Abstract
Adenine-DNA-glycosylase MutY is a monofunctional enzyme and catalyzes hydrolysis of N-glycosidic bonds with adenine residues located opposite 8-oxonuanine residues in DNA. Rational design was carried out to construct mutant enzyme forms with altered catalytic activity. Structures of the MutY mutants were calculated by molecular dynamics (MD). Their analysis showed that some of the MutY mutants may have AP lyase activity in addition to hydrolyzing the N-glycosidic bond, as is the case with bifunctional DNA glycosylases. MutY mutants with the A120K or S124K substitution were obtained by site-directed mutagenesis, and their catalytic activities were determined. The S120K substitution was shown to confer additional AP lyase activity, while the A124K substitution completely inactivated the enzyme.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Constructing enzymes with changed properties is of importance for biotechnological and medical applications. Promising current approaches used to obtain enzymes with changed or improved properties include experimental screening of enzymes from various sources, construction of gene libraries coding for target enzyme variants with random mutations and their screening for enzymatic properties, and rational design of enzymes with the use of molecular modeling and property predictions [1, 2]. It is safe to state, in fact, that modern experimental and computational techniques make it possible to create improved forms of enzymes [3]. For example, substantial progress has been made in improving the fidelity, processivity, thermal stability, and other parameters of DNA polymerases, which are broadly employed in diagnostic applications [2].
However, substantial problems are still encountered in creating enzymes with changed properties because not only the relatively rigid general structure of a protein, but also, what is more important, its dynamics properties determine its functional properties [4, 5]. Concerted movements of particular parts of a protein molecule relative to its other parts can take place in certain regions of the protein globule in the course of enzyme–substrate interactions. Some regions may be highly dynamics, while other regions remain relatively static [6–8]. These sophisticated interactions form a dynamic cross-correlation network, which may be of importance for allosteric transitions [9, 10] and may act as an effector of catalytic activity during the formation of protein–protein complexes [11, 12].
Protein engineering has intensely developed over the past 20 years. Its progress is due to new technologies, such as directed evolution using a phage display method [13, 14] and rational design utilizing site-directed mutagenesis and construction of chimeric enzymes [2, 15]. Rapid progress in genetic technologies renders the nucleic acid-modifying enzymes, which are necessary for main genetic-engineering manipulations, promising objects of rational design.
Structure-oriented engineering is still an important approach to construction of enzymes with changed properties. In fact, a high substrate specificity or a certain type of catalytic activity, which are characteristic of the majority of enzymes, may prevent their use in particular applications. It is therefore of importance to change the substrate specificity or catalytic properties of individual nucleic acid-processing enzymes in a directional manner.
Adenine-DNA glycosylase MutY catalyzes excision of adenine from its noncanonical pair with 7,8-dihydro-8-oxoguanine (oxoG) [16–20]. Apart from excising adenine from the oxoG:A pair, MutY can remove adenine from the G:A pair, but at a far lower rate [22]. Multiple contacts of the enzyme with oxoG:A and G:A pairs must be responsible for its unique specificity. In this work, we performed rational design of amino acid residues in the active center of the enzyme and constructed a mutant MutY form with changed catalytic activity. Molecular dynamics (MD) was used to obtain models of the MutY mutants that are capable not only of hydrolyzing the N-glycosidic bond with adenine, but also of exerting new catalytic activity, namely, catalyzing the cleavage of the 2'-deoxyribose phosphate backbone of DNA at the enzyme recognition site. Site-directed mutagenesis was performed on the basis of the simulation data to obtain the Escherichia coli MutY mutants with the A120K or S124K substitution, and their catalytic activities were determined. The S120K substitution was shown to confer additional activity, while the A124K substitution completely inactivated the enzyme.
