Abstract
DNA damage contributes to the aging of hematopoietic stem cells (HSCs), yet the underlying molecular mechanisms are not fully understood. In this study, we identified a heterogeneous functional role of microcephalin (MCPH1) in the nucleus and cytoplasm of mouse HSCs. In the nucleus, MCPH1 maintains genomic stability, whereas in the cytoplasm, it prevents necroptosis by binding with p-RIPK3. Aging triggers MCPH1 translocation from cytosol to nucleus, reducing its cytoplasmic retention and leading to the activation of necroptosis and deterioration of HSC function. Mechanistically, we found that KAT7-mediated lysine acetylation within the NLS motif of MCPH1 in response to DNA damage facilitates its nuclear translocation. Targeted mutation of these lysines inhibits MCPH1 translocation and, consequently, compromises necroptosis. The dysfunction of necroptosis signaling, in turn, improves the function of aged HSCs. In summary, our findings demonstrate that DNA damage-induced redistribution of MCPH1 promotes HSC aging and could have broader implications for aging and aging-related diseases.







Similar content being viewed by others
Data availability
RNA-seq data are deposited in the Gene Expression Omnibus under accession code GSE217101. Proteomics data from mass spectrometry are deposited in the ProteomeXchange Consortium through iProX65 under dataset identifier PXD048580. Further information and requests for resources and reagents should be directed to and will be fulfilled by J.W.
References
de Haan, G. & Lazare, S. S. Aging of hematopoietic stem cells. Blood 131, 479–487 (2018).
Rossi, D. J. et al. Deficiencies in DNA damage repair limit the function of haematopoietic stem cells with age. Nature 447, 725–729 (2007).
Beerman, I., Seita, J., Inlay, M. A., Weissman, I. L. & Rossi, D. J. Quiescent hematopoietic stem cells accumulate DNA damage during aging that is repaired upon entry into cell cycle. Cell Stem Cell 15, 37–50 (2014).
Flach, J. et al. Replication stress is a potent driver of functional decline in ageing haematopoietic stem cells. Nature 512, 198–202 (2014).
Mohrin, M. et al. Hematopoietic stem cell quiescence promotes error-prone DNA repair and mutagenesis. Cell Stem Cell 7, 174–185 (2010).
Schumacher, B., Pothof, J., Vijg, J. & Hoeijmakers, J. H. J. The central role of DNA damage in the ageing process. Nature 592, 695–703 (2021).
McNeely, T., Leone, M., Yanai, H. & Beerman, I. DNA damage in aging, the stem cell perspective. Hum. Genet. 139, 309–331 (2020).
Tiwari, V. & Wilson, D. M. DNA damage and associated DNA repair defects in disease and premature aging. Am. J. Hum. Genet. 105, 237–257 (2019).
Wang, J., Lu, X., Sakk, V., Klein, C. A. & Rudolph, K. L. Senescence and apoptosis block hematopoietic activation of quiescent hematopoietic stem cells with short telomeres. Blood 124, 3237–3240 (2014).
Lin, S.-Y. & Elledge, S. J. Multiple tumor suppressor pathways negatively regulate telomerase. Cell 113, 881–889 (2003).
Jackson, A. P. et al. Identification of microcephalin, a protein implicated in determining the size of the human brain. Am. J. Hum. Genet. 71, 136–142 (2002).
Gruber, R. et al. MCPH1 regulates the neuroprogenitor division mode by coupling the centrosomal cycle with mitotic entry through the Chk1–Cdc25 pathway. Nat. Cell Biol. 13, 1325–1334 (2011).
Liang, Y. et al. BRIT1/MCPH1 is essential for mitotic and meiotic recombination DNA repair and maintaining genomic stability in mice. PLoS Genet. 6, e1000826 (2010).
