Abstract—
Planctomycetes of the class Phycisphaerae are aerobic and anaerobic heterotrophic bacteria that colonize a wide range of marine and terrestrial habitats. Their functional roles in the environment, however, are still poorly understood. Humisphaera borealis M1803T is one of the very few characterized planctomycetes of this class. It is also the first described representative of the previously uncultured group WD2101, which is commonly detected in soils and peatlands. This work analyzed the genetic determinants that define the ability of Humisphaera borealis M1803T to grow on xylan, one of the plant cell wall polymers. The whole genome sequence analysis of this planctomycete resulted in identification of five genes encoding the proteins homologous to previously described endo-β-xylanases. For two of these proteins, evolutionarily closer experimentally characterized homologs with other substrate specificities were found. In a member of the GH10 family of glycoside hydrolases, the active center of the enzyme was destroyed. We consider two proteins from GH62 and GH141 families as the most likely candidates for the role of β-xylanase responsible for xylan utilization. Phylogenetic analysis of proteins of GH10, GH62, and GH141 families was carried out. The role of lateral transfers in the evolution of the genes for glycoside hydrolases and their close homologs is discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The class Phycisphaerae includes aerobic and anaerobic heterotrophic planctomycetes that utilize a broad range of organic compounds and inhabit marine and terrestrial ecosystems (Fukunaga et al., 2009). Their key difference from the other class of planctomycetes, Planctomycetia, is the way of reproduction: members of Planctomycetia are budding bacteria, whereas Phycisphaerae propagate by binary fission (Ward and Dedysh, 2022). The number of cultured and characterized Planctomycetia members, as well as the body of data on their biology and organization of their cells, exceed considerably the amount of information available on Phycisphaerae. The latter class includes three orders: Phycisphaerales (Fukunaga et al., 2009), Sedimentisphaerales (Spring et al., 2018), and Tepidisphaerales (Kovaleva et al., 2015), with the total number of characterized members of less than a dozen.
Until recently, the order Tepidisphaerales included only one genus, Tepidisphaera, with a single described species, Tepidisphaera mucosa, represented by thermophilic polysaccharide-utilizing planctomycetes from terrestrial thermal springs (Kovaleva et al., 2015). One year ago, a mesophilic species isolated from an eutrophic lake of the boreal zone, Humisphaera borealis M1803T, was added to the diversity of cultured members of the order Tepidisphaerales (Dedysh et al., 2021). This planctomycete is the first representative of the phylogenetic group WD2101 within the order Tepidisphaerales that has been obtained in culture. According to molecular studies of microbial diversity, planctomycetes of group WD2101 are typical inhabitants of peatlands and soils rich in organic matter (Ivanova et al., 2016; Dedysh and Ivanova, 2019; Dedysh et al., 2021). Genome analysis showed that the H. borealis M1803T possesses a complete set of genes responsible for the synthesis of planctomycete-type microcompartments, specialized cell structures involved in utilization of monosaccharides of plant cell walls, e.g., rhamnose (Dedysh et al., 2021). Analysis of cellular ultrastructure by electron microscopy confirmed the presence of these compartments in cells grown on rhamnose, suggesting that these planctomycetes participate in degradation of plant debris in wetland ecosystems. It was found that a culture of H. borealis M1803T can also grow on xylan, a polymeric constituent of the plant cell wall; however, its growth on this substrate has not been studied in any detail.
Planctomycetes have a great glycolytic potential, and their genomes encode a broad range of glycoside hydrolases (Ivanova et al., 2017; Kulichevskaya et al., 2020; Drula et al., 2022). Glycoside hydrolases, or glycosidases, (EC 3.2.1), are a vast group of enzymes that cleave O-glycoside bonds in various substrates, including heteropolysaccharides, glycoproteins, and glycolipids (Naumoff, 2011). Depending on substrate specificity, more than 200 variants of enzymatic activity are recognized: EC 3.2.1.1–3.2.1.217 (McDonald et al., 2009). In the CAZy database, most of the currently known glycoside hydrolases are combined into 165 families based on amino acid sequence homology, GH1–GH173 (except GH21, GH40, GH41, GH60, GH61, GH69, GH145, and GH155) (Lombard et al., 2014; Terrapon et al., 2017; Garron and Henrissat, 2019; Drula et al., 2022). As a rule, one family may include enzymes with different activities, while members of different families may exhibit the same substrate specificity. Using the results of phylogenetic analysis, families can be divided into largely monofunctional subfamilies, making it possible to predict fairly reliably the biological functions of proteins that have not been characterized experimentally. In the present work, this approach was applied to search for genes that encode putative β-xylanases in H. borealis M1803T.
Endo-β-xylanases (EC 3.2.1.8 and EC 3.2.1.32) are a widespread group of glycoside hydrolases with a substantial biotechnological potential (Kulkarni et al., 1999; Beg et al., 2001; Pollet et al., 2010; Naumoff, 2016; Linares-Pasten et al., 2018; Nguyen et al., 2018; Nordberg Karlsson et al., 2018; Capetti et al., 2021; Gupta et al., 2021). They are responsible for degradation of diverse heteroxylans: a structurally heterogeneous group of heteropolysaccharides that basically form the hemicellulose of cell walls of higher plants. In the CAZy database, the known endo-β-xylanases are classified into 17 glycoside hydrolase families: GH3, GH5, GH6, GH8, GH9, GH10, GH11, GH16, GH18, GH26, GH30, GH43, GH44, GH51, GH62, GH98, and GH141 (Drula et al., 2022). However, GH10 and GH11 are the only two families where enzymes with endo-β-xylanase activity clearly dominate among the biochemically characterized members.
