Abstract
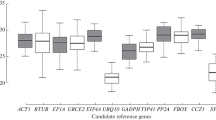
Reference genes are frequently used as a normalization standard to obtain reliable data during quantitative real-time polymerase chain reaction (qRT-PCR). However, recent studies showed that most traditional reference genes were not stable under different treatments or environmental stresses, which may lead to misinterpret expression of the target genes. In this study, 7 candidate reference genes in tea plant (Camellia sinensis (L.) Kuntze cv. Yingshuang) were selected and their expression stability under different abiotic stresses was analyzed using geNorm, NormFinder, and BestKeeper methods. Our results suggest that TUA1 (alpha-1 tubulin) has the most stable expression under damage stresses according to 3 methods of analysis. For drought stresses, 18S rRNA, and GAPDH (glyceraldehyde-3-phosphate dehydrogenase) were the most stable genes. For cold, Al, and NaCl stresses, GAPDH and TUA1 may be the alternative options. Our results may provide an insight for identification of the optimal reference genes for tea plants under various treatments.
Similar content being viewed by others
Abbreviations
- Ct:
-
cycle threshold
- CV:
-
percentage covariance
- EF1-α:
-
elongation factor 1-alpha
- GAPDH:
-
glyceraldehyde-3-phosphate dehydrogenase
- MMCS:
-
maximum moisture content of soil
- qRT-PCR:
-
quantitative real-time polymerase chain reaction
- T m :
-
melting temperature
- TUA1:
-
alpha-1 tubulin
- TUA2:
-
alpha-2 tubulin
- UBI:
-
ubiquitin
References
Wong, M.L. and Medrano, J.F., Real-time PCR for mRNA quantitation, Biotechniques, 2005, vol. 39, pp. 75–85.
Bustin, S.A., Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays, J. Mol. Endocrinol., 2000, vol. 25, pp. 169–193.
Lee, J.M., Roche, J.R., Donaghy, D.J., Thrush, A., and Sathish, P., Validation of reference genes for quantitative RT-PCR studies of gene expression in perennial ryegrass (Lolium perenne L.), BMC Mol. Biol., 2010, vol. 11: 8.
Maroufi, A., van Bockstaele, E., and de Loose, M., Validation of reference genes for gene expression analysis in chicory (Cichorium intybus) using quantitative real-time PCR, BMC Mol. Biol., 2010, vol. 11: 15.
Pfaffl, M.W., Tichopad, A., Prgomet, C., and Neuvians, T.P., Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—Excel-based tool using pairwise correlations, Biotech. Lett., 2004, vol. 26, pp. 509–515.
Bas, A., Forsberg, G., Hammarström, S., and Hammarström, M.L., Utility of the Housekeeping Genes 18S rRNA, β-actin and glyceraldehyde-3-phosphatedehydrogenase for normalization in real-time quantitative reverse transcriptase-polymerase chain reaction analysis of gene expression in human T lymphocytes, Scand. J. Immunol., 2004, vol. 59, pp. 566–573.
Andersen, C.L., Jensen, J.L., and Ørntoft, T.F., Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets, Cancer Res., 2004, vol. 64, pp. 5245–5250.
Aerts, J.L., Gonzales, M.I., and Topalian, S.L., Selection of appropriate control genes to assess expression of tumor antigens using real-time RT-PCR, Biotechniques, 2004, vol. 36, pp. 84–97.
Selvey, S., Thompson, E.W., Matthaei, K., Lea, R.A., Irving, M.G., and Griffiths, L., β-Actin—an unsuitable internal control for RT-PCR, Mol. Cell. Probes, 2001, vol. 15, pp. 307–311.
Iskandar, H.M., Simpson, R.S., Casu, R.E., Bonnett, G.D., Maclean, D.J., and Manners, J.M., Comparison of reference genes for quantitative realtime polymerase chain reaction analysis of gene expression in sugarcane, Plant Mol. Biol. Rep., 2004, vol. 22, pp. 325–337.