Lin, S. Y., Rai, R., Li, K., Xu, Z. X. & Elledge, S. J. BRIT1/MCPH1 is a DNA damage responsive protein that regulates the Brca1-Chk1 pathway, implicating checkpoint dysfunction in microcephaly. Proc. Natl Acad. Sci. USA 102, 15105–15109 (2005).
Peng, G. et al. BRIT1/MCPH1 links chromatin remodelling to DNA damage response. Nat. Cell Biol. 11, 865–872 (2009).
Alderton, G. K. et al. Regulation of mitotic entry by microcephalin and its overlap with ATR signalling. Nat. Cell Biol. 8, 725–733 (2006).
Tibelius, A. et al. Microcephalin and pericentrin regulate mitotic entry via centrosome-associated Chk1. J. Cell Biol. 185, 1149–1157 (2009).
Brown, J. A. L., Bourke, E., Liptrot, C., Dockery, P. & Morrison, C. G. MCPH1/BRIT1 limits ionizing radiation-induced centrosome amplification. Oncogene 29, 5537–5544 (2010).
Journiac, N. et al. Cell metabolic alterations due to Mcph1 mutation in microcephaly. Cell Rep. 31, 107506 (2020).
Wood, J. L., Liang, Y., Li, K. & Chen, J. Microcephalin/MCPH1 associates with the condensin II complex to function in homologous recombination repair. J. Biol. Chem. 283, 29586–29592 (2008).
Wu, X. et al. Microcephalin regulates BRCA2 and Rad51-associated DNA double-strand break repair. Cancer Res. 69, 5531–5536 (2009).
Rai, R. et al. BRIT1 regulates early DNA damage response, chromosomal integrity, and cancer. Cancer Cell 10, 145–157 (2006).
Liu, X. et al. The N-terminal BRCT domain determines MCPH1 function in brain development and fertility. Cell Death Dis. 12, 143 (2021).
Jeffers, L. J., Coull, B. J., Stack, S. J. & Morrison, C. G. Distinct BRCT domains in Mcph1/Brit1 mediate ionizing radiation-induced focus formation and centrosomal localization. Oncogene 27, 139–144 (2008).
Xu, X., Lee, J. & Stern, D. F. Microcephalin is a DNA damage response protein involved in regulation of CHK1 and BRCA1. J. Biol. Chem. 279, 34091–34094 (2004).
Ling, J. P. et al. ASCOT identifies key regulators of neuronal subtype-specific splicing. Nat. Commun. 11, 137 (2020).
Liu, X., Zhou, Z. W. & Wang, Z. Q. The DNA damage response molecule MCPH1 in brain development and beyond. Acta Biochim. Biophys. Sin. 48, 678–685 (2016).
Wang, J. et al. A differentiation checkpoint limits hematopoietic stem cell self-renewal in response to DNA damage. Cell 148, 1001–1014 (2012).
Koulnis, M. et al. Identification and analysis of mouse erythroid progenitors using the CD71/TER119 flow-cytometric assay. J. Vis. Exp. https://doi.org/10.3791/2809 (2011).
Liu, M., Li, H., Yang, R., Ji, D. & Xia, X. GSK872 and necrostatin-1 protect retinal ganglion cells against necroptosis through inhibition of RIP1/RIP3/MLKL pathway in glutamate-induced retinal excitotoxic model of glaucoma. J. Neuroinflammation 19, 262 (2022).
He, S. et al. Receptor interacting protein kinase-3 determines cellular necrotic response to TNF-alpha. Cell 137, 1100–1111 (2009).
Yu, X., Chini, C. C. S., He, M., Mer, G. & Chen, J. The BRCT domain is a phospho-protein binding domain. Science 302, 639–642 (2003).
Chen, W. et al. Diverse sequence determinants control human and mouse receptor interacting protein 3 (RIP3) and mixed lineage kinase domain-like (MLKL) interaction in necroptotic signaling. J. Biol. Chem. 288, 16247–16261 (2013).
Xie, Y. et al. Gut epithelial TSC1/mTOR controls RIPK3-dependent necroptosis in intestinal inflammation and cancer. J. Clin. Invest. 130, 2111–2128 (2020).
