Abstract
Recurrent gene fusions between the genes TMPRSS2 and ERG have been described in prostate cancer (PCa) and are found in 27% to 79% of radical prostatectomy. This fusion transcription results in ERG overexpression, which can be detected by immunohistochemistry (IHC) and provide a potential diagnostic marker for PCa. Three tissue microarrays (TMAs) containing samples from 98 patients with PCa and one TMA of 27 samples from individuals without PCa were tested for ERG immunostaining and the presence of TMPRSS2:ERG transcripts was confirmed by quantitative real time PCR (qRT-PCR). The results showed that 46.9% of tumors tested positive for ERG immunostaining and this finding was consistent with the results of qRT-PCR testing (k = 0.694, p < 0.001). IHC had a specificity of 83.3% and a sensitivity of 81% in detecting TMPRSS2:ERG fusion. Patients with PSA < 4.0 ng/mL showed positive immunoreactivity for ERG (p = 0.031). Kaplan-Meier analysis suggested that ERG expression did not influence the time of biochemical recurrence. This study demonstrates that both IHC and qRT-PCR are useful tools in detecting TMPRSS2:ERG fusions. A correlation between ERG expression and clinical and pathological parameters was not found, but the frequency, specificity and recurrence of ERG in PCa suggests that it may be a potential adjunct diagnostic tool.
Similar content being viewed by others
Introduction
Prostate cancer (PCa) remains the most common non-cutaneous cancer and the most common cause of cancer-related death among men in the USA, with 29 720 deaths estimated for 20131,2. In Brazil, there were 12 778 registered deaths from PCa in 2010 and an estimated 60 180 new cases in 20123.
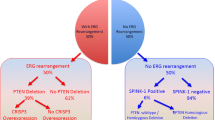
In 2005, genomic rearrangement between androgen-regulated transmembrane protease (TMPRSS2, 21q22.3) and v-ets erythroblastosis virus E26 oncogene homolog (ERG, 21q22.2) was described as the most common genetic alteration in PCa cells, occurring in approximately 50% of cases4. During chromosomal rearrangement, one TMPRSS2 promoter allele is lost and one ERG allele gain it, which results in the overexpression of ERG protein in tumor cells5.
This fusion can be detected by fluorescence in situ hybridization (FISH) across deletions or chromosomal translocations6,7, real time PCR (qRT-PCR) across fusion transcripts8,9 and immunohistochemistry by ERG protein overexpression10,11,12.
TMPRSS2 promoter analysis revealed the presence of a non-canonical androgen receptor (AR) as a CIS-regulatory target of AR action13. Mutations, amplifications or overexpression of the AR and deletions or inactivation of the tumor suppressor gene PTEN (phosphatase and tensin homolog) are frequently identified in PCa cells14,15. ERG gene rearrangements and PTEN loss often occur concomitantly in PCa cells and promote tumor progression through the PI3K pathway, which increases the advantages of pre-malignant cells compared to normal cells16,17.
This study aimed to evaluate the presence of ERG protein overexpression and TMPRSS2:ERG gene fusion in a cohort of patients with PCa who have undergone radical prostatectomy or radiotherapy and to determine whether there is a correlation between these events. This analysis is particularly interesting in Brazilian patients, where there is a lack of studies evaluating gene fusion in PCa and the population is ethnically heterogeneous, which is a consequence of centuries of miscegenation among Europeans, Africans and Amerindians.
Methods
Study population
The study was designed in accordance with the Guidelines and Standards Regulation Research Involving Human Beings (Resolution 196/1996 of the National Health Council) and was approved by the Ethics Committee of Barretos Cancer Hospital, SP, Brazil (425/2010). In this Institution is requested of all patients completing an informed consent at the first appointment, getting attached to records.
