Abstract—
This study provides an overview of scientific results on the feasibility of using type III interferons against SARS-CoV-2. We have analyzed data obtained from the PubMed electronic database for the period 2020‒2022. The results of our own studies of pharmacological substances based on recombinant IFN-λ1 and its pegylated form are also presented. Completed and ongoing investigations allow us to position IFN-λ as an effective therapeutic agent against SARS-CoV-2.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Interferons (IFNs) are natural antiviral cytokines that confer effective early immune response. IFNs are secreted by host cells in response to viral infection and promote an antiviral state by inducing the expression of IFN-stimulated genes (ISGs). ISGs reduce viral load by suppressing virus entry into host cells, viral replication and/or transmission [1, 2].
IFNs of three types have been discovered, I (IFN‑α/β), II (IFN-γ) and III (IFN -λ). IFN-λ is the predominant IFN produced by mucosal epithelial cells and is pivotal to antiviral immunity against viral infection in the respiratory tract [3‒5]. Since the discovery of IFN-λ in 2003 significant knowledge has been accumulated about this group of cytokines and their functions in the body [6]. The study of IFN-λ functions confirmed a significant similarity of its antiviral activity with type I IFN, which is implemented via the same signaling pathways, as well as distinctive effects of IFN-λ on the body associated with the limited expression of its receptors. The type I IFN receptor is ubiquitously expressed by nearly all cell types, while the expression of the IFN-λ receptor, IFNLR1, is mainly limited to mucosal surfaces that act as anatomical barriers against entry and colonization of many microbes [7]. By contrast to type I IFNs, IFN-λ confers antiviral protection without driving a systemic proinflammatory state and exhibits tissue-protective and anti-inflammatory effects [8]. The lack of proinflammatory responses in the lung tissues [9] is an important advantage of IFN-λ over type I IFNs.
Thus, IFN-λ constitutes the first line of defense against microbial infections and together with IFN-α drives an antiviral state. IFN-λ functions as a sentinel molecule in host defense, since IFN-α starts its action only 48 h post-infection [10]. In general, type III IFNs control infection locally at barrier surfaces whereas type I IFNs control infection systemically.
POTENTIAL PATHWAYS OF SARS-CoV-2 EVASION OF HOST ANTIVIRAL ACTION
Coronavirus-2 has caused a large global outbreak of severe acute respiratory syndrome (SARS-CoV-2) and is a major public health issue worldwide. New tools and tactics to help stop the COVID-19 pandemic and treat COVID-19 are necessary.
SARS-CoV-2 is highly sensitive to IFN inhibition, but at the same time the virus can evade the immune detection.Several immune evasion strategies developed by SARS-CoV-2 are currently known [11]. First, during SARS-CoV infection, coronavirus isolates its double‒stranded RNA (dsRNA) inside double membrane vesicles, which shields the viral dsRNA from detection by the pattern recognition receptor (PRR) [12]. Another strategy of CoV is the inhibition of IFN production and signaling. Several studies showed that IFN-β production is suppressed by a range of SARS-CoV-2 proteins: NSP1, NSP3, NSP5, NSP12, NSP13, NSP14, NSP15, ORF3a, ORF3b, ORF6, ORF7a, ORF7b, ORF8, ORF9b, N, and M [13‒15]. It was shown that these proteins reduce the RIG-I (retinoic acid-inducible gene I)-mediated induction of IFN-β-promoter (IFNB1) activity, which suggests that they may suppress RLR-mediated (RIG-I-like receptor) signaling [13, 16, 17]. Xia et al. [17] compared the inhibitory activity of the Middle East respiratory syndrome coronavirus (MERS-CoV), severe acute respiratory coronavirus-1 (SARS-CoV-1), and SARS-CoV-2 in terms of type I IFN production and transmission of its signal and revealed that the NSP1 and NSP6 proteins of SARS-CoV-2 inhibit signal transmission of type I IFNs more effectively than the other two coronaviruses. Furthermore, coronavirus may antagonize IFN signaling for immune evasion through blocking phosphorylation of transcription factor STAT1 [18]. Miorin et al. [19] showed that SARS-CoV-2 is able to efficiently block STAT1 and STAT2 nuclear translocation in order to impair ISG transcription. Shin et al. [20] found that IFNs may be antagonized by the SARS-CoV-2 papain-like protease (PLpro) which cleaves the ubiquitin-like interferon-stimulated gene 15 protein (ISG15) to evade host innate immunity.
SARS-CoV-2 proteins suppress production of the endogenous interferons that affect COVID-19 severity [21‒23]. Scagnolari et al. [24] revealed that critically-ill patients that required invasive mechanical ventilation had a decreased expression of the IFN-λ(1‒3), type I IFN, and ISGs genes. In addition, it was reported that the presence of anti-IFN autoantibodies predicted critical COVID-19 [25, 26]. At the same time, some patients with severe COVID-19 had high levels of IFN-α, and hence the severe illness in these patients may be explained by reduced sensitivity of the virus to IFNs [27].