Sun, L. et al. Mixed lineage kinase domain-like protein mediates necrosis signaling downstream of RIP3 kinase. Cell 148, 213–227 (2012).
Lange, A. et al. Classical nuclear localization signals: definition, function, and interaction with importin α. J. Biol. Chem. 282, 5101–5105 (2007).
Horton, P. et al. WoLF PSORT: protein localization predictor. Nucleic Acids Res. 35, W585–W587 (2007).
Cao, X. et al. Acetylation promotes TyrRS nuclear translocation to prevent oxidative damage. Proc. Natl Acad. Sci. USA 114, 687–692 (2017).
Narita, T., Weinert, B. T. & Choudhary, C. Functions and mechanisms of non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 20, 156–174 (2019).
MacPherson, L. et al. HBO1 is required for the maintenance of leukaemia stem cells. Nature 577, 266–270 (2020).
Newton, K., Sun, X. Q. & Dixit, V. M. Kinase RIP3 is dispensable for normal NF-KBs, signaling by the B-cell and T-cell receptors, tumor necrosis factor receptor 1, and toll-like receptors 2 and 4. Mol. Cell. Biol. 24, 1464–1469 (2004).
Li, D. et al. RIPK1-RIPK3-MLKL-dependent necrosis promotes the aging of mouse male reproductive system. Elife 6, e27692 (2017).
Leist, M., Single, B., Castoldi, A. F., Kühnle, S. & Nicotera, P. Intracellular adenosine triphosphate (ATP) concentration: a switch in the decision between apoptosis and necrosis. J. Exp. Med. 185, 1481–1486 (1997).
Ferrari, D., Stepczynska, A., Los, M., Wesselborg, S. & Schulze-Osthoff, K. Differential regulation and ATP requirement for caspase-8 and caspase-3 activation during CD95- and anticancer drug–induced apoptosis. J. Exp. Med. 188, 979–984 (1998).
Elmore, S. Apoptosis: a review of programmed cell death. Toxicol. Pathol. 35, 495–516 (2007).
Eguchi, Y., Shimizu, S. & Tsujimoto, Y. Intracellular ATP levels determine cell death fate by apoptosis or necrosis. Cancer Res. 57, 1835–1840 (1997).
Nombela-Arrieta, C. et al. Quantitative imaging of haematopoietic stem and progenitor cell localization and hypoxic status in the bone marrow microenvironment. Nat. Cell Biol. 15, 533–543 (2013).
Takubo, K. et al. Regulation of glycolysis by Pdk functions as a metabolic checkpoint for cell cycle quiescence in hematopoietic stem cells. Cell Stem Cell 12, 49–61 (2013).
Nakamura-Ishizu, A., Ito, K. & Suda, T. Hematopoietic stem cell metabolism during development and aging. Dev. Cell 54, 239–255 (2020).
Benayoun, B. A., Pollina, E. A. & Brunet, A. Epigenetic regulation of ageing: linking environmental inputs to genomic stability. Nat. Rev. Mol. Cell Biol. 16, 593–610 (2015).
Schwer, B. et al. Calorie restriction alters mitochondrial protein acetylation. Aging Cell 8, 604–606 (2009).
Newman, J. C. et al. Ketogenic diet reduces midlife mortality and improves memory in aging mice. Cell metabolism 26, 547–557 (2017).
Roberts, M. N. et al. A ketogenic diet extends longevity and healthspan in adult mice. Cell metabolism 26, 539–546 (2017).
Klein, H.-U. et al. Epigenome-wide study uncovers large-scale changes in histone acetylation driven by tau pathology in aging and Alzheimer’s human brains. Nat. Neurosci. 22, 37–46 (2019).
Krishnan, V. et al. Histone H4 lysine 16 hypoacetylation is associated with defective DNA repair and premature senescence in Zmpste24-deficient mice. Proc. Natl Acad. Sci. USA 108, 12325–12330 (2011).