EXPERIMENTAL
Molecular modeling. MD was performed using the structure of Geobacillus stearothermophilus MutY in complex with an uncleavable 11-bp substrate DNA containing 2'-fluoro-2'-deoxyadenosine (FLRC, PDB ID 3G0Q) [23]. Based on the structure, initial models were constructed for the wild-type G. stearothermophilus MutY and its mutant forms carrying the substitutions Y126K and A130K, which correspond to the substitutions S120K and A124K in E. coli MutY. We used the AMBER ff99SB-ildn to describe the protein [24–26], the bsc1 force field to describe DNA [27, 28] with additional parameters used for the iron-sulfur cluster [4Fe-4S]2+ and 8-oxoguanine [29, 30], the TIP3P water model, and the respective ion parameters [31, 32]. Protonation of amino acid residues was evaluated using published data and the H++ web server [33]. Calculations were carried out in GROMACS [34, 35]. A complex was placed in a dodecahedral periodic box with a minimal distance of 1.1 nm to a box boundary; the box was filled with water molecules and ions. The cut-off radius of long-range interactions was 1 nm; electrostatic interactions were calculated by the PME method [36]; covalent bonds with hydrogen atoms were processed by the LINCS method [37]; the integration step was 2 fs. System relaxation was performed consecutively for NVT and NPT ensembles with harmonic restraints on heavy atoms of the enzyme–substrate complex; the duration of each step was 0.5 ns. After relaxation, independent molecular-dynamics trajectories of 100 ns were generated three times for each variant of the complex. The modified Berendsen thermostat with a temperature set to 300 K and the Parrinello–Rahman barostat were used [38].
Oligodeoxyribonucleotides were synthesized by the phosphoramidite method, using an ASM-700 synthesizer (Biosset, Russia) and monomers from Glen Research (United States). The oligodeoxyribonucleotides were purified by HPLC on an ion-exchange column (PRP-X500 (Hamilton Company), 3.9 × 300 mm, particle size 12–30 µm) and subsequent reversed-phase chromatography (Nucleoprep 100-20 C18, 10 × 250 mm (Macherey-Nagel, Germany)). Oligonucleotide purity was checked by PAGE in denaturing 20% gel. The oligonucleotide concentration was measured by the optical density of their solutions at 260 nm in electron absorption spectra and calculated according to the Bouguer–Beer–Lambert law, taking the molar extinction coefficient estimates obtained by the nearest neighbor method [39]. An oligodeoxyribonucleotide complex with the sequence CTCTC(oxoG)CCTTCC, which contained adenine opposite to 8-oxoguanine, was used as a substrate DNA.
Enzymes. MutY was isolated from E. coli BL21(DE3) cells transformed with pET28c-MutY as described previously [40]. The pET28c-MutY plasmid, which carried MutY, was kindly provided by M.K. Saparbaev (Groupe “Réparation de l’AND”, Université Paris-Sud XI, Institut Gustave Roussy, France).
Mutant MutY forms with the substitutions S120K and A124K were obtained by site-directed mutagenesis. Correct introduction of the substitutions in MutY was verified by sequencing. The MutY mutants were purified as follows. An E. coli BL21(DE3) culture was grown in LB (1 L) supplemented with 25 µg/mL kanamycin at 37°C to an optical density of 0.6–0.7 at 600 nm. Transcription was induced by adding isopropyl-β-D-thiogalactopyranoside to 0.2 mM, and the culture was incubated for 16 h. Cells were collected by centrifugation (10 000 g, 10 min) and suspended in 30 mL of 20 mM HEPES-NaOH (pH 7.8), 40 mM NaCl supplemented with a protease inhibitor cocktail (Complete, Germany). Cells were lysed using a French pressure cell press. All subsequent procedures were carried out at 4°C. The cell lysate was centrifuged (40 000 g, 40 min), and the enzyme was isolated chromatographically. The supernatant was applied onto column I (Q-Sepharose Fast Flow, Amersham Biosciences, Sweden), and elution was carried out using 20 mM HEPES-NaOH (pH 7.8), 200 mM NaCl. MutY-containing fractions were collected and applied onto column II (HiTrap Chelating, Amersham Biosciences) in 20 mM HEPES-NaOH (pH 7.8), 500 mM NaCl, 20 mM imidazole. Elution was performed with a linear gradient of imidazole concentration (20 → 500 mM); the optical density was measured at 280 nm. Protein purity was checked by gel electrophoresis. Fractions containing the MutY mutants were dialyzed against 20 mM HEPES-NaOH (pH 7.5) supplemented with 1 mM EDTA, 1 mM DTT, 250 mM NaCl, and 50% glycerol and stored at –20°C. The enzyme concentration was calculated from the optical density at 280 nm and the molar extinction coefficient 77328 M–1 cm–1 [41].