Bustin, S., Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): trends and problems, J. Mol. Endocrinol., 2002, vol. 29, pp. 23–39.
Andersen, C., Ledet-Jensen, J., and Orntoft, T., Normalization of real-time quantitative RT-PCR data: a mode-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets, Cancer Res., 2004, vol. 64, pp. 5245–5250.
Vandesompele, J., de Preter, K., Pattyn, F., Poppe, B., van Roy, N., de Paepe, A., and Speeleman, F., Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes, Genome Biol., 2002, vol. 3.
Ye, X., Zhang, F., Tao, Y., Song, S., and Fang, J., Reference gene selection for quantitative real-time PCR normalization in different cherry genotypes, developmental stages and organs, Sci. Hortic., 2015, vol. 181, pp. 182–188.
Gu, C.S., Liu, L.Q., Xu, C., Zhao, Y.H., Zhu, X.D., and Huang, S.Z., Reference gene selection for quantitative real-time RT-PCR normalization in Iris lactea var. chinensis roots under cadmium, lead, and salt stress conditions, Sci. World J., 2014, vol. 2014. http://dx.doi.org/10.1155/2014/532713
Sun, M., Wang, Y., Yang, D., Wei, C., Gao, L., Xia, T., Shan, Y., and Luo, Y., Reference genes for real-time fluorescence quantitative PCR in Camellia sinensis, Chin. Bull. Bot., 2010, vol. 45, pp. 579–587.
Løvdal, T. and Lillo, C., Reference gene selection for quantitative real-time PCR normalization in tomato subjected to nitrogen, cold, and light stress, Anal. Biochem., 2009, vol. 387, pp. 238–242.
Jain, M., Nijhawan, A., Tyagi, A.K., and Khurana, J.P., Validation of housekeeping genes as internal control for studying gene expression in rice by quantitative real-time PCR, Biochem. Biophys. Res. Commun., 2006, vol. 345, pp. 646–651.
Wan, H., Zhao, Z., Qian, C., Sui, Y., Malik, A.A., and Chen, J., Selection of appropriate reference genes for gene expression studies by quantitative real-time polymerase chain reaction in cucumber, Anal. Biochem., 2010, vol. 399, pp. 257–261.
Brunner, A.M., Yakovlev, I.A., and Strauss, S.H., Validating internal controls for quantitative plant gene expression studies, BMC Plant Biol., 2004, vol. 4: 14.
Gohain, B., Bandyopadhyay, T., Borchetia, S., Bharalee, R., Gupta, S., Bhorali, P., Agarwala, N., and Das, S., Identification and validation of stable reference genes in Camellia species, J. Biotechnol. Pharm. Res., 2011, vol. 2: 009–018.
Hao, X., Horvath, D.P., Chao, W.S., Yang, Y., Wang, X., and Xiao, B., Identification and evaluation of reliable reference genes for quantitative real-time PCR analysis in tea plant (Camellia sinensis (L.) O. Kuntze), Int. J. Mol. Sci., 2014, vol. 15, pp. 22155–22172.
Kim, J.W. and Dang, C.V., Multifaceted roles of glycolytic enzymes, Trends Biochem. Sci., 2005, vol. 30, pp. 142–150.
Stürzenbaum, S.R. and Kille, P., Control genes in quantitative molecular biological techniques: the variability of invariance, Comp. Biochem. Physiol. B. Biochem. Mol. Biol., 2001, vol. 130, pp. 281–289.
Hochstrasser, M., Evolution and function of ubiquitinlike protein-conjugation systems, Nat. Cell Biol., 2000, vol. 2, pp. E153–E157.
Author information
Authors and Affiliations
Corresponding author
Additional information
The article is published in the original.
Both authors contributed eqaully to this work.
Rights and permissions
About this article
Cite this article
Ma, Q.P., Hao, S., Chen, X. et al. Validation of reliability for reference genes under various abiotic stresses in tea plant. Russ J Plant Physiol 63, 423–432 (2016). https://doi.org/10.1134/S1021443716030080
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1021443716030080