Abstract
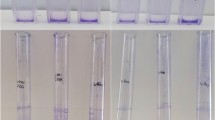
The anti-biofilm effects of the following probiotic LAB strains: Lactobacillus plantarum M16 (producer of plantarisin), Lactobacillus sake M17 (producer of sakacin), Pediococcus acidilactici M20 (producer of pediocin), Bifidobacterium longum M23 (bifidosin A producer strain), Bifidobacterium bifidum M24 (bifidocin B producer strain), and Pediococcus pentosaceus M46 (bacteriocin like substance producer strain) on the formation and eradication of mature Salmonella Typhimurium 14 028 biofilms were investigated. The strains were previously isolated from fermented foods traditionally produced in Turkey and identified. Biofilm formation by Salmonella Typhimurium 14 028 was completely inhibited after treatment with the cell-free filtrates of lactic acid bacteria. The amount of mature biofilms dropped at statistically significant levels after six hours (p < 0.05) only as the result of treatment with Pediococcus pentosaceus M46 cell-free filtrate, rather than with other cell-free filtrates. On the other hand, the cell-free filtrates of the other LAB strains promoted biofilm maturation. Biofilm assays were carried out with neutralized cell-free filtrates and neutralized cell-free filtrates treated with proteinase K to determine the source of the antibiofilm activity. The antibiofilm activities that were initially determined dropped statistically significantly (p < 0.05) as a result of both treatments. Based on these findings, the probiotic strains isolated from Turkey exhibited particularly significant antibiofilm capabilities, which were mostly due to the bacteriocins produced by these strains.







Similar content being viewed by others
REFERENCES
Afdora, P.T., Ardiyati, T., Sjofjan, O., and Kalsum, U., Potential antibacterials compounds of lactic acid bacteria (LAB) from quail intestine (Coturnix japonica) in inhibition growth of Escherichia coli and Salmonella typhimurium, J. Trop. Life Sci., 2010, vol. 1, pp. 28‒31.
Barzegari, A., Kheyrolahazadeh, K., Khatibi, S.M.H., Sharifi, S., Memar, My., and Vahed, S.Z., The battle of Probiotics and their derivatives against biofilms, Infect. Drug Resist., 2020, vol. 13, pp. 659‒672.
Costerton, J.W., Lewandowski, Z., Caldwell, D.E., Korber, D.R., and Lappin-Scott, H.M., Microbial biofilms, Annu. Rev. Microbiol.,1995, vol. 49, pp. 711‒745.
Cui, X., Shi, Y., Gu, S., Yan, X., and Chen, H., Antibacterial and antibiofilm activity of lactic acid bacteria isolated from traditional artisanal milk cheese from northeast China against enteropathogenic bacteria, Probiotics and Antimicrobial Proteins, 2018, vol. 10, pp. 601‒610.
Davey, M.E. and O’toole, G.A., Microbial biofilms: from ecology to molecular genetics, Microbiol. Mol. Biol. Rev., 2000, vol. 64, pp. 847‒867.
De Vuyst, L. and Leroy, F., Bacteriocins from lactic acid bacteria: production, purification, and food applications, J. Mol. Microbiol. Biotechnol., 2007, vol. 13, pp. 194‒199.
Forde, A. and Fitzgerald, G.F., Molecular organization of exopolysaccharide (EPS) encoding genes on the lactococcal bacteriophage adsorption blocking plasmid, pCI658, Plasmid, 2003, vol. 49, pp. 130‒142.
Galvez, A., Abriouel, H., Benomar, N., and Lucas, R., Microbial antagonists to food-borne pathogens and biocontrol, Curr. Opin. Biotechnol., 2010, vol. 21, pp. 142‒148.
Ge, J., Sun, Y., Xin, X., Wang, Y., and Ping, W., Purification and partial characterization of a novel bacteriocin synthesized by Lactobacillus paracasei HD1-7 isolated from Chinese sauerkraut juice, Sci. Rep., 2016, vol. 6, pp. 1‒7.
