Abstract
Many insects and other animals carry microbial endosymbionts that influence their reproduction and fitness. These relationships only persist if endosymbionts are reliably transmitted from one host generation to the next. Wolbachia are maternally transmitted endosymbionts found in most insect species, but transmission rates can vary across environments. Maternal transmission of wMel Wolbachia depends on temperature in natural Drosophila melanogaster hosts and in transinfected Aedes aegypti, where wMel is used to block pathogens that cause human disease. In D. melanogaster, wMel transmission declines in the cold as Wolbachia become less abundant in host ovaries and at the posterior pole plasm (the site of germline formation) in mature oocytes. Here, we assess how temperature affects maternal transmission and underlying patterns of Wolbachia localization across 10 Wolbachia strains diverged up to 50 million years—including strains closely related to wMel—and their natural Drosophila hosts. Many Wolbachia maintain high transmission rates across temperatures, despite highly variable (and sometimes low) levels of Wolbachia in the ovaries and at the developing germline in late-stage oocytes. Identifying strains like closely related wMel-like Wolbachia with stable transmission across variable environmental conditions may improve the efficacy of Wolbachia-based biocontrol efforts as they expand into globally diverse environments.
Similar content being viewed by others
Introduction
Microbes form diverse relationships with host organisms that span the tree of life, including animals, plants, and protists. Endosymbiosis is an especially close relationship where microbes occupy eukaryotic host cells. Heritable endosymbionts are particularly common in insects1,2,3, and these relationships alter fundamental aspects of host biology, including reproduction4,5, protection for natural enemies6,7,8,9, and nutrient acquisition10,11,12.
Many endosymbionts rely on vertical-maternal transmission to spread and persist in insect populations13,14,15. In obligate relationships, these endosymbionts must be maternally transmitted to ensure host survival (e.g., Buchnera and aphids16). For facultative endosymbionts like Wolbachia, imperfect maternal transmission decreases the prevalence of Wolbachia-positive individuals within the host population each generation17,18,19,20,21. Thus, maternal transmission is a fundamental determinant of how endosymbionts spread, persist, and evolve in host populations17,22,23. In addition, endosymbiont-based programs to control human diseases24,25,26 and agricultural pests27,28 explicitly depend on efficient maternal transmission of Wolbachia in transinfected insect vectors and pest populations. This includes introducing the virus-blocking wMel Wolbachia strain from Drosophila melanogaster into mosquito populations to block dengue and other arboviruses on multiple continents29,30,31,32. Nonetheless, it is largely unknown why maternal transmission may break down under certain circumstances, especially for non-model systems15,21,33,34,35,36.
Maternal transmission is predicted to depend on endosymbiont density (titer) in female reproductive tissues, and specifically at the site of germline formation during oogenesis14. In insects, endosymbionts often achieve vertical transmission by occupying the female ovaries. Endosymbionts are observed in germline stem cells in female hosts where they maintain a tight association with the germline throughout oogenesis, thereby ensuring a germline-to-germline route of vertical transmission through the host matriline14,15,36,37. In a more circuitous route, some endosymbionts may also colonize the germline from neighboring somatic cells via cell-to-cell migration through a soma-to-germline route of transmission4,14,15,38,39. Our canonical understanding of these cellular processes—and their contributions to maternal transmission—are based almost entirely in the context of standard laboratory conditions where maternal transmission rates are near perfect or perfect18,20,21,40. However, maternal transmission of facultative endosymbionts is often imperfect in natural host populations19,20,41,42 and the cellular processes that lead to transmission breakdown under natural conditions are generally unresolved21,36. Understanding why transmission breaks down under certain conditions is critical for predicting endosymbiont spread in nature and improving Wolbachia-based biocontrol programs25,33,34,43.
Maternally transmitted Wolbachia are the most common endosymbionts on earth, associating with about half of all insect species, as well as other arthropods and nematodes4,44,45,46. Wolbachia are primarily maternally (vertically) transmitted via the host germline within host species; although, host switching via horizonal transfer between host species is common on evolutionary timescales47,48,49,50,51,52,53,54. Wolbachia generally form facultative relationships (from the host perspective) in insects, with some individuals in the host population that do not carry Wolbachia due to imperfect maternal transmission. Maternal transmission is perhaps the most important determinant of Wolbachia prevalence in host populations: theory demonstrates that Wolbachia frequency dynamics and equilibria are approximated by the degree of imperfect maternal transmission (μ), the relative fitness (e.g., fecundity) of females with Wolbachia (F), and the strength of Wolbachia-induced cytoplasmic incompatibility (sh)17,55. Some Wolbachia strains (e.g., wRi in D. simulans) cause cytoplasmic incompatibility (CI), a crossing incompatibility that generates a frequency-dependent advantage for females with Wolbachia5,56.
The prevalence of different Wolbachia strains varies widely among host systems, which may be partially explained by differences in maternal transmission rates. For instance, the strong CI-causing wRi strain is observed at high frequencies in global D. simulans populations17,18,57, whereas the weak CI-causing wMel strain is found at intermediate, fluctuating frequencies in D. melanogaster58. wRi frequencies can be plausibly explained by strong CI that causes spread to high frequencies, whereas lower, variable wMel frequencies can be plausibly explained by maternal transmission breakdown as temperatures decrease, in addition to a minimal influence of CI21. The thermal sensitivity of maternal transmission may be a feature of wMel, given that heat stress also reduces wMel transmission in transinfected Aedes aegypti hosts33,34,43. Identifying Wolbachia variants that are efficiently transmitted across environmental conditions could potentially contribute to improving the efficacy of Wolbachia based biocontrol of human diseases and agricultural pests25,26.
What might underlie variable maternal transmission rates? Maternal transmission breakdown is predicted to stem from reductions in Wolbachia abundance and localization throughout host development and specifically at the critical stages of oogenesis thought to facilitate germline-to-germline maternal transmission14,15,21,36,59,60,61,62,63. wMel cells are present in the germline stem cells of adult females, and over the course of oogenesis, wMel localize via kinesin-mediated transport to the pole plasm—the site of germline formation—at the posterior cortex of the developing oocyte. In late oogenesis (stage 10), wMel Wolbachia display a pattern of posterior localization in oocytes, which is considered to be a key step in the process of germline-to-germline maternal transmission under standard laboratory conditions14,50,59,60,61,62,63. In support of this, we recently demonstrated that declining wMel transmission in the cold covaries with a significant reduction in cellular Wolbachia abundance at the posterior cortex of stage 10 oocytes, which can plausibly explain why maternal transmission is thermally sensitive21.
Outside of the wMel strain, it is unclear how temperature influences Wolbachia maternal transmission and posterior localization in stage 10 oocytes for other diverse Wolbachia-host associations15,20. Under standard 25 °C conditions, divergent Wolbachia strains exhibit highly variable patterns of localization to the pole plasm at the posterior cortex of stage 10 oocytes of Drosophila species, deviating from our canonical understanding of wMel15. These patterns range from strongly localized (e.g., wAna in D. anannassae) to a complete lack of localization (e.g., wTri in D. triauraria) at the site of germline formation. Some strains, particularly the divergent B-group wMau strain, also exhibit evidence for an alternative soma-to-germline route of transmission, whereby Wolbachia in somatically derived follicle cells may invade the oocyte via cell-to-cell migration. It is entirely unknown whether the diverse localization patterns of these Wolbachia strains influence rates of maternal transmission and/or are sensitive to thermal perturbation like wMel.