Sequencing (Dedysh et al., 2021) and subsequent annotation (Drula et al., 2022) of the genome of H. borealis M1803T showed that it encodes five proteins representing five glycoside hydrolase families from the above list: GH5, GH10, GH18, GH62, and GH141. In this context, the goal of the present work was to verify experimentally the ability of H. borealis M1803T to grow on xylan and to identify the most likely candidates for the role of β-xylanase among the proteins encoded in its genome.
MATERIALS AND METHODS
Strain M1803T and its growth on xylan. The study was conducted with the planctomycete H. borealis M1803T (=KCTC 25269T) isolated and described previously by our research group (Dedysh et al., 2021). The culture was maintained on the MPYVG medium (modification of DSMZ no. 621) that contained (g/L distilled water): peptone (Fluka), 0.1; yeast extract, 0.25; NH4NO3, 0.1; glucose, 0.5; Hutner’s basal salts, 20 mL, and Hutner’s vitamin solution, 1 mL (Staley et al., 1992). To assess the ability of strain M1803T to grow on xylan, it was cultured at 22°С in 160-mL serum bottles with 30 mL liquid nutrient medium containing 20 mL/L of Hutner’s basal salts, 0.1 g/L NH4-NO3, 1 mL/L vitamin solution, and 0.05 g/L yeast extract. Xylan (Sigma-Aldrich) was added to the medium at the concentration of 0.1%. At the same time, the growth of an M1803T culture was assessed in a similar medium with glucose (0.05%) and in a control medium containing no carbon source. Experiments were performed in three replicates. To determine the cell numbers, suspension aliquots were collected at the beginning of the experiment and after incubation for 10 and 20 days. Planctomycete cells were counted by phase contrast microscopy on Teflon-laminated slides coated with 0.1% gelatin solution. Bacterial abundance in the samples was determined by counting cells in 50 fields of vision for each experimental variant and calculating cell numbers per 1 mL of culture.
Search for putative β-xylanases and their phylogenetic analysis. Using the CAZy database (Drula et al., 2022), the genome of H. borealis M1803T was searched for the presence of genes that encode proteins from glycoside hydrolase families known to include experimentally characterized enzymes with the β-xylanase activity. The closest homologs of the identified proteins were searched for using the blastp algorithm on the NCBI site (http://www.ncbi.nlm.nih.gov/) and subsequently analyzed using the PSI Protein Classifier program (Naumoff and Carreras, 2009). Lists of experimentally characterized enzymes for the analyzed families of glycoside hydrolases were compiled based on the information available in the CAZy database.
To perform phylogenetic analysis, several dozens of closest homologs were selected for each of the candidate proteins from H. borealis M1803T. Multiple alignment of amino acid sequences was performed manually with BioEdit (http://www.mbio.ncsu.edu/ BioEdit/bioedit.html) taking into account the results of pairwise comparisons obtained with blastp.
The outgroup used to study phylogeny of the GH62 family included the closest to IPV69_21845 (H. borealis) experimentally characterized enzyme (GenPept, DAC80243.1) and its 26 nearest homologs. For the GH10 family, the outgroup included nine proteins with the closest phylogenetic relationship to the planctomycete protein cluster (according to Naumoff et al., 2014), as well as additional 12 closest homologs. No special outgroup was selected for the GH141 family; to visualize the tree, the most divergent group was compiled of the proteins included in the analysis.
The results of multiple sequence alignment (after removal of the most variable sequence fragments) were used to construct phylogenetic trees with the PROTPARS program (Protein Sequence Parsimony method, MP) of the PHYLIP software package (http://evolution.gs.washington.edu/phylip.html). Statistical significance of branching nodes was evaluated using bootstrap analysis (1000 pseudoreplicates for each tree). Graphic representations of the trees were obtained using TreeView Win32 (http://taxonomy.zoology.gla.ac.uk/rod/treeview.html).
RESULTS AND DISCUSSION
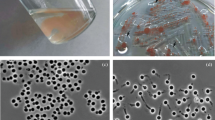
Growth of Humisphaera borealis M1803T on xylan. Microscopy of H. borealis M1803T cultures after 20 days of incubation in bottles with xylan revealed active development of planctomycetes on the surface of biopolymer particles (Fig. 1a). After 20 days of incubation with xylan, the number of M1803T cells increased by more than an order of magnitude, whereas no significant change in the cell number was observed in the control culture (Fig. 1b). The specific growth rate for the strain M1803T on xylan was 0.006 h–1. As expected, the growth on glucose was more rapid.
(a) Micrograph of Humisphaera borealis M1803T cells after 20 days of incubation with xylan as a growth substrate. Scale bar, 10 μm. (b) Dynamics of cell numbers in strain M1803T cultures growing on xylan (1), glucose (2), and in control (3) at the beginning of the experiment and after 10 and 20 days of incubation.