The association of IFN gene polymorphisms with COVID-19 susceptibility and severity is also of interest. Some diseases are known to be caused by genetic variants within components of the immune system, for example, in IFN-mediated signaling pathways. In 2009, three independent genome-wide association studies revealed that the rs12979860 single-nucleotide polymorphism in the promoter region of the IFN-λ3 (IL28B) gene is significantly associated with spontaneous clearance of the hepatitis C virus and a sustained virological response to chronic hepatitis [28‒30]. It was later revealed that IL28B genetic variants lead to variable immune responses to acute respiratory infections and SARS-CoV-2 infection. Homozygous variants of the genes encoding IFN-λ3/4 may correlate with reduced clearance of viruses in children with acute respiratory infections [31]. Saponi-Cortes et al. [32] revealed a relationship of the rs12979860 polymorphism of the IFN-λ4 (IFNL4) gene with symptomatic COVID-19 infection. With regard to the general population without this disease, the T allele of rs12979860 was overexpressed in COVID-19 patients, suggesting that this allele may be a risk factor for COVID-19. It was also found that the gene polymorphisms in the host (IL28B rs1297860 C/T and IFNL4 rs368234815 TT/ΔG) can affect the ability of the host to modulate viral infection without a clear impact on the outcome of COVID-19 [33]. Rahimi et al. [34] demonstrated that more severe COVID-19 was associated with unfavorable genotypes of IL28B and IFNL4 SNPs than favorable genotypes. The relationship of COVID-19 severity with certain allelic variants of IFN genes has a considerable clinical value and can be used to predict and optimize individual antiviral therapy regimens.
IFN-λ AS SARS-CoV-2 INHIBITOR
The innate immune system of the airway epithelium is the first line of defense against respiratory viruses, it produces IFN-λ (IFN- λ1, -λ2, -λ3 and -λ4), molecules eliciting a rapid antiviral response [35].
Davidson et al. [9] and Fox et al. [36] showed that primary tracheal epithelial cells and mouse lung epithelial cell lines induce the expression of antiviral genes in response to type I and type III IFNs. Viral infection of the epithelium stimulates the production of type I and type III IFNs, but type III IFN is the main IFN expressed in response to influenza A infection, respiratory syncytial virus, SARS-CoV-2 and others [37, 38]. New strains of coronaviruses that have emerged in recent decades cause severe upper and lower respiratory tract infection. Beucher et al. [39] reconstituted primary bronchial epithelia from adult and child donors to follow the SARS-CoV-2 infection dynamics using imaging methods and RT-PCR. Some reconstituted epithelia limited virus replication and spread. This phenotype was more frequent in epithelia derived from children versus adults and correlated with an accelerated release of type III IFN, while type III IFN gene knockout using CRISPR/Cas9 promoted infection. These data indicate the important role of IFN-λ in restriction of SARS-CoV-2 infection in the respiratory tract. The expression of type I INF (α and β) was undetectable in the supernatant and transcriptome signatures.
Currently, there is evidence of the antiviral activity of IFN-λ, both in vitro and in vivo, in relation to SARS-CoV-2. These data highlight a key role of IFN-λ in shaping immunity against SARS-CoV-2 infection and preventing clinical outcomes. The ability of IFN-λ to induce a narrower set of ISGs than type I IFN does in a more targeted set of cells expressing IFNLR1 and lack of systemic inflammation suggests possible therapeutic antiviral applications for IFN-λ. The critical stage in interferon therapy is the time when the treatment begins. A recent retrospective study of 446 patients with COVID-19 showed that early administration of IFN-α was associated with favorable outcome and recovery, whereas late administration of IFN-α, when multiple organs are affected, was associated with increased mortality [40].
Felgenhauer et al. [41] demonstrated an antiviral effect of IFN I (IFN-α) and III (IFN-λ) types against SARS-CoV-2. The researchers used two mammalian epithelial cell lines and found that both IFNs dose-dependently inhibit SARS-CoV-2. It was concluded that SARS-CoV-2 is sensitive to exogenously added IFNs [41]. Vanderheiden et al. [42] showed that pretreated human airway epithelial cultures with type I or III IFN 24 h prior to infection reduced viral RNA titers (3-fold) resulting in a 90% reduction in virus replication. Posttreatment of infected cells also reduced viral load: a significant effect was seen on day 3 of the experiment. In the above-mentioned study, Beucher et al. [39] demonstrated that treatment of bronchial adult epithelia with exogenous type III IFN restricted SARS-CoV-2 infection. Sohn et al. [43] revealed that IFN-λ1 significantly limited SARS-CoV-2 production in primary human bronchial epithelial cells in culture. Pretreatment of human lung cells with type I IFN completely blocked infectious virus production, and treatment with IFN-λ1 at the time of infection inhibited virus production more than 10-fold. In experiments with transgenic mice expressing the human angiotensin-converting enzyme 2 (ACE2), it was shown that one dose of IFN-λ1 administered intranasally reduced animal morbidity and mortality. Dijkman et al. [44] conducted a search for effective IFN-λ1 treatment regimen to control MERS-CoV infection. Cultures of human primary airway epithelial cells (hAECs) treated with a single dose of IFN-α and IFN-λ showed similar ISG expression, whereas cells treated with two doses of IFNs expressed elevated levels of ISGs only in IFN-λ-treated cells. Similarly, mice treated with two doses of IFN-λ were better protected than mice that received a single dose, and a combination of prophylactic and delayed therapeutic regimens completely protected mice from MERS-CoV infection.