PAGE. Oligonucleotides 5'-end-labeled with 32P were used in experiments with reaction product separation by PAGE. All experiments were carried out at 25°C. The reaction buffer contained 50 mM Tris-HCl (pH 7.5), 50 mM KCl, 1 mM EDTA, 1 mM DTT, and 7% glycerol. Time dependences of substance conversion were obtained as follows. A 32P-labeled substrate DNA in 10 µL of the buffer was combined with 10 µL of 2.0–8.2 µM enzyme in the same buffer. The reaction mixture was quickly stirred. Aliquots (2 µL) were collected at certain time points and transferred into tubes with 2 µL of 7 M urea, 0.1% Bromphenol Blue, and 0.1% xylene cyanol. Each sample was divided in half. One half was combined with 1 µL of 1 M NaOH and incubated at 56°C for 15 min to hydrolyze the phosphodiester bonds in AP sites. The solution was neutralized with 1 µL of HCl and applied onto a gel; PAGE was carried out at 50 V/cm. The accumulation of the reaction products was assessed as in [40].
To quantitate the product, autoradiographs were scanned using a Molecular Imager FX phosphorimager (Bio-Rad, United States) and the data were processed using Gel-Pro Analyzer 4.0 software (Media Cybernetics, United States). The product accumulation was calculated by dividing the total area of product peaks to by the sum of the total product peak area and the peak area of the initial oligonucleotide. The error of measuring the percent conversion was usually no more than 20%.
RESULTS AND DISCUSSION
Degree of Sugar-Phosphate Backbone Bending in Enzyme–Substrate Complexes
Several crystal structures are available for MutY: structures have been solved for a free fragment of the E. coli enzyme [42, 43], mutant forms G. stearothermophilus MutY in complex with various oligonucleotide duplexes [23, 44–47], and human and mouse MUTYH fragments 48, 49]. X-ray analyses of MutY in isolation and in complex with DNA have shown that the DNA duplex is substantially bent in the catalytically active complex. However, the bending angle and the extent of deformation of DNA bound in the active center of the enzyme differ between the structure of the complex of catalytically active MutY with a 2'-F-dA DNA substrate (Fig. 1, FLRC, 3G0Q) and the structure of the complex of catalytically inactive mutant MutY D144N and a DNA substrate (Fig. 1, LRC, 1RRQ). The difference might be explained by a MutY mechanism of specific substrate binding via electron transfer and oxidation of the [4Fe-4S]2+ cluster to [4Fe-4S]3+ by damaged DNA as well as by an effect of 2'-fluoro nucleotide modification in the FLRC complex. It is of interest to note that the DNA bending angle in our MD experiments was intermediate between the angles observed in the FLRC and LRC complexes (Fig. 1).
Comparative bending of the sugar-phosphate backbone of a substrate as inferred from the distance between the P atoms that flank the nucleotide base flipped out of the duplex. Yellow, X-ray structure of the complex of catalytically active MutY with 2′-F-dA DNA substrate (FLRC, 3G0Q); purple, catalytically inactive MutY D144N in complex with a DNA substrate (LRC, 1RRQ); blue, a representative snapshot of the MD trajectory of the MutY–DNA complex.
Structure of the OxoG Recognition Site
The oxoG recognition site of MutY is formed by a conserved loop, which harbors the amino acid residues Phe307-Ser308-His309. The residues Gln48, Thr49, Leu86, Tyr88, and Ser308 form direct contacts with oxoG. A substitution of G for oxoG changes the conformation of Ser308, which forms two hydrogen bonds with N7 and C8 of oxoG, but not a single bond with guanine (Fig. 2). The substitution S308A has been shown to decrease enzyme affinities for oxoG:A and G:A substrates in a proportionate manner without affecting the selectivity, while the double substitution F307A/S308A is required for loss of specificity toward oxoG-containing DNA [47]. However, our MD data indicate that Ser308 is also capable of hydrogen bonding with N7 of guanine because the peptide backbone of the loop is motile in the MutY complex with DNA containing a G:A pair. In this case, the conformation of Phe307 changes in both wild-type enzyme and its mutant forms so that its side chain is rotated by ~90° relative to the positions characteristic of the models with the oxoG:A pair and X-ray structures. Thus, our MD calculations implicate Phe307 in recognizing the target substrate.
Structure of the 2 '-Deoxyadenosine Recognition Site and Mechanism of N-Glycoside Bond Hydrolysis
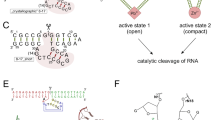
In the enzyme–substrate complexes, 2′-deoxyadenosine is flipped out of the double helix into the active center, which is formed by Arg26, Leu28, Trp30, Arg31, Leu46, Glu43, Val51, Tyr126, Glu188, Ile191, and Glu192 (the amino acid residues are numbered as in G. stearothermophilus MutY) (Fig. 3a).
(a) Structure of the 2′-deoxyadenosine recognition site (FLRC, 3G0Q) and (b) the mechanism of N-glycosidic bond hydrolysis ([53], Creative Commons Attribution License).