Glycoside hydrolase families GH5 and GH18. According to the CAZy database (Drula et al., 2022), protein IPV69_13055 (GenPept, QOV92224.1) from GH5 family of glycoside hydrolases belongs to GH5_19 subfamily, which includes two experimentally characterized enzymes. One of them (QFQ13828.1) is an exo-mannanase (EC 3.2.1.100), and the other one (AEH51033.1) is a β-mannosidase (EC 3.2.1.25). Endo-β-1,4-xylanase activity (EC 3.2.1.8) was experimentally demonstrated for 16 GH5 proteins that belong to five other subfamilies: GH5_2, GH5_4, GH5_21, GH5_25, and GH5_35. According to the phylogenetic tree of GH5 family published by Aspeborg et al. (2012), GH5_4 and GH5_25 subfamilies form a common monophyletic cluster, and so do GH5_21 and GH5_35. GH5_2 and GH5_26 subfamilies are close to the latter of these clusters. However, GH5_19 subfamily lies far from both of them and forms a common cluster with GH5_18 and GH5_42 subfamilies. All five experimentally characterized members of GH5_18 subfamily (AAN25133.1, ACU71175.1, AEE72695.1, AEW98905.1, and EEI79871.1) exhibit β-mannosidase activity (EC 3.2.1.25), and none of the proteins of GH5_42 subfamily has been studied experimentally (Drula et al., 2022). Based on these data, it can be supposed that glycosidase IPV69_13055 should be able to cleave some substrates containing β-D-mannose residues but does not possess β-xylanase activity.
According to the CAZy database (Drula et al., 2022), among 486 experimentally characterized enzymes of GH18 family of glycoside hydrolases, there is only one β-xylanase, which belongs to the fungus Purpureocillium lilacinum (GenPept, XP_018173934.1). Its comparison to protein IPV69_11765 (QOV91983.1) showed that their amino acid sequences had only 26% identity, which means that they belong to different subfamilies (according to the 30% criterion; Naumoff, 2005). This suggests that IPV69_11765 glycosidase does not possess β-xylanase activity.
Glycoside hydrolase family GH62. Together with GH43 and GH117 families, glycoside hydrolase family GH62 forms the GH-F clan (Drula et al., 2022). On a higher hierarchical level, it belongs to the furanosidase superfamily (Naumoff, 2001, 2011, 2012; Pons et al., 2004), characteristically featuring catalytic domains with the five-bladed β-propeller structure and three highly conservative acidic amino acid residues (Asp/Glu) that form the active center. It should be noted that GH43 glycosidases are predominantly characterized by two types of enzymatic activity: α-L-arabinofuranosidase (EC 3.2.1.55) and β-xylosidase (EC 3.2.1.37). Both types of activity are frequently exhibited by members of the same subfamily, and sometimes even by one and the same protein (Mewis et al., 2016). These facts suggest that β-propeller catalytic domains can recognize both α-L-arabinofuranoside and β-D-xylopyranoside monosaccharide residues of their substrates. A vast majority of experimentally characterized members of GH62 family possess only α-L-arabinofuranosidase activity (Drula et al., 2022). However, two multidomain proteins exhibit several types of activity. In particular, a metagenomic glycosidase (GenPept, DAC80243.1) contains three catalytic domains that belong to the families CE1 (carbohydrate esterase with feruloyl-esterase activity, EC 3.1.1.73), GH62 (α-L-arabinofuranosidase), and GH10 (β-xylanase). Furthermore, subcloning and subsequent expression of its isolated GH62 domain showed that it also has a certain level of β-xylanase activity (Holck et al., 2019). Glycosidase from Ruminiclostridium josui (WP_024831741.1) contains two catalytic domains that belong to the families of GH62 glycoside hydrolases and CE6 carbohydrate esterases. Their subcloning showed that one of them mediated α-L-arabinofuranosidase and endo-β-xylanase activities, and the other one acted as an acetylxylan esterase (EC 3.1.1.72) (Wang et al., 2018). That is, GH62 domains of at least two known proteins possess the β‑xylanase activity. Protein IPV69_21845 (QOV88842.1) from H. borealis contains only one domain, which represents GH62 family of glycoside hydrolases. A comparison of this protein to all known experimentally characterized members of this family showed that it had 45% sequence identity to the metagenomic glycosidase DAC80243.1 and only 30–39% identity to the other 24 proteins. Based on these data, IPV69_21845 glycosidase can be expected to possess not only arabinofuranosidase but also endo-β-xylanase activity, which makes it a likely candidate for the role of β-xylanase in H. borealis.
We performed a phylogenetic analysis of the amino acid sequences of the GH62 domains of putative glycosidase IPV69_21845 and metagenomic glycosidase DAC80243.1, as well as of their closest homologs (altogether, 103 proteins) (Fig. 2). All proteins included in the analysis were found to feature the triad of catalytically important amino acid residues (in case of IPV69_21845, Asp-64, Asp-179, and Glu-227), which indicates that they possess a glycosidase activity. The closest homologs of the metagenomic glycosidase are a vast group of proteins encoded in the genomes of various planctomycetes. Moreover, two proteins have a very high level of sequence identity to its GH62 domain fragment (positions 406–714): NLK41752.1 (100%) and HDS84024.1 (98%). This invites a hypothesis that the polyfunctional metagenomic glycosidase DAC80243.1 also belongs to some planctomycete of the class Phycisphaerae.