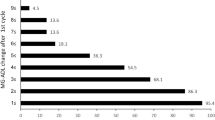
Miorin et al. [19] evaluated the susceptibility of SARS-CoV-2 to IFN pretreatment. Vero E6 cells were treated with different concentrations of type I, type II, or type III IFNs prior to infection. It was found that IFN-I pretreatment of Vero E6 cells drastically reduced the percentage of infected cells. Pretreatment with IFN-II also inhibited viral infection, whereas pretreatment of Vero E6 cells with IFN-III resulted in only a minor reduction of infection (Fig. 1). As IFN-III pretreatment effectively controls SARS-CoV-2 infection in vitro and in vivo [45, 46], it was suggested that Vero E6 cells express only low levels of the IFN-III receptor.
The effect of types I, II, and III interferons on infected SARS-CoV-2 Vero E6 cells. (a) The percentage of infected Vero E6 cells estimated as the ratio of cells positive for the viral nucleoprotein (NP) to the total number of cells (the data are shown as mean value ± standard deviation, n = 3). (b) Analysis of virus titers expressed as TCID50 in Vero E6 cell supernatants (data are shown as mean ± SEM, n = 3) [17, Creative Commons Attribution License 4.0 (CC BY)].
In experiments with three conventional mouse cell lines and transgenic mice expressing the human version of ACE2 Chong et al. [47] revealed that nasally-delivered IFN-λ confers pre- and post-exposure protection against infection by several SARS-CoV-2 strains without causing extensive inflammation.
ELABORATION OF IFN-λ COVID-19 THERAPEUTICS
Biopharmaceuticals based on recombinant human IFN-λ1 targeting COVID-19 hold promise as antiviral countermeasures. Nevertheless, despite the urgent need to find such antiviral interventions, there are still no effective antiviral drugs available for COVID-19. Recombinant protein therapeutics exhibit a good pharmacokinetic profile and are resistant to enzymes in blood.
PEGylation, a chemical modification of target proteins and peptides with polyethylene glycol (PEG), is one of the most common solutions to enhance the stability of protein drugs and prolong their therapeutic half-life [48]. Many such drugs are available in the pharmaceutical industry (Pegasys, PEG-Intron, Neulasta, Esperoct, Plegridy etc.) and numerous similar compounds are at the stage of development and preclinical studies. ZymoGenetics and Bristol-Myers Squibb (United States) have a candidate product of pegylated IFN-λ for parenteral administration to be used in prophylaxis of hepatitis virus. This pegylated IFN-λ showed a greater safety profile than pegylated IFN-α [49]. However, PEG-IFN-λ is an experimental agent and has not yet been approved.
An electron beam may be used for protein immobilization instead of chemical attachment [50]. Electron beam immobilization uses laser-driven acceleration of electrons to MeV-scale energies (1‒5 MeV) at doses of 0.5 to 6 Mrad and γ-radiation for immobilization of biologically active molecules on low molecular weight water-soluble carriers. The use of this technology makes it possible to obtain protein therapeutics with high bioavailability and low toxicity while preserving their biological activity.
We also used electron beam immobilization to prepare PEG-IFN-λ1 in order to obtain an active pharmaceutical substance to inhibit SARS-CoV-2. The PEG-IFN-λ1 therapeutic agent was created at the Institute of Cytology and Genetics (Novosibirsk, Russia) and the Goldberg Research Institute of Pharmacology and Regenerative Medicine (Tomsk, Russia) in cooperation with the Siberian Center of Pharmacology and Biotechnology (Russia). PEGylated products were obtained with electron beam usage [51]. PEG-IFN-λ1 was cytotoxic on cultured Vero E6 cells only at its maximum concentration, i.e., 108 μg/mL, while a three times lower concentration was not cytotoxic. In terms of the antiviral activity of both forms of IFN-λ1 against SARS-CoV-2 in Vero E6 cells, the half maximal inhibitory concentration (IC50) for PEG-IFN-λ1 was 25.0 ± 6.5 ng/mL, while it was 7.3 ± 3.1 ng/mL for recombinant IFN-λ1 (p < 0.05, Mann‒Whitney test) (Table 1).
Therefore, the inhibitory activity of PEG-IFN-λ1 on SARS-CoV-2 replication in Vero E6 cells was three times lower than that of unpegylated IFN-λ1. It is known that modifications of protein molecules often attenuate their functional activity because of an increased stereospecific blockade, which interferes with receptor binding [53, 54]. At the same time, a significant increase in stability and extended serum persistence of modified proteins often counteracts the reduced biological activity of target proteins.
Currently, PEG-IFN-λ is being investigated as a drug for the treatment of COVID-19. Dinnon et al. [55] demonstrated in a mouse model that the human PEG-IFN-λ1a potently inhibits SARS-CoV-2 replication in epithelial cells and both prophylactic and therapeutic administration of PEG-IFN-λ1a diminished SARS-CoV-2 replication in the lungs.