We evaluated 98 patients who underwent radical prostatectomy at the Barretos Cancer Hospital between 01/2006 and 12/2007. Additionally, 27 samples containing cells negative for PCa were evaluated. A review of patient medical records provided the following information: age at the time of treatment, race, pre-treatment PSA level, Gleason score, clinical stage, D'Amico progression risk group, surgical margin status, perineural infiltration of the tumor and time to PSA recurrence or follow-up time after treatment. Biochemical recurrence of PSA was defined as two consecutive PSA tests with levels greater than or equal to 0.2 ng.mL−1 for patients who underwent radical prostatectomy, with follow-up tests occurring for 5 to 6 years. By evaluating PSA levels at the time of treatment, Gleason scores of patient biopsies and clinical stages of PCa, patients were given a low, intermediate, or high risk D'Amico classification18. None of the patients in this study were treated with neoadjuvant therapy.
Tissue microarray (TMA) construction
The pathologist reviewed hematoxylin and eosin stained sections from each patient and identified the most representative regions of tumor samples. To construct a TMA, two representative cores 1 mm in diameter were taken from tissue samples and arrayed on an individual paraffin block. A total of four TMAs were constructed using an MTA-1 Manual Tissue Arrayer (Beecher Instruments, Silver Spring, MD, USA); three of the TMAs contained PCa samples and one contained non-PCa samples.
Evaluation of ERG protein expression by immunohistochemistry
One 4 μm section of each sample was prepared for antibody testing. Immunohistochemistry staining was performed at room temperature using an Autostainer Link 48 (Dako, Copenhagen, DK). Slides were incubated for 20 minutes with Flex Ready-to-Use Monoclonal Rabbit Anti-Human ERG Clone EP111 (Dako, Copenhagen, DK). Slides were incubated with secondary antibody Envision Flex + Mouse ™ (LINKER) (Dako) for 15 minutes. Staining was performed using 3,3′-diaminobenzidine (DAB+, Dako) and the samples were counterstained with Harris' hematoxylin.
Immunohistochemical evaluation
Immunoreactions were evaluated semiquantitatively using the criteria previously described19,20. The immunoreaction extent was scored semiquantitatively as follows: 0: 0% presence of immunoreactive cells; 1: <5% presence of immunoreactive cells; 2: 5–50% presence of immunoreactive cells; and 3: >50% presence of immunoreactive cells. In addition, the intensity of staining was scored semiquantitatively as: 0: negative; 1: weak; 2: intermediate; and 3: strong. The final immunoreaction score was defined as the sum of both parameters (extent and intensity) and grouped as negative (score 0 and 1) or positive (3–6).
Immunohistochemistry validation by qRT-PCR
To confirm the findings obtained from immunohistochemistry testing, qRT-PCR was performed in a subset of 45 samples that tested positive or negative for ERG protein expression. Total RNA was extracted from four 10 μm sections of these samples using a RNeasy FFPE kit (Qiagen, Gaithersburg, MD, USA), with 200 ng of RNA converted into cDNA using a SuperScript® VILO™ Master Mix (Invitrogen, Carlsbad, CA, USA), both according to the manufacturer's instructions. We used TaqMan® inventoried assays (Applied Biosystems, Carlsbad, CA, USA) for ERG (Hs03063375_ft) and the endogenous control HPRT1 (hypoxanthine phosphoribosyltransferase 1, Hs99999909_m1). The qRT-PCR reaction was performed using 7900HT System equipment (Applied Biosystems) and a TaqMan® Fast Advanced Master Mix (Applied Biosystems), according to the manufacturer's instructions. To determine the relative expression level, the 2−ΔCT values were calculated.
Statistical analysis
To check the accuracy of immunohistochemistry in determining the presence of TMPRSS2:ERG, gene fusion, sensitivity, sensibility, kappa and ROC curve analyses were performed. The Chi-Square or Fisher exact test was used to compare immunohistochemistry data and clinico-pathological parameters. Joint association of clinical characteristics with ERG expression was performed by multiple logistic regression. A biochemical recurrence-free survival curve was estimated using the Kaplan-Meier method and a Cox regression model was used to identify associated risk factors. For the entire study, a p-value < 0.05 was considered to be statistically significant. All analyses were performed using IBM-SPSS 21.0 (IBM Corporation, New York, NY, USA).