Phylogenetic tree of the GH62 family of glycoside hydrolases constructed using the maximum parsimony method. Statistical significance of the tree was assessed by bootstrap analysis; the number of supporting pseudoreplicates (out of 1000) is indicated next to each branching site. Letters indicate the taxonomic affiliation of the source organisms: A, Armatimonadetes; B, Bacteroidetes; C, Chloroflexi; D, Deltaproteobacteria; E, Eukaryota (Alveolata); I, Acidobacteria; K, candidate division KSB1; M, bacterial metagenome; N, Candidatus Nealsonbacteria; O, Candidatus Omnitrophica; P, Planctomycetes; R, Candidatus Poribacteria; and V, Verrucomicrobia. Protein IPV69_21845 (QOV88842.1) from Humisphaera borealis M1803T and the experimentally characterized bifunctional β-xylanase/α-L-arabinofuranosidase (DAC80243.1) encoded in a bacterial anaerobic digester metagenome are shown in red. Strain names are indicated for IPV69_21845 protein and its two closest homologs. A blue triangle indicates the cluster (94.9% bootstrap support) including 22 proteins from Armatimonadetes (MBB6053333.1), Planctomycetes (HFG32237.1, HFJ73202.1, HGS95713.1, MBM4035677.1, and MBM4043100.1), and Verrucomicrobia (HFG26915.1, HGF95708.1, HGS23301.1, HGT44975.1, MBB5032842.1, MBE2282078.1, MBE7495167.1, MBI2929915.1, MBI4660194.1, MBI5383639.1, MBI5690282.1, MBL9117542.1, MBL9177803.1, MBL9182578.1, MBN8420223.1, and MBP8261673.1).
Putative glycosidase IPV69_21845 from H. borealis and its closest homolog from an uncharacterized planctomycete SH-PL14 (AMV21057.1) are single-domain proteins, and their amino acid sequences have 61% identity. They belong to a large cluster that mainly includes proteins from planctomycetes and a significantly lower number of proteins from Armatimonadetes and Verrucomicrobia (Fig. 2); other bacterial phyla are represented by very few members (Acidobacteria, Chloroflexi, Proteobacteria, and other) or are not represented at all (e.g., Actinobacteria, Bacteroidetes, and Firmicutes). The results of the phylogenetic analysis (Fig. 2) suggest that the last common ancestor of the genes encoding all GH62 proteins studied belonged to some planctomycete. Subsequent evolution of this gene involved multiple lateral transfers into genomes of other bacterial phyla and even reverse transfers from Verrucomicrobia back into planctomycetes (one or two cases in the cluster marked in blue; Fig. 2).
Glycoside hydrolase family GH10. Almost all enzymatically characterized members of GH10 family possess endo-β-xylanase activity (EC 3.2.1.8 and EC 3.2.1.32). In our previous work, a comparative analysis of planctomycete proteins of this family showed that they could be divided in two principal groups (Naumoff et al., 2014; Naumoff, 2014, 2016). Proteins of the first group were present only in some planctomycetes, usually as several paralogs, and in the phylogenetic tree they did not form a separate cluster among proteins from other prokaryotes, which is a typical pattern for many glycoside hydrolase families (if members of one given high-rank bacterial taxon are considered). Proteins of the other group were represented in all heterotrophic planctomycetes, always as only one paralog, and they formed a single cluster in the phylogenetic tree where their positions were consistent with the taxonomy of the respective bacterial species (i.e., the gene inheritance was strictly vertical without lateral transfers, deletions, or duplications). Multiple alignment of amino acid sequences showed that all proteins of this group had a disrupted active center that lacked the two key Glu residues acting as a proton donor and a nucleophile (see Fig. S1 in Rakitin et al., 2021); therefore, it is impossible for them to exhibit any enzymatic activity. For this reason, we indicated the lack of enzymatic activity for the corresponding proteins when annotating the genomes of the planctomycetes Paludisphaera borealis (GenPept, APW63099.1), Limnoglobus roseus (QEL20001.1), Frigoriglobus tundricola (QJX00392.1), and Telmatocola sphagniphila (QVL33104.1). Disrupted active centers were also detected in a number of other prokaryotic GH10 proteins (unpublished data), but they were not in a close evolutionary relationship to the proteins of the planctomycete cluster. This fact suggests that the loss of enzymatic function occurred repeatedly in the course of evolution of this protein family.
In the present work, a phylogenetic analysis of the amino acid sequences of GH10 domains (altogether, 181 proteins; Fig. 3) provided more detailed information on the structure of the planctomycete cluster, which corresponds to the second group of GH10 proteins. It is composed of two subclusters corresponding to the classes Phycisphaerae (49.1% bootstrap support) and Planctomycetia (99.9%). Furthermore, the subcluster Planctomycetia contains very few proteins from other bacterial taxa (see caption to Fig. 3) and is in turn composed of four clusters corresponding to the orders Gemmatales (82.5%), Isosphaerales (99.9%), Pirellulales (93.7%), and Planctomycetales (87.4%). Subcluster Phycisphaerae includes several metagenomic proteins annotated as belonging to other bacterial phyla: Acidobacteria (MBE3132059.1), Actinobacteria (NIA06888.1), Chloroflexi (NIA21059.1), Cyanobacteria (KAA0216956.1 and MBC6952665.1), Gemmatimonadetes (TFG86988.1), and Proteobacteria (KPJ65876.1), as well as Candidatus Dependentiae (MBN2267663.1). This can be explained either by multiple lateral gene transfer events or by imperfection of algorithms used for annotation of metagenomes of the class Phycisphaerae. Similarly to the other proteins of the planctomycete cluster, protein IPV69_18965 (QOV88313.1) from H. borealis lacks both catalytically important Glu residues, which means it can be ruled out as a candidate for the role of β-xylanase in this organism.
