Abstract
Reproductive ageing is one of the earliest human ageing phenotypes, and mitochondrial dysfunction has been linked to oocyte quality decline; however, it is not known which mitochondrial metabolic processes are critical for oocyte quality maintenance with age. To understand how mitochondrial processes contribute to Caenorhabditis elegans oocyte quality, we characterized the mitochondrial proteomes of young and aged wild-type and long-reproductive daf-2 mutants. Here we show that the mitochondrial proteomic profiles of young wild-type and daf-2 worms are similar and share upregulation of branched-chain amino acid (BCAA) metabolism pathway enzymes. Reduction of the BCAA catabolism enzyme BCAT-1 shortens reproduction, elevates mitochondrial reactive oxygen species levels, and shifts mitochondrial localization. Moreover, bcat-1 knockdown decreases oocyte quality in daf-2 worms and reduces reproductive capability, indicating the role of this pathway in the maintenance of oocyte quality with age. Notably, oocyte quality deterioration can be delayed, and reproduction can be extended in wild-type animals both by bcat-1 overexpression and by supplementing with vitamin B1, a cofactor needed for BCAA metabolism.







Similar content being viewed by others
Data availability
All RNA-seq data are deposited under the PRJNA966212 BioProject. Microarray data are available on the PUMA database at http://puma.princeton.edu as experiment set 7,368. Proteomics data are available on PRIDE under accession code PXD048296. Source data are provided with this paper.
Code availability
No custom code was used for this study.
References
Duncan, F. E., Confino, R. & Pavone, M. E. in Conn’s Handbook of Models for Human Aging. 2nd Edn (eds Ram, J. L. & Conn, P. M.) 109–130 (Academic Press, 2018).
Huber, S. & Fieder, M. Evidence for a maximum ‘shelf-life’ of oocytes in mammals suggests that human menopause may be an implication of meiotic arrest. Sci. Rep. 8, 14099 (2018).
Mathews, T. J. & Hamilton, B. E. Mean age of mothers is on the rise: United States, 2000–2014. NCHS Data Brief. 232, 1–8 (2016).
Tilly, J. L. & Sinclair, D. A. Germline energetics, aging, and female infertility. Cell Metab. 17, 838–850 (2013).
van der Reest, J., Nardini Cecchino, G., Haigis, M. C. & Kordowitzki, P. Mitochondria: their relevance during oocyte ageing. Ageing Res. Rev. 70, 101378 (2021).
Broekmans, F. J., Soules, M. R. & Fauser, B. C. Ovarian aging: mechanisms and clinical consequences. Endocr. Rev. 30, 465–493 (2009).
te Velde, E. R. & Pearson, P. L. The variability of female reproductive ageing. Hum. Reprod. Update 8, 141–154 (2002).
Hughes, S. E., Evason, K., Xiong, C. & Kornfeld, K. Genetic and pharmacological factors that influence reproductive aging in nematodes. PLoS Genet. 3, e25 (2007).
Luo, S., Kleemann, G. A., Ashraf, J. M., Shaw, W. M. & Murphy, C. T. TGF-β and insulin signaling regulate reproductive aging via oocyte and germline quality maintenance. Cell 143, 299–312 (2010).
Hamatani, T. et al. Age-associated alteration of gene expression patterns in mouse oocytes. Hum. Mol. Genet. 13, 2263–2278 (2004).
Steuerwald, N. M., Bermúdez, M. G., Wells, D., Munné, S. & Cohen, J. Maternal age-related differential global expression profiles observed in human oocytes. Reprod. Biomed. Online 14, 700–708 (2007).
Mishina, T. et al. Single‐oocyte transcriptome analysis reveals aging‐associated effects influenced by life stage and calorie restriction. Aging Cell 20, e13428 (2021).
