Abstract
The induction of cellular reprogramming via expression of the transcription factors Oct4, Sox2, Klf4 and c‐Myc (OSKM) can drive dedifferentiation of somatic cells and ameliorate age-associated phenotypes in multiple tissues and organs. However, the benefits of long-term in vivo reprogramming are limited by detrimental side‐effects. Here, using complementary genetic approaches, we demonstrated that continuous induction of the reprogramming factors in vivo leads to hepatic and intestinal dysfunction resulting in decreased body weight and contributing to premature death (within 1 week). By generating a transgenic reprogrammable mouse strain, avoiding OSKM expression in both liver and intestine, we reduced the early lethality and adverse effects associated with in vivo reprogramming and induced a decrease in organismal biological age. This reprogramming mouse strain, which allows longer-term continuous induction of OSKM with attenuated toxicity, can help better understand rejuvenation, regeneration and toxicity during in vivo reprogramming.




Similar content being viewed by others
Data availability
The authors confirm that data supporting the findings of this study are available within the article and its Supplementary Information, or are available from the corresponding author upon reasonable request. Raw sequencing reads related to microbiome analysis were deposited in the European Nucleotide Archive (ENA) under the project number PRJEB66275.
References
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Lapasset, L. et al. Rejuvenating senescent and centenarian human cells by reprogramming through the pluripotent state. Genes Dev. 25, 2248–2253 (2011).
Olova, N., Simpson, D. J., Marioni, R. E. & Chandra, T. Partial reprogramming induces a steady decline in epigenetic age before loss of somatic identity. Aging Cell 18, e12877 (2019).
Sarkar, T. J. et al. Transient non-integrative expression of nuclear reprogramming factors promotes multifaceted amelioration of aging in human cells. Nat. Commun. 11, 1545 (2020).
Roux, A. et al. Partial reprogramming restores youthful gene expression through transient suppression of cell identity. Cell Syst. https://doi.org/10.1016/j.cels.2022.05.002 (2021).
Abad, M. et al. Reprogramming in vivo produces teratomas and iPS cells with totipotency features. Nature 502, 340–345 (2013).
Ocampo, A. et al. In vivo amelioration of age-associated hallmarks by partial reprogramming. Cell 167, 1719–1733 e1712 (2016).
Doeser, M. C., Scholer, H. R. & Wu, G. Reduction of fibrosis and scar formation by partial reprogramming in vivo. Stem Cells 36, 1216–1225 (2018).
Chen, Y. et al. Reversible reprogramming of cardiomyocytes to a fetal state drives heart regeneration in mice. Science 373, 1537–1540 (2021).
Wang, C. et al. In vivo partial reprogramming of myofibers promotes muscle regeneration by remodeling the stem cell niche. Nat. Commun. 12, 3094 (2021).
de Lazaro, I. et al. Non-viral, tumor-free induction of transient cell reprogramming in mouse skeletal muscle to enhance tissue regeneration. Mol. Ther. 27, 59–75 (2019).
Chondronasiou, D. et al. Multi-omic rejuvenation of naturally aged tissues by a single cycle of transient reprogramming. Aging Cell 21, e13578 (2022).
Hishida, T. et al. In vivo partial cellular reprogramming enhances liver plasticity and regeneration. Cell Rep. 39, 110730 (2022).
Rodriguez-Matellan, A., Alcazar, N., Hernandez, F., Serrano, M. & Avila, J. In vivo reprogramming ameliorates aging features in dentate gyrus cells and improves memory in mice. Stem Cell Rep. https://doi.org/10.1016/j.stemcr.2020.09.010 (2020).
Chiche, A. et al. Injury-induced senescence enables in vivo reprogramming in skeletal muscle. Cell Stem Cell 20, 407–414 e404 (2017).
Lu, Y. et al. Reprogramming to recover youthful epigenetic information and restore vision. Nature 588, 124–129 (2020).
Browder, K. C. et al. In vivo partial reprogramming alters age-associated molecular changes during physiological aging in mice. Nature Aging https://doi.org/10.1038/s43587-022-00183-2 (2022).
Alle, Q. et al. A single short reprogramming early in life initiates and propagates an epigenetically related mechanism improving fitness and promoting an increased healthy lifespan. Aging Cell 21, e13714 (2022).
Mosteiro, L. et al. Tissue damage and senescence provide critical signals for cellular reprogramming in vivo. Science https://doi.org/10.1126/science.aaf4445 (2016).
Ohnishi, K. et al. Premature termination of reprogramming in vivo leads to cancer development through altered epigenetic regulation. Cell 156, 663–677 (2014).
Okamoto, O. K. et al. Whole transcriptome analysis of the hippocampus: toward a molecular portrait of epileptogenesis. BMC Genomics 11, 230 (2010).
