Abstract
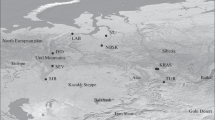
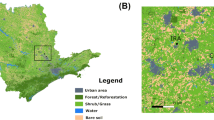
The objectives of conservation and sustainable forest management require in depth study of genomes of woody plants and definition of their intraspecific genetic diversity. In recent years, an approach was developed based on the study of “candidate genes” that can potentially be involved in the formation of adaptive traits. In this study, we investigated nucleotide polymorphism of several adaptive candidate genes in the populations of Siberian larch (Larix sibirica Ledeb.) in the Urals. Representatives of this genus are among the most valuable and widely distributed forest tree species in Russia. From ten selected gene loci in the genome of L. sibirica, we isolated and investigated three loci, one of which (ABA-WDS) was sequenced in L. sibirica for the first time. The total length of the analyzed sequence in each individual amounted to 2865 bp. The length of locus alignment was from 360 bp to 1395 bp. In total, we identified 200 polymorphic positions. The most conservative is locus 4CL1-363, and the most polymorphic is locus sSPcDFD040B03103-274. The studied populations of L. sibirica are characterized by a high level of nucleotide polymorphism in comparison with other species and genuses (Picea, Pinus, Pseudotsuga, Abies) conifers plants (Hd = 0.896; π = 0.007; θW = 0.015). The most selectively neutral polymorphism (D T =–0.997) was attributed to locus 4CL1-363, and polymorphism with high probability of adaptability (D T =–1.807) was determined for the ABA-WDS locus. We identified 54 SNP markers, only five of which were nonsynonymous (9.26%) replacements. The average frequency of SNPs in the three studied loci of L. sibirica was one SNP in 53 bp. We detected unique SNP markers for eight populations, which could potentially be used to identify populations. Populations that are characterized by the highest number of unique SNP markers can be recommended for selection in order to preserve the gene pool of the species.
Similar content being viewed by others
References
Vetchinnikova, L.V., Titov, A.F., and Kuznetsova, T.Yu., Karel’skaya bereza: biologicheskie osobennosti, dinamika resursov i vosproizvodstvo (Karelian Birch: Biological Features, Dynamic Resource and Reproduction), Petrozavodsk: Karelskii Nauchnyi Tsentr Rossiiskoi Akademii Nauk, 2013.
Nemova, N.N., Mechanisms of biochemical adaptations of aquatic organisms: ecological and evolutionary aspects, in Sovremennye problemy fiziologii i biokhimii vodnykh organizmov (Modern Challenges in Physiology and Biochemistry of Aquatic Organisms), vol. 1: Ekologicheskaya fiziologiya i biokhimiya vodnykh organizmov (Ecological Physiology and Biochemistry of Aquatic Organisms), Petrozavodsk: Karelskii Nauchnyi Tsentr Rossiiskoi Akademii Nauk, 2010, pp. 198–215.
Krutovsky, K.V. and Neale, D.B, Nucleotide diversity and linkage disequilibrium in cold-hardiness- and wood quality-related candidate genes in Douglas fir, Genetics, 2005, vol. 171, no. 4, pp. 2029–2041.
Neale, D.B. and Ingvarsson, P.K, Population,quantitative and comparative genomics of adaptation in forest trees, Curr. Opin. Plant Biol., 2008, no. 11, pp. 149–155.
Politov, D.V, Application of molecular markers in forestry for identification, inventory and assessment of the genetic diversity of forest resources, Lesokh. Inf., 2008, nos. 3–4, pp. 24–27.
Krutovskii, K.V, From population genetics to population genomics of forest trees: integrated population genomics approach, Russ. J. Genet., 2006, vol. 42, no. 10, pp. 1088–1100.
Eckert, A.J., Wegrzyn, J.L., Pande, B., et al., Multilocus patterns of nucleotide diversity and divergence reveal positive selection at candidate genes related to cold hardiness in coastal Douglas fir (Pseudotsuga menziesii var. menziesii), Genetics, 2009, vol. 183, no. 1, pp. 289–298.
Krutovskii, K.V, Perspectives of using of genomic research in forestry, Sib. Lesnoi Zh., 2014, no. 4, pp. 11–15.