Phylogenetic tree of the GH10 family of glycoside hydrolases constructed using the maximum parsimony method. Statistical significance of the tree was assessed by bootstrap analysis; the number of supporting pseudoreplicates (out of 1000) is indicated next to each branching site. Letters indicate the taxonomic affiliation of the source organisms: C, Chloroflexi; D, Gammaproteobacteria; G, Gemmatimonadetes; I, Acidobacteria; J, Candidatus Dependentiae; M, bacterial metagenome; T, Actinobacteria; and Y, Cyanobacteria. All unmarked proteins belong to planctomycetes of the class Phycisphaerae. Protein IPV69_18965 (QOV88313.1) from Humisphaera borealis M1803T is shown in red. Strain names are indicated for protein IPV69_18965 and four its closest homologs. Yellow triangles indicate the clusters corresponding to the orders Gemmatales, Isosphaerales, Pirellulales, and Planctomycetales. The bootstrap support of each cluster is given inside the triangle, and the number of proteins it includes is indicated to the right. The cluster Pirellulales (marked with an asterisk) additionally includes a protein from an aquatic metagenome annotated as Candidatus Nealsonbacteria (NQU20226.1). The outgroup (not shown) includes 21 proteins representing five prokaryotic phyla: Crenarchaeota (RLF19374.1), Cyanobacteria (ABW28526.1, ACK64415.1, ACU99283.1, ADN16869.1, NHC56590.1, and NJN38475.1), Firmicutes (AGC67515.1, EEQ57087.1, EGN35770.1, EMS72420.1, HIR57335.1, HIS30199.1, HIS30200.1, HIS30214.1, MBQ1860455.1, MBQ3079900.1, MBQ6240474.1, and WP_148410326.1), Spirochaetes (AEF81843.1), and Verrucomicrobia (HGY63858.1).
Glycoside hydrolase family GH141. According to the CAZy database (Drula et al., 2022), only two proteins of GH141 family have been characterized enzymatically: α-L-fucosidase (EC 3.2.1.51) from Bacteroides thetaiotaomicron (GenPept, AAO76109.1) and β-xylanase (EC 3.2.1.8) from Hungateiclostridium thermocellum (ABN53397.1). These proteins belong to different subfamilies but both feature two Asp residues in homologous positions; these residues acting as proton donors and nucleophiles are highly conserved across GH141 family (Heinze et al., 2017; Ndeh et al., 2017). Protein IPV69_20890 (QOV88669.1) from H. borealis does not belong to either of these subfamilies and has less than 27% amino acid sequence identity with the above proteins, but it possesses both conserved Asp residues (positions 489 and 525). This makes it another possible candidate for the role of β‑xylanase in this species.
We performed a phylogenetic analysis of the amino acid sequences of the putative glycosidase IPV69_20890 and its 109 closest homologs (Fig. 4). Both enzymatically characterized GH141 proteins were not included in the analysis, because their considerable divergence made it impossible to obtain an unambiguous multiple sequence alignment. Protein IPV69_20890 was placed in a stable cluster composed mainly of proteins belonging to bacteria of the phylum Armatimonadetes (100% bootstrap support; top right in Fig. 4). The fact that proteins representing other phyla (Lentisphaerae and Planctomycetes) did not exhibit a trend to co-clusterization within this cluster suggests that the corresponding genes might have appeared due to a series of independent lateral transfers from Armatimonadetes. The remaining GH141 members included in the analysis formed eight more clusters in the tree. One of them was formed exclusively from planctomycete proteins, mainly representing the genus Paludisphaera (100% bootstrap support; top left in Fig. 4). All other clusters (I–VII) also included a certain number of planctomycete proteins. Cluster IV included a stable subcluster (100% bootstrap support) of seven planctomycete proteins (MBC8871159.1, MBM4047734.1, NLF73417.1, NLY00979.1, NOX55446.1, NOZ20755.1, and NUQ63536.1); however, all these proteins are encoded by metagenomes, and for this reason it was difficult to determine more accurately the taxonomic affiliation of their species of origin.
Phylogenetic tree of the GH141 family of glycoside hydrolases constructed using the maximum parsimony method. Statistical significance of the tree was assessed by bootstrap analysis; the number of supporting pseudoreplicates (out of 1000) is indicated next to each branching site. Letters indicate the taxonomic affiliation of the source organisms: A, Armatimonadetes; L, Lentisphaerae; N, Candidatus Nealsonbacteria; P, Planctomycetes; V, Verrucomicrobia; W, Candidatus Hydrogenedentes; X, Kiritimatiellaeota; and Z, Nitrospirae. Protein IPV69_20890 (QOV88669.1) from Humisphaera borealis M1803T is shown in red. Species names are indicated for IPV69_20890, as well as for five proteins of the Paludisphaera cluster (see text). Seven yellow triangles indicate protein clusters I–VII. The bootstrap support of the each cluster is given within the corresponding triangle, and the number of proteins it contains and the taxonomic affiliation of the species are indicated next to it.
Five of these seven clusters (II–V, and VII) included several proteins of the phylum Lentisphaerae; Armatimonadetes were represented in four clusters (II, III, VI, and VII), and so were Verrucomicrobia (II, IV, V, and VII); Kiritimatiellaeota were found in three clusters (II, VI, and VII). Three bacterial phyla were represented in only one cluster each: Nitrospirae (I), Candidatus Hydrogenedentes (I), and Candidatus Nealsonbacteria (V). These facts indicate that the last common ancestor of the genes encoding all GH141 proteins studied in this work must have belonged to some planctomycete. Subsequent evolution of this gene involved multiple lateral transfers into bacterial genomes of several other phyla and even a reverse transfer from Armatimonadetes back into planctomycetes.
