Abstract
The successful determination of reliable protein interaction networks (PINs) in several species in the post-genomic era has hitherto facilitated the quest to understanding systems and structural properties of such networks. It is envisaged that a clearer understanding of their intrinsic topological properties would elucidate evolutionary and biological topography of organisms. This, in turn, may inform the understanding of diseases' aetiology. By analysing sub-networks that are induced in various layers identified by zones defined as distance from central proteins, we show that zones of human PINs display self-similarity patterns. What is observed at a global level is repeated at lower levels of inducement. Furthermore, it is observed that these levels of strength point to refinement and specialisations in these layers. This may point to the fact that various levels of representations in the self-similarity phenomenon offer a way of measuring and distinguishing the importance of proteins in the network. To consolidate our findings, we have also considered a gene co-expression network and a class of gene regulatory networks in the same framework. In all cases, the phenomenon is significantly evident. In particular, the truly unbiased regulatory networks show finer level of articulation of self-similarity.
Similar content being viewed by others
Introduction
Recently, self-repeating phenomena has been observed in remarkably many systems, both natural as well as man made. What piques man's interest in them is often their aesthetic value more than their organising principles. In particular, long-range power-law correlations depicting self-similarities have been discovered in a remarkably wide variety of systems1. There have been attempts to identify self-similarity phenomenon in biological complex systems2,3 through some kind of re-normalisation. For instance, in biology the observation of the self-similarity phenomenon has been observed in surface areas and vesicular distributions of tissues4,5.
In respect of self-similarity of the general complex systems to which biological networks belong, the work of Song et al6 is seminal. They analysed a variety of real complex networks and found that these systems consist of self-repeating patterns. This result was achieved by the application of a re-normalisation procedure that coarse-grains the system into boxes containing nodes within a given neighbourhood size. They identified a power-law relation between the number of boxes needed to cover the network and the size of the box, defining a finite self-similar exponent. In the precise terminology of graph theory, they found out that quotients of complex networks defined by covering neighbourhoods of certain distances were also power-law. Others have used variations of the method with some notable improvements7,8.
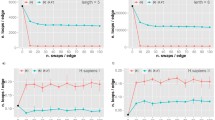
However, it is not surprising that coarse-grain self-similarity was weak in PINs. It has been shown that the majority of nodes (over 90% in all cases that have been considered) lie within 3 distances away from the centre9. It is therefore not surprising that any coarse-graining beyond 2 distances from the centre would completely destroy the intrinsic power-law behaviour of the system. Coarse-graining requires that the network has a reasonable diameter and nodes are reasonably spread around the centre.
We have, on the other hand, looked at power-law properties of networks from a different perspective; incomparable to the seminal work of Song et al. As has been shown elsewhere, PINs display a certain recognisable structure9, which for brevity, we call the stingray structure with quills in this sequel. This structure has been both our point of departure and our focus. We contend that PINs are self-repeating from the stingray structural point of view.
There has been an intense and deliberate effort to determine PINs of many organisms with notable successes. The determination of these networks is to help uncover the generic organising principles of functional cellular networks10,11,12,13,14,15,16,17,18,19. This progress is an important step in our understanding of the evolution and behaviour of such systems.
It is envisaged that an understanding of the organizing principles at systems level of biological networks would elucidate many of the perplexing questions including that of finding therapeutic targets20,21,22. Such effort is under way in many fronts. Whilst this has been the general aim, much of the recent effort has focused on finding functional dependencies amongst the so-called hubs and their topological importance and positions in the network23,24.
There has been serious undertaking to understand both structural and functional systems level of protein-protein interaction (PPI) networks through graph visualisation and drawing. The most important piece of information that is required in visualisation is spatial distribution of the network. Yet, such information is calculable if networks are treated as metric spaces. Recently it has been shown that, treated as metric spaces, PINs of various organisms are what we have coined, as alluded to, a stingray structure with quills. That is, proteins with high degree coagulate in the centre of the network whilst those in the periphery have low degree and in the fringes we have nodes of single degrees9.