Peleg, S. et al. Altered histone acetylation is associated with age-dependent memory impairment in mice. Science 328, 753–756 (2010).
Peleg, S. et al. Lifespan extension by targeting a link between metabolism and histone acetylation in Drosophila. Embo Rep. 17, 455–469 (2016).
Wang, W. et al. A genome-wide CRISPR-based screen identifies KAT7 as a driver of cellular senescence. Sci. Transl. Med. 13, eabd2655 (2021).
Yang, Y. et al. The histone lysine acetyltransferase HBO1 (KAT7) regulates hematopoietic stem cell quiescence and self-renewal. Blood 139, 845–858 (2022).
Imai, S. -I. & Guarente, L. NAD+ and sirtuins in aging and disease. Trends Cell Biol. 24, 464–471 (2014).
Hall, J. A., Dominy, J. E., Lee, Y. & Puigserver, P. The sirtuin family’s role in aging and age-associated pathologies. J. Clin. Invest. 123, 973–979 (2013).
He, H., Wang, Y. & Wang, J. ALKBH3 is dispensable in maintaining hematopoietic stem cells but forced ALKBH3 rectified the differentiation skewing of aged hematopoietic stem cells. Blood Sci. 2, 137–143 (2020).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
He, H. et al. Aging-induced IL27Ra signaling impairs hematopoietic stem cells. Blood 136, 183–198 (2020).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2021).
Acknowledgements
We thank X. Wang (National Institute of Biological Sciences) for providing Ripk3−/− mice. We thank G. -H. Liu (Institute of Zoology, Chinese Academy of Sciences) for providing Kat7 cDNA. We thank S. -J. Chen (Shanghai Jiao Tong University School of Medicine) for providing the 32D cell line. This work was supported by grants Z200022, 82250002, 92249305 and 2018YFA0800200 to J.W. from the National Key R&D Program of China or the Beijing Municipal Science & Technology Commission and the National Natural Science Foundation of China.
Author information
Authors and Affiliations
Contributions
Conceptualization, J.W.; Methodology, H.H., Y.W., B.T., Q.D. and C.W.; Investigation, H.H., Y.W., B.T., Q.D., C.W. and W.S.; Formal Analysis, J.W. and H.H.; Resources, J.W.; Writing, J.W.; Funding acquisition, J.W.; Supervision, J.W.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Zhaoqi Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Dysfunction of Mcph1 mildly disturbs hematopoietic homeostasis at young age.
(a) This histogram shows the expression of MCPH1 in various immune cells. Data were download from ASCOT. (b) The schematic diagram showing the targeting strategy to generate Mcph1flox mice. (c) The schematic diagram showing the breeding strategy to generate Mcph1−/− (Vav1-Cre; Mcph1flox/flox) mice. (d-j) 2-5 months old Mcph1−/− or WT mice were analyzed for hematopoietic progenitor cells and HSCs. (d-e) The histogram depicts the frequency (d) and the absolute numbers (e) of HSCs (CD34- CD150+ KSL). n = 7 mice per group. (f-g) The histogram shows the frequency (f) and the absolute numbers (g) of CMPs, GMPs, and MEPs. n = 7 mice per group. (h-i) The histogram shows the frequency (h) and the absolute numbers (i) of CLPs. n = 4 mice per group. (j) The scatter plots show the bone marrow (BM) cellularity of Mcph1−/− and WT mice. n = 7 mice per group. (k-n) 3-4 months old Mcph1−/− or WT mice were analyzed for thymocyte. (k) Representative flow cytometry of thymocytes for CD4/CD8 expression. (l) The scatter plots show the total cell number of thymus. (m-n) The scatter plots depict the percentage (m) and absolute numbers (n) of DN (CD4−CD8−), DP (CD4+CD8+), CD4+, and CD8+ T cells in thymus. n = 6-7 mice per group (WT: n = 7; Mcph1−/−: n = 6). (o-r) 3-4 months old Mcph1−/− or WT mice were analyzed for splenocytes. (o) Representative flow cytometry. (p) The scatter plots show the total cell number of the spleen. (q-r) The scatter plots depict the percentage (q) and absolute numbers (r) of myeloid, B, CD4+ T and CD8+ T cells in spleen. n = 6-7 mice per group (WT: n = 7; Mcph1−/−: n = 6). Two-tailed unpaired Student’s t-test was used for statistical analysis in l-n and q-r. Data are shown as mean ± SD in d-j, l-n and p-r.
