Abstract
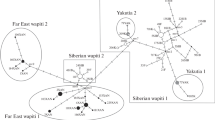
Analysis of MtDNA cytochrome b gene (1140 bp) polymorphism of 106 samples red deer (Cervus elaphus) from different regions of Eurasia was performed; the phylogenetic relationships of groups throughout the entire inhibiting area (including North America) were reconstructed. Totally 75 haplotypes were detected, 33 of which were found in the European and 42 in the Asian part of the area. There were no identical haplotypes for these two parts of the area found. The close relatedness between Siberian red deer (C. e. sibirica) and North American wapiti (C. e. canadensis) was confirmed. Red deer inhibiting Yakutia were close to the Siberian red deer from Altai and Tuva, whereas red deer inhibiting Krasnoyarsk and Irkutsk regions formed a separate clade. Overall, the reconstructed phylogeographic pattern of the species was significantly different from the subspecies differentiation based on morphological traits.
Similar content being viewed by others
References
Bandelt, H.-J., Forster, P., and Ruhl, A., Median-Joining Networks for Inferring Intraspecific Phylogenies, Mol. Biol. Evol., 1999, vol. 16, pp. 37–48.
Danilkin, A.A., Mlekopitayushchie Rossii i sopredel’nykh regionov. Olen’i (Mammals of Russia and Adjacent Regions: Cervidae), Moscow: GEOS, 1999.
Ellerman, J.R. and Morrison-Scot, T.C.S., Checklist of Palaearctic and Indian Mammals 1758 to British Museum (Nat. Hist.), L.: Brit. Mus. Press, 1946.
Excoffier, L. and Lischer, H., Arlequin Suite ver. 3.5: A New Series of Programs to Perform Population Genetics Analyses under Linux and Windows, Mol. Ecol. Res., 2010, vol. 10, pp. 564–567.
Geptner, V.G. and Tsalkin, V.I., Oleni SSSR (sistematika i zoogeografiya) (Deer of the USSR (Systematics and Zoogeography)), Moscow: MOIP, 1947.
Geptner, V.G., Nasimovich, A.A., and Bannikov, A.G., Mlekopitayushchie Sovetskogo Soyuza. Parnokopytnye i neparnokopytnye (Mammals of the Soviet Union. Artiodactyls and Solipeds), Moscow: Vyssh. Shkola, 1961, vol. 1.
Groves, C., The Genus Cervus in Eastern Eurasia, Eur. J. Wildl. Res., 2006, vol. 52, pp. 14–22.
Grubb, P. and Gardner, A.L., List of Species and Subspecies of the Families Tragulidae, Moschidae, and Cervidae, in Deer Status Survey and Conservation Action Plan, IUCN/SSC Deer Specialist Group, Oxford: Inform. Press, 1998.
Hall, T.A., BioEdit: A User-Friendly Biological Sequence Alignment Editor and Analysis Program for Windows 95/98/NT, Nucl. Acids Symp., 1999, vol. 41, pp. 95–98.
Kimura, M., A Simple Method for Estimating Evolutionary Rate of Base Substitutions through Comparative Studies of Nucleotide Sequences, J. Mol. Evol., 1980, vol. 16, pp. 111–120.
Kocher, T.D., Thomas, W.K., Meyer, A., et al., Dynamics of Mitochondrial DNA Evolution in Animals: Amplification and Sequencing with Conserved Primers, Proc. Natl. Acad. Sci. USA, 1989, vol. 86, pp. 6196–6200.
Kumar, S., Dudley, J., Nei, M., and Tamura, K., MEGA: A Biologist-Centric Software for Evolutionary Analysis of DNA and Protein Sequences, Brief. Bioinform., 2008, vol. 9, pp. 299–306.
Kuznetsova, M.V., Volokh, A.M., Domnich, V.I., et al., Molecular-Genetic Study of the Red Deer, Cervus elaphus (Cervidae) in Eastern Europe, Vestn. Zool., 2007, vol. 41, no. 6, pp. 505–509.
Lemmon, A.R. and Milinkovitch, M.C., The Metapopulation Genetic Algoritm: An Efficient Solution for the Problem of Large Phylogeny Estimation, Proc. Natl. Acad. Sci. USA, 2002, vol. 99, pp. 10516–10521.
Lowe, V.W. and Gardiner, A.S., A Re-Examination of the Subspecies of Red Deer (Cervus elaphus) with Particular Reference to the Stocks in Britain, J. Zool., 1974, vol. 174, pp. 185–201.
Ludt, C.J., Schroeder, W., Rottmann, O., and Kuehn, R., Mitochondrial DNA Phylogeography of Red Deer (Cervus elaphus), Mol. Phylog. Evol., 2004, vol. 31, pp. 1064–1083.
Mahmut, H., Masuda, R., Onuma, M., et al., Molecular Phylogeography of the Red Deer (Cervus elaphus) Populations in Xinjiang of China: Comparison with Other Asian, European, and North American Populations, Zool. Sci., 2002, vol. 19, no. 4, pp. 485–495.
Pérez-Espona, S., Pérez-Barbería, F.J., Goodall-Copestake, W.P., et al., Genetic Diversity and Population Structure of Scottish Highland Red Deer (Cervus elaphus) Populations: A Mitochondrial Survey, Heredity (Edinb.), 2009, vol. 102, no. 2, pp. 199–210.
Wada, K., Okumura, K., Nishibori, M., et al., The Complete Mitochondrial Genome of the Domestic Red Deer (Cervus elaphus) of New Zealand and Its Phylogenic Position within the Family Cervidae, Anim. Sci. J., 2010, vol. 81, no. 5, pp. 551–557.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © M.V. Kuznetsova, A.A. Danilkin, M.V. Kholodova, 2012, published in Izvestiya Akademii Nauk, Seriya Biologicheskaya, 2012, No. 4, pp. 391–398.
Rights and permissions
About this article
Cite this article
Kuznetsova, M.V., Danilkin, A.A. & Kholodova, M.V. Phylogeography of red deer (Cervus elaphus): Analysis of MtDNA cytochrome b polymorphism. Biol Bull Russ Acad Sci 39, 323–330 (2012). https://doi.org/10.1134/S1062359012040048
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1062359012040048