Luo, S., Shaw, W. M., Ashraf, J. & Murphy, C. T. TGF-β Sma/Mab signaling mutations uncouple reproductive aging from somatic aging. PLoS Genet. 5, e1000789 (2009).
Kenyon, C., Chang, J., Gensch, E., Rudner, A. & Tabtiang, R. A C. elegans mutant that lives twice as long as wild type. Nature 366, 461–464 (1993).
Hahm, J.-H. et al. C. elegans maximum velocity correlates with healthspan and is maintained in worms with an insulin receptor mutation. Nat. Commun. 6, 8919 (2015).
Garigan, D. et al. Genetic analysis of tissue aging in Caenorhabditis elegans: a role for heat-shock factor and bacterial proliferation. Genetics 161, 1101–1112 (2002).
Huang, C., Xiong, C. & Kornfeld, K. Measurements of age-related changes of physiological processes that predict lifespan of Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 101, 8084–8089 (2004).
Templeman, N. M. et al. Insulin signaling regulates oocyte quality maintenance with age via cathepsin B activity. Curr. Biol. 28, 753–760.e4 (2018).
Athar, F. & Templeman, N. M. C. elegans as a model organism to study female reproductive health. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 266, 111152 (2022).
Llonch, S. et al. Single human oocyte transcriptome analysis reveals distinct maturation stage-dependent pathways impacted by age. Aging Cell 20, e13360 (2021).
Zhang, D., Keilty, D., Zhang, Z. & Chian, R. Mitochondria in oocyte aging: current understanding. Facts Views Vis. ObGyn 9, 29–38 (2017).
Rodríguez-Nuevo, A. et al. Oocytes maintain ROS-free mitochondrial metabolism by suppressing complex I. Nature 607, 756–761 (2022).
Cota, V., Sohrabi, S., Kaletsky, R. & Murphy, C. T. Oocyte mitophagy is critical for extended reproductive longevity. PLoS Genet. 18, e1010400 (2022).
Meldrum, D. R. et al. Aging and the environment affect gamete and embryo potential: can we intervene? Fertil. Steril. 105, 548–559 (2016).
Simsek-Duran, F. et al. Age-associated metabolic and morphologic changes in mitochondria of individual mouse and hamster oocytes. PLoS ONE 8, e64955 (2013).
Min, H., Lee, M., Cho, K. S., Lim, H. J. & Shim, Y.-H. Nicotinamide supplementation improves oocyte quality and offspring development by modulating mitochondrial function in an aged Caenorhabditis elegans model. Antioxidants 10, 519 (2021).
Song, C. et al. Melatonin improves age-induced fertility decline and attenuates ovarian mitochondrial oxidative stress in mice. Sci. Rep. 6, 35165 (2016).
Narayan, V. et al. Deep proteome analysis identifies age-related processes in C. elegans. Cell Syst. 3, 144–159 (2016).
Meissner, B. et al. Determining the sub-cellular localization of proteins within Caenorhabditis elegans body wall muscle. PLoS ONE 6, e19937 (2011).
Bratic, I. et al. Mitochondrial DNA level, but not active replicase, is essential for Caenorhabditis elegans development. Nucleic Acids Res. 37, 1817–1828 (2009).
Gitschlag, B. L. et al. Homeostatic responses regulate selfish mitochondrial genome dynamics in C. elegans. Cell Metab. 24, 91–103 (2016).
Tsang, W. Y. & Lemire, B. D. Mitochondrial genome content is regulated during nematode development. Biochem. Biophys. Res. Commun. 291, 8–16 (2002).
Murphy, C. T. et al. Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans. Nature 424, 277–283 (2003).
Guan, Y. et al. SKP1 drives the prophase I to metaphase I transition during male meiosis. Sci. Adv. 6, eaaz2129 (2020).
Kinterova, V., Kanka, J., Petruskova, V. & Toralova, T. Inhibition of Skp1-Cullin-F-box complexes during bovine oocyte maturation and preimplantation development leads to delayed development of embryos. Biol. Reprod. 100, 896–906 (2019).