Feld et al. [56] described the advantage of a single subcutaneous injection of PEG-IFN-λ in the treatment of mild to moderate COVID-19 within 7 days of symptom onset or first positive swab if asymptomatic. Meanwhile, in another randomized placebo-controlled study of 120 patients with mild to moderate COVID-19, a subcutaneous dose of PEG-IFN-λ1 within 72 hours of diagnosis neither shortened the duration of SARS-CoV-2 viral shedding nor improved symptoms [57]. Hence, further investigations of PEG-IFN-λ use for prophylaxis and treatment of COVID-19 are necessary due to ambiguous results.
CONCLUSIONS
To date, the basic understanding of the antiviral role of IFN-λ has been obtained. The concept that IFN-λ is a sentinel molecule of epithelial barriers and a potent contributor to urgent antiviral protection appears reasonable. In our opinion, this concept is now at the stage of integration into practical usage. Evidently, exogenous IFNs instead of IFN inducers will be more effective in enhancing antiviral protection. The high risks of viral pandemics necessitate the urgent development and manufacture of medicines based on IFN-λ. Based on the experience of using IFN types I and II in medicine, it is argued that modified forms of IFN-λ will be the most advantageous, including pegylated forms obtained by electron beam or chemical immobilization with optimal pharmacokinetics.
REFERENCES
Schneider W.M., Chevillotte M.D., Rice C.M. 2014. Interferon-stimulated genes: a complex web of host defenses. Annu. Rev. Immunol. 32, 513–545.
Totura A.L., Baric R.S. 2012. SARS coronavirus pathogenesis: host innate immune responses and viral antagonism of interferon. Curr. Opin. Virol. 2 (3), 264–275.
Andreakos E., Salagianni M., Galani I.E., Koltsida O. 2017. Interferon-lambdas: front-line guardians of immunity and homeostasis in the respiratory tract. Front. Immunol. 8, 1232.
Dellgren C., Gad H.H., Hamming O.J., Melchjorsen J., Hartmann R. 2009. Human interferon-λ3 is a potent member of the type III interferon family. Genes Immun. 10 (2), 125–131.
Galani I.E., Triantafyllia V., Eleminiadou E.E., Koltsida O., Stavropoulos A., Manioudaki M., Thanos D., Doyle S.E., Kotenko S.V., Thanopoulou K., Andreakos E. 2017. Interferon-λ mediates nonredundant front-line antiviral protection against influenza virus infection without compromising host fitness. Immunity. 46 (5), 875–890. e6.
Donnelly R.P., Kotenko S.V. 2010. Interferon-lambda: a new addition to an old family. J. Interferon Cytokine Res. 30 (8), 555–564. https://doi.org/10.1089/jir.2010.0078
Mordstein M., Neugebauer E., Ditt V., Jessen B., Rieger T., Falcone V., Sorgeloos F., Ehl S., Mayer D., Kochs G., Schwemmle M., Günther S., Drosten C., Michiels T., Staeheli P. 2010. Lambda interferon renders epithelial cells of the respiratory and gastrointestinal tracts resistant to viral infections. J. Virol. 84 (11), 5670–5677.
Ye L., Schnepf D., Staeheli P. 2019. Interferon-λ orchestrates innate and adaptive mucosal immune responses. Nat. Rev. Immunol. 19 (10), 614–625.
Davidson S., McCabe T.M., Crotta S., Gad H.H., Hessel E.M., Beinke S., Hartmann R., Wack A. 2016. IFNλ is a potent anti-influenza therapeutic without the inflammatory side effects of IFNα treatment. EMBO Mol. Med. 8 (9), 1099–1112.
Jewell N.A., Cline T., Mertz S.E., Smirnov S.V., Flaño E., Schindler C., Grieves J.L., Durbin R.K., Kotenko S.V., Durbin J.E. 2010. Lambda interferon is the predominant interferon induced by influenza A virus infection in vivo. J. Virol. 84 (21), 11515–11522. https://doi.org/10.1128/JVI.01703-09
Znaidia M., Demeret C., van der Werf S., Komarova A.V. 2022. Characterization of SARS-CoV-2 evasion: interferon pathway and therapeutic options. Viruses. 14 (6), 1247.
Kumar S., Nyodu R., Maurya V.K., Saxena S.K. 2020. Host immune response and immunobiology of human SARS-CoV-2 infection. In Coronavirus Disease 2019 (COVID-19). Saxena, S., Ed. Medical Virology: From Pathogenesis to Disease Control. Singapore: Springer. 43–53. https://doi.org/10.1007/978-981-15-4814-7_5
Yuen C.K., Lam J.Y., Wong W.M., Mak L.F., Wang X., Chu H., Cai J.P., Jin D.Y., To K.K., Chan J.F., Yuen K.Y., Kok K.H. 2020. SARS-CoV-2 Nsp13, Nsp14, Nsp15 and Orf6 function as potent interferon antagonists. Emerg. Microbes Infect. 9 (1), 1418–1428.
Lei X., Dong X., Ma R., Wang W., Xiao X., Tian Z., Wang C., Wang Y., Li L., Ren L., Guo F., Zhao Z., Zhou Z., Xiang Z., Wang J. 2020. Activation and evasion of type I interferon responses by SARS-CoV-2. Nat. Commun. 11 (1), 3810.