Further, in that sequel it was shown that the observed stingray structure has significant biological implications. Amongst others, it was observed that proteins involved in sensing pathways tend to be more expressed in central zones and those in the periphery specialise in routine metabolic pathways. Second, it was observed that some zones are uniquely-enriched and represent a far more pronounced specialisation. Third, it was shown that cancer pathways are significantly over represented in zone 225.
In this article, we have analysed substructures that are defined by zones from the centre. In other words, we have statistically visualised the human PINs at both global as well as at subsystems level. What has been revealed is as startling as is aesthetic. These substructures display the same phenomenon that is played out on a global scale. The core of human PINs are imposing self-similarity structures. The systems structures and the ensuing organising principles of these human PINs are repeated at macro as well as at lower levels. In other words, if one would appreciate the beauty of the structure and considered it as a flower with many petals; these very petals would also have petals, which would have more petals of the same kind. Moreover, in most cases, central proteins of various levels from human PINs are from same families, playing the same biological role possibly at every level of consideration. This repetition in similarity of centres is observed in gene regulatory networks, albeit with a finer level of articulation26.
When pathway and function enrichment analysis are applied to various layers of the induced subgraphs, our results show that there is reinforcement and refinement of these phenomena in various levels of consideration. Moreover, it is clear that there is increased strength in specialisation. Overall, therefore, this self-similarity phenomenon offer a natural way to understanding the biological systems mechanics of the human PINs.
As molecular networks may be biased, we also tested our method and hypothesis on truly unbiased networks such as gene co-expression network and transcriptional regulatory networks. Both cases strongly support the case; and in the case of regulatory network, it is even more pronounced than in PINs.
In other words, we propose that at the core of human PINs, proteins assemble in the same manner of coagulation as systems structures at all levels defined by distance throughout a given network. The key organizing features of the central zones of human PINs are repeated at the level of induced subgraphs defined by distances from the centre. Proteins interact in the same manner, varying only in scale and refinement of functionality. This recurrence may point to another way of identifying important proteins that may have utility as target drugs.
Results
The general structure of the human PINs
We modeled the human functional protein interaction network (HFPIN)27, which consists of 9448 nodes and 181706 interactions and the highly curated and currently largest available human signaling network (HSN)28,29, which consists of 6305 nodes and 62937 interactions. We also looked at the combination of both HFPIN and HSN and produced what we have called the combined human network (CHN), which consists of 10573 nodes and 210689 interactions. Also, a new human protein interaction set based on three-dimensional information with other functional tools has recently been predicted (NHPIS)30, which consists of 7863 nodes and 23779 interactions. It was equally subjugated to our method. We have also modelled truly unbiased datasets: gene co-expression31 and regulatory networks26.
We used a formal method that finds the protein(s) that has the smallest maximal distance to other proteins in the network. The starting point is that in all the networks under consideration, the centres were identified and nodes were grouped (in zones) according to the distances they are from these central proteins. With this classification, functional enrichment was performed and biological hypotheses were drawn9,25.
Here, we follow the same approach in our consideration of subgraphs of the networks we consider. Before we present self-similarity we are alluding to, let us first summarize the pertinent features of the structure in all the biological networks that were considered. We will argue that the same pattern is evident in induced subgraphs of these networks, determined by distances from central nodes.
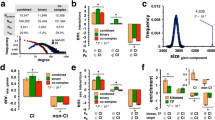
The essence of the structure is in the following manner. First, the centres consist of single nodes, all heavily involved in signalling pathway9. As for the HFPIN, the centre is MAPK14 and that of the HSN the centre is MAPK1. The combined human network has MAPK3 as the centre. Second, nodes in the central positions have higher degrees than those in the periphery. Moreover, degrees distribution is power law. The third feature is that while the diameters are generally large, the majority of proteins are located in the central positions (zone 1 to zone 3). Fourthly, proteins in the periphery are of low degree. They display the quill structure (node with degree 1) in the fringes of the network. To aid in visualising these networks, we have called these imposing structures stingray structures with quills.
The structures of the HFPIN, HSN, CHN and the NHPIS are summarized in Tables 1 to 234.