Nadarajan, S., Govindan, J. A., McGovern, M., Hubbard, E. J. A. & Greenstein, D. MSP and GLP-1/Notch signaling coordinately regulate actomyosin-dependent cytoplasmic streaming and oocyte growth in C. elegans. Dev. Camb. Engl. 136, 2223–2234 (2009).
Vanorny, D. A. & Mayo, K. E. The role of Notch signaling in the mammalian ovary. Reproduction 153, R187–R204 (2017).
Agarwal, S. & Ganesh, S. Perinuclear mitochondrial clustering, increased ROS levels, and HIF1 are required for the activation of HSF1 by heat stress. J. Cell Sci. https://doi.org/10.1242/jcs.245589 (2020).
Mor, D. E. et al. Metformin rescues Parkinson’s disease phenotypes caused by hyperactive mitochondria. Proc. Natl Acad. Sci. USA 117, 26438–26447 (2020).
Mansfeld, J. et al. Branched-chain amino acid catabolism is a conserved regulator of physiological ageing. Nat. Commun. 6, 10043 (2015).
Luo, S. & Murphy, C. T. Caenorhabditis elegans reproductive aging: regulation and underlying mechanisms. Genesis 49, 53–65 (2011).
Pinkston-Gosse, J. & Kenyon, C. DAF-16/FOXO targets genes that regulate tumor growth in Caenorhabditis elegans. Nat. Genet. 39, 1403–1409 (2007).
Austin, M. U. et al. Knockout of the folate transporter folt-1 causes germline and somatic defects in C. elegans. BMC Dev. Biol. 10, 46 (2010).
Strandgaard, T. et al. Maternally contributed folate receptor 1 is expressed in ovarian follicles and contributes to preimplantation development. Front. Cell Dev. Biol. 5, 89 (2017).
Mann, G., Mora, S., Madu, G. & Adegoke, O. A. J. Branched-chain amino acids: catabolism in skeletal muscle and implications for muscle and whole-body metabolism. Front. Physiol. 12, 702826 (2021).
Palmieri, F., Monné, M., Fiermonte, G. & Palmieri, L. Mitochondrial transport and metabolism of the vitamin B-derived cofactors thiamine pyrophosphate, coenzyme A, FAD and NAD+, and related diseases: a review. IUBMB Life 74, 592–617 (2022).
Marrs, C. & Lonsdale, D. Hiding in plain sight: modern thiamine deficiency. Cells 10, 2595 (2021).
Ravanelli, S. et al. Reprograming of proteasomal degradation by branched chain amino acid metabolism. Aging Cell 21, e13725 (2022).
Watson, E. et al. Metabolic network rewiring of propionate flux compensates vitamin B12 deficiency in C. elegans. eLife 5, e17670 (2016).
Walker, M. D. et al. WormPaths: Caenorhabditis elegans metabolic pathway annotation and visualization. Genetics 219, iyab089 (2021).
Yu, D. et al. The adverse metabolic effects of branched-chain amino acids are mediated by isoleucine and valine. Cell Metab. 33, 905–922 (2021).
Weaver, K. J., Holt, R. A., Henry, E., Lyu, Y. & Pletcher, S. D. Effects of hunger on neuronal histone modifications slow aging in Drosophila. Science 380, 625–632 (2023).
Yao, V. et al. An integrative tissue-network approach to identify and test human disease genes. Nat. Biotechnol. https://doi.org/10.1038/nbt.4246 (2018).
Li, H. et al. Defect of branched-chain amino acid metabolism promotes the development of Alzheimer’s disease by targeting the mTOR signaling. Biosci. Rep. 38, BSR20180127 (2018).
Gao, A. W. et al. Identification of key pathways and metabolic fingerprints of longevity in C. elegans. Exp. Gerontol. 113, 128–140 (2018).