Shemesh M., Aktepe T.E., Deerain J.M., McAuley J.L., Audsley M.D., David C.T., Purcell D.F.J., Urin V., Hartmann R., Moseley G.W., Mackenzie J.M., Schreiber G., Harari D. 2021. SARS-CoV-2 suppresses IFNβ production mediated by NSP1, 5, 6, 15, ORF6 and ORF7b but does not suppress the effects of added interferon. PLoS Pathog. 17 (8), e1009800.
Kouwaki T., Nishimura T., Wang G., Oshiumi H. 2021. RIG-I-like receptor-mediated recognition of viral genomic RNA of severe acute respiratory syndrome coronavirus-2 and viral escape from the host innate immune responses. Front. Immunol. 12, 700926.
Xia H., Cao Z., Xie X., Zhang X., Chen J.Y., Wang H., Menachery V.D., Rajsbaum R., Shi P.Y. 2020. Evasion of type I interferon by SARS-CoV-2. Cell Rep. 33 (1), 108234.
Lokugamage K.G., Hage A., de Vries M., Valero-Jimenez A.M., Schindewolf C., Dittmann M., Rajsbaum R., Menachery V.D. 2020. Type I interferon susceptibility distinguishes SARS-CoV-2 from SARS-CoV. J. Virol. 94 (23), e01410-20.
Miorin L., Kehrer T., Sanchez-Aparicio M.T., Zhang K., Cohen P., Patel R.S., Cupic A., Makio T., Mei M., Moreno E., Danziger O., White K.M., Rathnasinghe R., Uccellini M., Gao S., Aydillo T., Mena I., Yin X., Martin-Sancho L., Krogan N.J., Chanda S.K., Schotsaert M., Wozniak R.W., Ren Y., Rosenberg B.R., Fontoura B.M.A., Garcia-Sastre A. 2020. SARS-CoV-2 Orf6 hijacks Nup98 to block STAT nuclear import and antagonize interferon signaling. Proc. Natl. Acad. Sci. U. S. A. 117 (45), 28344–28354.
Shin D., Mukherjee R., Grewe D., Bojkova D., Baek K., Bhattacharya A., Schulz L., Widera M., Mehdipour A.R., Tascher G., Geurink P.P., Wilhelm A., van der Heden van Noort G.J., Ovaa H., Müller S., Knobeloch K.P., Rajalingam K., Schulman B.A., Cinatl J., Hummer G., Ciesek S., Dikic I. 2020. Papain-like protease regulates SARS-CoV-2 viral spread and innate immunity. Nature. 587 (7835), 657‒662.
Hadjadj J., Yatim N., Barnabei L., Corneau A., Boussier J., Smith N., Pere H., Charbit B., Bondet V., Chenevier-Gobeaux C., Breillat P., Carlier N., Gauzit R., Morbieu C., Pene F., Marin N., Roche N., Szwebel T.A., Merkling S.H., Treluyer J.M., Veyer D., Mouthon L., Blanc C., Tharaux P.L., Rozenberg F., Fischer A., Duffy D., Rieux-Laucat F., Kerneis S., Terrier B. 2020. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science. 369 (6504), 718‒724.
Blanco-Melo D., Nilsson-Payant B.E., Liu W.C., Uhl S., Hoagland D., Moller R., Jordan T.X., Oishi K., Panis M., Sachs D., Wang T.T., Schwartz R.E., Lim J.K., Albrecht R.A., tenOever B.R. 2020. Imbalanced host response to SARS-CoV2 drives development of COVID-19. Cell. 181 (5), 1036–1045. e9.
Schultze J.L., Aschenbrenner A.C. 2021. COVID-19 and the human innate immune system. Cell. 184 (7), 1671–1692.
Scagnolari C., Pierangeli A., Frasca F., Bitossi C., Viscido A., Oliveto G., Scordio M., Mazzuti L., Di Carlo D., Gentile M., Solimini A., Ceccarelli G., Pugliese F., Mastroianni C.M., d’Ettorre G., Turriziani O., Antonelli G. 2021. Differential induction of type I and III interferon genes in the upper respiratory tract of patients with coronavirus disease 2019 (COVID-19). Virus Res. 295, 198283.
Bastard P., Orlova E., Sozaeva L., Levy R., James A., Schmitt M.M., Ochoa S., Kareva M., Rodina Y., Gervais A., Le Voyer T., Rosain J., Philippot Q., Neehus A.L., Shaw E., Migaud M., Bizien L., Ekwall O., Berg S., Beccuti G., Ghizzoni L., Thiriez G., Pavot A., Goujard C., Fremond M.L., Carter E., Rothenbuhler A., Linglart A., Mignot B., Comte A., Cheikh N., Hermine O., Breivik L., Husebye E.S., Humbert S., Rohrlich P., Coaquette A., Vuoto F., Faure K., Mahlaoui N., Kotnik P., Battelino T., Trebušak Podkrajšek K., Kisand K., Ferré E.M.N., DiMaggio T., Rosen L.B., Burbelo P.D., McIntyre M., Kann N.Y., Shcherbina A., Pavlova M., Kolodkina A., Holland S.M., Zhang S.Y., Crow Y.J., Notarangelo L.D., Su H.C., Abel L., Anderson M.S., Jouanguy E., Neven B., Puel A., Casanova J.L., Lionakis M.S. 2021. Preexisting autoantibodies to type I IFNs underlie critical COVID-19 pneumonia in patients with APS-1. J. Exp. Med. 218 (7), e20210554.