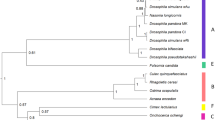
Here, we examine temperature effects on Wolbachia maternal transmission and localization in host tissues across 10 divergent Wolbachia strains naturally found in eight Drosophila host species within the melanogaster species group (Fig. 1, Table S1). These Wolbachia comprise nine A-group Wolbachia—including closely related wMel-like strains (wMel, wMelCS, wSeg in D. seguyi, and wCha in D. chauvacae) and closely related wRi-like strains (wRi, wAna in D. ananassae, wAura in D. auraria, and wTri in D. triauraria)20,51,53—and divergent B-group wMau in D. mauritiana that diverged up to 46 million years ago from the other A-group strains40. We reared flies at 25° and 20 °C and then measured maternal transmission rates in conjunction with Wolbachia densities (titer) at the tissue level in female ovaries and the remaining somatic tissues. We then used confocal microscopy to examine how temperature impacts the cellular abundance of Wolbachia at the site of germline formation at the posterior cortex of stage 10 oocytes during oogenesis. Our results reveal diverse patterns of localization in ovary tissues and late-stage oocytes that are temperature-dependent, but only weakly predictive of maternal transmission rates. We also identify Wolbachia strains that are transmitted efficiently regardless of temperature.
a Estimated Bayesian phylogram of the 10 A- and B-group Wolbachia strains included in the study. The phylogram was estimated using 170 single-copy genes of identical length in all genomes, spanning 135,105 bp. b Estimated Bayesian phylogram of the eight Drosophila species using 20 single-copy genes. All nodes on both trees are supported with Bayesian posterior probabilities of 1. Estimates of Wolbachia and Drosophila divergence reported in millions of years ago (MYA) are reproduced from Meany et al.40 and Suvorov et al.135, respectively. The Wolbachia and host trees are generally discordant, as expected with frequent Wolbachia host switching47,48,51,53,54.
Results
We estimated the mean rate of maternal transmission (±BCa confidence intervals) for each Wolbachia-host genotype at 25° and 20 °C by pairing individual virgin females with Wolbachia-free males (see Methods), allowing the females to lay for 8 days, and then measuring maternal transmission (1 – μ) to the newly emerged offspring of each subline (Fig. 2, Table S2)20,21. The final dataset comprised 365 F0 females and 3612 F1 offspring. Overall, we found a significant interaction effect between the Wolbachia-host genotype and temperature (GLMM LRT, χ2(9) = 23.49, P = 0.005), such that temperature effects on maternal transmission varied depending on the Wolbachia-host genotype. For most Wolbachia-host genotypes, mean rates of maternal transmission were perfect (e.g., wRi-D. simulans) or near-perfect (e.g., wMel-like wSeg-D. seguyi) and invariant across the two temperatures (Fig. 2, Table S3), with the exception of a few specific Wolbachia-host genotypes. Maternal transmission of wMel by D. melanogaster (W = 251.5, P = 0.003) and wAura by D. auraria (W = 154, P = 0.038) decreased significantly at 20 °C; although, wMel experienced a much greater decline in the average rate of transmission (14.9%) than wAura (2.8%). In contrast, B-group wMau transmission by D. mauritiana increased slightly by 3.4% in the cold at 20 °C (W = 117, P = 0.019). Notably, our analysis also revealed a significant effect of sex (GLMM LRT, χ2(1) = 63.71, P < 0.001), whereby rates of maternal transmission tended to be lower to female offspring than males (Table S2). This finding is consistent with our previous findings for wMel21 and the wMel-like wYak strain in D. yakuba20, but counter to theoretical expectations that selection should favor faithful transmission to female offspring64,65.
Mean maternal transmission rates (±BCa confidence intervals) of different Wolbachia strains in their naturally associated host species (N = 364 sublines). Asterisks indicate the rates of maternal transmission differ between 25° and 20 °C for a given Wolbachia strain according to a Wilcoxon rank-sum test at P < 0.05. Below, the cladogram depicts evolutionary relationships among Wolbachia strains.
To test for temperature effects on Wolbachia at the tissue level, we dissected out the ovaries from a subset of the same F0 females assessed for maternal transmission above (N = 183). For this analysis we used qPCR to measure Wolbachia densities in the ovaries and the remaining somatic carcass tissue (Fig. 3). We found a significant interaction effect between the Wolbachia-host genotype and temperature on Wolbachia density in ovaries (Two-way ANOVA; F(9,161) = 4.286, P < 0.001) and the remaining carcasses (F(9,161) = 10.612, P < 0.001), such that temperature effects on Wolbachia density varied depending on the Wolbachia-host genotype. While we uncovered considerable variation depending on the Wolbachia-host genotype and temperature, Wolbachia ovary densities generally tended to decline at 20 °C for the majority of systems (Fig. 3, Table S4). Five of the Wolbachia strains (wMel, wRi, wAna, wHa, and wMau) decreased in density in the ovaries in the cold. In contrast, carcasses exhibited variation in the directionality of changes in Wolbachia density. wMel, wMelCS, wSeg, and wAura densities decreased significantly in the cold, whereas wCha and wMau significantly increased. wCha densities in carcasses were roughly an order of magnitude lower than all other strains, regardless of temperature, suggesting a relatively high degree of localization to the ovaries. wMel, the strain that experienced the most dramatic decline in transmission in the cold (Fig. 2), was also the only Wolbachia strain that experienced a significant decline in Wolbachia density in both the ovaries and the carcasses in the cold. This included a particularly large decline in density in the carcasses (a 4.7-fold reduction) compared to the other strains.
Wolbachia densities are shown for dissected female ovaries and the remaining somatic carcass tissue (N = 183 F0 females). Asterisks indicate the Wolbachia density differs between 25° and 20 °C for a given Wolbachia strain according to a Wilcoxon rank-sum test at P < 0.05. Boxplots show medians, first and third quartiles (hinges), and the smallest/largest values within 1.5*IQR of the hinges (whiskers).
For the three Wolbachia-host systems with temperature-dependent transmission (wMel, wAura, and wMau), we also tested for a correlation between transmission rates and Wolbachia density in the ovaries and carcasses of the same individual females (Fig. S1). Declining wMel transmission in the cold was significantly correlated with Wolbachia density in the ovaries (Spearman’s ρ = 0.537, P = 0.022), but not the carcasses (ρ = 0.392, P = 0.108), providing additional support that reduced wMel density in reproductive tissues contributes to transmission breakdown in the cold. Maternal transmission of wAura by D. auraria did not strongly correlate with wAura density in the ovaries (ρ = −0.244, P = 0.380) or the carcasses (ρ = 0.035, P = 0.902). Finally, increasing maternal transmission of wMau by D. mauritiana in the cold did not correlate with wMau density in the ovaries (ρ = −0.119, P = 0.649), but was significantly correlated with density in the carcasses (ρ = 0.549, P = 0.022).