Results
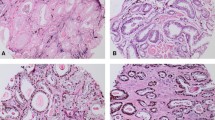
Immunohistochemistry analysis detected ERG protein expression in 46 of the 98 samples (46.9%); all 27 negative controls tested negative for ERG protein. Endothelial tissue usually expresses ERG protein and was used as an internal positive control for all reactions (Figure 1).
The total RNA quality was assessed using a 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA) and the majority of samples had an RNA integrity number (RIN) of 2.4. Of the 45 samples tested, 21 (46.67%) showed amplification and 24 (53.33%) did not show amplification (Figure 2). There was no significant difference in the detection rates between immunohistochemistry and qRT-PCR (k = 0.643, p < 0.001). Accuracy analysis showed that immunohistochemistry had high sensitivity (81.0%) and high specificity (83.3%). Accuracy analysis showed that immunohistochemistry had high sensitivity (81.0%) and high specificity (83.3%) and technique had in common with qRT-PCR 17 positive and 19 negative cases (Figure 3).
Patients who had PSA levels less than or equal to 4.0 ng.mL−1 had positive ERG expression compared with patients with PSA levels greater than or equal to 10.0 ng/mL (p = 0.031, Table 1). There was no statistical association between ERG expression and the other clinico-pathologic parameters. Logistic regression was performed by pre-treatment PSA level, clinical stage and surgical margin status and only PSA levels less than or equal to 4.0 ng.mL−1 (OR 4.37; 95% CI: 1.19–16.04; p = 0.026) and PSA levels between 4.0 and 10.0 ng.mL−1 (OR 3.12; 95% CI: 1.15–8.49; p = 0.026) was statistically significant.
In Kaplan-Meier analysis, ERG expression was not predictive for biochemical recurrence-free survival (Figure 4). Univariate Cox regression was performed by D'Amico score and surgical margin status and only the patients in the intermediate or high-risk D'Amico groups were further likely to experience biochemical recurrence than the low risk group (HR 5.00; 95% CI: 1.51–16.55; p = 0.008, Table 2).
Discussion
This study evaluated the presence and potential prognostic value of the ERG protein in patients with PCa tumors.
Our study of samples from Brazilian patients with PCa found that immunohistochemistry tests were positive for ERG expression in 45.8% of cases, which is consistent with other studies demonstrating a frequency of approximately 50%4,21,22. Detection of TMPRSS2:ERG gene fusion by immunohistochemistry had a sensitivity of 81.0% and specificity of 83.3%. Chaux et al validated ERG immunohistochemistry using FISH and reported a sensitivity and specificity of 86% and 89%, respectively and an area under the ROC curve of 0.87 (p < 0.00001)23. These results corroborate the fact that TMPRSS2:ERG gene fusion, which leads to overexpression of the ERG protein, is the most frequent oncogene in this type of cancer4.
Several studies have attempted to evaluate TMPRSS2:ERG gene fusion as a prognostic indicator of some risk factors, including the Gleason score, PSA level, metastatic PCa or death by PCa and the results have been variable24,25,26,27,28. Our findings showed no relationship between the presence of the ERG protein and a patient's clinical stage, Gleason score or biochemical recurrence, which corroborates the results of other studies24,28,29,30,31. However, we found an increased likelihood of ERG positive immunoreactions when PSA levels were less than 10.0 ng.ml−1 (p = 0.031)31,32. Sun et al concluded, using cell and animal models, that ERG protein overexpression increases the regulation of the c-MYC oncogene. This occurs because ERG is capable of interacting with ETS binding elements in the oncogene promoter region. This c-MYC upregulation blocks the expression of the MSMB, SLC45A3 and PSA genes, which affect prostate cell differentiation by altering epithelial differentiation. This event might explain the association between the presence of ERG protein and lower levels of PSA33.
As PCa appears years after androgen levels peak, hormone exposure for a long period may be necessary for the development of cancer. This relationship also appears to be associated with the appearance of gene fusion. It has been proposed that AR and TMPRSS2:ERG jointly promote the development of invasive adenocarcinoma because the transcription of the gene TMPRSS2 is dependent on androgen24,34,35.