Carey, B. W., Markoulaki, S., Beard, C., Hanna, J. & Jaenisch, R. Single-gene transgenic mouse strains for reprogramming adult somatic cells. Nat. Methods 7, 56–59 (2010).
Postic, C. et al. Dual roles for glucokinase in glucose homeostasis as determined by liver and pancreatic beta cell-specific gene knock-outs using Cre recombinase. J. Biol. Chem. 274, 305–315 (1999).
Madison, B. B. et al. Cis elements of the villin gene control expression in restricted domains of the vertical (crypt) and horizontal (duodenum, cecum) axes of the intestine. J. Biol. Chem. 277, 33275–33283 (2002).
Singh, P. B. & Zhakupova, A. Age reprogramming: cell rejuvenation by partial reprogramming. Development https://doi.org/10.1242/dev.200755 (2022).
de Lázaro, I., Orejón-Sánchez, T. L., Tringides, C. M. & Mooney, D. J. Induced reprogramming of adult murine cardiomyocytes to pluripotency in vivo. Preprint at bioRxiv https://doi.org/10.1101/2021.12.22.473302 (2021).
Senis, E. et al. AAV vector-mediated in vivo reprogramming into pluripotency. Nat. Commun. 9, 2651 (2018).
Macip, C. C. et al. Gene therapy mediated partial reprogramming extends lifespan and reverses age-related changes in aged mice. Preprint at bioRxiv https://doi.org/10.1101/2023.01.04.522507 (2023).
Tetteh, P. W., Farin, H. F. & Clevers, H. Plasticity within stem cell hierarchies in mammalian epithelia. Trends Cell Biol. 25, 100–108 (2015).
Yanger, K. et al. Robust cellular reprogramming occurs spontaneously during liver regeneration. Genes Dev. 27, 719–724 (2013).
Belteki, G. et al. Conditional and inducible transgene expression in mice through the combinatorial use of Cre-mediated recombination and tetracycline induction. Nucleic Acids Res. 33, e51 (2005).
Mozhui, K. et al. Genetic loci and metabolic states associated with murine epigenetic aging. eLife https://doi.org/10.7554/eLife.75244 (2022).
Arneson, A. et al. A mammalian methylation array for profiling methylation levels at conserved sequences. Nat. Commun. 13, 783 (2022).
Zhou, W., Triche, T. J. Jr., Laird, P. W. & Shen, H. SeSAMe: reducing artifactual detection of DNA methylation by Infinium BeadChips in genomic deletions. Nucleic Acids Res. 46, e123 (2018).
zAMP. GitHub https://github.com/metagenlab/zAMP (2022).
Acknowledgements
The authors thank all members of the Ocampo laboratory for helpful discussions. We thank the teams of mouse facilities at the University of Lausanne including Francis Derouet (head of the animal facility at Epalinges), L. Arlandi, I. Grandjean (head of the animal facility of Agora) and L. Lecomte (head of the animal facility of the Department of Biomedical Sciences). We thank A. Ablasser for the kind donation of the 4Fs-B rtTA mice. We thank the Service de Chimie Clinique du CHUV for mouse plasma analysis and the Mouse Pathology Facility of the University of Lausanne for tissue processing and immunostainings. This work was supported by the Milky Way Research Foundation (MWRF), the Eccellenza grants from the Swiss National Science Foundation (SNSF), the University of Lausanne, and the Canton Vaud. G.D.-M. was supported by the EMBO postdoctoral fellowship (EMBO ALTF 444-2021 to G.D.-M.). Microbiota analysis was supported as a part of NCCR Microbiomes, a National Centre of Competence in Research, funded by the Swiss National Science Foundation (grant number 180575).
Author information
Authors and Affiliations
Contributions
A.P., A.V.-A. and G.D-M were involved in the design of the study, in all experiments, data collection and statistical analysis. C.R., C.M., M.C.M., S.P. and C.Y.M. contributed to RNA and protein extraction, qRT–PCR and western blot analysis. S.P. helped with figure generation and revision. K.P. made intellectual contributions. C.V.B. and F.B. were involved in genotyping and sample collection. C.S performed H&E staining of the samples. R.B., A.H. and S.H. performed epigenetic clock analysis. E.M.-B. and C.B. performed microbiota analysis. A.O. directed and supervised the study and designed the experiments. A.P. and A.O wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
K.P. and A.O. are co-founders and shareholders of EPITERNA SA (non-financial interests). A.O. is co-founder of Longevity Consultancy Group (non-financial interests). S.H. is a founder of the non-profit Epigenetic Clock Development Foundation which licenses several patents from his former employer UC Regents. These patents list S.H. as inventor. The rest of the authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks Yasuhiro Yamada and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
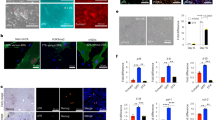
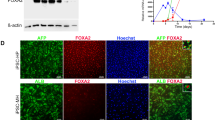
Extended Data Fig. 1 In vivo reprogramming induces liver and intestinal damage.