Finally, we examined how temperature impacts cellular Wolbachia abundance at the site of germline formation in stage 10 oocytes during host oogenesis. At this stage, wMel Wolbachia localization at the posterior cortex of the oocyte is considered to be a critical step in the process of germline-to-germline Wolbachia maternal transmission14,15,21,36,59,60,61,62,63. We imaged a total of 283 stage 10 oocytes from 8-day-old females reared at the two different temperatures and quantified cellular Wolbachia abundance (measured as corrected total cell fluorescence; CTCF) in the whole oocytes, the posterior region, and the posterior cortex (Fig. S2)14,15,21. We defined the posterior region as the posterior 1/8th portion of the oocyte and the posterior cortex as the narrow cortical region of Vasa expression (a germline protein component)15,63. For the sake of clarity and consistency with the literature, we refer to our Wolbachia quantifications with confocal microscopy here as “cellular Wolbachia abundance” in oocytes, as opposed to the quantification of “Wolbachia density” at the tissue level using qPCR (described above).
Cellular Wolbachia abundance in whole oocytes (two-way ANOVA; F(9,263) = 2.9, P = 0.003), at the posterior region (F(9,263) = 4.16, P < 0.001), and at the posterior cortex (F(9,263) = 2.41, P = 0.012) depended on significant two-way interactions between the Wolbachia-host genotype and temperature. As with Wolbachia density, cellular Wolbachia abundance in oocytes varied considerably depending on the Wolbachia-host genotype and temperature (Fig. 4, Fig. S3). The cellular abundance of six of the Wolbachia strains (wMelCS, wSeg, wAna, wAura, and wTri, and wHa) declined significantly throughout whole oocytes in the cold (Table S5). Temperature affected fewer strains (only wMel, wMelCS, wRi, and wAura) at the posterior region and the posterior cortex, the location of the developing germline. wMel and wAura, the two strains transmitted at significantly lower rates in the cold, both declined significantly in abundance in the posterior oocyte region (W = 113, P = 0.015; W = 172, P = 0.041). wMel cellular abundance also significantly declined at the posterior cortex (W = 122, P = 0.002). Interestingly, wMelCS also exhibited a significant decline in cellular abundance in the posterior region (W = 203, P < 0.001) and the cortex (W = 205, P < 0.001), despite the fact that wMelCS transmission by D. melanogaster was not altered in the cold (Fig. 2). This can perhaps be explained by the fact that, of all 10 strains, wMelCS had the highest cellular abundance at the posterior region and cortex, regardless of temperature. Finally, wRi was perfectly transmitted by D. simulans at both temperatures, but wRi abundance increased at the oocyte posterior region (W = 45, P = 0.004) and the cortex (W = 49, P = 0.008) in the cold. Also of note, wAura and wTri occurred at particularly low abundances in the oocyte posterior region and the posterior cortex, regardless of temperature, despite both being transmitted at relatively high rates by their female hosts (1 – μ > 0.967).
Cellular Wolbachia abundance in stage 10 oocytes (measured as fluorescence due to propidium iodide; CTCF), measured in a the whole oocyte, b the posterior region, and c the posterior cortex (N = 283 oocytes). Asterisks indicate that Wolbachia abundance differs between 25° and 20 °C for a given Wolbachia strain according to a Wilcoxon rank-sum test at P < 0.05. d Representative confocal images of Wolbachia strains that have decreased abundance at the posterior cortex in the cold (wMel), increased abundance at the cortex in the cold (wRi), and no change in the cold (wCha). Confocal micrographs are DNA-stained with PI (red) and actin-stained with phalloidin (green). The second column depicts a single channel image of the PI stain and the third column depicts an enlarged PI-stained image at the posterior cortex of each oocyte. Scale bars are set to 25 µm.
Our analyses revealed that the diverse Wolbachia strains and Drosophila host species exhibit extensive variation in Wolbachia density in host tissues (Fig. 3) and abundance in host oocytes (Fig. 4). We hypothesized that this diversity may be explained by factors in the Wolbachia or host genomes (or both). If so, we expect closely related Wolbachia strains (or host species) to exhibit similar patterns of Wolbachia localization in host tissues15,66,67. We used the Wolbachia and host trees (Fig. 1) to test whether patterns of Wolbachia localization in tissues and oocytes exhibit phylogenetic signal using Pagel’s λ68. A value of 1 is consistent with trait evolution that entirely agrees with the Wolbachia (or host) phylogeny, suggesting factors in the Wolbachia genome (or host) contribute to Wolbachia localization patterns (i.e., strong phylogenetic signal). In contrast, a value of 0 is consistent with trait evolution that occurs independently of phylogenetic relationships68,69. We found no evidence that Wolbachia density in the ovaries and carcasses, nor cellular abundance in the oocytes, exhibit phylogenetic signal on the Wolbachia phylogeny (Table S6, Fig. S4), implying that factors in the Wolbachia genome do not account for observed diversity. In contrast, we found that Wolbachia density in the ovaries at 25 °C exhibits strong, significant phylogenetic signal on the host phylogeny (λ = 1.000, P = 0.001), implying that factors in the host genome may help determine observed diversity of Wolbachia densities in host ovaries. In this instance, related species in the montium species group tended to have lower Wolbachia densities in the ovaries relative to the melanogaster subgroup (see Fig. S5). Divergent D. ananassae also had a higher Wolbachia density than all the other host species. We also found that cellular Wolbachia abundance in the oocyte posterior region (λ = 1.000, P = 0.037) and the posterior cortex (λ = 1.000, P = 0.036) exhibited a significant signature of phylogenetic signal at 20 °C on the host phylogeny. This pattern is largely driven by the relatively low abundance of Wolbachia in the posterior oocytes of the two closely related species D. triauraria and auraria.
Discussion
Maternal transmission rates are a key determinant of Wolbachia prevalence in host populations17,55. For many Wolbachia strains and host species, we found that maternal transmission is generally high and stable across two ecologically relevant temperatures, 25° and 20 °C, that flies are likely to experience in natural environments21,70. Two Wolbachia strains (wAura and B-group wMau) exhibited minor temperature-dependent changes in transmission, but wMel stood out with a relatively large decrease in transmission as temperature dropped from 25° to 20 °C (Fig. 2). Maternal transmission is generally expected to depend on Wolbachia density in host reproductive tissue20,34,35,43, and specifically within developing oocytes14,15,21,36,59,60,61,62,63. Our analyses uncovered considerable, significant variation in Wolbachia densities at the tissue level and cellular abundances in oocytes, which also often varied depending on temperature. With the exception of wMel, Wolbachia densities in ovary tissue generally did not predict maternal transmission rates (Fig. 3, Fig. S1). In stage 10 oocytes, declines in the cellular abundance of wMel and wAura at the posterior region coincided with declining transmission in the cold (Fig. 4), but the wMelCS and wRi strains also experienced changes in cellular abundance that did not correlate with differences in maternal transmission. Together, these results suggest that maternal transmission rates cannot be accurately predicted from measurements of Wolbachia quantities in ovary tissue or late-stage oocytes. First, we explore the diverse patterns of Wolbachia localization in relation to maternal transmission in our comparative analyses. Then, we examine why maternal transmission of some strains, particularly wMel, may be impacted by temperature.
