Abstract
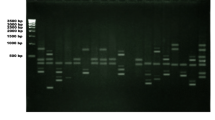
Species-specific LTR retrotransposons were first cloned in five rare relic species of drug plants located in the Perm’ region. Sequences of LTR retrotransposons were used for PCR analysis based on amplification of repeated sequences from LTR or other sites of retrotransposons (IRAP). Genetic diversity was studied in six populations of rare relic species of plants Adonis vernalis L. by means of the IRAP method; 125 polymorphic IRAP-markers were analyzed. Parameters for DNA polymorphism and genetic diversity of A. vernalis populations were determined.
Similar content being viewed by others

References
McClintok, B., Controlling Elements and the Gene, Cold Spring Harbor Symp. Quant. Biol., 1956, vol. 21, pp. 197–216.
Ilyin, Y.V., Tchurikov, N.A., Ananiev, E.V., et al., Studies on the DNA Fragments and Adjacent Sequences, Cold Spring Harbor Symp. Quant. Biol., 1978, vol. 42, pp. 959–969.
Lyubomirskaya, N.V. and Il’in, Yu.V., Mobile Genetic Elements of Eukaryotes: Past, Present, Future, Mol. Biol. (Moscow), 1999, vol. 33, no. 6, pp. 958–968.
Kalendar, R., Tanskanen, J., Immonen, S., et al., Genome Evolution of Wild Barley (Hordeum spontaneum) by BARE-1 Retrotransposon Dynamics in Response to Sharp Microclimatic Divergence, Proc. Natl. Acad. Sci. USA, 2000, vol. 97, no. 12, pp. 6603–6607.
Evgen’ev, M.B., Mobile Elements and Genome Evolution, Mol. Biol. (Moscow), 2007, vol. 41, no. 2, pp. 234–245.
Kumar, A. and Bennetzen, J., Plant Retrotransposons, Annu. Rev. Genet., 1999, vol. 33, pp. 479–532.
Wicker, T., Sabot, F., Hua-Van, A., et al., A Unified Classification System for Eukaryotic Transposable Elements, Nat. Rev. Genet., 2007, no. 8, pp. 973–982.
Kalendar, R., Grob, T., Regina, M., et al., IRAP and REMAP: Two New Retrotransposon-Based DNA Fingerprinting Techniques, Theor. Appl. Genet., 1999, vol. 98, pp. 704–711.
Kalendar, R. and Schulman, A.H., IRAP and REMAP for Retrotransposon-Based Genotyping and Fingerprinting, Nat. Protocols, 2006, vol. 1, no. 5, pp. 2478–2484.
Leigh, F., Kalendar, R., Lea, V., et al., Comparison of the Utility of Barley Retrotransposon Families for Genetic Analysis by Molecular Marker Techniques, Mol. Gen. Genomics, 2003, vol. 269, no. 3, pp. 464–474.
Bannikova, A.A., Matveev, V.A., and Kramerov, D.A., Using Inter-SINE PCR to Study Mammalian Phylogeny, Russ. J. Genet., 2002, vol. 38, no. 6, pp. 714–724.
Ryabinina, N.L., Bannikova, A.A., Sheremet’eva, V.A., et al., Analysis of DNA of Higher Primates Using Inter-SINE PCR, Russ. J. Genet., 2008, vol. 44, no. 3, pp. 266–272.
Kalendar, R.N. and Glazko, V.I., Types of Molecular Genetic Markers and Their Application, Fiziol. Biokhim. Kulturnyh Rasteniy, 2002, vol. 34, no. 4, pp. 279–296.
Krasnaya kniga Permskogo kraya (Red Book of Perm Krai), Shepel’, A.I., Ed., Perm: Knizhnyi mir, 2008.
Torres, A.M., Weeden, N.F., and Martin, A., Linkage among Isozyme, RFLP and RAPD Markers in Vicia faba, Theor. Appl. Genet., 1993, vol. 5, pp. 937–945.
Kalendar, R., Tanskanen, J., Antonius-Klemola, K., et al., Cassandra Retrotransposons Carry Independently Transcribed 5S RNA, Proc. Natl. Acad. Sci. USA, 2008, vol. 105, no. 15, pp. 5833–5838.
Kalendar, R., Lee, D., and Schulman, A.H., FastPCR Software for PCR Primer and Probe Design and Repeat Search, Genes Genomes Genomics, 2009, vol. 3, no. 1, http://www.biocenter.helsinki.fi/bi/programs/fastper.htm.
Poshkurlat, A.P., Rod goritsvet-Adonis L.: Sistematika, rasprostranenie, biologiya (Genus Adonis: Systematics, Distribution, Biology), Moscow: Nauka/Interperiodika, 2000.
Nei, M., Molecular Population Genetics and Evolution, Amsterdam, 1975.
Kalendar, R.N. and Boronnikova, S.V., Molecular Genetic Polymorphism of Natural Populations of Ural Rare Plant Species Using Retrotransposons, Biotekhnologiya: sostoyanie i perspektivy razvitiya (Biotechnology: Current State and Perspectives of Development), Proc. 4th Moscow Int. Congress, Moscow: Ekspobiokhim-tekhnologii, 2007, part 2, p. 121.
Boyko, E., Kalendar, R., Korzun, V., et al., A HighDensity Cytogenetic Map of the Aegilops tauschii Genome Incorporating Retrotransposons and Defense Related Genes: Insights into Cereal Chromosome Structure and Function, Plant Mol. Biol., 2002, vol. 48, pp. 767–790.
Baumel, A., Ainouch, M., Kalendar, R., et al., Retrotransposons and Genomic Stability in Populations of the Young Allopolyploid Species Spartina anglica Hubard (Poaceae), Mol. Biol. Evol., 2002, vol. 19, no. 8, pp. 1218–1227.
Kokaeva, Z.G., Boronnikova, S.V., Tikhomirova, N.N., et al., Genetic Diversity Analysis of Population Systems of Rare and Resource Plant Species, Biotekhnologiyaokhrane okruzhayushchei sredy (Biotechnology to Environment Conservation), Proc. 4th Int. Sci. Conf., in Dokl. MOIP, Moscow, 2006, pp. 75–79.
Boronnikova, S.V., Molekulyarno-geneticheskaya identifikatsiya i pasportizatsiya redkikh i nakhodyashchikhsya pod ugrozoi unichtozheniya vidov rastenii (Molecular Genetic Identification and Certification of Rare and Endangered Plant Species), Perm: Perm Univ., 2008.
Boronnikova, S.V., Genetic Certification of Population of Rare Plant Species of the Genus Adonis Using ISSRand IRAP-Markers, Izv. Mosk. S-kh. Akad., 2009, no. 1, pp. 83–89.
Boronnikova, S.V., Svetlakova, T.N., and Boboshina, I.V., Genetic Polymorphism Study of Populus tremula L. Using ISSRand IRAP-Markers, Agrarnaya Rossiya, 2009, no. 2, pp. 20–22.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © S.V. Boronnikova, R.N. Kalendar, 2010, published in Genetika, 2010, Vol. 46, No. 1, pp. 44–50.
Rights and permissions
About this article
Cite this article
Boronnikova, S.V., Kalendar, R.N. Using IRAP markers for analysis of genetic variability in populations of resource and rare species of plants. Russ J Genet 46, 36–42 (2010). https://doi.org/10.1134/S1022795410010060
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795410010060