It is still uncertain whether the presence of ERG protein is a predictor of other risk factors, but the frequency and specificity of ERG in PCa make it a potential adjunct tool for the diagnosis of prostate cancer, regardless of the population type.
References
Siegel, R., Naishadham, D. & Jemal, A. Cancer statistics, 2012. CA Cancer J Clin 62, 10–29 (2012).
Siegel, R., Naishadham, D. & Jemal, A. Cancer statistics, 2013. CA Cancer J Clin 63, 11–30 (2013).
INCA. Edn. First. (ed. CEDC. ) (Rio de Janeiro - RJ; 2011).
Tomlins, S. A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005).
Mwamukonda, K. et al. Quantitative expression of TMPRSS2 transcript in prostate tumor cells reflects TMPRSS2-ERG fusion status. Prostate Cancer Prostatic Dis 13, 47–51 (2010).
Yoshimoto, M. et al. Three-color FISH analysis of TMPRSS2/ERG fusions in prostate cancer indicates that genomic microdeletion of chromosome 21 is associated with rearrangement. Neoplasia 8, 465–469 (2006).
Perner, S. et al. TMPRSS2:ERG fusion-associated deletions provide insight into the heterogeneity of prostate cancer. Cancer Res 66, 8337–8341 (2006).
Lu, B. et al. Detection of TMPRSS2-ERG Fusion Gene Expression in Prostate Cancer Specimens by a Novel Assay Using Branched DNA. Urology 74, 1156–1161 (2009).
Tu, J. J. et al. Gene fusions between TMPRSS2 and ETS family genes in prostate cancer: frequency and transcript variant analysis by RT-PCR and FISH on paraffin-embedded tissues. Mod Pathol 20, 921–928 (2007).
Park, K. et al. Antibody-based detection of ERG rearrangement-positive prostate cancer. Neoplasia 12, 590–598 (2010).
Braun, M. et al. ERG protein expression and genomic rearrangement status in primary and metastatic prostate cancer--a comparative study of two monoclonal antibodies. Prostate Cancer Prostatic Dis 15, 165–169 (2012).
Falzarano, S. M. et al. ERG gene rearrangement status in prostate cancer detected by immunohistochemistry. Virchows Arch 459, 441–447 (2011).
Wang, Q. et al. A hierarchical network of transcription factors governs androgen receptor-dependent prostate cancer growth. Mol Cell 27, 380–392 (2007).
Yoshimoto, M. et al. FISH analysis of 107 prostate cancers shows that PTEN genomic deletion is associated with poor clinical outcome. Br J Cancer 97, 678–685 (2007).
Taylor, B. S. et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 18, 11–22 (2010).
Yoshimoto, M. et al. PTEN genomic deletions that characterize aggressive prostate cancer originate close to segmental duplications. Genes Chromosomes Cancer 51, 149–160 (2012).
King, J. C. et al. Cooperativity of TMPRSS2-ERG with PI3-kinase pathway activation in prostate oncogenesis. Nat Genet 41, 524–526 (2009).
D'Amico, A. V. et al. Biochemical outcome after radical prostatectomy, external beam radiation therapy, or interstitial radiation therapy for clinically localized prostate cancer. JAMA 280, 969–974 (1998).
Pinheiro, C. et al. Increased expression of monocarboxylate transporters 1, 2 and 4 in colorectal carcinomas. Virchows Archiv: an international journal of pathology 452, 139–146 (2008).
de Oliveira, A. T. et al. Co-expression of monocarboxylate transporter 1 (MCT1) and its chaperone (CD147) is associated with low survival in patients with gastrointestinal stromal tumors (GISTs). Journal of bioenergetics and biomembranes 44, 171–178 (2012).
Mehra, R. et al. Comprehensive assessment of TMPRSS2 and ETS family gene aberrations in clinically localized prostate cancer. Mod Pathol 20, 538–544 (2007).