a, Experimental design indicating doxycycline administration protocol, activity measure by open field (OF) test, and organ and plasma collection. b, Superficial temperature of 4Fj and 4Fs-B mice upon continuous administration of doxycycline, as wells as untreated 4F control mice (Ctrl). c, Sox2 mRNA and d, OCT4 protein levels in the indicated organs of untreated 4F controls, and 4Fj and 4Fs-B mice after induction of 4 days. e, Immunostaining and quantification of Ki67 positive cells in the liver, small intestine and pancreas (n = 3 regions from four untreated controls, five 4Fj, and four 4Fs-B mice). Scale bars, 100 μm. f, Potassium, phosphate, uric acid and lipase pancreatic levels in plasma of untreated control mice (Ctrl), and 4Fj and 4Fs-B mice after 4 days of induction of in vivo reprogramming. Data show mean ± SEM. Statistical significance was assessed by (b-f) one-way ANOVA followed by Tukey’s or Games-Howells post hoc tests.
Extended Data Fig. 2 Instestinal analysis after in vivo reprogramming.
a, Small intestine hematoxylin and eosin staining for an untreated controls and 4Fj mice and treated 4Fj and 4Fs-B mice during 4 days; white arrowhead points apoptotic figures, black arrowhead points an example of globet cell, black arrows points dedifferentiated enterocytes. b, Alpha diversity of the microbiota at the AVS level of the 3 groups at each time point. The table inside the Chao1 plot represents the number of mice samples per group at each time point. c, Results of the Bonferroni-corrected pairwise Wilcoxon test of the Chao1 (bottom diagonal matrix) and Shannon (upper diagonal matrix) indexes. P-values intervals: *** (0, 0.001], ** (0.001, 0.01], * (0.01, 0.05], * (0.05, 0.1]. d, Distance-based redundacy analysis by Bray-Curtis distance. Variance explained by the model: 0.453. Axis1 and 2 explain 0.689 and 0.180 of the model variance, respectively. Arrows show the effect and direction of the explanatory variables. Samples are represented by dots and their colour indicates their group plus time point. Diamonds represents differentially abundant genera identified by NBZIMM with the same formula as the dbRDA model.
Extended Data Fig. 3 Liver and intestine-specific reprogramming lead to decrease body weight and early mortality.
a, Expression of Cre recombinase in the liver, small intestine, and kidney of control (4Fj LSL-rtTA), 4F-Liver (4Fj LSLrtTA AlbCre), and 4F-Intestine (4Fj LSLrtTA Vil1Cre) mice. b, Sox2 mRNA levels in control, 4F-Liver, and 4F-Intestine mice after induction of reprogramming for 4 days. c, Superficial body temperature upon continuous administration of doxycycline in controls and 4F tissue-specific reprogramming mice. Data show mean ± SEM. Statistical significance was assessed by (b-c) one-way ANOVA followed by Tukey’s post hoc test.
Extended Data Fig. 4 Avoiding the expression of OSKM in liver and intestine diminishes adverse phenotypes and premature death associated with in vivo reprogramming.
a, Sox2 mRNA levels in in the liver, small intestine, and kidney of 4F-Flox (OSKM whole body), 4F Non-Liver, and 4F Non-Intestine mice, after 4 days of induction of in vivo reprogramming. b, Superficial body temperature upon continuous administration of doxycycline in whole body (4F-Flox), 4F Non-liver, and 4F Non-intestine mice. Data show mean ± SEM. Statistical significance was assessed by (a-b) one-way ANOVA followed by Tukey’s post hoc test.
Extended Data Fig. 5 Safe and long-term continuous induction of in vivo reprogramming in a novel reprogrammable strain.
a, Sox2 mRNA levels in control and 4F non-Liver/Intestine mice after induction of in vivo reprogramming for 4 days. b, Body temperature of control and 4F Non-Liver/Intestine mice upon continuous administration of doxycycline. c, mRNA levels of Sox2 in multiple organs of 4F whole body reprogrammable and 4F Non-Liver/Intestine mice after 4 days and 4 weeks of continuous doxycycline treatment. d, Glucose, liver enzymes (ASAT, ALAT, ALP) and bilirubin levels in plasma of untreated 4F controls, treated no reprogrammable mice, 4F whole body after 4 days of treatment, and 4F Non-Liver/Intestine mice after 4 weeks of induction of in vivo reprogramming. Data show mean ± SEM. (a, c) Two-sided unpaired t-test and Mann-Whitney-Wilcoxon test, (b, d) one-way ANOVA followed by Tukey’s post hoc test.