Bastard P., Rosen L.B., Zhang Q., Michailidis E., Hoffmann H.H., Zhang Y., Dorgham K., Philippot Q., Rosain J., Béziat V., Manry J., Shaw E., Haljasmägi L., Peterson P., Lorenzo L., Bizien L., Trouillet-Assant S., Dobbs K., de Jesus A.A., Belot A., Kallaste A., Catherinot E., Tandjaoui-Lambiotte Y., Le Pen J., Kerner G., Bigio B., Seeleuthner Y., Yang R., Bolze A., Spaan A.N., Delmonte O.M., Abers M.S., Aiuti A., Casari G., Lampasona V., Piemonti L., Ciceri F., Bilguvar K., Lifton R.P., Vasse M., Smadja D.M., Migaud M., Hadjadj J., Terrier B., Duffy D., Quintana-Murci L., van de Beek D., Roussel L., Vinh D.C., Tangye S.G., Haerynck F., Dalmau D., Martinez-Picado J., Brodin P., Nussenzweig M.C., Boisson-Dupuis S., Rodríguez-Gallego C., Vogt G., Mogensen T.H., Oler A.J., Gu J., Burbelo P.D., Cohen J.I., Biondi A., Bettini L.R., D’Angio M., Bonfanti P., Rossignol P., Mayaux J., Rieux-Laucat F., Husebye E.S., Fusco F., Ursini M.V., Imberti L., Sottini A., Paghera S., Quiros-Roldan E., Rossi C., Castagnoli R., Montagna D., Licari A., Marseglia G.L., Duval X., Ghosn J., Tsang J.S., Goldbach-Mansky R., Kisand K., Lionakis M.S., Puel A., Zhang S.Y., Holland S.M., Gorochov G., Jouanguy E., Rice C.M., Cobat A., Notarangelo L.D., Abel L., Su H.C., Casanova J.L. 2020. Autoantibodies against type I IFNs in patients with life-threatening COVID-19. Science. 370 (6515), eabd4585.
Lucas C., Wong P., Klein J., Castro T.B.R., Silva J., Sundaram M., Ellingson M.K., Mao T., Oh J.E., Israelow B., Takahashi T., Tokuyama M., Lu P., Venkataraman A., Park A., Mohanty S., Wang H., Wyllie A.L., Vogels C.B.F., Earnest R., Lapidus S., Ott I.M., Moore A.J., Muenker M.C., Fournier J.B., Campbell M., Odio C.D., Casanovas-Massana A., Herbst R., Shaw A.C., Medzhitov R., Schulz W.L., Grubaugh N.D., Dela Cruz C., Farhadian S., Ko A.I., Omer S.B., Iwasaki A. 2020. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature. 584 (7821), 463–469.
Ge D., Fellay J., Thompson A.J., Simon J.S., Shi-anna K.V., Urban T.J., Heinzen E.L., Qiu P., Be-rtelsen A.H., Muir A.J., Sulkowski M., McHutchison J.G., Goldstein D.B. 2009. Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature. 461 (7262), 399–401.
Suppiah V., Moldovan M., Ahlenstiel G., Berg T., Weltman M., Abate M.L., Bassendine M., Spengler U., Dore G.J., Powell E., Riordan S., Sheridan D., Smedile A., Fragomeli V., Muller T., Bahlo M., Stewart G.J., Booth D.R., George J. 2009. IL28B is associated with response to chronic hepatitis C interferon-α and ribavirin therapy. Nat. Genet. 41 (10), 1100–1104.
Tanaka Y., Nishida N., Sugiyama M., Kurosaki M., Matsuura K., Sakamoto N., Nakagawa M., Korenaga M., Hino K., Hige S., Ito Y., Mita E., Tanaka E., Mochida S., Murawaki Y., Honda M., Sakai A., Hiasa Y., Nishiguchi S., Koike A., Sakaida I., Imamura M., Ito K., Yano K., Masaki N., Sugauchi F., Izumi N., Tokunaga K., Mizokami M. 2009. Genome-wide association of IL28B with response to pegylated interferon-α and ribavirin therapy for chronic hepatitis C. Nat. Genet. 41 (10), 1105–1109.
Rugwizangoga B., Andersson M.E., Kabayiza J.C., Nilsson M.S., Ármannsdóttir B., Aurelius J., Nilsson S., Hellstrand K., Lindh M., Martner A. 2019. IFNL4 genotypes predict clearance of RNA viruses in Rwandan children with upper respiratory tract infections. Front. Cell Infect. Microbiol. 9, 340.
Saponi-Cortes J.M.R., Rivas M.D., Calle-Alonso F., Sanchez J.F., Costo A., Martin C., Zamorano J. 2021. IFNL4 genetic variant can predispose to COVID-19. Sci. Rep. 11 (1), 21185.