The Paludisphaera cluster included a protein from Paludisphaera borealis PX4T (APW58834.1), which was initially annotated as “putative beta-solenoid-type carbohydrate-active enzyme (GH, PL, or CE) of unknown function” based on remote evolutionary relationships (family PF13229 in the Pfam database; Mistry et al., 2021), since the GH141 family of glycoside hydrolases was not recognized at that time (Ivanova et al., 2017). All proteins of this family analyzed in our work (with an only exception of protein HFH06191.1 from Candidatus Hydrogenedentes; cluster I) possess both highly conserved Asp residues, which suggests that all of them probably exhibit some enzymatic activity.
Evolution of glycoside hydrolases of the families studied. Glycoside hydrolases are among those proteins that perform functions of facultative significance and therefore are not obligate components of the proteome (none of the glycoside hydrolase families is present in most biological species, and some microorganisms do not possess them at all). Vertical inheritance is not typical for prokaryotic genes of glycoside hydrolases: they are easily acquired by lateral transfer and subsequent duplications and also rapidly lost as soon as their presence provides no selective advantages (Naumoff, 2011).
In this aspect, the group of planctomycete proteins within family GH10 analyzed in our work represents a highly unusual case (Fig. 3), which requires an explanation. Apparently, these proteins lack enzymatic activity due to the loss of the active center. However, this group of orthologs must have acquired a currently unidentified novel function that is obviously obligate for bacteria of this phylum, which underlies the consistent vertical inheritance of their genes.
Two other groups of proteins studied, those representing GH62 and GH141 families, exhibited the pattern of evolution typical for glycoside hydrolases and characterized by numerous events of lateral gene transfer among different bacterial phyla. Apparently, for both families, the transfer events originated in planctomycetes and were directed exclusively to other gram-negative bacteria: predominantly members of the phyla Armatimonadetes and Verrucomicrobia in case of GH62 family (Fig. 2); and members of Armatimonadetes, Lentisphaerae, and Verrucomicrobia in case of GH141 family (Fig. 4).
It should be mentioned that the two other families studied, GH5 and GH18, also followed the general pattern; however, in this case, planctomycetes more frequently acted as acceptors of laterally transferred genes. The closest homologs of protein IPV69_13055 from GH5 family belonged to bacteria of the phylum Chloroflexi, and the closest homologs of protein IPV69_11765 (GH18) predominantly belonged to Verrucomicrobia (data not shown).
On the whole, the data obtained in this work are consistent with our previous observations suggesting that lateral transfer of glycoside hydrolase genes among various groups of prokaryotes occurred in nonrandom directions (Naumoff and Dedysh, 2018). In particular, gene exchange between gram-negative and gram-positive bacteria is a relatively rare event, and Proteobacteria are separated from other members of the gram-negative group. The highest intensity of transfers was observed, on the one hand, within a group of phyla including Armatimonadetes, Planctomycetes, and Verrucomicrobia, and on the other hand, among members of the phyla Acidobacteria, Bacteroidetes, and Balneolaeota.
The results of our work indicate that proteins IPV69_21845 (family GH62) and IPV69_20890 (GH141) are the most probable candidates for the role of β-xylanase in Humisphaera borealis M1803T. Protein IPV69_21845 can be expected to exhibit both α‑L-arabinofuranosidase and β-xylanase activity, whereas IPV69_20890 probably possesses either α-L-fucosidase or β-xylanase activity. Further investigation of their characteristics is of substantial interest for biochemical research and can provide deeper insights into the functional role that planctomycetes of the class Phycisphaerae play in degradation of plant debris in natural ecosystems.
REFERENCES
Aspeborg, H., Coutinho, P.M., Wang, Y., Brumer, H. III, and Henrissat, B., Evolution, substrate specificity and subfamily classification of glycoside hydrolase family 5 (GH5), BMC Evol. Biol., 2012, vol. 12, art. 186.
Beg, Q.K., Kapoor, M., Mahajan, L., and Hoondal, G.S., Microbial xylanases and their industrial applications: a review, Appl. Microbiol. Biotechnol., 2001, vol. 56. pp. 326–338.
Capetti, C.C.M., Vacilotto, M.M., Dabul, A.N.G., Sepulchro, A.G.V., Pellegrini, V.O.A., and Polikarpov, I., Recent advances in the enzymatic production and applications of xylooligosaccharides, World J. Microbiol. Biotechnol., 2021, vol. 37, art. 169.
Dedysh, S.N., Beletsky, A.V., Ivanova, A.A., Kulichev-ska-ya, I.S., Suzina, N.E., Philippov, D.A., Rakitin, A.L., Mardanov, A.V., and Ravin, N.V., Wide distribution of Phycisphaera-like planctomycetes from WD2101 soil group in peatlands and genome analysis of the first cultivated representative, Environ. Microbiol., 2021, vol. 23, pp. 1510–1526.
Dedysh, S.N. and Ivanova, A.A., Planctomycetes in boreal and subarctic wetlands: diversity patterns and potential ecological functions, FEMS Microbiol. Ecol., 2019, vol. 95, art. fiy227.
Drula, E., Garron, M.-L., Dogan, S., Lombard, V., Henrissat, B., and Terrapon, N., The carbohydrate-active enzyme database: functions and literature, Nucleic Acids Res., 2022, vol. 50 (Database issue), pp. D571–D577. http://www.cazy.org/.