A catalytic mechanism of N-glycosidic bond hydrolysis in 2'-deoxyadenosine has been assumed on the basis of X-ray data and quantum-mechanical calculations [23, 47, 50, 51]. It is thought that Glu43 is in a neutral form in the active center of the enzyme [50] (Fig. 3b). Thus, published data indicate that the Ade base is protonated at N7 by Glu43 in the catalytic complex and that the N-glycosidic bond is consequently cleaved by the SN1 mechanism [52, 53]. The existence of the mechanism is supported by the data that an adenosine analog with N7 replaced with the C7-H group [54] or a Glu43 substitution [23, 42] completely abolish MutY catalytic activity.
Selection of the Amino Acid Residues of the MutY Catalytic Center That Are Necessary for AP Lyase Activity
To extend the catalytic properties of the enzyme without affecting its high substrate specificity, we proceeded from structural homology of the active center between monofunctional MutY and bifunctional DNA glycosylases (Fig. 4). For example, the MutY S120K mutant (Y126K in G. stearothermophilus MutY) has been proposed by analogy with bifunctional DNA glycosylases hOGG1 and E. coli Nth, which also belong to the helix-hairpin-helix (HhH) structural family and possess not only glycosylase, but also AP lyase activity [56]. The position of Ser120 in E. coli MutY corresponds indeed to the positions of catalytic Lys249 in hOGG1 and Lys120 in Nth.
Proceeding from steric considerations, we proposed the substitution A124K (A130K in G. stearothermophilus MutY), in which the side chain of the lysine residue also occur in the catalytic center of the enzyme. It should be noted that the position of Ala130 in G. stearothermophilus MutY corresponds to the spatial position of Cys253, which stabilizes the side chain of catalytic Lys249 in DNA glycosylase hOGG1. The hOGG1 enzyme retains part of its catalytic activity when containing the double substitution K249C/C253K, which preserves the stabilizing interaction between the lysine and cysteine residues in the active center [57].
Construction and Analysis of the MutY Y126K and MutY A130K Model Structures
To construct the model structures of the MutY Y126K and MutY A130K mutants in complex with DNA, we used the X-ray structure of G. stearothermophilus MutY in complex with a DNA substrate containing 2'-fluoro-2'-deoxyadenosine (PDB ID 3G0Q) [23].
The active center structure with coordination of the adenine N7 atom by the side chains of Tyr126 and Glu43В was preserved in our MD models of the wild-type enzyme; this agreed well with the reaction mechanism assumed for MutY [50] (Fig. 5a). Estimation of pKa of amino acid residues by the H++ algorithm showed additionally that Glu43 is protonated in the DNA complexes with the wild-type enzyme and the MutY Y126K mutant, but not with the MutY A130K mutant.
The model structure of the MutY A130K mutant with DNA indicated that the orientations of key amino acid residues are changed in the active center because the side chain of Lys130 is hydrogen bonded with Glu43 and Glu188 (Fig. 5b). The formation of these contacts get the flipped-out adenine farther away from the catalytic amino acid residues and must decrease N-glycosylase activity.
It is of interest to note that the orientation of amino acid residues was optimal for N-glycosylase activity within a relatively short interval, ~5% of the total simulation time, in MD trajectories of the complex formed by the MutY Y126K mutant form (Fig. 6a). For a major part of the simulation time, the flipped-out adenosine is far away from the Lys126–Glu43 pair and thus cannot form the direct contacts that are necessary for N-glycosylase activity according to the mechanism assumed for MutY [50]. However, the possible mutual arrangements included one that was similar to the arrangement in the complex of hOGG1 DNA glycosylase with damaged DNA, where the amino group of the lysine side chain is oriented so that its position is suitable for N1 protonation in the flipped-out nitrogenous base. The finding makes it possible to assume that hydrolysis of the N-glycosidic bond by MutY Y126K may proceed via the mechanism described for hOGG1 [58] (Fig. 6b).
The active center of the MutY Y126K mutant. (a) Superimposition of an active center configuration with the arrangement of amino acid residues that is optimal for catalysis (green, 5% of the MD trajectory time) and a more stable configuration (yellow). (b) Superimposition of the hOGG1 complex with flipped-out oxoG in the active center (green, 3KTU) and a similar structural organization of the MutY Y126K complex from its MD trajectory (yellow).