Central zones of human PINs as induced subgraphs repeat the structure that is observed by the whole network
The key feature of self-similarity is the self-repeating patterns at various levels of consideration. In our case, we reveal that all the networks we dealt with splits into smaller parts that resemble the whole from a structural point of view of graphs. We split the graphs into parts that are defined by the zones from the centre, i.e., we look at the graphs induced by nodes that are zone i from the centre, where i is 1, 2 and 3. We examine their structure as was done in the global graphs, following closely what was done in our recent work9. We show that the structures we observe have similar patterns. What is striking is that centres of these substructures have similar functions and belong to the same families.
When we now examine the repeating substructures of the giant graphs, in all cases, the induced subgraphs of zones 1 and 2, there is a single node for the centres, which are from the same family of the centres of the human PINs. In the first zones, they are from MAPK family, both in the HFPIN and the HSN. In zone 2, the centres of the induced subgraphs of the HFPIN are from the RAS family; those of HSN are from the general kinase family.
The next natural consideration was to look at zones formed from the zones in the first instant to describe the self-similarity phenomenon. We considered a subset of proteins that form a particular zone and their interactions amongst themselves as a separate induced subgraph. Again, the same phenomenon was observed with varying degree of connectivity and expressed level of manifestation of this organizing principle, depending of the distance of the zone from the centre.
In all the induced subgraphs, we observed the same organizing principles. Nodes with high degree coagulate in central positions and those with low degree are in the periphery of the graphs. Of particular importance, the degree distribution of proteins in these induced subgraphs follow similar patterns (see supplementary figures S1 to S7). The centre of the whole graph is MAPK14 for the HFPIN and MAPK1 for the HSN. As for the HFPIN, at the centre of the induced subgraph of nodes in the first zone is MAPK3. When one considers the zone 1 nodes at MAPK3, the centre is MAPK1 of which its zone 1 subgraph has centre MAPK11 (table 1). In which case, we repeatedly look at induced subgraphs of induced subgraphs. While the level of expression may weaken as we consider the induced subgraphs of these subgraphs, the centres at zones 1 all belong to the MAPK family, a critical family of proteins in signalling. The same is observed for the HSN (table 2).
It is not particularly surprising that, considering that the combined human network has more data, the features of the self-similarity is more pronounced (table 3).
This repeatedness is also observed in zones 2 of the human PINs. Centres are from KRAS family for HFPIN and AKT for the HSN respectively (tables 1 and 2). Both of these families are heavily implicated in cancer pathways32,33.
Biological ramifications of the self-similarity structure in the HFPIN and similar networks
It has recently been observed that there is some level of specialization by proteins in various zones of the HFPIN25. Also, while some pathways cut across zones, of importance is that sensing pathways are far more pronounced in central zones than in periphery. Zones in the periphery tend to be involved in gene expression and metabolic pathways more than those in the centre. In addition, it was also observed that zone 2 bear the significant burnt of pathways involved in cancers.
It is therefore natural that we understand how this phenomenon is played out from the point of view of the self-repeating topology we have alluded to in this article in biological terms. What is made clear is that there seem to be some level of strengthening in terms of pathways.
Four issues are worthy noting. First, the fact that some zones have uniquely-enriched pathways is a clear indication that in those zones, there is a strong representation of proteins that are associated with such pathways. Consider for instance the TRAF6 Mediated Induction of Proin-flammatory cytokines pathway, which is uniquely-enriched in zone 1 in the entirety of the network in the HFPIN. In zone 1 of the induced subgraph of zone 1, as a percentage of proteins involved in this pathway, there is an increase to 20.5% from 10.4%. In the second layer, (zone 1 of zone 1 of zone 1), the percentage incereases to 26.2%. In the next level, it increases to 28.1%. This points to the fact that as one moves into deeper levels, one sees that there is a coagulation of proteins that are highly specialised in specific pathways (table 5).
Second, this phenomenon of strengthening is not restricted to uniquely-enriched pathways. Consider the top 4 pathways in zone 1: signal transduction (38.1%), immune system (31.3%), MAPK (26.6%), pathways in cancer (22%). In the third level of consideration (zone 1 of zone 1 of zone 1), the order changes: immune system (55.3%), signal transduction (52%), MAPK (48.5%), pathway in cancer (31%). By the time the next level is considered, the MAPK signalling pathway dominates, with 54.6% (table 5).