Extended Data Fig. 2 Mcph1 dysfunction severely impairs the reconstitution capacity of HSCs.
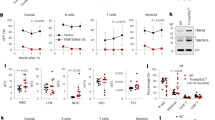
(a-f) Freshly isolated 20 HSCs from 4-month old Mcph1−/− mice or age-matched WT mice were transplanted into lethally irradiated recipients together with 3 × 105 competitor cells. Chimera in peripheral blood was checked every month until the 3rd month. (a) This histogram displays the percentage of donor cell reconstitution in overall (CD45.2+), B (B220+), T (CD3+) and myeloid (Mac-1+) cell every month after transplantation. (b and d) The scatter plots show the percentage of donor cell reconstitution in bone marrow (b) and HSC (d). (c) This histogram displays the lineage distribution in donor-derived bone marrow at the 3rd month. (e) This histogram displays the percentage of donor cell reconstitution in erythroid progenitors (Ter119+) every month after transplantation. (f) the scatter plots show the percentage of donor cell reconstitution in erythroid progenitors of bone marrow. n = 6-7 mice per group in a and e (n =6 for WT, n =7 for Mcph1−/−). n = 6 mice per group in b, c, d, and f. Data were shown as mean ± SD. Two-tailed unpaired Student’s t-test was used for statistical analysis.
Extended Data Fig. 3 MCPH1 antagonizes necroptosis through interaction with RIPK3.
(a-b) 4 months old Mcph1−/− or WT mice were analyzed for cell death of fresh HSCs without ex vivo culture by Annexin-V and DAPI staining. (a) Representative flow cytometry. (b) Percentage of early stage of cell death (DAPI− Annexin-V+) in Mcph1−/− or WT HSCs. n = 4-5 mice per group (WT: n = 5; Mcph1−/−: n = 4). (c) The images show morphological alteration of WT and Mcph1−/− HSCs (3 months). The experiment was repeated 3 times. (d) Representative western blot shows the interaction between MCPH1 and RIPK3 in response to necroptotic stimuli. L929 cells stably expressing Flag-tagged MCPH1 were treated with T+Z for 1 hour. Cell lysates were immunoprecipitated with anti-Flag antibody, and western blot was performed with indicated antibodies. The experiment was repeated 3 times. (e) Interaction between MCPH1 and p-RIPK3 in HEK293T cells co-transfected with SFB-tagged MCPH1 and Myc-tagged RIPK3, demonstrated by co-immunoprecipitation using S beads and subsequent western blot analysis with specific antibodies. The experiment was repeated 3 times. (f) SFB-RIPK3 were transfected into HEK293T cells together with GFP-MLKL and Myc-MCPH1-WT or Myc-MCPH1-Δ624–882 constructs as indicated. Cells were subjected to immunoprecipitated with S beads and western blot analysis was performed with indicated antibodies. Representative western blot shows that MCPH1-WT but not MCPH1-Δ624-882 inhibits interaction between p-RIPK3 and MLKL. The experiment was repeated 3 times. (g) Co-immunoprecipitation assay in HEK293T cells transiently transfected with SFB- MCPH1 and either Myc-tagged wild-type or T231A-S232A RIPK3, revealing a reduced interaction between MCPH1 and the RIPK3T231A-S232A mutant, as indicated by western blot analysis. The experiment was repeated 3 times. (h) The mRNA expression of Ripk3 in aged (23 months) Mcph1−/− and WT HSCs. n = 3 independent experiments. (i) Representative western blot showing the expression of p-S6, S6, p-4EBP1, 4EBP1, and MCPH1 in aged (21–26 months) Mcph1−/− and WT HSCs. The experiment was repeated twice. Two-tailed unpaired Student’s t-test was used for statistical analysis in b. Data are shown as mean ± SD in b and h.