Chhatre, V.E., Byram, T.D., Neale, D.B., et al., Genetic structure and association mapping of adaptive and selective traits in the East Texas loblolly pine (Pinus taeda L.) breeding populations, Tree Genet. Genomes, 2013, vol. 9, no. 5, pp. 1161–1178.
Koralewski, T.E., Brooks, J.E., and Krutovsky, K.V, Molecular evolution of drought tolerance and wood strength related candidate genes in loblolly pine (Pinus taeda L.), Silvae Genet., 2014, vol. 63, nos. 1–2, pp. 59–66.
Semerikov, V.L., Semerikova, S.A., and Polezhaeva, M.A, Nucleotide diversity and linkage disequilibrium of adaptive significant genes in Larix (Pinaceae), Russ. J. Genet., 2013, vol. 49, no. 9, pp. 915–923.
Gonzalez-Martinez, S.C., Wheeler, N.C., and Ersoz, E, Association genetics in Pinus taeda L.: 1. Wood property traits, Genetics, 2007, vol. 175, pp. 399–409.
Gonzalez-Martinez, S.C., Ersoz, E., Brown, G.R., et al., DNA sequence variation and selection of tag SNPs at candidate genes for drought-stress response in Pinus taeda L., Genetics, 2006, vol. 172, pp. 1915–1926.
Pyhäjärvi, T., García-Gil, M.R., Knürr, T., et al., Demographic history has influenced nucleotide diversity in European Pinus sylvestris populations, Genetics, 2007, vol. 177, pp. 1713–1724.
Wachowiak, W., Balk, P.A., and Savolainen, O, Search for nucleotide diversity patterns of local adaptation in dehydrins and other cold-related candidate genes in Scots pine (Pinus sylvestris L.), Tree Genet. Genomes, 2009, vol. 5, no. 1, pp. 117–132.
Vangestel, C., Vázquez-Lobo, A., Martínez-García, P.J., et al., Patterns of neutral and adaptive genetic diversity across the natural range of sugar pine (Pinus lambertiana Dougl.), Tree Genet. Genomes, 2016, vol. 12, p. 51.
Heuertz, M., De Paoli, E., Källman, T., et al., Multilocus patterns of nucleotide diversity, linkage disequilibrium and demographic history of Norway spruce (Picea abies (L.) Karst), Genetics, 2006, vol. 174, pp. 2095–2105.
Pavy, N., Namroud, M.C., Gagnon, F., et al., The heterogeneous levels of linkage disequilibrium in white spruce genes and comparative analysis with other conifers, Heredity, 2012, vol. 108, pp. 273–284.
Mosca, E., Eckert, A.J., Liechty, J.D., and Wegrzyn, J.L, Contrasting patterns of nucleotide diversity for four conifers of Alpine European forests, Evol. Appl., 2012, no. 5, pp. 762–775.
Altukhov, Yu.P., The dynamics of the population gene pools under anthropogenic influences, Inf. Vestn. Vavilovskogo O-va Genet. Sel., 2004, vol. 8, no. 2, pp. 40–59.
Vidyakin, A.I., Boronnikova, S.V., Nechaeva, Yu.S., et al., Genetic variation, population structure,and differentiation in scots pine (Pinus sylvestri L.) from the northeast of the Russian plain as inferred from the molecular genetic analysis data, Russ. J. Genet., 2015, vol. 51, no. 12, pp. 1213–1220.
Putenikhin, V.P., Farukshina, G.G., and Shigapov, Z.Kh., Listvennitsa Sukacheva na Urale: izmenchivost’ i populyatsionno- geneticheskaya struktura (Sukachev Larch in the Urals: Variation and Population-Genetic Structure), Moscow Nauka, 2004.
Dylis, N.V., Sibirskaya listvennitsa: materialy k sistematike, geografii i istorii (Siberian Larch: Materials to Systematics, Geography, and History), Moscow: Mosk. O-vo Ispytatekei Prirody, 1947.