Amodio E., Pipitone R.M., Grimaudo S., Immordino P., Maida C.M., Prestileo T., Restivo V., Tramuto F., Vitale F., Craxì A., Casuccio A. 2020. SARS-CoV-2 viral load, IFNλ polymorphisms and the course of COVID-19: an observational study. J. Clin. Med. 9 (10), 3315.
Rahimi P., Tarharoudi R., Rahimpour A., Mosayebi Amroabadi J., Ahmadi I., Anvari E., Siadat S.D., Aghasadeghi M., Fateh A. 2021. The association between interferon lambda 3 and 4 gene single-nucleotide polymorphisms and the recovery of COVID-19 patients. Virol. J. 18 (1), 221.
Kim H.J., Jo A., Jeon Y.J., An S., Lee K.M., Yoon S.S., Choi J.Y. 2019. Nasal commensal Staphylococcus epidermidis enhances interferon-λ-dependent immunity against influenza virus. Microbiome. 7 (1), 80.
Fox J.M., Crabtree J.M., Sage L.K., Tompkins S.M., Tripp R.A. 2015. Interferon lambda upregulates IDO1 expression in respiratory epithelial cells after influenza virus infection. J. Interferon Cytokine Res. 35 (7), 554–562.
Iwasaki A., Pillai P.S. 2014. Innate immunity to influenza virus infection. Nat. Rev. Immunol. 14 (5), 315–328.
Wang J., Oberley-Deegan R., Wang S., Nikrad M., Funk C.J., Hartshorn K.L., Mason R.J. 2009. Human alveolar type II cells secrete antiviral IL-29 (IFN-λ1) in response to influenza A infection. J. Immunol. 182 (3), 1296–1304.
Beucher G., Blondot M.L., Celle A., Pied N., Recordon-Pinson P., Esteves P., Faure M., Métifiot M., Lacomme S., Dacheux D., Robinson D.R., Längst G., Beaufils F., Lafon M.E., Berger P., Landry M., Malvy D., Trian T., Andreola M.L., Wodrich H. 2022. Bronchial epithelia from adults and children: SARS-CoV-2 spread via syncytia formation and type III interferon infectivity restriction. Proc. Natl. Acad. Sci. U. S. A. 119 (28), e2202370119.
Wang N., Zhan Y., Zhu L., Hou Z., Liu F., Song P., Qiu F., Wang X., Zou X., Wan D., Qian X., Wang S., Guo Y., Yu H., Cui M., Tong G., Xu Y., Zheng Z., Lu Y., Hong P. 2020. Retrospective multicenter cohort study shows early interferon therapy is associated with favorable clinical responses in COVID-19 patients. Cell Host Microbe. 28 (3), 455–464.
Felgenhauer U., Schoen A., Gad H.H., Hartmann R., Schaubmar A.R., Failing K., Drosten C., Weber F. 2020. Inhibition of SARS-CoV-2 by type I and type III interferons. J. Biol. Chem. 295 (41), 13958‒13964.
Vanderheiden A., Ralfs P., Chirkova T., Upadhyay A.A., Zimmerman M.G., Bedoya S., Aoued H., Tharp G.M., Pellegrini K.L., Manfredi C., Sorscher E., Mainou B., Lobby J.L., Kohlmeier J.E., Lowen A.C., Shi P.Y., Menachery V.D., Anderson L.J., Grakoui A., Bosinger S.E., Suthar M.S. 2020. Type I and type III interferons restrict SARS-CoV-2 infection of human airway epithelial cultures. J. Virol. 94 (19), e00985-20.
Sohn S.Y., Hearing J., Mugavero J., Kirillov V., Gorbunova E., Helminiak L., Mishra S., Mackow E., Hearing P., Reich N.C., Kim H.K. 2021. Interferon-lambda intranasal protection and differential sex pathology in a murine model of SARS-CoV-2 infection. mBio. 12 (6), e0275621.
Dijkman R., Verma A.K., Selvaraj M., Ghimire R., Gad H.H., Hartmann R., More S., Perlman S., Thiel V., Channappanavar R. 2022. Effective interferon lambda treatment regimen to control lethal MERS-CoV infection in mice. J. Virol. 96 (11), e0036422. https://doi.org/10.1128/jvi.00364-22
Stanifer M.L., Kee C., Cortese M., Zumaran C.M., Triana S., Mukenhirn M., Kraeusslich H.G., Alexandrov T., Bartenschlager R., Boulant S. 2020. Critical role of type III interferon in controlling SARS-CoV-2 infection in human intestinal epithelial cells. Cell Rep. 32 (1), 107863.