At the tissue level, we found that Wolbachia densities in the ovaries generally did not predict rates of maternal transmission, with the exception of wMel. wMel density in the ovaries of individual females was tightly correlated with declining transmission rates in the cold (ρ = 0.537, Fig. S1). Temperature did not alter wAura density in the ovaries despite declining wAura transmission in the cold, and wMau density counterintuitively declined at 20 °C despite transmission increasing. Other strains (e.g., wRi and wHa in D. simulans) experienced large declines in Wolbachia ovary density at 20 °C with no impact on maternal transmission. Wolbachia densities in the remaining somatic carcass tissue also generally did not predict declining transmission in the cold; however, wMau densities in the carcasses of individual females were strongly correlated with increased transmission in the cold (ρ = 0.549; discussed further below). Notably, wCha was exceptional in terms of its very low density in carcass tissue relative to other strains, despite perfect transmission at both temperatures. Changes to host ploidy in the ovaries could plausibly contribute to the observed differences in relative Wolbachia density71; however, we did not find evidence for significant changes in host gene copy number in the ovaries across temperature treatments (Fig. S6).
At the cellular level, Wolbachia abundance in stage 10 oocytes may help explain why wMel and wAura experienced declining transmission in the cold. Many strains experienced declines in cellular Wolbachia abundance throughout the whole oocyte at 20 °C (Fig. 4a), but this was generally unrelated to transmission rates. wMel was one of the few strains that had a significant reduction in cellular abundance at the posterior region (Fig. 4b) and cortex (Fig. 4c), the site of germline formation. Similarly, wAura had a reduced abundance in the posterior region at 20 °C (but not at the cortex). wMelCS (closely related to wMel) was the only additional strain with significant reductions in abundance at the posterior region and the cortex. However, wMelCS occurred at a much higher absolute abundance than wMel, wAura, and all other Wolbachia strains, which may explain why transmission remained high at 20 °C (discussed further below). We conjecture that there is presumably a minimum cellular abundance required for efficient transmission, and Wolbachia strains like wMelCS that are generally very abundant are less likely to fall below this threshold. A reduction in cellular abundance at the oocyte posterior implies that fewer Wolbachia cells will be incorporated into the germline later during host development, perturbing germline-to-germline maternal transmission21. The fact that both wMel and wAura had significantly reduced abundances in the posterior region at 20 °C implies that perturbations of cellular Wolbachia abundance in this region may help explain why some Wolbachia strains have reduced transmission in the cold. wRi was the only other Wolbachia strain with an altered abundance at the oocyte posterior region and cortex, but cellular abundance increased at 20 °C. This may help explain why maternal transmission of wRi remained perfect at 20 °C. Interestingly, a low absolute abundance of Wolbachia was not necessarily associated with declining maternal transmission. The wTri strain (closely related to wAura) occurred at a lower posterior abundance than all other strains, despite maintaining a high rate of transmission at both temperatures.
It is important to note that our analysis of Wolbachia in ovary tissues and stage 10 oocytes of 8-day-old females represents only a snapshot of host development. Many other unexplored factors may contribute to transmission rate variation. Other stages of development, like embryogenesis, could also be involved in mediating maternal transmission. For instance, temperature could impact Wolbachia localization at the newly formed germline of late blastoderm and cellularized embryos, which could influence maternal transmission15,60. Cellular Wolbachia abundance changes over the course of oogenesis and embryogenesis15,61,71,72 and a number of studies report complex, age-related changes in Wolbachia densities over the course of the host’s lifespan73,74,75,76,77,78,79,80. Maternal age and the order of egg-laying could also conceivably influence Wolbachia transmission and densities in offspring81. Our work motivates future studies on how temperature affects the distribution of Wolbachia in host tissues over the course of host development in relation to maternal transmission.
Our comparative analysis highlights how wMel seems to be especially susceptible to cooling temperatures21, which is consistent with our previous work suggesting the strain is less prevalent in temperate host populations due to declining transmission in the cold21,58. In transinfected mosquitoes, wMel maternal transmission and densities are also more susceptible to heat stress relative to another Wolbachia strain, wAlbB25,26,33,34,43. A number of factors could plausibly help explain why wMel maternal transmission is especially impacted by the cool temperature, relative to other strains. At the tissue level, wMel was the only Wolbachia we examined that exhibited a significant decline in Wolbachia density in both the ovaries and the carcasses at 20 °C (Fig. 3). Declining transmission was highly correlated with declining wMel density in the ovaries, plus wMel stood out with a dramatic decline in Wolbachia density in carcasses at 20 °C compared to other strains. While wMel is generally thought to follow a germline-to-germline route of maternal transmission during oogenesis, our recent work and others suggests that the strain may also rely on a soma-to germline mode of transmission15,82,83. wMel Wolbachia are capable of cell-to-cell migration83 and wMel injected directly into the abdomen of adult D. melanogaster females can migrate to and occupy the germline and follicle stem cells82. The declines of wMel densities throughout host tissues could potentially impact routes of transmission originating in somatic tissue. At the cellular level, wMel is also one of the few strains (in addition to wAura) to experience a decline in abundance at the posterior of stage 10 oocytes, which is expected to impact a germline-to-germline route of transmission. The concurrent reduction in Wolbachia density in female tissues and oocytes may be a one-two punch that impacts both soma- and germline-to-germline routes of wMel transmission.
The juxtaposition between wMel and wMelCS is interesting, because the two strains diverged recently in only the last 3000–14,000 years84. The wMel and wMelCS strains we used in our study are highly similar (0.007% third-position pairwise differences). Despite the close relationship, wMelCS transmission was not impacted by temperature and the strain occurred at a higher cellular abundance at the oocyte posterior than any of the other strains that we examined, regardless of temperature. Some oocytes contained especially large quantities of wMelCS throughout the whole oocyte and at the posterior (Fig. S7), as compared to all the other oocytes we examined (e.g., Fig. S3). wMelCS is only found at low frequencies in a few populations of D. melanogaster74,84, which is attributed to the fact that ancestral wMelCS was largely replaced by wMel in global populations of D. melanogaster in roughly the last 50 to 100 years85,86,87,88. Perhaps deleterious host effects associated with a high cellular wMelCS abundance in oocytes may have contributed to the global replacement of wMelCS by wMel. To ensure maternal transmission, Wolbachia cells must maintain an association with the host germline without perturbing highly conserved processes of germline and oocyte development15,62,63,72. Migration of wMel to the posterior germ plasm is coincident with recruitment of host factors required for germline formation and anterior/posterior and dorsal/ventral axis formation62 and an excessively high Wolbachia abundance can disrupt dorsal/ventral axis determination60,62,72,89. Perhaps similar deleterious effects may help explain why the highly abundant wMelCS strain was replaced by wMel, despite a higher rate of transmission in the cold. Previous works has shown that higher densities of wMelCS in D. melanogaster often covaries with a reduced lifespan74. The cellular abundance and fitness consequences of wMelCS (as compared to wMel) may also be influenced by interactions with the host genome and the environment20,21,90. Indeed, our phylogenetic analyses suggest that the host genome contributes to Wolbachia abundance in oocytes (Table S6). Given that interactions with Wolbachia and the environment influence host fitness (e.g., fecundity and longevity), it is difficult to point to a single explanation for why wMelCS was globally replaced by wMel.