Third, some pathways are more highly represented in the periphery of central zones. For instance, it is interesting to note that signal transduction has an ebbing effect as one moves deeper into central zones of central zones; it still leads in zone 2 of induced subgraph of zone 1. In zone 2 of zone 1 of the induced subgraph, the percentage of proteins involved in signal transduction is highest with 52.5% of proteins involved in this pathway (table 6).
Finally, while it was noted that zones in periphery have a tendency to diversify in metabolic functions, it is important to note that such pathways are ubiquitous. However, there are more enriched in periphery of zones of central zones. Consider for instance, gene expression, metabolism and membrane trafficking. In the induced subgraph of zone 1, the gene expression pathway is uniquely-enriched in zone 2, whilst in the induced subgraph of zone 2, it is significant in zones 2. In the induced subgraph of zone 3, it is the main theme of central zones.
These observations are equally evident in the HSN, CHN and NHPIS (see supplementary tables S1 to S6).
In summary, therefore, we see that the self-repeating structure is played out even from the biological point of view. Signalling pathways continue to be significant in central zones; routine metabolic pathways are significant in the periphery of the network, at all levels of consideration. However, the consideration of the self-repeating structure renders specialisation even more prominent: there are cases where pathways are highly distinguished or uniquely-enriched. Using the self-similarity structure, it is possible to group proteins in some order of importance, a theme we discuss below.
Cancer pathways' zonal distribution in self-similarity terms
In our recent work when we considered the distribution of proteins that consistently expressed in 13 types of cancer25, it was shown that most of these proteins are prominent in zone 2 of the HFPIN, HSN and CHN (tables 7 to 89). Here, the same methods were applied as we analysed each of the subgraphs from each zone. While on the whole network, cancer proteins are in zone 2, the critical compartment is zone 3 of zone 2 for the HFPIN (table 10) and zone 2 of zone 2 for both HSN and CHN referred to in Tables 11 and 12.
Distinguishing proteins using the self-similarity edifice
It is generally accepted that the degree of the node is a strong indicator of the importance and/or essentiality of the protein in the network23,24. As one looks at various layers of zones, central zones of central zones tend to have higher degree in the entirety of the network than the other zones. For instance, proteins from zone 1 of zone 1 in HFPIN have an average degree of 118 and that of zone 1 of zone 2 is 85 (table 1).
It has also been shown that, in general, both sensing pathways and proteins implicated in diseases tend to be pronounced in central positions25. While there is some disagreements about what is more important between sensing pathways and metabolic ones, we contend that sensing pathways are more important as they are likely to elicit a metabolic response to facilitate homeostasis.
In view of the foregoing, we propose that proteins in zone 1 have a higher weighting than those in zone 2 and so on. So, for instance, nodes in zone 3 of zone 1 would have more weight than those in zone 1 of zone 2.
Self-similarity in other biological networks
Both gene co-expression and regulatory networks show stingray structures. When gene co-expression network is subjugated to sub-structure analysis, the majority of the induced subgraphs have single centres. However, as we delve further, we do not obtain single centres. Also, that centres are from the same family cannot fully be established (table 13).
However, the gene regulatory networks we looked at, despite that the networks have small orders, show a much more pronounced articulation of the phenomenon (see supplementary tables S7 to S10).
Methods
Evaluation of biological networks as metric spaces
We considered human PINs (HFPIN, HSN, NHPIS) and gene co-expression and regulatory networks as metric spaces by defining the usual graph theoretic distance between nodes of a graph. Using a python wrapper around the C++ BOOST graph library (http://www.boost.org/), we used the Dijkstra algorithm to compute the shortest distances between all pairs of nodes and then identifyied the node or all nodes whose greatest distance to other nodes is/are smallest. This is the network center(s). From here, nodes were classified according to their distances from the centre and divided into zones based on distance from the topological centre(s). From each distance class, we calculated their degree distributions and also considered their connectivity of the graphs induced for each zone.