Juricic, P., Grönke, S. & Partridge, L. Branched-chain amino acids have equivalent effects to other essential amino acids on lifespan and aging-related traits in Drosophila. J. Gerontol. A Biol. Sci. Med. Sci. 75, 24–31 (2020).
Richardson, N. E. et al. Lifelong restriction of dietary branched-chain amino acids has sex-specific benefits for frailty and lifespan in mice. Nat. Aging 1, 73–86 (2021).
Nishi, K. et al. Branched-chain keto acids inhibit mitochondrial pyruvate carrier and suppress gluconeogenesis in hepatocytes. Cell Rep. 42, 112641 (2023).
Ferguson, D. et al. Mitochondrial pyruvate carrier inhibition initiates metabolic crosstalk to stimulate branched chain amino acid catabolism. Mol. Metab. 70, 101694 (2023).
Liemburg-Apers, D. C., Willems, P. H. G. M., Koopman, W. J. H. & Grefte, S. Interactions between mitochondrial reactive oxygen species and cellular glucose metabolism. Arch. Toxicol. 89, 1209–1226 (2015).
Fernhoff, P. M. et al. Thiamine response in maple syrup urine disease. Pediatr. Res. 19, 1011–1016 (1985).
Danner, D. J., Davidson, E. D. & Elsas, L. J. Thiamine increases the specific activity of human liver branched chain α-ketoacid dehydrogenase. Nature 254, 529–530 (1975).
Mouchiroud, L. et al. Pyruvate imbalance mediates metabolic reprogramming and mimics lifespan extension by dietary restriction in Caenorhabditis elegans. Aging Cell 10, 39–54 (2011).
Chin, R. M. et al. The metabolite α-ketoglutarate extends lifespan by inhibiting ATP synthase and TOR. Nature 510, 397–401 (2014).
Cabreiro, F. et al. Metformin retards aging in C. elegans by altering microbial folate and methionine metabolism. Cell 153, 228–239 (2013).
Li-Leger, E. et al. Identification of essential genes in Caenorhabditis elegans through whole-genome sequencing of legacy mutant collections. G3 11, jkab328 (2021).
C. elegans Deletion Mutant Consortium. large-scale screening for targeted knockouts in the Caenorhabditis elegans genome. G3 2, 1415–1425 (2012).
Tsuji, A., Nakamura, T. & Shibata, K. Effects of mild and severe vitamin B1 deficiencies on the meiotic maturation of mice oocytes. Nutr. Metab. Insights 10, 1178638817693824 (2017).
Ishii, N. et al. Coenzyme Q10 can prolong C. elegans lifespan by lowering oxidative stress. Mech. Ageing Dev. 125, 41–46 (2004).
Institute of Medicine (US) Standing Committee on the Scientific Evaluation of Dietary Reference Intakes and its Panel on Folate, Other B Vitamins, and Choline. Dietary Reference Intakes for Thiamin, Riboflavin, Niacin, Vitamin B6, Folate, Vitamin B12, Pantothenic Acid, Biotin, and Choline. (National Academies Press, 1998).
Blair, M. C., Neinast, M. D. & Arany, Z. Whole-body metabolic fate of branched-chain amino acids. Biochem. J. 478, 765–776 (2021).
Sieber, M. H., Thomsen, M. B. & Spradling, A. C. Electron transport chain remodeling by GSK3 during oogenesis connects nutrient state to reproduction. Cell 164, 420–432 (2016).
Li, M. et al. Integrative proteome analysis implicates aberrant RNA splicing in impaired developmental potential of aged mouse oocytes. Aging Cell 20, e13482 (2021).
Mu, L. et al. PPM1K-regulated impaired catabolism of branched-chain amino acids orchestrates polycystic ovary syndrome. eBioMedicine 89, 104492 (2023).
Ntostis, P. et al. The impact of maternal age on gene expression during the GV to MII transition in euploid human oocytes. Hum. Reprod. 37, 80–92 (2021).