Catalytic Properties of E. coli MutY Mutants Containing the S120K and A124K Substitutions
To check the assumptions based on our MD calculations, we experimentally studied activities of the wild-type MutY and its mutant forms toward a DNA substrate containing an oxoG:A pair. To detect the N-glycosylase reaction products, which contained an AP site, the reaction mixtures were treated with an alkaline solution to cleave DNA. Direct PAGE separation of the reaction mixture without alkaline treatment was used to detect AP lyase activity of the mutant forms.
The results showed that the A124K substitution completely inactivated the enzyme. The MutY S120K mutant acquired the properties of a bifunctional DNA glycosylase, excising adenine and cleaving the resulting AP site. Figure 7 shows a typical electrophoretic pattern and kinetic curves of product accumulation in the N-glycosylase and AP lyase reactions for the interaction of MutY S120K with the DNA substrate. A comparison with the wild-type enzyme [40, 59] demonstrated a decrease in N-glycosylase activity of the enzyme toward the oxoG:A pair. A decrease in catalytic activity is possible to explain by assuming that the S120K substitution simultaneously decreases pKa of catalytic Glu43 so that Lys120 is no longer capable of stabilizing the oxacarbenium-cation intermediate [53]. It should also be noted that the initial orientation of the active-center residues that is necessary for the reaction to proceed via the mechanism assumed for the wild-type MutY was preserved over only ~5% of the trajectory in MD simulations. The more stable alternative arrangement of the lysine side chain and flipped-out nitrogenous base may lead to N-glycosidic bond cleavage via a mechanism similar to that of hOGG1 glycosylase, though with a lower efficiency.
Accumulation of the reaction product during interaction of MutY with DNA substrates as revealed by PAGE. (a) Electrophoretic pattern characterizes the cleavage of an oxoG:A substrate by MutY S120K and was obtained without KOH treatment. Sample collection time (s) is indicated at the top of each lane. (b, c) Kinetic curves characterize the accumulation of the products of oxoG:A substrate cleavage and were obtained (b) with and (c) without alkaline treatment.
CONCLUSIONS
To summarize, our MD analysis revealed the nature of key interactions that occur in the active center to ensure specific recognition of a lesion and catalytic hydrolysis of the N-glycosidic bond by MutY. Based on the results, we proposed the amino acid substitutions of the active center that were potentially capable to allow the enzyme not only to hydrolyze the N-glycosidic bond of adenine, but also to break the 2′‑deoxyribophosphate backbone at the recognition site, thus conferring new AP lyase activity. This means that MutY monofunctional DNA glycosylase is possible to convert to a bifunctional DNA glycosylase via site-directed mutagenesis.
Site-directed mutagenesis was performed to construct the E. coli MutY mutants carrying the S120K or A124K substitution, and their catalytic activities were assayed. The A124K substitution was found to fully inactivate the enzyme by distorting the network of contacts in the active center and thus getting the catalytic residues farther away from the flipped-out nitrogenous base. The MutY S120K mutant acquired additional AP lyase activity and displayed the properties of a bifunctional DNA glycosylase, excising adenine and cleaving the resulting AP site.
REFERENCES
Vanella R., Kovacevic G., Doffini V., de Santaella J.F., Nash M.A. 2022. High-throughput screening, next generation sequencing and machine learning: advanced methods in enzyme engineering. Chem. Commun. 58, 2455.
Nikoomanzar A., Chim N., Yik E.J., Chaput J.C. 2020. Engineering polymerases for applications in synthetic biology. Q. Rev. Biophys. 53, 1–31.
Siedhoffa N.E., Schwaneberg U., Davari M.D. 2020. Machine learning-assisted enzyme engineering. in Methods in Enzymology. 643, 281–315.
Kuznetsov N.A., Fedorova O.S. 2020. Kinetic milestones of damage recognition by DNA glycosylases of the helix–hairpin–helix structural superfamily. Adv. Exp. Biol. Med. 1241, 1–18.
Kuznetsova A.A., Fedorova O.S., Kuznetsov N.A. 2022. Structural and molecular kinetic features of activities of DNA polymerases. Int. J. Mol. Sci. 23, 6373.
Yu H., Dalby P.A. 2020. A beginner’s guide to molecular dynamics simulations and the identification of cross-correlation networks for enzyme engineering. Methods Enzymol. 643, 15–49.
Bulygin A.A., Kuznetsova A.A., Vorobjev Y.N., Fedorova O.S., Kuznetsov N.A. 2020. The role of active-site plasticity in damaged-nucleotide recognition by human apurinic/apyrimidinic endonuclease APE1. Molecules. 25, 3940.