Fukunaga, Y., Kurahashi, M., Sakiyama, Y., Ohuchi, M., Yokota, A., and Harayama, S., Phycisphaera mikurensis gen. nov., sp. nov., isolated from a marine alga, and proposal of Phycisphaeraceae fam. nov., Phycisphaerales ord. nov. and Phycisphaerae classis nov. in the phylum Planctomycetes, J. Gen. Appl. Microbiol., 2009, vol. 55, pp. 267–275.
Garron, M.-L. and Henrissat, B., The continuing expansion of CAZymes and their families, Curr. Opin. Chem. Bio-l., 2019, vol. 53, pp. 82–87.
Gupta, G.K., Dixit, M., Kapoor, R.K., and Shukla, P., Xylanolytic enzymes in pulp and paper industry: new technologies and perspectives, Mol. Biotechnol., 2022, vol. 64, pp. 130–143.
Heinze, S., Mechelke, M., Kornberger, P., Liebl, W., Schwarz, W.H., and Zverlov, V.V., Identification of endoxylanase XynE from Clostridium thermocellum as the first xylanase of glycoside hydrolase family GH141, Sci. Rep., 2017, vol. 7, art. 11178.
Holck, J., Djajadi, D.T., Brask, J., Pilgaard, B., Krogh, K.B.R.M., Meyer, A.S., Lange, L., and Wilkens, C., Novel xylanolytic triple domain enzyme targeted at feruloylated arabinoxylan degradation, Enzyme Microb. Technol., 2019, vol. 129, art. 109353.
Ivanova, A.A., Kulichevskaya, I.S., Merkel, A.Y., Toshchakov, S.V., and Dedysh, S.N., High diversity of Planctomycetes in soils of two lichen-dominated sub-arctic ecosystems of northwestern Siberia, Front. Microbiol., 2016, vol. 7, art. 2065.
Ivanova, A.A., Naumoff, D.G., Miroshnikov, K.K., Liesack, W., and Dedysh, S.N., Comparative genomics of four Isosphaeraceae planctomycetes: a common pool of plasmids and glycoside hydrolase genes shared by Paludisphaera borealis PX4T, Isosphaera pallida IS1BT, Singulisphaera acidiphila DSM 18658T, and strain SH-PL62, Front. Microbiol., 2017, vol. 8, art. 412.
Kovaleva, O.L., Merkel, A.Yu., Novikov, A.A., Baslerov, R.V., Toshchakov, S.V., and Bonch-Osmolov-skaya, E.A., Tepidisphaera mucosa gen. nov., sp. nov., a moderately thermophilic member of the class Phycisphaerae in the phylum Planctomycetes, and proposal of a new family, Tepidisphaeraceae fam. nov., and a new order, Tepidisphaerales ord. nov., Int. J. Syst. Evol. Microbiol., 2015, vol. 65, pp. 549–555.
Kulichevskaya, I.S., Naumoff, D.G., Miroshnikov, K.K., Ivanova, A.A., Philippov, D.A., Hakobyan, A., Rijp-stra, W.I.C., Damsté, J.S.S., Liesack, W., and De-dysh, S.N., Limnoglobus roseus gen. nov., sp. nov., a novel freshwater planctomycete with a giant genome from the family Gemmataceae, Int. J. Syst. Evol. Microbiol., 2020, vol. 70, pp. 1240–1249.
Kulkarni, N., Shendye, A., and Rao, M., Molecular and biotechnological aspects of xylanases, FEMS Microbiol. Rev., 1999, vol. 23, pp. 411–456.
Linares-Pasten, J.A., Aronsson, A., and Karlsson, E.N., Structural considerations on the use of endo-xylanases for the production of prebiotic xylooligosaccharides from biomass, Curr. Protein Pept. Sci., 2018, vol. 19, pp. 48–67.
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P.M., and Henrissat, B., The carbohydrate-active enzymes database (CAZy) in 2013, Nucleic Acids Res., 2014, vol. 42 (Database issue), pp. D490–D495.
McDonald, A.G., Boyce, S., and Tipton, K.F., ExplorEnz: the primary source of the IUBMB enzyme list, Nucleic Acids Res., 2009, vol. 37 (Database issue), pp. D593–D597.
Mewis, K., Lenfant, N., Lombard, V., and Henrissat, B., Dividing the large glycoside hydrolase family 43 into subfamilies: a motivation for detailed enzyme characterization, Appl. Environ. Microbiol., 2016, vol. 82, pp. 1686–1692.
Mistry, J., Chuguransky, S., Williams, L., Qureshi, M., Salazar, G.A., Sonnhammer, E.L.L., Tosatto, S.C.E., Paladin, L., Raj, S., Richardson, L.J., Finn, R.D., and Bateman, A., Pfam: the protein families database in 2021, Nucleic Acids Res., 2021, vol. 49 (Database issue), pp. D412–D419.
Naumoff, D.G., β-Fructosidase superfamily: homology with some α-L-arabinases and β-D-xylosidases, Proteins, 2001, vol. 42, pp. 66–76.
Naumoff, D.G., GH97 is a new family of glycoside hydrolases, which is related to the α-galactosidase superfamily, BMC Genomics, 2005, vol. 6, art. 112.
Naumoff, D.G., Hierarchical classification of glycoside hydrolases, Biochemistry (Moscow), 2011, vol. 76, pp. 622–635.
Naumoff, D.G., Furanosidase superfamily: search of homologues, Mol. Biol. (Moscow), 2012, vol. 46, pp. 322–327.
Naumoff, D.G., Bioinformatic analysis of endo-β-xylanases from Planctomycetes, Proc. 9th Int. Conf. on the Bioinformatics of Genome Regulation and Structure\Systems Biology, Novosibirsk, Russia, 2014, p. 112. http://conf.bionet.nsc. ru/bgrssb2016/wp-content/uploads/sites/2/2015/11/BGRS2014.pdf.