The divergent B-group wMau strain was the only strain where transmission significantly increased at 20 °C (Fig. 2). Transmission was high at 25 °C and increased to perfect at 20 °C. Our previous work suggests the wMau strain may also be an outlier in its mode of transmission during oogenesis relative to A-group Wolbachia, which tend to exhibit evidence of a germline-to-germline route of transmission15. wMau occurs at relatively low abundance at the posterior cortex of stage 10 oocytes (Fig. 4), but wMau cells are frequently found within the somatically derived follicle cells surrounding the oocytes (Fig. S7)15, which is consistent with a soma-to-germline route of transmission. The presence of Wolbachia in the follicle cells is generally not the case for A-group Wolbachia (with the exception of wMel, discussed above). We found that wMau was also one of the few strains (in addition to wCha) where Wolbachia density in the somatic carcass tissues significantly increased at 20 °C (Fig. 3), which may help explain why transmission increased slightly at 20 °C. wMau transmission was also strongly correlated with Wolbachia densities in the carcasses of individual females. Notably, our previous analysis of two other wMau-D. mauritiana genotypes using 3- to 5-day-old females found perfect rates of transmission at 25 °C40, unlike our results here (Fig. 2). This suggests that differences in fly age80, the Wolbachia and host genomes21, or other unknown factors may also contribute to variation in rates of maternal transmission.
The wMau findings also may inform our previous work on the thermoregulatory behavior of Drosophila host species. We found that Drosophila species carrying A-group Wolbachia, including wRi and wHa, tend to prefer cooler temperatures on a thermogradient (relative to flies without Wolbachia), whereas D. mauritiana flies carrying wMau prefer warmer temperatures66. Taken together, these results suggest that fly movement to cool temperatures reduces the density of A-group Wolbachia in host tissues, whereas movement to warm temperatures reduces B-group wMau density. This is consistent with the inference that flies may thermoregulate as a behavioral response to ameliorate negative effects of infection and a high Wolbachia density67,91,92,93,94. Movement to cooler temperatures with A-group Wolbachia (i.e., behavioral chill) and to warmer temperatures with B-group wMau (behavioral fever) are both behaviors that would reduce Wolbachia density in host tissues according to our results here (Fig. 3). This finding motives further work dissecting the relationships among thermoregulatory behavior, Wolbachia density in host tissues, and maternal transmission.
Our phylogenomic analyses revealed that Wolbachia density in the ovaries at 25 °C and cellular Wolbachia abundance in the oocyte posterior region and cortex at 20 °C all exhibit phylogenetic signal on the host phylogeny (Table S6, Fig. S5), implying that factors in the host genome help explain the observed diversity in Wolbachia localization across species. For example, interspecific variation in the host proteins that Wolbachia engage during oogenesis could plausibly influence Wolbachia localization patterns at the posterior cortex in stage 10 oocytes15. Unfortunately, it is not yet known how Wolbachia engage host factors like kinesin during oogenesis63. Other work also suggests the host genome can influence Wolbachia abundance in oocytes21 and embryos60. Similarly, genome-wide screens reveal that host factors play an important role in determining cellular Wolbachia abundance95,96. The findings here complement our previous comparative analyses that suggest the Wolbachia genome also contributes to variation in Wolbachia localization patterns in oocytes15 and effects on host thermoregulatory behavior66. While we did not identify any traits that exhibit phylogenetic signal on the Wolbachia phylogeny, it is worth noting that relatively few Wolbachia genomic changes (e.g., in the ampliconic “Octomom” gene region) are required to influence density and tissue distributions74,97,98,99,100. This highlights the potential for rapid changes in these traits and their effects on transmission. DNA analyses reveal discordant Wolbachia and host phylogenies, with recently diverged Wolbachia found in distantly related hosts47,48,49,50,51,52,53,54,101. This includes evidence of the rapid spread of wRi-like Wolbachia across Drosophila flies diverged about 50 MYA51 and wMel-like Wolbachia across holometabolous insects102 diverged about 350 MYA53. Given the short persistence of Wolbachia with hosts in our study (see Fig. S8), it is unlikely that the observed patterns of Wolbachia transmission and tissue localization result from coevolution between Wolbachia and host genomes.
The rate of Wolbachia maternal transmission, in conjunction with Wolbachia effects on host fitness and cytoplasmic compatibility (CI), ultimately determine Wolbachia prevalence in host populations17,55. In this study, we uncovered limited variation in maternal transmission rates across temperatures, despite dramatic variation in Wolbachia localization patterns within host tissues. Nonetheless, a few Wolbachia strains, particularly wMel, had variable transmission rates that are likely to impact Wolbachia prevalence in natural host populations. For instance wMel tends to cause weak CI80,103,104 and occurs at lower frequencies in temperate host populations in Australia and North America, which can plausibly be explained by declining maternal transmission at cool temperatures (Fig. 2)21. Similarly, wMau does not cause CI and occurs at intermediate frequencies in D. mauritiana on Mauritius40,105. Our results suggest temperature-related variation in wMau transmission may perhaps help contribute to intermediate frequencies. Supporting our prior observations20,21, Wolbachia transmission rates to female offspring were lower than to male offspring (Table S2), which could influence sex-specific Wolbachia frequencies observed in nature. Future work focused on understanding the causes of sex-specific transmission rates and Wolbachia densities will be important for understanding this pattern.
Other Wolbachia strains like wRi17,18,19,106, wHa107,108,109, and wAna110,111 cause strong CI and tend to occur at high equilibrium frequencies in nature. In addition to strong CI, our results suggest that stable transmission rates may help these strains maintain high frequencies in the face of fluctuating conditions (Fig. 2). However, this may not always be the case, as we found that maternal transmission the strong CI-causing strain wAura51,110 can be perturbed by temperature. Unlike CI and Wolbachia maternal transmission, the role of Wolbachia effects on host fitness in natural Drosophila populations is poorly understood, although recent work suggests Wolbachia block viruses in their native Drosophila hosts8,9,112,113.
Understanding how temperature influences Wolbachia maternal transmission, especially in relation to other key factors like CI and host fitness effects, is critical for explaining global Wolbachia prevalence in host populations. Research on this front will also inform Wolbachia-based biocontrol programs, as thermally stable Wolbachia strains can be leveraged to improve the efficacy of Wolbachia-based biocontrol applications in mosquito populations that experience extreme environments25,114. In this respect, temperature and the environment are emerging as key factors that mediate interactions between Wolbachia, their hosts, and pathogens.
Methods
Fly lines
We evaluated 10 different Wolbachia strains that naturally occur in eight different Drosophila species (Table S1). For two of these host species, we tested multiple Wolbachia-host genotypes: wMel and wMelCS in D. melanogaster and wRi and wHa in D. simulans. With the exception of the wMelCS-D. melanogaster line, all the Wolbachia-host genotypes were sampled from nature to form isofemale lines, such that single gravid females were collected from the field and placed individually into vials (see Hague et al.66 for further discussion of the wMelCS genotype). Each isofemale line was stably maintained in the lab for at least four years prior to the experiments. For the maternal transmission experiments explained below, we paired each Wolbachia-positive female with Wolbachia-free males of the same host species (Table S1). Here, we used males from naturally Wolbachia-free isofemale lines whenever possible; however, in some cases, we were unable to obtain natural genotypes without Wolbachia. For these species, Wolbachia-free genotypes were created by treating the Wolbachia-positive genotype with 0.03% tetracycline for at least four generations. After the fourth generation, we used quantitative PCR (qPCR) to confirm that flies were cleared of Wolbachia66. We then reconstituted the gut microbiome of the tetracycline-cleared flies by rearing them on food where Wolbachia-positive males of the same genotype had been fed and defecated for the prior 48 h. Tetracycline-cleared flies were given at least three more generations before we conducted experiments to avoid detrimental effects of the antibiotic treatment on mitochondrial function115.