Gandre, S. & van der Bliek, A. M. Mitochondrial division in Caenorhabditis elegans. Methods Mol. Biol. 372, 485–501 (2007).
Wieckowski, M. R., Giorgi, C., Lebiedzinska, M., Duszynski, J. & Pinton, P. Isolation of mitochondria-associated membranes and mitochondria from animal tissues and cells. Nat. Protoc. 4, 1582–1590 (2009).
Annunziata, I., Patterson, A. & d’Azzo, A. Mitochondria-associated ER membranes (MAMs) and glycosphingolipid enriched microdomains (GEMs): isolation from mouse brain. J. Vis. Exp. https://doi.org/10.3791/50215 (2013).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ *. Mol. Cell. Proteom. 13, 2513–2526 (2014).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Kwon, Y. J., Guha, S., Tuluc, F. & Falk, M. J. High-throughput BioSorter quantification of relative mitochondrial content and membrane potential in living Caenorhabditis elegans. Mitochondrion 40, 42–50 (2018).
Charmpilas, N. & Tavernarakis, N. Mitochondrial maturation drives germline stem cell differentiation in Caenorhabditis elegans. Cell Death Differ. 27, 601–617 (2020).
Sarasija, S. & Norman, K. R. Analysis of mitochondrial structure in the body wall muscle of Caenorhabditis elegans. Bio-Protoc. 8, e2801 (2018).
Templeman, N. M., Cota, V., Keyes, W., Kaletsky, R. & Murphy, C. T. CREB non-autonomously controls reproductive aging through Hedgehog/Patched signaling. Dev. Cell 54, 92–105 (2020).
Sohrabi, S., Cota, V. & Murphy, C. T. CeLab, a microfluidic platform for the study of life history traits, reveals metformin and SGK-1 regulation of longevity and reproductive span. Lab Chip 23, 2738–2757 (2023).
Sohrabi, S., Moore, R. S. & Murphy, C. T. CeAid: a smartphone application for logging and plotting Caenorhabditis elegans assays. G3 11, jkab259 (2021).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Shaw, W. M., Luo, S., Landis, J., Ashraf, J. & Murphy, C. T. The C. elegans TGF-β Dauer pathway regulates longevity via insulin signaling. Curr. Biol. 17, 1635–1645 (2007).
Acknowledgements
We thank H. Shwe and S. Kyin (Princeton Proteomics and Mass Spectrometry Core) for their assistance in sample processing and MS instrumentation, J. Miller and J. Volmar at the Genomics Core Facility of Princeton University for performing library preparation and next-generation sequencing, G. Laevsky and S. Wang (Confocal Imaging Facility, a Nikon Center of Excellence, in the Department of Molecular Biology at Princeton University), The De Botton Protein Profiling Institute of the Nancy and Stephen Grand Israel National Center for Personalized Medicine Weizmann Institute of Science (Y. Levin), the Petry Laboratory and the Gitai Laboratory (the Department of Molecular Biology at Princeton University) for help with equipment, the CGC for strains and Murphy Laboratory members for comments on the manuscript. The study was supported by funding to C.T.M. from the Global Consortium for Reproductive Longevity and Equality (AWD1006679), The Simons Foundation (811235SPI), The Glenn Foundation for Medical Research and Pioneer (DP1AG077430) and Transformative (R01AT011963) grants from the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Conceptualization was the responsibility of C.L., R.K. and C.T.M. Methodology was the responsibility of C.L., R.K. and C.T.M. Investigation was the responsibility of C.L., J.M.A., R.K., S.S., V.C., T.S., W.K. and S.L. Writing was the responsibility of C.L., R.K. and C.T.M. Funding acquisition was the responsibility of C.T.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks Patrick Bradshaw, Andrea Jurisicova and Michael Ristow for their contribution to the peer review of this work. Primary Handling Editor: Yanina-Yasmin Pesch, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Highly purified mitochondria are isolated from C. elegans.