Bulygin A.A., Fedorova O.S., Kuznetsov N.A. 2022. Insights into mechanisms of damage recognition and catalysis by APE1-like enzymes. Int. J. Mol. Sci. 23, 4361.
Bowerman S., Wereszczynski J. 2016. Detecting allosteric networks using molecular dynamics simulation. Methods Enzymol. 578, 429–447.
Tekpinar M., Neron B., Delarue M. 2021. Extracting dynamical correlations and identifying key residues for allosteric communication in proteins by correlation plus. J. Chem. Inf. Model. 61, 4832–4838.
Kladova O.A., Bazlekowa-Karaban M., Baconnais S., Piétrement O., Ishchenko A.A., Matkarimov B.T., Iakovlev D.A., Vasenko A., Fedorova O.S., Le Cam E., Tudek B., Kuznetsov N.A., Saparbaev M. 2018. The role of the N-terminal domain of human apurinic/apyrimidinic endonuclease 1, APE1, in DNA glycosylase stimulation. DNA Repair (Amst.). 64, 10–25.
Kladova O.A., Alekseeva I.V., Saparbaev M., Fedorova O.S., Kuznetsov N.A. 2020. Modulation of the apurinic/apyrimidinic endonuclease activity of human APE1 and of its natural polymorphic variants by base excision repair proteins. Int. J. Mol. Sci. 21, 7174.
Smith G.P., Petrenko V.A. 1997. Phage display. Chem. Rev. 97, 391–410.
Ghadessy F., Ong J., Holliger P. 2001. Directed evolution of polymerase function by compartmentalized self-replication. Proc. Natl. Acad. Sci. U. S. A. 98, 4552–4557.
Choi J., Kim H.-S. 2020. Structure-guided rational design of the substrate specificity and catalytic activity of an enzyme. Methods Enzymol. 643, 181–202.
Au K.G., Cabrera M., Miller J.H., Modrich P. 1988. Escherichia coli MutY gene-product is required for specific A-G-]C.G mismatch correction. Proc. Natl. Acad. Sci. U. S. A. 85, 9163–9166.
Slupska M.M., Baikalov C., Luther W.M., Chiang J.-H., Wei Y.-F., Miller J.H. 1996. Cloning and sequencing a human homolog (hMYH) of the Escherichia coli mutY gene whose function is required for the repair of oxidative DNA damage. J. Bacteriol. 178, 3885–3892.
Back J.H., Park J.H., Chung J.H., Kim D.S.H.L., Han Y.S. 2006. A distinct TthMutY bifunctional glycosylase that hydrolyzes not only adenine but also thymine opposite 8-oxoguanine in the hyperthermophilic bacterium, Thermus thermophilus. DNA Repair. 5, 894–903.
Kunrath-Lima M., Repolês B.M., Alves C.L., Furtado C., Rajão M.A., Macedo A.M., Franco G.R., Pena S.D.J., Valenzuela L., Wisnovsky S., Kelley S.O., Galanti N., Cabrera G., Machado C.R. 2017. Characterization of Trypanosoma cruzi MutY DNA glycosylase ortholog and its role in oxidative stress response. Infect. Genet. Evol. 55, 332–342.
Li X., Lu A.L. 2001. Molecular cloning and functional analysis of the MutY homolog of Deinococcus radiodurans. J. Bacteriol. 183, 6151–6158.
Au K.G., Clark S., Miller J.H., Modrich P. 1989. Escherichia coli mutY gene encodes an adenine glycosylase active on G-A mispairs. Proc. Natl. Acad. Sci. U. S. A. 86, 8877–8881.
Bulychev N.V, Varaprasad C.V, Dorman G., Miller J.H., Eisenberg M., Grollman A.P., Johnson F. 1996. Substrate specificity of Escherichia coli MutY protein. Biochemistry. 35, 13147–13156.
Lee S., Verdine G.L. 2009. Atomic substitution reveals the structural basis for substrate adenine recognition and removal by adenine DNA glycosylase. Proc. Natl. Acad. Sci. U. S. A. 106, 18497–18502.
Cornell W.D., Cieplak P., Bayly C.I., Gould I.R., Merz K.M., Ferguson D.M., Spellmeyer D.C., Fox T., Caldwell J.W., Kollman P.A. 1995. A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J. Am. Chem. Soc. 117, 5179–5197.