Maternal transmission
We tested how temperature influences maternal transmission and Wolbachia distributions in host tissues by rearing females of each genotype at a moderate (25 °C) and low (20 °C) temperatures. Prior to experiments, the isofemale lines were reared for at least two generations in incubators set to each temperature and a 12L:12D light cycle (Percival Model I-36LL) on a standard food diet66. We estimated maternal transmission at each temperature by aspirating individual virgin females into vials with two 3–5-day-old virgin males of the Wolbachia-free genotype and then allowing females to lay eggs for 8 days20,21. We crossed Wolbachia-positive females to males without Wolbachia, because in crosses with Wolbachia-positive males, Wolbachia-free ova produced by Wolbachia-positive females could be lost if they are susceptible to cytoplasmic incompatibility, potentially leading to an overestimation of maternal transmission18. The males in our experiments were all collected from the Wolbachia-free genotypes that were maintained at 25 °C.
Newly emerged adult F1 offspring were collected in the first 48 h of emergence, preserved in 95% ethanol, and stored at −20 °C for later analysis of Wolbachia status. For each subline, we measured maternal transmission by screening five female and five male F1s for Wolbachia using PCR. We extracted DNA from individual F1s using 96-well plates and a “squish” extraction buffer (10 mL Tris-HCl [1 M], 0.0372 g EDTA, 0.1461 g NaCl, 90 mL dH2O, followed by 150 mL Proteinase K after autoclaving) that allows for high-throughput and cost-effective DNA extraction. We screened each F1 for Wolbachia status using previously described PCR primers for the Wolbachia surface protein (wsp) and a second set of primers for the arthropod-specific 28S rDNA that served as a positive control20,40,116.
We computed maternal transmission for each genotype at each temperature as the mean proportion of Wolbachia-positive offspring produced by Wolbachia-positive mothers in each subline. We then estimated 95% bias-corrected and accelerated bootstrap (BCa) confidence intervals using the “boot.ci” function and 5000 iterations in the boot package in R117,118. We analyzed whether maternal transmission rates vary by Wolbachia-host genotype (e.g., wMel-D. melanogaster) and temperature using a generalized linear mixed model (GLMM) logistic regression with the “glmer” function in the lme4 R package119. We treated the Wolbachia status of F1s as the dependent variable and then included sex, the Wolbachia-host genotype, temperature, and a two-way interaction between the Wolbachia-host genotype and temperature as independent variables. We included sex in the model because we previously found that wMel-like Wolbachia (wMel and wYak) are transmitted more efficiently to male offspring20,21. Finally, we included the subline of each F1 as a random effect. We assessed significance of the fixed effects with a likelihood ratio test (LRT) and type III sum of squares using the “mixed” function in the afex package120. Lastly, we used two-sided Wilcoxon rank-sum tests to evaluate whether transmission rates differ between 25° and 20 °C for each strain.
Wolbachia density in host ovaries
After each female was allowed to lay for 8 days, we immediately dissected out her ovaries in chilled 1X PBS. Both the ovaries and remaining carcass were then frozen at −80 °C. We then used qPCR to quantify Wolbachia density in the dissected ovaries and carcasses of each individual female. DNA was first extracted using a DNeasy Blood and Tissue Kit (Qiagen) and then amplified the Wolbachia-specific locus ftsZ and the Drosophila-specific locus nAcRalpha-34E. Preliminary analyses indicated that separate primers were needed for Wolbachia in the divergent A-Group (F: 5’-ATCCTTAACTGCGGCTCTTG-3’, R: 5’-TTCATCACAGCAGGAATGGG-3’) and B-Group lineages (F: 5’-CAGAGAAGCAAGAGCGGTAG-3’, R: 5’-TCTTCAAGTCCAAGCTCTGC-3’). We were able to use a single set of primers for all the Drosophila hosts (F: 5’-CTATGGTCGTTGACAGACT-3’, R: 5’-GTAGTACAGCTATTG TGGC-3’). We generated efficiency curves to confirm that each primer pair amplified with adequate efficiency for each Wolbachia strain and host species. The individual 10 μl reactions included 5 μl PowerUp SYBR Green Master Mix (Applied Biosystems), 0.25 μl of the forward and reverse primers, 0.5 μl of water, and 4 μl of gDNA. All qPCR reactions were amplified using the following cycling conditions: 50 °C for 2 min, 95 °C for 2 min, and then 40 cycles, with one cycle consisting of 95 °C for 15 s, 58 °C for 15 s, and 72 °C for 1 min. We used the average cycle threshold (Ct) value of three technical replicates for each sample. We then estimated relative Wolbachia density as 2∆Ct, where ∆Ct = CtnAcRalpha-34E − CtftsZ121. We used a two-way ANOVA and type III sums of squares to test whether log-transformed Wolbachia density varied among Wolbachia-host genotypes and temperature for both the ovaries and the carcasses. We also used two-sided Wilcoxon rank-sum tests to test whether densities differed between 25° and 20 °C for each tissue and Wolbachia-host genotype. If endoreplication in nurse cells is altered by temperature, changes to host ploidy in ovary tissues could plausibly influence our estimates of relative Wolbachia density (2∆Ct) at each temperature71. We evaluated the raw Ct values for evidence of temperature differences in copy number at the host locus nAcRalpha-34E (Fig. S6) and found that host ovary Ct values did not change significantly between temperature treatments, implying that differences in relative Wolbachia density are due to changes in absolute abundance of the Wolbachia gene.
Cellular Wolbachia abundance in host oocytes
We used previously described protocols and confocal imaging to quantify cellular Wolbachia in host oocytes at each temperature15,21. Briefly, we paired 20 newly emerged virgin Wolbachia-positive females en masse with ten 3–5-day-old virgin Wolbachia-negative males in individual vials for 8 days to stimulate oocyte production. Females were then separated using CO2 and ovaries were dissected in a chilled dish of 1X PBS. Ovaries were fixed in 200 μl of devitellinizing solution (2% paraformaldehyde and 0.5% v/v NP40 in 1X PBS) mixed with 600 μl of heptane for 20 min on a shaker at room temperature. We then removed the organic layer using a brief centrifugation and washed the ovaries three times with PBS-T (0.1% Triton X-100 in 1X PBS), followed by three additional 5 min washes. Samples were treated with RNAse A (10 mg/ml) and left overnight at room temperature. The samples were then washed in PBS-T multiple times over 2 h and then stained with a dilute solution of Alexa 488 phalloidin on a shaker for 2 h for actin staining. Subsequently, the samples were washed again multiple times over the course of two more hours. Finally, the wash solution was removed and we added 60 μl of propidium iodide (PI) in mounting media (VECTASHIELD) and left again overnight. Ovaries were then mounted and carefully separated out again for ease of imaging. Slides were sealed with a coat of nail polish and stored at −20 °C until imaging.
We used a Zeiss Olympia confocal microscope and the 40× Plan-Apochromat oil objective for image acquisition. Alexa 488 was imaged using the 488 nm laser line and emissions were collected from 493 to 584 nm. Propidium iodide was imaged using the 488 and 561 nm lines and emissions were collected from 584 to 718 nm. Stage 10 oocytes were identified based on two criteria. First, the egg chamber had to have two compartments, one with nurse cells and the other with the developing oocyte. Second, the oocyte had to account for ~50% of the egg chamber. The top and bottom of the oocyte were identified by scanning through the egg chamber and then a 0.8-μm slice was taken around the midplane. The pixel dwell time was 8.19 μsec.