Representative western blot of the different cellular fractions. The mitochondrial marker ATP5A is prominently enriched in the membranous fraction, unlike the cytosolic marker αTUB. The membranous fraction was then loaded on a gradient to highly purify the mitochondrial fraction. T- total homogenate, C- cytosolic fraction, M- crude membranes fraction. αTUB- αtubulin, HISH3- histone H3. The asterisk represents a non-specific band detected when using the anti-ATP5A antibody. Dashed lines mark where membranes were cut prior to incubation with the indicated antibody. 3 biological replicates were performed.
Extended Data Fig. 2 Differentially expressed oocyte genes in day 8 daf-2;control and daf-2;bcat-1(RNAi) worms.
Volcano plot of differentially expressed genes identified between daf-2;control and daf-2;bcat-1(RNAi) Day 8 oocytes. Red dots denote upregulated genes in daf-2;control oocytes, and orange dots denote genes downregulated in daf-2;control oocytes relative to daf-2;bcat-1(RNAi) (0.5 < log2FC < −0.5).
Extended Data Fig. 3 Controls for mitochondrial assays.
(a) mtROS levels increase in oocytes after paraquat treatment. A white dashed outline marks the most mature oocytes (−1 oocyte). Right: mtROS levels quantified in mature oocytes either with or without paraquat treatment. Control n = 20, Paraquat n = 24. (b) Left: FCCP treatment reduces mitochondrial membrane potential in oocytes. A white dashed outline marks the most mature oocytes (−1 oocyte). Right: TMRE levels quantified in mature oocytes under control or FCCP treatment conditions. Control n = 15, FCCP n = 22. (a, b) Two-tailed, unpaired t-test. ****p ≤ 0.0001. Dashed lines on the graphs represent mean intensity of autofluorescence. (a, b) Box plots: centre line, median; box range, 25–75th percentiles; whiskers denote minimum–maximum values. 3 biological replicates were performed.
Extended Data Fig. 4 Deletion of acdh-1 in daf-2 or wild-type worms does not affect mitochondrial function.
(a) Left: mtROS levels do not change in daf-2 mature oocytes on Day 5 upon acdh-1 deletion. Right: quantification of mtROS levels in daf-2;acdh-1 mature oocytes on day 5. N ≥ 45 for each condition. Graph shows results of pooled three biological replicates. (b) Left: acdh-1 deletion does not affect mitochondrial membrane potential in daf-2 mature oocytes on day 5. Right: quantification of TMRE signal in daf-2 mature oocytes on Day 5 upon acdh-1 deletion. N > 10 for all conditions. (c) Left: mtROS levels do not change in wild-type mature oocytes on Day 5 upon acdh-1 deletion. Right: quantification of mtROS levels in wild-type vs. acdh-1 mutant mature oocytes on day 5. N ≥ 17 for each condition. Graph shows results of pooled three biological replicates. (a–c) A white dashed outline marks the most mature oocytes (−1 oocytes). Two-tailed, unpaired t-test. ns- nonsignificant. For all panels, representative images are shown. 3 biological replicates were performed.
Extended Data Fig. 5 Downregulating bcat-1 in wild-type worms does not affect lifespan.
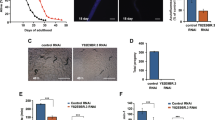
(a) Downregulation of bcat-1 reduces the late-mated reproduction of adult wild-type worms at day 6 of adulthood. n = 69 for control, n = 58 for bcat-1(RNAi). Chi-square test. ****p ≤ 0.0001. 3 biological replicates were performed. (b) GFP expression in control (left) and in the bcat-1-overexpression strain (GFP-BCAT-1 protein fusion). 3 biological replicates were performed. (c) Adult-only knockdown of bcat-1 does not affect the lifespan of wild-type animals. Lifespan data from 6 biological replicates are shown. Results are plotted as Kaplan–Meier survival curves, and the p-values were calculated using Mantel–Cox log-ranking. All replicates are shown. Statistical data is detailed in Table S2.