Naumoff, D.G., GH10 family of glycoside hydrolases: structure and evolutionary connections, Mol. Biol. (Moscow), 2016, vol. 50, pp. 132–140.
Naumoff, D.G. and Carreras, M., PSI protein classifier: a new program automating PSI-BLAST search results, Mol. Biol. (Moscow), 2009, vol. 43, pp. 652–664.
Naumoff, D.G. and Dedysh, S.N., Bacteria from poorly studied phyla as a potential source of new enzymes: β-galactosidases from Planctomycetes and Verrucomicrobia, Microbiology (Moscow), 2018, vol. 87, pp. 796–805.
Naumoff, D.G., Ivanova, A.A., and Dedysh, S.N., Phylogeny of β-xylanases from Planctomycetes, Mol. Biol. (Moscow), 2014, vol. 48, pp. 439–447.
Nguyen, S.T.C., Freund, H.L., Kasanjian, J., and Berlemont, R., Function, distribution, and annotation of characterized cellulases, xylanases, and chitinases from CAZy, Ap-pl. Microbiol. Biotechnol., 2018, vol. 102, pp. 1629–1637.
Nordberg Karlsson, E., Schmitz, E., Linares-Pastén, J.A., and Adlercreutz, P., Endo-xylanases as tools for production of substituted xylooligosaccharides with prebiotic properties, Appl. Microbiol. Biotechnol., 2018, vol. 102, pp. 9081–9088.
Pollet, A., Delcour, J.A., and Courtin, C.M., Structural determinants of the substrate specificities of xylanases from different glycoside hydrolase families, Crit. Rev. Biotechnol., 2010, vol. 30, pp. 176–191.
Pons, T., Naumoff, D.G., Martínez-Fleites, C., and Hernández, L., Three acidic residues are at the active site of a β-propeller architecture in glycoside hydrolase families 32, 43, 62, and 68, Proteins, 2004, vol. 54, pp. 424–432.
Rakitin, A.L., Naumoff, D.G., Beletsky, A.V., Kulichevskaya, I.S., Mardanov, A.V., Ravin, N.V., and Dedysh, S.N., Complete genome sequence of the cellulolytic planctomycete Telmatocola sphagniphila SP2T and characterization of the first cellulolytic enzyme from planctomycetes, Syst. Appl. Microbiol., 2021, vol. 44, art. 126276.
Spring, S., Bunk, B., Spröer, C., Rohde, M., and Klenk, H.P., Genome biology of a novel lineage of planctomycetes widespread in anoxic aquatic environments, Environ. Microbiol., 2018, vol. 20, pp. 2438–2455.
Staley, J.T., Fuerst, J.A., Giovannoni, S., and Schles-ner, H., The Order Planctomycetales and the Genera Planctomyces, Pirellula, Gemmata, and Isosphaera, in The Prokaryotes: A Handbook on the Biology of Bacteria: Ecophysiology, Isolation, Identification, Applications, Balows, A., Trüper, H.G., Dworkin, M., Harder, W., and Schleifer, K.-H., Eds., New York: Springer New York, 1992, pp. 3710–3731.
Terrapon, N., Lombard, V., Drula, E., Coutinho, P.M., and Henrissat, B., The CAZy database/the Carbohydrate-Active Enzyme (CAZy) database: principles and usage guidelines, in A Practical Guide to Using Glycomics Databases, Aoki-Kinoshita, K.F., Ed., Tokyo: Springer, 2017, Ch. 6, pp. 117–131.
Wang, Y., Sakka, M., Yagi, H., Kaneko, S., Katsuzaki, H., Kunitake, E., Kimura, T., and Sakka, K., Ruminiclostridium josui Abf62A-Axe6A: a tri-functional xylanolytic enzyme exhibiting α-L-arabinofuranosidase, endoxylanase, and acetylxylan esterase activities, Enzyme Microb. Technol., 2018, vol. 117, pp. 1–8.
Ward, N.L. and Dedysh, S.N., Planctomycetes, in Bergey’s Manual of Systematics of Archaea and Bacteria, Trujillo, M.E., Dedysh, S., DeVos, P., Hedlund, B., Kämpfer, P., Rainey, F.A., and Whitman, W.B., Eds., 2022. https://doi.org/10.1002/9781118960608.pbm00021.pub2
Funding
This work was supported by the Ministry of Science and Higher Education of the Russian Federation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
Conflict of interests. The authors declare that they have no conflict of interest.
Statement on the welfare of animals. This article does not contain any studies involving animals or human participants performed by any of the authors.
AUTHORS’ CONTRIBUTIONS
The idea of the study was proposed by S.N.D. Experiments were designed by S.N.D. and D.G.N. Planctomycete cultures and cell count were performed by I.S.K. Identification of putative β-xylanase genes and their phylogenetic analysis were conducted by D.G.N. The manuscript was prepared by D.G.N. and S.N.D. All authors participated in the discussion of the results.
Additional information
Translated by D. Timchenko
Rights and permissions
Open Access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Naumoff, D.G., Kulichevskaya, I.S. & Dedysh, S.N. Genetic Determinants of Xylan Utilization in Humisphaera borealis M1803T, a Planctomycete of the Class Phycisphaerae. Microbiology 91, 249–258 (2022). https://doi.org/10.1134/S002626172230004X
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S002626172230004X