Extended Data Fig. 4 Dysfunction of Ripk3 partially rescues the deleterious impact of Mcph1 deletion on HSCs during short-term ex vivo culture.
(a-c) Freshly isolated KSL cells either from Ripk3−/− mice (2 months old) or age-matched WT mice were infected by lentivirus carrying Mcph1 shRNA or scramble control (shRNA expressing lentiviruses coexpressed GFP), and 72 hours later, FACS-purified 100 GFP+ CD48− Sca1+ cells were cultured in SFEM medium. 7 days later, the proliferation capacity was evaluated by examining the colony size and cell number. (a) Experimental design. (b) Representative images of ex vivo-cultured HSCs carrying the indicated shRNA on day 7. Scale bar, 50 μm. (c) The scatter plots depict the cell numbers of ex vivo-cultured HSCs carrying the indicated shRNA on day 7. Statistical significance was measured by two-way ANOVA followed by Sidak′s test for post hoc comparison. n = 18–20 independent experiments (n = 20 for WT: shCon, n = 19 for WT: shMcph1, n = 18 for Ripk3−/−: shCon, n = 20 for Ripk3−/−: shMcph1). Data were shown as mean±SD.
Extended Data Fig. 5 Nuclear translocation of MCPH1 in response to DNA damage.
(a-b) Cells were treated with 100 μM etoposide and 1 hour later, cell lysates were separated into cytoplasmic and nuclear fractions. The subcellular distribution of MCPH1 were analyzed by western blot. GAPDH and H3 were used to detect the purity of cytoplasmic and nuclear fractions respectively. (a) EL4 cells. (b) 32D cells. The experiment was repeated 3 times. (c) KSL cells were treated with 100 μM etoposide and 1 hour later, cell lysates were separated into cytoplasmic and nuclear fractions. The subcellular distribution of RIPK3 were analyzed by western blot. The experiment was repeated 3 times.
Extended Data Fig. 6 KAT7-dependent acetylation of MCPH1 facilitating nuclear translocation in response to DNA damage.
(a) HEK293T cells were either mock treated or treated with 100 μM etoposide with or without 1 μM KAT7 inhibitor for 1 hours. cell lysates were separated into cytoplasmic and nuclear fractions. The subcellular distribution of MCPH1 were analyzed by western blot. The experiment was repeated 3 times. (b) HEK293T cells were transiently transfected with SFB-tagged wild-type MCPH1 or indicated K/R mutants together with Myc-tagged KAT7. Cell lysates were immunoprecipitated with S beads, and western blot analysis was performed with indicated antibodies. Representative western blot shows the acetylation level of wild-type or various K/R mutants of MCPH1. The experiment was repeated twice. (c) HEK293T cells were transiently transfected with SFB-tagged wild-type MCPH1 or MCPH1K348/350R, MCPH1K360/362R, MCPH1K364/365R, MCPH1K360/362/364/365R together with Myc-tagged KAT7. Cell lysates were immunoprecipitated with S beads, and western blot analysis was performed with indicated antibodies. Representative western blot shows the acetylation level of wild-type or indicated mutants of MCPH1. The experiment was repeated 3 times. (d) HEK293T cells were transiently transfected with SFB-tagged wild-type MCPH1 or K348R-K350R-K360R-K362R mutant together with Myc-tagged KAT7. 24 hours later, cell lysates were immunoprecipitated with S beads, and western blot analysis was performed with indicated antibodies. Representative western blot shows the interaction between MCPH1K348/350/360/362R and MCPH1 with KAT7. The experiment was repeated 3 times. (e) HEK293T cells were transiently transfected with SFB-tagged wild-type MCPH1 or K348R-K350R-K360R-K362R mutant together with Myc-tagged RIPK3. 24 hours later, cell lysates were immunoprecipitated with S beads, and western blot analysis was performed with indicated antibodies. Representative western blot shows the interaction between MCPH1K348/350/360/362R and MCPH1 with RIPK3. The experiment was repeated twice. (f) MCPH1- and MCPH1K348R-K350R-K360R-K362R-expressed L929 cells were treated with etoposide + zVAD (100 μM etoposide, 20 μM zVAD) for 20 hours. Representative western blot shows the expression of p-MLKL, MLKL, p-RIPK3, RIPK3, and MCPH1 in WT or K348R-K350R-K360R-K362R mutant MCPH1 overexpressed cells upon etoposide + zVAD treatment. The experiment was repeated 3-4 times.