Pathway and function enrichment analysis
In order to determine whether zones of the human PINs we considered have biological significance, we divided proteins into subsets based on their distance from the true topological centre. Protein sets representing each zone were then subjected to a pathway over-representation analysis in order to determine whether the zones were specialised for specific functions. The Comparative Toxigenomics Databases Gene Set Enricher web service (http://ctdbase.org/tools/enricher.go and Gene Ontology enrichment (http://geneontology.org/page/go-enrichment-analysis) was used to perform the enrichment analysis and a corrected P-value of 0.01 was chosen as a statistical significance cutoff. Lastly, when such enrichment was observed, we calculated the proportion of proteins involved in each enriched pathway as a way to assess whether any zone displayed functional specialization.
Cancer gene expression data sources
We considered gene expression absence/presence calls from the following cancers types: breast, lung, kidney, pancreas, liver, cervix, ovary, glioblastoma, pituitary, glioma, fallopian, endometrium and rectum, which was downloaded from Gene Expression Barcode database (http://barcode.luhs.org/index.php?page=genesexp). Genes expressed in at least 99% of samples of a cancer of interest based on the Human HGU133 platform were downloaded. Gene expression was used as a proxy for protein expression and was mapped onto the PINs of interest in order to identify the zones in which gene product is located in.
Testing the difference between proportions
We performed a z-test for the difference between two population proportions p1 and p2. We identified the null and alternative hypotheses and we specified the level of significance to be P < 0.01. After that we determined the critical value(s) from the statistic table. Finally we found the standardized test statistic as shown below.
Statistical significance of the proportional analysis of pathway representation of zones
To test differences between proportions among zones, we need a statistical comparison of observed differences. A two-sample z-test for the differences between proportions for the top statistically enriched REACTOME pathways among zones was conducted. We defined the null hypothesis H0 to be: classification proportions of zones in the periphery in human PINs have as high proportion significance as zones closest to the centre, i.e the accuracy of the sensing functions in zones closest to the centre and the accuracy of metabolic functions in zones in the periphery. If the P < 0.01, we rejected H0 and concluded that the proportions support our claim that zones closest to the centre have high proportion significance than the zones in the periphery. In the other words, we have enough evidence at the 1% level to conclude that zones closest to the centre have high proportion significance than the zones in the periphery.
References
Havlin, S. et al. Fractals in biology and medicine. Chaos, Solitons and Fractals 6, 171–201 (1995).
Tao, S. & Zhang, Y. Self-similarity formed of complex networks. In: Circuits, Communications and Systems, 2009. PACCS'09. Pacific-Asia Conference on, 155–158 (IEEE, 2009).
Serrano, M. A., Krioukov, D. & Boguná, M. Self-similarity of complex networks and hidden metric spaces. Phy. Rev. Lett. 100, 078701 (2008).
Avnir, D., Farin, D. & Pfeifer, P. Molecular fractal surfaces. Nature 308, 261–263 (1984).
Huet, S. et al. Relevance and limitations of crowding, fractal and polymer models to describe nuclear architecture. International Review of Cell and Molecular Bio 307, 443–79 (2014).
Song, C., Havlin, S. & Makse, H. A. Self-similarity of complex networks. Nature 433, 392–395 (2005).
Zhou, W.-X., Jiang, Z.-Q. & Sornette, D. Exploring self-similarity of complex cellular networks: The edge-covering method with simulated annealing and log-periodic sampling. Physica A: Statistical Mechanics and its Applications 375, 741–752 (2007).
Gallos, L. K., Song, C. & Makse, H. A. A review of fractality and self-similarity in complex networks. Physica A: Statistical Mechanics and its Applications 386, 686–691 (2007).
Fadhal, E., Gamieldien, J. & Mwambene, E. C. Protein interaction networks as metric spaces: a novel perspective on distribution of hubs. BMC Sys Bio 8, 6 (2014).
Hartwell, L. H., Hopfield, J. J., Leibler, S. & Murray, A. W. From molecular to modular cell biology. Nature 402, C47–C52 (1999).
Ouzounis, C. A. & Karp, P. D. Global properties of the metabolic map of escherichia coli. Genome research 10, 568–576 (2000).