Dinnon K.H., Leist S.R., Schäfer A., Edwards C.E., Martinez D.R., Montgomery S.A., West A., Yount B.L., Hou Y.J., Adams L.E., Gully K.L., Brown A.J., Huang E., Bryant M.D., Choong I.C., Glenn J.S., Gralinski L.E., Sheahan T.P., Baric R.S. 2020. A mouse-adapted SARS-CoV-2 model for the evaluation of COVID-19 medical countermeasures. bioRxiv. 2020.05.06.081497. https://doi.org/10.1101/2020.05.06.081497
Chong Z., Karl C.E., Halfmann P.J., Kawaoka Y., Winkler E.S., Yu J., Diamond M.S. 2022. Nasally-delivered interferon-λ protects mice against upper and lower respiratory tract infection of SARS-CoV-2 variants including Omicron. bioRxiv. 2022.01.21.477296. https://doi.org/10.1101/2022.01.21.477296
Yadav D., Dewangan H.K. 2021. PEGYLATION: An important approach for novel drug delivery system. J. Biomater. Sci. Polym. Ed. 32 (2), 266–280.
Chan H.L.Y., Ahn S.H., Chang T.T., Peng C.Y., Wong D., Coffin C.S., Lim S.G., Chen P.J., Janssen H.L.A., Marcellin P., Serfaty L., Zeuzem S., Cohen D., Critelli L., Xu D., Wind-Rotolo M., Cooney E. 2016. Peginterferon lambda for the treatment of HBeAg-positive chronic hepatitis B: a randomized phase 2b study (LIRA-B). J. Hepatol. 64 (5), 1011–1019.
Madonov P.G., Ershov K.I., Dubrovin A.V., Za-polotsky E.N., Miroshnikov P.N., Shilova, M.A. Kinsht D.N. 2013. Electron beam modification of protein preparations to reveal their pharmacological properties. Med. Obraz. Sib. 4, 83.
Artamonov A.V., Bekarev A.A., Dygai A.M., Z-hdanov V.V., Kinsht D.N., Madonov P.G., Sherstoboev E.Yu. 2019. Pegylated lambda interferon with high oral bioavailability and method for its production. RF Patent no. RU2678332C1, 2019.
Madonov P.G., Svyatchenko V.A., Legostaev S.S., Kikhtenko N.A., Kotlyarova A.A., Oleynik L.A., Baykalov G.I., Udut B.B. 2021. Antiviral activity against SARS-CoV-2 of a pharmaceutical substance based on immobilized recombinant human interferon lambda-1. Eksp. Klin. Farmakol. 84 (7), 15‒20.
Kubetzko S., Sarkar C.A., Plückthun A. 2005. Protein PEGylation decreases observed target association rates via a dual blocking mechanism. Mol. Pharmacol. 68 (5), 1439–1454.
Grace M.J., Lee S., Bradshaw S., Chapman J., Spond J., Cox S., Delorenzo M., Brassard D., Wylie D., Cannon-Carlson S., Cullen C., Indelicato S., Voloch M., Bordens D. 2005. Site of pegylation and polyethylene glycol molecule size attenuate interferon-alpha antiviral and antiproliferative activities through the JAK/STAT signaling pathway. J. Biol. Chem. 280 (8), 6327–6336.
Dinnon K.H., Leist S.R., Schäfer A., Edwards C.E., Martinez D.R., Montgomery S.A., West A., Yount B.L. Jr., Hou Y.J., Adams L.E., Gully K.L., Brown A.J., Huang E., Bryant M.D., Choong I.C., Glenn J.S., Gralinski L.E., Sheahan T.P., Baric R.S. 2020. A mouse-adapted model of SARS-CoV-2 to test COVID-19 countermeasures. Nature. 586 (7830), 560–566.
Feld J.J., Kandel C., Biondi M.J., Kozak R.A., Zahoor M.A., Lemieux C., Borgia S.M., Boggild A.K., Powis J., McCready J., Tan D.H.S., Chan T., Coburn B., Kumar D., Humar A., Chan A., O’Neil B., Noureldin S., Booth J., Hong R., Smookler D., Aleyadeh W., Patel A., Barber B., Casey J., Hiebert R., Mistry H., Choong I., Hislop C., Santer D.M., Lorne Tyrrell D., Glenn J.S., Gehring A.J., Janssen H.L.A., Hansen B.E. 2021. Peginterferon lambda for the treatment of outpatients with COVID-19: A phase 2, placebo-controlled randomised trial. Lancet Respir. Med. 9 (5), 498–510.
Jagannathan P., Andrews J.R., Bonilla H., Hedlin H., Jacobson K.B., Balasubramanian V., Purington N., Kamble S., de Vries C.R., Quintero O., Feng K., Ley C., Winslow D., Newberry J., Edwards K., Hislop C., Choong I., Maldonado Y., Glenn J., Bhatt A., Blish C., Wang T., Khosla C., Pinsky B.A., Desai M., Parsonnet J., Singh U. 2021. Peginterferon lambda-1α for treatment of outpatients with uncomplicated COVID-19: a randomized placebo-controlled trial. Nat. Commun. 12 (1), 1967.
Funding
This study was not funded by a specific project grant.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by M. Novikova
Abbreviations: IFN, interferon; ISGs, interferon-stimulated genes.
Rights and permissions
About this article
Cite this article
Oleinik, L.A., Madonov, P.G. & Pykhtina, M.B. Potential of Interferon Lambda as an Inhibitor of SARS-CoV-2. Mol Biol 57, 291–298 (2023). https://doi.org/10.1134/S0026893323020152
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893323020152