Lindorff-Larsen K., Piana S., Palmo K., Maragakis P., Klepeis J.L., Dror R.O., Shaw D.E. 2010. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins. 78 (8), 1950–1958. https://doi.org/1002/prot.22711
Hornak V., Abel R., Okur A., Strockbine B., Roitberg A., Simmerling C. 2006. Comparison of multiple amber force fields and development of improved protein backbone parameters. Proteins. 65, 712–725.
Ivani I., Dans P.D., Noy A., Pérez A., Faustino I., Hospital A., Walther J., Andrio P., Goñi R., Balaceanu A., Portella G., Battistini F., Gelpi J.L., Gonzalez C., Vendruscolo M., Laughton C.A., Harris S.A., Case D.A., Orozco M. 2016. PARMBSC1: a refined force-field for DNA simulations. Nat. Methods. 13, 55–58.
Pérez A., Marchán I., Svozil D., Sponer J., Cheatham T.E., Laughton C.A., Orozco M. 2007. Refinement of the AMBER force field for nucleic acids: improving the description of α/γ conformers. Biophys. J. 92, 3817–3829.
Cheng X., Kelso C., Hornak V., De Los Santos C., Grollman A.P., Simmerling C. 2005. Dynamic behavior of DNA base pairs containing 8-oxoguanine. J. Am. Chem. Soc. 127 (40), 13906‒13918. https://doi.org/10.1021/ja052542s
Smith D.M.A., Xiong Y., Straatsma T.P., Rosso K.M., Squier T.C. 2012. Force-field development and molecular dynamics of [NiFe] hydrogenase. J. Chem. Theory Comput. 8, 2103–2114.
Jorgensen W.L., Chandrasekhar J., Madura J.D., Impey R.W., Klein M.L. 1983. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926.
Joung I.S., Cheatham T.E. 2008. Determination of alkali and halide monovalent ion parameters for use in explicitly solvated biomolecular simulations. J. Phys. Chem. B. 112, 9020–9041.
Anandakrishnan R., Aguilar B., Onufriev A.V. 2012. H++ 3.0: automating PK prediction and the preparation of biomolecular structures for atomistic molecular modeling and simulations. Nucleic Acids Res. 40, 537–541.
Abraham M.J., Murtola T., Schulz R., Páll S., Smith J.C., Hess B., Lindah E. 2015. Gromacs: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX. https://doi.org/10.1016/j.softx.2015.06.001
Berendsen H.J.C., van der Spoel D., van Drunen R. 1995. GROMACS: a message-passing parallel molecular dynamics implementation. Comput. Phys. Commun. 91, 43–56.
Essmann U., Perera L., Berkowitz M.L., Darden T., Lee H., Pedersen L.G. 1995. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577.
Hess B., Bekker H., Berendsen H.J.C., Fraaije J.G.E.M. 1997. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. https://doi.org/10.1002/(SICI)1096-987X(199709)18:12<1463::AID-JCC4>3.0.CO;2-H
Bussi G., Donadio D., Parrinello M. 2007. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101.
Fasman G.D. 1975. Handbook of Biochemistry and Molecular Biology. Cleveland: CRC Press, 3rd ed.
Tyugashev T.E., Kuznetsova A.A., Kuznetsov N.A., Fedorova O.S. 2017. Interaction features of adenine DNA glycosylase MutY from E. coli with DNA substrates. Russ. J. Bioorg. Chem. 43, 13–22.
Gill S.C., von Hippel P.H. 1989. Calculation of protein extinction coefficients from amino acid sequence data. Anal. Biochem. 182, 319–326.
Guan Y., Manuel R.C., Arvai A.S., Parikh S.S., Mol C.D., Miller J.H., Lloyd R.S., Tainer J.A. 1998. MutY catalytic core, mutant and bound adenine structures define specificity for DNA repair enzyme superfamily. Nat. Struct. Biol. 5, 1058–1064.
Zharkov D.O., Gilboa R., Yagil I., Kycia J.H., Gerchman S.E., Shoham G., Grollman A.P. 2000. Role for lysine 142 in the excision of adenine from A:G mispairs by MutY DNA glycosylase of Escherichia coli. Biochemistry. 39, 14768–14778.
Fromme J.C., Banerjee A., Huang S.J., Verdine G.L. 2004. Structural basis for removal of adenine mispaired with 8-oxoguanine by MutY adenine DNA glycosylase. Nature. 427, 652–656.
Wang L., Lee S.J., Verdine G.L. 2015. Structural basis for avoidance of promutagenic DNA repair by MutY adenine DNA glycosylase. J. Biol. Chem. 290, 17096–17105.