Yoshimoto, M. et al. Absence of TMPRSS2:ERG fusions and PTEN losses in prostate cancer is associated with a favorable outcome. Mod Pathol 21, 1451–1460 (2008).
Chaux, A. et al. Immunohistochemistry for ERG expression as a surrogate for TMPRSS2-ERG fusion detection in prostatic adenocarcinomas. Am J Surg Pathol 35, 1014–1020 (2011).
Nam, R. K. et al. Expression of TMPRSS2:ERG gene fusion in prostate cancer cells is an important prognostic factor for cancer progression. Cancer Biol Ther 6, 40–45 (2007).
Demichelis, F. et al. TMPRSS2:ERG gene fusion associated with lethal prostate cancer in a watchful waiting cohort. Oncogene 26, 4596–4599 (2007).
Gopalan, A. et al. TMPRSS2-ERG gene fusion is not associated with outcome in patients treated by prostatectomy. Cancer Res 69, 1400–1406 (2009).
Leinonen, K. A. et al. Association of SPINK1 expression and TMPRSS2:ERG fusion with prognosis in endocrine-treated prostate cancer. Clin Cancer Res 16, 2845–2851 (2010).
Rubio-Briones, J. et al. Clinical implications of TMPRSS2-ERG gene fusion expression in patients with prostate cancer treated with radical prostatectomy. J Urol 183, 2054–2061 (2010).
Miyagi, Y. et al. ETS family-associated gene fusions in Japanese prostate cancer: analysis of 194 radical prostatectomy samples. Mod Pathol 23, 1492–1498 (2010).
Hoogland, A. M. et al. ERG immunohistochemistry is not predictive for PSA recurrence, local recurrence or overall survival after radical prostatectomy for prostate cancer. Mod Pathol 25, 471–479 (2012).
Pettersson, A. et al. The TMPRSS2:ERG rearrangement, ERG expression and prostate cancer outcomes: a cohort study and meta-analysis. Cancer Epidemiol Biomarkers Prev 21, 1497–1509 (2012).
Schaefer, G. et al. Distinct ERG rearrangement prevalence in prostate cancer: higher frequency in young age and in low PSA prostate cancer. Prostate Cancer Prostatic Dis 16, 132–138 (2013).
Sun, C. et al. TMPRSS2-ERG fusion, a common genomic alteration in prostate cancer activates C-MYC and abrogates prostate epithelial differentiation. Oncogene 27, 5348–5353 (2008).
Leshem, O. et al. TMPRSS2/ERG promotes epithelial to mesenchymal transition through the ZEB1/ZEB2 axis in a prostate cancer model. PLoS One 6, e21650 (2011).
Yu, J. et al. An integrated network of androgen receptor, polycomb and TMPRSS2-ERG gene fusions in prostate cancer progression. Cancer Cell 17, 443–454 (2010).
Acknowledgements
The authors thank the Pathological Anatomy Department and Molecular Oncology Research Center of the Barretos Cancer Hospital for technical assistance. This study was funded by the São Paulo State Research Foundation (FAPESP, No. 2011/14934-0, fellow-ship to FCE), the CNPq Brazilian National Research and Development Council (CNPq, No. 473142/2010-4, fellowship to SGPC) and the Department of Teaching and Research, Barretos Cancer Hospital.
Author information
Authors and Affiliations
Contributions
F.C.E. and S.G.P.C. designed the study and wrote the manuscript. F.C.E. collected the data and interpreted it with E.F.F., S.R.T. and C.Z.O. C.S.N. reviewed the histological slides of all patients (histological analyses) and A.L.F. performed the immunohistocehmistry analyses. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Eguchi, F., Faria, E., Neto, C. et al. The role of TMPRSS2:ERG in molecular stratification of PCa and its association with tumor aggressiveness: a study in Brazilian patients. Sci Rep 4, 5640 (2014). https://doi.org/10.1038/srep05640
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05640
- Springer Nature Limited
This article is cited by
-
Profile of chimeric RNAs and TMPRSS2-ERG e2e4 isoform in neuroendocrine prostate cancer
Cell & Bioscience (2022)