Extended Data Fig. 6 Dynamics of expression and safety induction of in vivo reprogramming in 4F-Non Liv/Int mice.
a, Experimental design indicating doxycycline administration protocol, activity measure by open field (OF) test, and organ and blood collection. b, Survival of 4F-Flox after 4 days of administration of doxycycline and 4F Non-Liver/Intestine mice after 4, 7, 10 or 14 days administration of doxycycline. c, Body weight of 4F Non-Liver/Intestine mice after 4, 7 or 10 days of reprogramming induction. d, mRNA levels of Oct4 in control, 4F Non-Liver/Intestine mice after 4, 7 or 10 days of reprogramming induction and 4F-Flox after 4 days of reprogramming induction. e, Activity measured in distance travelled in open field test in control and 4F Non-Liver/Intestine mice after 4, 7 or 10 days of administration of doxycycline. f, Red blood cells (RBC), hemoglobine (HGB) and hematocrit (HCT) levels in blood of control and 4F Non-Liver/Intestine mice after 4, 7 or 10 days of administration of doxycycline. Data show mean ± SEM. Statistical significance was assessed by (c) two-sided unpaired t-test and Mann-Whiney - Wilcoxon test, (b, d-f) one-way ANOVA followed by Tukey’s post hoc test and c, log-rank (Mantel-Cox) test.
Extended Data Fig. 7 Expression and safety of induction of in vivo reprogramming in 4F-Non Liv/int mice.
a, mRNA levels of Sox2 in control, 4F Non-Liver/Intestine mice after 4, 7 or 10 days of reprogramming induction and 4F-Flox after 4 days of reprogramming induction. b, Spleen, kidney and skeletal muscle hematoxylin and eosin staining of an untreated wildtype and 4F Non-Liver/Intestine mice after 4, 7 or 10 days of reprogramming induction. Data show mean ± SEM. (a) one-way ANOVA followed by Tukey’s post hoc test.
Extended Data Fig. 8 Rejuvenation in middle-aged 4F-Non Liv/Int mice upon in vivo reprogramming.
a, Experimental design indicating doxycycline administration protocol, activity measure by open field (OF) test, and organ and blood collection. b, Survival of 4F-Flox after 4 days of administration of doxycycline and middle-aged 4F Non-Liver/Intestine mice after 2 cycles of 10 days of doxycycline administration. c, mRNA levels of Oct4 and d, Epigenetic clock analysis in control and middle-aged 4F Non-Liver/Intestine mice after 2 cycles of 10 days of doxycycline administration. Data show mean ± SEM. Statistical significance was assessed by (c, d) two-sided unpaired t-test and Mann-Whiney - Wilcoxon test and b, log-rank (Mantel-Cox) test.
Extended Data Fig. 9 Safety reprogramming in middle-aged 4F-Non LIv/Int mice.
a, Body weight, b, Activity measured in distance travelled in open field test in control and middle-aged 4F Non-Liver/Intestine mice after 2 cycles of 10 days of doxycycline administration. c, Blood analysis of red blood cells (RBC), hemoglobin (HGB), hematocrit (HCT) and platelets (PLT) in control and middle aged 4F Non-Liver/Intestine mice after 1 or 2 cycles of 10 days od doxycycline administration. d, Spleen, kidney and skeletal muscle hematoxylin and eosin staining of control and middle 4F Non-Liver/Intestine mice after 2 cycles of 10 days of doxycycline administration. e, mRNA levels of Sox2 in control and middle-aged 4F Non-Liver/Intestine mice after 2 cycles of 10 days of doxycycline administration f, Epigenetic clock analysis in spleen, heart, brain and kidney in control and middle-aged 4F Non-Liver/Intestine mice after 2 cycles of 10 days of doxycycline administration. Data show mean ± SEM. (a-c, f) Two-sided unpaired t-test and Mann-Whitney-Wilcoxon test and (e) one-way ANOVA followed by Tukey’s post hoc test.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed gel.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Parras, A., Vílchez-Acosta, A., Desdín-Micó, G. et al. In vivo reprogramming leads to premature death linked to hepatic and intestinal failure. Nat Aging 3, 1509–1520 (2023). https://doi.org/10.1038/s43587-023-00528-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-023-00528-5
- Springer Nature America, Inc.
This article is cited by
-
Partial reprogramming of the mammalian brain
Nature Aging (2024)
-
Restoration of neuronal progenitors by partial reprogramming in the aged neurogenic niche
Nature Aging (2024)
-
The long and winding road of reprogramming-induced rejuvenation
Nature Communications (2024)
-
The Information Theory of Aging
Nature Aging (2023)