Extended Data Fig. 6 Differentially expressed oocyte genes in day 8 control and bcat-1 overexpressing worms.
Volcano plot of differentially expressed genes identified between Day 8 oocytes from bcat-1-overexpressing worms compared to the control (DESeq2, padj < 0.05). Pink dots denote upregulated genes in oocytes from bcat-1 overexpressing worms, and gold dots denote downregulated genes (0.5 <log2FC <−0.5).
Supplementary information
Supplementary Tables 1–5
Supplementary Table 1 Proteomic data. LFQ intensity of all peptides detected by LC–MS/MS proteomic analysis in mitochondria fractions isolated from young (day 1 adult) and reproductively aged (day 5 adult) wild-type worms and from daf-2(e1370) mutants. a–c, All peptides detected, and the significant proteins (proteins with more than one peptide, FC > 2 and q value < 0.05) (a) for comparing wild-type proteomic data of day 1 versus day 5 (b), as well as the significant proteins for comparing proteomic data of Supp daf-2 versus wild-type day 5 (c). Supplementary Table 2 Statistical data. a–c, Statistical analysis data of all biological replicates for reproductive span (a), lifespan (b) and late-mated reproduction experiments (c). Supplementary Table 3 RNA sequencing data for day 8 oocytes isolated from daf-2(e1370);control versus daf-2(e1370);bcat-1(RNAi) animals. a,b, Oocyte genes significantly upregulated (a) or downregulated (b) in daf-2(e1370);control(RNAi) versus daf-2(e1370);bcat-1(RNAi) animals. DESeq2 results were filtered for daf-2;control(RNAi) upregulated genes (bcat-1-dependent daf-2 oocyte genes) and daf-2;control(RNAi) downregulated genes (Padj < 0.05). c, Significant Gene Ontology terms are listed for the upregulated and downregulated gene sets. Supplementary Table 4 RNA sequencing data for day 8 oocytes isolated from fem-1(hc17) IV vs fem-1(hc17) IV; bcat-1 overexpressing worms. a,b, Significantly upregulated (a) and downregulated (b) genes in isolated oocytes of fem-1(hc17) IV versus fem-1(hc17) IV; risIs3 worms. DESeq2 results were filtered for differentially expressed genes (Padj < 0.05). Supplementary Table 5 C. elegans strains used in this study. Strain names and genotypes are listed.
Supplementary Video 1
Supplementary Video 1: daf-2 control germline mitochondria. 3D projections of z-slices of germline-labelled mitochondria in daf-2 animals treated with control RNAi. Related to data in Fig. 4a.
Supplementary Video 2
Supplementary Video 2: daf-2;bcat-1(RNAi) germline mitochondria. 3D projections of z-slices of germline-labelled mitochondria in daf-2 animals treated with bcat-1 RNAi. Related to data in Fig. 4a.
Supplementary Video 3
Supplementary Video 3: daf-2;bcat-1(RNAi) germline mitochondria. 3D projections of z-slices of germline-labelled mitochondria in daf-2 animals treated with bcat-1RNAi. Related to data in Fig. 4a.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Source Data Fig. 7
Statistical Source Data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 4
Statistical Source Data.
Source Data Extended Data Fig. 5
Statistical Source Data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lesnik, C., Kaletsky, R., Ashraf, J.M. et al. Enhanced branched-chain amino acid metabolism improves age-related reproduction in C. elegans. Nat Metab 6, 724–740 (2024). https://doi.org/10.1038/s42255-024-00996-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-024-00996-y
- Springer Nature Limited
This article is cited by
-
Slowing reproductive ageing by preserving BCAT-1
Nature Metabolism (2024)