of the microbiome associated with different soil horizons of a moist tussock tundra in Alaska,” Frontiers in Microbiology 10, 1442 (2019). https://doi.org/10.3389/fmicb.2019.01442
E. Yergeau, K. K. Newsham, D. A. Pearce, and G. A. Kowalchuk, “Patterns of bacterial diversity across a range of Antarctic terrestrial habitats,” Environ. Microbiol. 9, 2670–2682 (2007). https://doi.org/10.1111/j.1462-2920.2007.01379.x
W. Zhang, P. A. Miller, C. Jansson, P. Samuelsson, J. Mao, B. Smith, “Self-amplifying feedbacks accelerate greening and warming of the Arctic,” Geophys. Rev. Lett. 45 (14), 7102–7111 (2018).
A. Zhelezova, T. Chernov, A. Tkhakakhova, N. Xenofontova, M. Semenov, O. Kutovaya, “Prokaryotic community shifts during soil formation on sands in the tundra zone,” Plos One 14 (4), e0206777 (2019). https://doi.org/10.1371/journal.pone.0206777
ACKNOWLEDGMENTS
The authors thank the Arctic Floating University project of the Lomonosov Northern Arctic Federal University and personally K.S. Zaikov for organizing field work on Novaya Zemlya. The authors thank the staff of the Department of Geography and Evolution of Soils of the Institute of Geography of the Russian Academy of Sciences and personally S.V. Goryachkin for their help in determining the taxonomic position of the studied soils.
Funding
This study was supported by the Russian Foundation for Basic Research, project no. 20-04-00328.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflicts of interest.
Additional information
Translated by D. Konyushkov
SUPPLEMENTARY MATERIALS
Table 1. Soil properties in the northern part of Novaya Zemlya
Supplementary Information
Rights and permissions
About this article
Cite this article
Nikitin, D.A., Lysak, L.V. & Badmadashiev, D.V. Molecular Biological Characteristics of Soil Microbiome in the Northern Part of the Novaya Zemlya Archipelago. Eurasian Soil Sc. 55, 1106–1115 (2022). https://doi.org/10.1134/S1064229322080130
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1064229322080130