Image analysis was conducted in the program Fiji122 following Russell et al.63. We first manually adjusted the contrast of each image to increase the threshold so that only white puncta corresponding to Wolbachia cells inside the oocyte were retained and all background noise was rendered black. We then used the polygon selection tool to select three different regions of the oocyte (see Fig. S1 in Russell et al.): the whole oocyte, the posterior region, and the posterior cortex. We defined the posterior region as the posterior 1/8th of the oocyte and the posterior cortex as the narrow cortical region where the pole plasm component Vasa is expressed15,63. The “area” and “integrated density” were measured for each region. Additionally, the average of a quadruplicate measure of “mean gray value” in background selections was measured to account for any remaining background fluorescence. The corrected total cell fluorescence (CTCF) was calculated as integrated density − (area × background mean gray value) for each region. We used two-way ANOVAs and type III sums of squares to test whether log-transformed CTCF values differed among Wolbachia-host genotypes and temperature in each region of the oocyte (whole oocyte, posterior region, and posterior cortex). We also used two-sided Wilcoxon rank-sum tests to test whether CTCF values differed between 25° and 20 °C for each oocyte region and Wolbachia-host genotype.
Phylogenomic analyses
We conducted phylogenomic analyses to characterize the evolutionary relationships among Wolbachia strains and Drosophila host species. We obtained Wolbachia sequences from publicly available genome assemblies for wMel21, wMelCS66, wSeg, wCha15, wRi123, wAna124, wAura, wTri51, wHa125, and wMau40. We used Prokka 1.11 to identify homologs to known bacterial genes in the assemblies126. We avoided pseudogenes and paralogs in our phylogenetic analyses by selecting only single-copy genes that uniquely matched a bacterial reference gene identified by Prokka. We removed loci with indels by requiring all homologs to have identical length in all Wolbachia genomes. A total of 170 full-length single-copy genes met these criteria, totaling 135,105 bp. We then estimated a Bayesian phylogram using RevBayes 1.0.8 under the GTR + Γ + I model partitioned by codon position127. Four independent runs were performed, which all converged on the same topology. All nodes were supported with Bayesian posterior probabilities >0.99.
To estimate an absolute chronogram for Wolbachia, we first estimated a relaxed-clock relative chronogram using RevBayes127 with the root age fixed to 1 using the GTR + Γ + I, partitioned by codon position, using the same birth-death prior as Turelli et al.51. Each partition had an independent rate multiplier with prior Γ(1,1), as well as stationary frequencies and exchangeability rates drawn from flat, symmetrical Dirichlet distributions. For each branch, the branch-rate prior was Γ(7,7), normalized to a mean of 1 across all branches. We performed four independent runs that all agreed. We converted the relative chronogram into an absolute chronogram using the scaled distribution Γ(7,7) × 6.87 × 10−9 substitutions per third position site per year, derived from the posterior distribution estimated by Richardson et al.84, assuming 10 generations per year. Absolute branch lengths were calculated as the relative branch length times the third position rate multiplier divided by the substitutions per third-position site per year estimate above.
We used similar methods to generate a phylogram for the Drosophila host species. Publicly available sequences were obtained for D. melanogaster128, D. simulans129, D. mauritiana40, D. seguyi, D. chauvacae130, D. auraria, D. triauraria51, and D. ananassae131. A host phylogeny was generated using the same nuclear genes implemented in Turelli et al.51: aconitase, aldolase, bicoid, ebony, enolase, esc, g6pdh, glyp, glys, ninaE, pepck, pgi, pgm, pic, ptc, tpi, transaldolase, white, wingless, and yellow. We used BLAST with the D. melanogaster coding sequences to extract orthologs from the genomes of each host species. Sequences were then aligned with MAFFT 7132. Finally, we used RevBayes and the GTR + Γ + I model partitioned by codon position and gene to accommodate potential variation in the substitution process among genes. All nodes were supported with Bayesian posterior probabilities of 1.
The resulting phylograms were used to test whether Wolbachia density and oocyte cellular abundance exhibit phylogenetic signal on either the Wolbachia or host phylogenies. For the host phylogeny, we pooled estimates of Wolbachia density and oocyte abundance by host species, because two of the hosts (D. melanogaster and D. simulans) had data for multiple genotypes carrying different Wolbachia strains. We used our estimates of Wolbachia density (2ΔCt) and cellular oocyte abundance (log-transformed CTCF) to test for phylogenetic signal using Pagel’s lambda (λ)68. A Pagel’s λ of 0 indicates that character evolution occurs independently of phylogenetic relationships, whereas λ = 1 is consistent with a Brownian motion model of character evolution. We used the “phylosig” function and a likelihood ratio test (LRT) in the phytools package133 to compare the fitted value to a model assuming no phylogenetic signal (λ = 0). We also used a Monte Carlo-based method to generate 95% confidence intervals surrounding our estimates using 1000 bootstrap replicates in the pmc package134.
Statistics and reproducibility
All statistical analyses were performed in R118. Specific details for each statistical analysis are reported above. We conducted distribution and leverage analyses to evaluate assumptions of normality and then used data transformations and non-parametric tests when applicable (described above). Samples sizes and test statistics for each Wolbachia-host genotype are reported in the Supplementary Information. For each experiment, we collected the largest sample size that was feasible within our experimental design. We do not report any issues with reproducibility.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
References
Sudakaran, S., Kost, C. & Kaltenpoth, M. Symbiont acquisition and replacement as a source of ecological innovation. Trends Microbiol. 25, 375–390 (2017).
McCutcheon, J. P., Boyd, B. M. & Dale, C. The life of an insect endosymbiont from the cradle to the grave. Curr. Biol. 29, R485–R495 (2019).
McCutcheon, J. P. The genomics and cell biology of host-beneficial intracellular infections. Annu. Rev. Cell Dev. Biol. 37, 115–142 (2021).
Werren, J. H., Baldo, L. & Clark, M. E. Wolbachia: master manipulators of invertebrate biology. Nat. Rev. Microbiol. 6, 741–751 (2008).
Shropshire, J. D., Leigh, B. & Bordenstein, S. R. Symbiont-mediated cytoplasmic incompatibility: what have we learned in 50 years? Elife 9, e61989 (2020).
Oliver, K. M., Russell, J. A., Moran, N. A. & Hunter, M. S. Facultative bacterial symbionts in aphids confer resistance to parasitic wasps. Proc. Natl Acad. Sci. USA 100, 1803–1807 (2003).
Jaenike, J., Unckless, R., Cockburn, S. N., Boelio, L. M. & Perlman, S. J. Adaptation via symbiosis: recent spread of a Drosophila defensive symbiont. Science 329, 212–215 (2010).
Cogni, R., Ding, S. D., Pimentel, A. C., Day, J. P. & Jiggins, F. M. Wolbachia reduces virus infection in a natural population of Drosophila. Commun. Biol. 4, 1–7 (2021).
Bruner-Montero, G. & Jiggins, F. M. Wolbachia protects Drosophila melanogaster against two naturally occurring and virulent viral pathogens. Sci. Rep. 13, 8518 (2023).
Baumann, P. Biology of bacteriocyte-associated endosymbionts of plant sap-sucking insects. Annu. Rev. Microbiol. 59, 155–189 (2005).