Wang L., Chakravarthy S., Verdine G.L. 2017. Structural basis for the lesion-scanning mechanism of the MutY DNA glycosylase. J. Biol. Chem. 292 (12), 5007‒5017. https://doi.org/10.1074/jbc.M116.757039
Russelburg P.L. O′Shea M., Valerie L. Demir M., Knutsen K.R., Sehgal S.L., Cao S., David S.S., Horvath M.P. 2020. Structural basis for finding OG lesions and avoiding undamaged G by the DNA glycosylase MutY. ACS Chem. Biol. 15, 93–102.
Luncsford P.J., Chang D.Y., Shi G., Bernstein J., Madabushi A., Patterson D.N., Lu A.L., Toth E. 2010. A structural hinge in eukaryotic MutY homologues mediates catalytic activity and Rad9-Rad1-Hus1 checkpoint complex interactions. J. Mol. Biol. 403, 351–370.
Nakamura T., Okabe K., Hirayama S., Chirifu M., Ikemizu S., Morioka H., Nakabeppu Y., Yamagata Y. 2021. Structure of the mammalian adenine DNA glycosylase MUTYH: insights into the base excision repair pathway and cancer. Nucleic Acids Res. 49, 7154–7163.
Kellie J.L., Wilson K.A., Wetmore S.D. 2013. Standard role for a conserved aspartate or more direct involvement in deglycosylation? An ONIOM and MD investigation of adenine–DNA glycosylase. Biochemistry. 52, 8753–8765.
Brunk E., Arey J.S., Rothlisberger U. 2012. Role of environment for catalysis of the DNA repair enzyme MutY. J. Am. Chem. Soc. 134, 8608–8616.
McCann J.A., Berti P.J. 2008. Transition-state analysis of the DNA repair enzyme MutY. J. Am. Chem. Soc. 130, 5789–5797.
Woods R.D., O′Shea V.L., Chu A., Cao S., Richards J.L., Horvath M.P., David S.S. 2016. Structure and stereochemistry of the base excision repair glycosylase MutY reveal a mechanism similar to retaining glycosidases. Nucleic Acids Res. 44, 801–810.
Porello S.L., Williams S.D., Kuhn H., Michaels M.L., David S.S. 1996. Specific recognition of substrate analogs by the DNA mismatch repair enzyme MutY. J. Am. Chem. Soc. 118, 10684–10692.
Kaur R., Nikkel D.J., Wetmore S.D. 2020. Computational studies of DNA repair: insights into the function of monofunctional DNA glycosylases in the base excision repair pathway. WIREs Comput. Mol. Sci. 10, e1471.
Ludwig D.L., MacInnes M.A., Takiguchi Y., Purtymun P.E., Henrie M., Flannery M., Meneses J., Pedersen R.A., Chen D.J. 1998. A murine AP-endonuclease gene-targeted deficiency with post-implantation embryonic progression and ionizing radiation sensitivity. Mutat. Res. 409 (1), 17‒29. https://doi.org/10.1016/S0921-8777(98)00039-1
Dalhus B., Forsbring M., Helle I.H., Vik E.S., Forstrom R.J., Backe P.H., Alseth I., Bjoras M. 2011. Separation-of-function mutants unravel the dual-reaction mode of human 8-oxoguanine DNA glycosylase. Structure. 19, 117–127.
Sebera J., Hattori Y., Sato D., Reha D., Nencka R., Kohno T., Kojima C., Tanaka Y., Sychrovsky V. 2017. The mechanism of the glycosylase reaction with hOGG1 base-excision repair enzyme: concerted effect of Lys249 and Asp268 during excision of 8-oxoguanine. Nucleic Acids Res. 45 (9), 5231‒5242. https://doi.org/10.1093/nar/gkx157
Williams S.D., David S.S. 2000. A single engineered point mutation in the adenine glycosylase MutY confers bifunctional glycosylase/AP lyase activity. Biochemistry. 39, 10098–10109.
Funding
This work was supported by the Ministry of Science and Higher Education (agreement no. 075-15-2021-1085).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflicts of interest. This article does not contain any studies involving animals or human subjects performed by any of the authors.
Additional information
Translated by T. Tkacheva
Rights and permissions
Open Access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tyugashev, T.E., Fedorova, O.S. & Kuznetsov, N.A. Modern Approaches to Protein Engineering to Create Enzymes with New Catalytic Properties. Mol Biol 57, 204–213 (2023). https://doi.org/10.1134/S0026893323020218
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893323020218