Urusov, V.M., Lobanova, I.I., and Varchenko, L.I., Khvoinye rossiiskogo Dal’nego Vostoka–tsennye ob”ekty izucheniya, okhrany, razvedeniya i ispol’zovaniya (Conifers of the Russian Far East–The Valuable Objects of Study, Conservation, Breeding and Use), Vladivostok: Dal’nauka, 2007.
Rogers, S.O. and Bendich, A.J, Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues, Plant Mol. Biol., 1985, vol. 1, no. 19, pp. 69–76.
Saleh, A. and Pagés, M, Plant AP2/ERF transcription factors, Genetika, 2003, vol. 35, no. 1, p. 37.
Goyal, K., Walton, L.J., and Tunnacliffe, A., LEA proteins prevent protein aggregation due to water stress, Biochem. J., 2005, vol. 388, pp. 151–157.
Tolleter, D., Jaquinod, M., Mangavel, C., et al., Structure and function of a mitochondrial late embryogenesis abundant protein are revealed by desiccation, Plant Cell, 2007, vol. 19, pp. 1580–1589.
Dixon, D.P., Skipsey, M., and Edwards, R, Role for glutathione transferases in plant secondary metabolism, Phytochemistry, 2010, vol. 71, pp. 338–350.
Zhang, J., Jia, W., Yang, J., and Ismail, A.M, Role of ABA in integrating plant responses to drought and salt stresses, Field Crops Res., 2006, vol. 97, pp. 111–119.
Gramzow, L. and Theissen, G., A hitchhiker’s guide to the MADS world of plants, Genome Biol., 2010, vol. 11, p. 214.
Hall, T.A., BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT, Nucleic Acids Symp., 1999, vol. 41, pp. 95–98.
Librado, P. and Rozas, J., DnaSP v5: a software for comprehensive analysis of DNA polymorphism data, Bioinformatics, 2009, vol. 25, pp. 1451–1452.
Nei, M, Analysis of gene diversity in subdivided populations, Proc. Natl. Acad. Sci. U.S.A., 1973, vol. 70, pp. 3321–3323.
Nei, M., Molecular Evolutionary Genetics, New York: Columbia Univ. Press, 1987.
Tajima, F, Statistical methods to test for nucleotide mutation hypothesis by DNA polymorphism, Genetics, 1989, vol. 123, pp. 585–595.
Nei, M. and Li, W.-H., Mathematical model for studying genetic variation in terms restriction endonucleases, Proc. Natl. Acad. Sci. U.S.A., 1979, vol. 76, pp. 5269–5273.
Watterson, G.A, On the number of segregating sites in genetical models without recombination, Theor. Popul. Biol., 1975, no. 7, pp. 256–276.
Brookes, A.J, The essence of SNP, Gene, 1999, vol. 234, pp. 177–186.
Wei, X.-X. and Wang, X.-Q, Recolonization and radiation in Larix (Pinaceae): evidence from nuclear ribosomal DNA paralogues, Mol. Ecol., 2004, no. 13, pp. 3115–3123.
Khatab, I.A., Ishiyama, H., Inomata, N., et al., Phylogeography of Eurasian Larix species inferred from nucleotide variation in two nuclear genes, Genes Genet. Syst., 2008, no. 83, pp. 55–56.
Igoshina, K.N, Larch in the Urals, in Materialy po istorii flory i rastitel’nosti SSSR (Materials on the History of the Flora and Vegetation in the Soviet Union), Leningrad: Akad. Nauk SSSR, 1963, issue 4, pp. 462–492.
Putenikhin, V.P. and Martinsson, O., Present Distribution of Larix sukaczewii Dyl. in Russia, Umea: Swed. Univ. Agric. Sci., 1995.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © Yu.S. Nechaeva, A.A. Julanov, S.V. Boronnikova, Ya.V. Prishnivskaya, 2017, published in Genetika, 2017, Vol. 53, No. 5, pp. 591–600.
Rights and permissions
About this article
Cite this article
Nechaeva, Y.S., Julanov, A.A., Boronnikova, S.V. et al. Nucleotide polymorphisms of candidate genes of adaptive significance in the ural populations of Larix sibirica Ledeb.. Russ J Genet 53, 587–595 (2017). https://doi.org/10.1134/S1022795417050064
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795417050064