Gomez, N.C., Rasmiro, J.M.P., Quecan, B.X.V., and Franco, B.D.G., Use of potential probiotic lactic acid bacteria (LAB) biofilms for the control of Listeria monocytogenes, Salmonella Typhimurium, and Escherichia coli O157:H7 biofilms formation, Front. Microbiol., 2016, vol. 12, pp. 1‒15.
Gunn, J.S., Marshall, J.M., Baker, S., Dongol, S., Charles, R.C., and Ryan, E.T., Salmonella chronic carriage: epidemiology, diagnosis, and gallbladder persistence, Trends Microbiol., 2014, vol. 22, pp. 648–655.
Hahn, M.M. and Gunn, J.S., Salmonella extracellular polymeric substances modulate innate phagocyte activity and enhance tolerance of biofilm-associated bacteria to oxidative stress, Microorganisms, 2020, vol. 8, pp. 253‒272.
Hall-Stoodley, L., Hu, F.Z., Gieseke, A., Nistico, L.N., Hayes, J., and Wackym, P.A., Direct detection of bacterial biofilms on the middle-ear mucosa of children with chronic otitis media, JAMA, 2006, vol. 296, pp. 202‒211.
Høiby, N., Bjarnsholt, T., Givskov, M., Molin, S., and Ciofu, O., Antibiotic resistance of bacterial biofilms, Int. J. Antimicrob. Agents, 2010, vol.35, pp. 322‒332.
Houndt, R.V. and Michiels, C.W., Biofilm formation and the food industry, a focus on the bacterial outer surface, J. Appl. Microbiol., 2010, vol. 109, pp. 1117‒1131.
Joseph, B., Otta, S.K., Karunasagar, I., and Karunasagar, J., Biofilm formation by Salmonella spp. on food contact surfaces and their sensitivity to sanitizers, Int. J. Food Microbiol., 2001, vol. 64, pp. 367‒372.
Karpanen, T.J., Worthington, T., Hendry, E.R., Conway, B.R., and Lambert, P.A., Antimicrobial efficacy of chlorhexidine digluconate alone and in combination with eucalyptus oil, tea tree oil, and thymol against planktonic and biofilm cultures of Staphylococcus epidermidis, J. Antimicrob. Chemother., 2008, vol. 62, pp. 1031‒1036.
Khan, A., Vu, K.D., Riedl, B., and Lacroix, M., Optimization of the antimicrobial activity of nisin, Na-EDTA, and pH against Gram-negative and Gram-positive bacteria, LWT-Food Sci. Technol., 2015, vol. 61, pp. 124‒129.
Kim, N.N., Kim, W.J., and Kang, S.S., Anti-biofilm effect of crude bacteriocin derived from Lactobacillus brevis DF01 on Escherichia coli and Salmonella Typhimurium, Food Control, 2019, vol.98, pp. 274‒280.
Lee, H.J., Jeong, S.E., Kim, P.J., Madsen, E.L., and Jeon, C.O., High resolution depth distribution of Bacteria, Archaea, methanotrophs, and methanogens in the bulk and rhizosphere soils of a flooded rice paddy, Front. Microbiol., 2015, vol. 6, pp. 1‒13.
Levkovich, T., Poutahidis, T., Smillie, C., Varian, B.J., Ibrahim, Y.M., Lakritz, J.R., and Erdman, S.E., Probiotic bacteria induce a ‘glow of health,’ PLoS One, 2013, vol. 8, e53867.
Li, P., Gu, Q., and Zhou Q., Complete genome sequence of Lactobacillus plantarum LZ206, a potential probiotic strain with antimicrobial activity against food-borne pathogenic microorganisms, J. Biotechnol., 2016, vol. 238, pp. 52‒55.
Mulet-Powell, N., Lacoste-Armynot, A.M., and De Bouchberg, M.S., Interactions between pairs of bacteriocins from lactic acid bacteria, J. Food Prot., 1998, vol. 61, pp. 1210‒1212.
Norhana, M.W., Poole, S.E., Deeth, H.C., and Dykes, G.A., Effects of nisin, EDTA and salts of organic acids on Listeria monocytogenes, Salmonella and native microflora on fresh vacuum packaged shrimps stored at 4°C, Food Microbiol., 2012, vol. 31, pp. 43‒50.