Extended Data Fig. 7 Enhanced susceptibility of HSCs to necroptosis during aging.
(a) The images show morphological alteration of in Mlkl−/− and WT HSCs at young (3 months) and old (29 months) age. (b) The scatter plots depict the percentage of necroptotic cell death in Mlkl−/− and WT HSCs at young (3 months) and old (29 months) age. Data were shown as mean ± SD. Statistical significance was measured by two-way ANOVA followed by Tukey’s test for post hoc comparison. n = 3 independent experiments. (c-f) KSL cells were freshly isolated from young (5 months) or old (18 months) mice were infected by lentivirus carrying the cDNA of WT MCPH1 or MCPH1- K348R-K350R-K360R-K362R mutant (MCPH14KR), and 72 hours later, 2 × 104 GFP+ cells were sorted and injected into lethally irradiated recipients together with 2 × 105 competitor cells. Peripheral blood of recipient mice was evaluated every month until the 4th month. This histogram displays the percentage of GFP+ cell in overall (CD45.2+), B (B220+), T (CD3+) and myeloid (Mac-1+) cell every month after transplantation. n = 6-7 mice per group (n = 6 for Young: Vector, Young: MCPH1-WT, Old: Vector, and Old: MCPH1-WT; n = 7 for Young: MCPH1-4KR and Old: MCPH1-4KR), data were shown as mean ± SD. Two-tailed unpaired Student’s t-test was used for statistical analysis.
Extended Data Fig. 8 Representative flow cytometry gating strategy used in this study.
(a) Representative flow cytometry showing the gating strategy used to isolate and analyze the hematopoietic stem and progenitor cell populations in mouse bone marrow presented on Figs. 1a,b,2a–h,3a–h,l,o,4a, b, j, k,5a, b,d–f,6n–o, F,7a–d, g–i, j, l and Extended Data Figs. 1d–h, 2a, d, 3a–c, h, i, 4a, b, 5c, 7a–f. (b) Strategy used to evaluate the the percentage of B, myeloid, CD4+ T, CD8+ T cells presented on Fig. 1d, e. (c-d) Strategy used to evaluate the the percentage (c) and lineage distribution (d) of test donor-derived cells (myeloid, B and T cells) presented on Fig. 2f–g, 4c–i, 7g, j, k, l, m, and Extended Data Figs. 2a–c, 7c–f. (e) Strategy used to evaluate the the percentage of test donor-derived erythroid progenitors (Ter119+) presented on Extended Data Fig. 2e, f.
Supplementary information
Supplementary Information
Working model
Supplementary Tables 1–3
Supplementary Table 1. GSEA gene list; Supplementary Table 2. Oligonucleotides; Supplementary Table 3. Materials;
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 7
Unprocessed western blots.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, H., Wang, Y., Tang, B. et al. Aging-induced MCPH1 translocation activates necroptosis and impairs hematopoietic stem cell function. Nat Aging 4, 510–526 (2024). https://doi.org/10.1038/s43587-024-00609-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-024-00609-z
- Springer Nature America, Inc.