McAdams, H. H. & Arkin, A. Gene regulation: Towards a circuit engineering discipline. Current Bio 10, R318–R320 (2000).
Elowitz, M. B. & Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 403, 335–338 (2000).
Savageau, M. A. Design principles for elementary gene circuits: Elements, methods and examples. Chaos: An Interdisciplinary Journal of Nonlinear Science 11, 142–159 (2001).
Bolouri, H. & Davidson, E. H. Modeling transcriptional regulatory networks. BioEssays 24, 1118–1129 (2002).
Hasty, J., McMillen, D., Isaacs, F. & Collins, J. J. Computational studies of gene regulatory networks: in numero molecular biology. Nature Reviews Genetics 2, 268–279 (2001).
Guet, C. C., Elowitz, M. B., Hsing, W. & Leibler, S. Combinatorial synthesis of genetic networks. Science 296, 1466–1470 (2002).
Newman, M. E. The structure and function of complex networks. SIAM review 45, 167–256 (2003).
Spirin, V. & Mirny, L. A. Protein complexes and functional modules in molecular networks. Proceedings of the National Academy of Sciences 100, 12123–12128 (2003).
Fry, D. C. Protein–protein interactions as targets for small molecule drug discovery. Peptide Science 84, 535–552 (2006).
White, A. W., Westwell, A. D. & Brahemi, G. Protein–protein interactions as targets for small-molecule therapeutics in cancer. Expert reviews in molecular medicine 10, e8 (2008).
Strosberg, A. D. Protein–protein interactions as targets for novel therapeutics. Drug Discov (2007).
He, X. & Zhang, J. Why do hubs tend to be essential in protein networks? PLoS Genetics 2, e88 (2006).
Jeong, H., Mason, S., Barabasi, A. & Oltvai, Z. Lethality and centrality in protein networks. Arxiv preprint cond-mat/0105306 (2001).
Fadhal, E., Mwambene, E. C. & Gamieldien, J. Modeling human protein interaction networks as metric spaces has potential in disease research and drug target discovery. BMC Sys Bio 8, 68 (2014).
Neph, S. et al. Circuitry and dynamics of human transcription factor regulatory networks. Cell 150, 1274–1286 (2012).
Wu, G., Feng, X. & Stein, L. A human functional protein interaction network and its application to cancer data analysis. Genome Bio 11, R53 (2010).
Li, L. et al. The human phosphotyrosine signaling network: evolution and hotspots of hijacking in cancer. Genome research 22, 1222–1230 (2012).
Awan, A. et al. Regulatory network motifs and hotspots of cancer genes in a mammalian cellular signalling network. IET Sys Bio 1, 292–297 (2007).
Zhang, Q. C. et al. Structure-based prediction of protein-protein interactions on a genomewide scale. Nature 490, 556–560 (2012).
Obayashi, T. et al. Atted-ii: a database of co-expressed genes and cis elements for identifying co-regulated gene groups in arabidopsis. Nucleic acids research 35, D863–D869 (2007).
Staal, S. P. Molecular cloning of the akt oncogene and its human homologues akt1 and akt2: amplification of akt1 in a primary human gastric adenocarcinoma. Proceedings of the National Academy of Sciences 84, 5034–5037 (1987).
Amado, R. G. et al. Wild-type kras is required for panitumumab efficacy in patients with metastatic colorectal cancer. J. of Clinical Oncology 26, 1626–1634 (2008).
Acknowledgements
This work was supported in part by a grant from South Africa National Research Foundation (NRF).
Author information
Authors and Affiliations
Contributions
E.F. implemented the algorithms, performed the analyses and drafted the original manuscript. E.C.M. proposed the concept of analyzing PINs as a self-similarity structure and oversaw the topological and statistical analyses. J.G. designed and oversaw and assisted in the functional evaluation tests and the biological interpretation of the results. E.C.M. and J.G. supervised the study and edited the manuscript. All authors have read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Fadhal, E., Gamieldien, J. & Mwambene, E. Self-similarity of human protein interaction networks: a novel strategy of distinguishing proteins. Sci Rep 5, 7628 (2015). https://doi.org/10.1038/srep07628
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07628
- Springer Nature Limited