Pelyuntha, W., Chaiyasut, C., Kantachote, D., and Sirilun, S., Cell-free supernatants from cultures of lactic acid bacteria isolated from fermented grape as biocontrol against Salmonella Typhi and Salmonella Typhimurium virulence via autoinducer-2 and biofilm interference, Peer J., 2019, vol. 7, pp. 7555‒7570.
Petrova, M.I., Imholz, N.C.E., Verhoeven, T.L.A., Balzarini, J., Van Damme, E.J.M., Schols, D., Vanderleyden, J., and Lebeer, S., Lectin-like molecules of Lactobacillus rhamnosus GG inhibit pathogenic Escherichia coli and Salmonella biofilm formation, PLos One, 2016, vol. 11, e0161337.
Post, J.C., Hall-Stoodley, P., and Ehrlich, G.D., The role of biofilms in otolaryngologic infections, Curr. Opin. Otolaryngol. Head Neck Surg., 2004, vol. 12, pp. 185‒190.
Roy, R., Tiwari, M., Donelli, G., and Tiwari, V., Strategies for combating bacterial biofilms: a focus on anti-biofilm agents and their mechanisms of action, Virulence, 2018, vol. 9, pp. 522‒554.
Stepanović, S., Vuković, D., Dakić, I., Savić, B., and Šv-abić-Vlahović, M.A., Modified microtiter-plate test for quantification of staphylococcal biofilm formation, J. Microbiol. Methods, 2000, vol.40, pp. 175–179.
Su, J., Wu, Y., Ma, X., Zhang, G., Feng, H., and Zhang, Y., Soil microbial counts and identification of culturable bacteria in an extreme by arid zone, Folia Microbiol., 2004, vol. 49, pp. 423‒429.
Teuber, M., Production of chymosin (EC 3.4.23.4) by microorganisms and its use for cheesemaking, Bull. Int. Dairy Fed., 1990, vol. 251, pp. 3‒15.
Van Belkum, M.J., Hayema, B.J., Geis, A., Kok, J., and Venema, G., Cloning of two bacteriocin genes from a lactococcal bacteriocin plasmid, Appl. Environ. Microbiol., 1989, vol. 55, pp. 1187‒1191.
Vestby, L.K., Møretrø, T., Langsrud, S., Heir, E., and Nesse, L.L., Biofilm forming abilities of Salmonella are correlated with persistence in fish meal-and feed factories, BMC Vet. Res., 2009, vol.5, pp. 20‒29.
Viedma, P.M., Abriouel, H., Lopez, A.S., Omar, N.B., Lopez, R.S., Valdiva, E., Belloso, O.V., and Galvez, A., Effect of enterocin AS-48 in combination with high-intensity pulsed-electric field treatment against the spoilage bacterium Lactobacillus diolivorans in apple juice, Food Microbiol., 2009, vol.26, pp. 491‒496.
ACKNOWLEDGMENTS
We would like to thank Prof. Hilmi Volkan Demir (National Nanotechnology Center, Bilkent University, UNAM), who let us work with a scanning electron microscope.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
Conflict of interests. The authors declare that they have no conflict of interest.
Statement on the welfare of animals. This article does not contain any studies involving animals or human participants performed by any of the authors.
AUTHORS CONTRIBUTION
M. Akçelik participated in designing and performing experiments, processing, and interpreting data, and preparing, writing, and revising the manuscript. Ş. Göksel participated in performing experiments, processing data, and preparation of the manuscript. N. Akçelik assisted in designing experiments, processing, and interpreting the data. C. Özdemir participated in biofilm assays, scanning electron microscopic examinations and processing data.
Rights and permissions
About this article
Cite this article
Göksel, Ş., Akçelik, N., Özdemir, C. et al. The Effects of Lactic Acid Bacteria on Salmonella Biofilms. Microbiology 91, 278–285 (2022). https://doi.org/10.1134/S0026261722300129
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261722300129