Abstract—
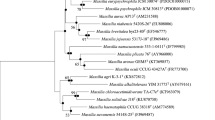
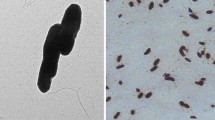
Two strains were isolated, Achromobacter sp. NE1 (GenBank MW132988) and Brevundimonas sp. 242 (GenBank MW132989), which possessed a unique ability to use lindane and the intermediates of its a-erobic biotransformation, 1,2,4-trichlorobenzene (strain NE1) and 2,5-dichlorophenol (strain 242) as a sole source of carbon and energy. The strains were found to contain the linA gene encoding dehydrochlorinase, the first enzyme of lindane degradation, exhibiting 99.65–100% sequence identity to the linA genes of Sphingobium degrader strains.


Similar content being viewed by others
REFERENCES
Camacho-Pérez, B., Ríos-Leal, E., Rinderknecht-Seijas, N., and Poggi-Varaldo, M., Enzymes involved in the biodegradation of hexachlorocyclohexane: a mini review, J. Environ. Manag., 2012, vol. 95, pp. S306–S318.
Dong, W.H., Zhang, P., Lin, X.Y., Zhang, Y., and Tabouré, A., Natural attenuation of 1,2,4-trichlorobenzene in shallow aquifer at the Luhuagang’s landfill site, Kaifeng, China, Sci. Total Environ., 2015, vol. 505, pp. 216–222.
Egorova, D.O., Gorbunova, T.I., Pervova, M.G., Kir’yanova, T.D., Demakov, V.A., Saloutin, V.I., and C-hupakhin, O.N., Biodegradability of hydroxylated derivatives of commercial polychlorobiphenyls mixtures by Rhodococcus-strains, J. Hazard. Mater., 2020, vol. 400, 123328.
Fang, L., Qin, H., Shi, T., Wu, X., Li, Q.X., and Hua, R., Ortho and para oxydehalogenation of dihalophenols catalyzed by the monooxygenase TcpA and NAD(P)H:FAD reductase Fre, J. Hazard. Mater., 2020, vol. 388, 121787.
Kumar, D. and Pannu, R., Perspectives of lindane (γ-hexachlorocyclohexane) biodegradation from the environment: a review, Bioresour. Bioprocs., 2018, vol. 5, p. 29.
Lal, R., Dogra, C., Malhotra, S., Sharma, P., and Pal, R., Diversity, distribution and divergence of lin genes in hexachlorocyclohexane-degrading sphingomonads, Trends Biotechnol., 2006, vol. 24, pp. 121–130.
Lal, R., Pandey, G., Sharma, P., Kumari, K., Malhotra, S., Pandey, R., Raina, V., Kohler, H.P., Holliger, C., Jackson, C., and Oakeshott, J.G., Biochemistry of microbial degradation of hexachlorocyclohexane and prospects for bioremediation, Microbiol. Mol. Biol. Rev., 2010, vol. 74, pp. 58–80.
Manual of Methods for General Bacteriology, Gerhardt, P., Murray, R.G.E., Costilow, R.N., Nester, E.W., Wood, W.A., Krieg, N.R., and Phillips, G.B., Eds., Washington: Amer. Soc. Microbiol., 1981.
Methods for General and Molecular Bacteriology, Gerhardt, P., Murray, R.G.E., Wood, W.A., and Krieg, N.R., Eds., Washington, D.C.: Amer. Soc. M-icrobiol., 1994.
Pannu, R. and Kumar, D., Process optimization of γ-hexachlorocyclohexane degradation using three novel Bacillus sp. strains, Biocat. Agric. Biothech., 2017, vol. 11, pp. 97–107.
Saez, J.M., García, V.C., and Benimeli, C.S., Improvenment of lindane removal by Streptomyces sp. M7 by using stable microemulsions, Ecotoxicol. Environ. Saf., 2017, vol. 144, pp. 351–359.
Song, Y., Wang, F., Kengara, F.O., Bian, Y.-R., Yang, X.-L., Liu, C.-Y., and Jiang, X., Improved biodegradation of 1,2,4-trichlorobenzene by adapted microorganisms in agricultural soil and in soil suspension cultures, Pedosphere, 2011, vol. 21, pp. 423–431.
Thomas, J.-C., Berger, F., Jacquier, M., Bernillon, D., Baud-Grasset, F., Truffaut, N., Norm, P., Vogel, T.M., and Simonet, P., Isolation and characterization of a novel γ-hexachlorocyclohexane-degrading bacterium, J. Bacteriol., 1996, vol. 178, pp. 6049–6055.
Weisburg, W.G., Barns, S.M., Pelletier, D.A., and Lane, D.J., 16S Ribosomal DNA amplification for phylogenetic study, J. Bacteriol., 1991, vol. 173, pp. 697–703.
Zaitsev, G.M., Tsoi, T.V., Grishenkov, V.G., Plotnikova, E.G., and Boronin, A.M., Genetic control of degradation of chlorinated benzoic acids in Arthrobacter globiformis, Corynebacterium sepedonicum and Pseudomonas cepacia strains, FEMS Microbiol. Lett., 1991, vol. 81, pp. 171–176.
Zheng, G., Selvam, A., and Wong, J.W.C., Rapid degradation of lindane (γ-hexachlorocyclohexane) at low temperature by Sphingobium strains, Int. Biodeterior. Biodegrad., 2011, vol. 65, pp. 612–618.
ACKNOWLEDGMENTS
We are grateful to research workers of the Molecular Genetic Laboratory of the Department of Botany and Plant Genetics, Perm State National Research University, as well as the Research of Materials and Substances Center for Collective Use, Perm Federal Research Center, for valuable assistance provided in using their equipment.
Funding
The work was supported by the R&D “Molecular Mechanisms of Adaptation of Microorganisms to Environmental Factors,” project no. AAAA-A19-119112290009-1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests. The authors declare that they have no conflict of interest.
Statement on the welfare of animals. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by E. Babchenko
Rights and permissions
About this article
Cite this article
Egorova, D.O., Nazarova, E.A. & Demakov, V.A. New Lindane-Degrading Strains Achromobacter sp. NE1 and Brevundimonas sp. 242. Microbiology 90, 392–396 (2021). https://doi.org/10.1134/S0026261721030036
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261721030036