Moran, N. A., McCutcheon, J. P. & Nakabachi, A. Genomics and evolution of heritable bacterial symbionts. Annu. Rev. Genet. 42, 165–190 (2008).
Douglas, A. E. The microbial dimension in insect nutritional ecology. Funct. Ecol. 23, 38–47 (2009).
Fisher, R. M., Henry, L. M., Cornwallis, C. K., Kiers, E. T. & West, S. A. The evolution of host-symbiont dependence. Nat. Commun. 8, 1–8 (2017).
Russell, S. L., Chappell, L. & Sullivan, W. A symbiont’s guide to the germline. Curr. Top. Dev. Biol. 135, 315–351 (2019).
Radousky, Y. A. et al. Distinct Wolbachia localization patterns in oocytes of diverse host species reveal multiple strategies of maternal transmission. Genetics 224, iyad038 (2023).
Dunbar, H. E., Wilson, A. C., Ferguson, N. R. & Moran, N. A. Aphid thermal tolerance is governed by a point mutation in bacterial symbionts. PLoS Biol. 5, e96 (2007).
Hoffmann, A. A., Turelli, M. & Harshman, L. G. Factors affecting the distribution of cytoplasmic incompatibility in Drosophila simulans. Genetics 126, 933–948 (1990).
Turelli, M. & Hoffmann, A. A. Cytoplasmic incompatibility in Drosophila simulans: dynamics and parameter estimates from natural populations. Genetics 140, 1319–1338 (1995).
Carrington, L. B., Lipkowitz, J. R., Hoffmann, A. A. & Turelli, M. A re-examination of Wolbachia-induced cytoplasmic incompatibility in California Drosophila simulans. PLoS ONE 6, e22565 (2011).
Hague, M. T., Mavengere, H., Matute, D. R. & Cooper, B. S. Environmental and genetic contributions to imperfect wMel-like Wolbachia transmission and frequency variation. Genetics 215, 1117–1132 (2020).
Hague, M. T. J. et al. Temperature effects on cellular host-microbe interactions explain continent-wide endosymbiont prevalence. Curr. Biol. 32, 878–888.e8 (2022).
Wade, M. J. The co-evolutionary genetics of ecological communities. Nat. Rev. Genet. 8, 185–195 (2007).
Weeks, A. R., Turelli, M., Harcombe, W. R., Reynolds, K. T. & Hoffmann, A. A. From parasite to mutualist: Rapid evolution of Wolbachia in natural populations of Drosophila. PLoS Biol. 5, e114 (2007).
Ross, P. A., Turelli, M. & Hoffmann, A. A. Evolutionary ecology of Wolbachia releases for disease control. Annu. Rev. Genet. 53, 93–116 (2019).
Ross, P. A. et al. Developing Wolbachia-based disease interventions for an extreme environment. PLoS Pathog. 19, e1011117 (2023).
Hien, N. T. et al. Environmental factors influence the local establishment of Wolbachia in Aedes aegypti mosquitoes in two small communities in central Vietnam. Gates Open Res. 5, 147 (2021).
Nikolouli, K. et al. Sterile insect technique and Wolbachia symbiosis as potential tools for the control of the invasive species Drosophila suzukii. J. Pest Sci. 91, 489–503 (2018).
Gong, J.-T. et al. Stable introduction of plant-virus-inhibiting Wolbachia into planthoppers for rice protection. Curr. Biol. 30, 4837–4845 (2020).
McMeniman, C. J. et al. Stable introduction of a life-shortening Wolbachia infection into the mosquito Aedes aegypti. Science 323, 141–144 (2009).
Hoffmann, A. A. et al. Successful establishment of Wolbachia in Aedes populations to suppress dengue transmission. Nature 476, 454–457 (2011).
Walker, T. et al. The wMel Wolbachia strain blocks dengue and invades caged Aedes aegypti populations. Nature 476, 450–453 (2011).
Utarini, A. et al. Efficacy of Wolbachia-infected mosquito deployments for the control of dengue. N. Engl. J. Med. 384, 2177–2186 (2021).
Ross, P. A. et al. Wolbachia infections in Aedes aegypti differ markedly in their response to cyclical heat stress. PLOS Pathog. 13, e1006006 (2017).
Ross, P. A., Ritchie, S. A., Axford, J. K. & Hoffmann, A. A. Loss of cytoplasmic incompatibility in Wolbachia-infected Aedes aegypti under field conditions. PLoS Negl. Trop. Dis. 13, e0007357 (2019).
Funkhouser-Jones, L. J., van Opstal, E. J., Sharma, A. & Bordenstein, S. R. The maternal effect gene Wds controls Wolbachia titer in Nasonia. Curr. Biol. 28, 1692–1702 (2018).
Porter, J. & Sullivan, W. The cellular lives of Wolbachia. Nat. Rev. Microbiol. 1, 17 (2023).
Bright, M. & Bulgheresi, S. A complex journey: transmission of microbial symbionts. Nat. Rev. Microbiol. 8, 218–230 (2010).
Landmann, F. et al. Both asymmetric mitotic segregation and cell-to-cell invasion are required for stable germline transmission of Wolbachia in filarial nematodes. Biol. Open 1, 536–547 (2012).
Correa, C. C. & Ballard, J. Wolbachia associations with insects: winning or losing against a master manipulator. Front. Ecol. Evol. 3, 153 (2016).
Meany, M. K. et al. Loss of cytoplasmic incompatibility and minimal fecundity effects explain relatively low Wolbachia frequencies in Drosophila mauritiana. Evolution 73, 1278–1295 (2019).
Olsen, K., Reynolds, K. T. & Hoffmann, A. A. A field cage test of the effects of the endosymbiont Wolbachia on Drosophila melanogaster. Heredity 86, 731–737 (2001).
Hamm, C. A. et al. Wolbachia do not live by reproductive manipulation alone: Infection polymorphism in Drosophila suzukii and D. subpulchrella. Mol. Ecol. 23, 4871–4885 (2014).
Ross, P. A. et al. Heatwaves cause fluctuations in wMel Wolbachia densities and frequencies in Aedes aegypti. PLoS Negl. Trop. Dis. 14, e0007958 (2020).
Ferri, E. et al. New insights into the evolution of Wolbachia infections in filarial nematodes inferred from a large range of screened species. PLoS One 6, e20843 (2011).
Zug, R. & Hammerstein, P. Still a host of hosts for Wolbachia: analysis of recent data suggests that 40% of terrestrial arthropod species are infected. PLoS ONE 7, e38544 (2012).
Weinert, L. A., Araujo-Jnr, E. V., Ahmed, M. Z. & Welch, J. J. The incidence of bacterial endosymbionts in terrestrial arthropods. Proc. R. Soc. B Biol. Sci. 282, 20150249 (2015).
O’Neill, S. L., Giordano, R., Colbert, A. M., Karr, T. L. & Robertson, H. M. 16S rRNA phylogenetic analysis of the bacterial endosymbionts associated with cytoplasmic incompatibility in insects. Proc. Natl Acad. Sci. USA 89, 2699–2702 (1992).
Raychoudhury, R., Baldo, L., Oliveira, D. C. & Werren, J. H. Modes of acquisition of Wolbachia: Horizontal transfer, hybrid introgression, and codivergence in the Nasonia species complex. Evolution 63, 165–183 (2009).
Conner, W. R. et al. Genome comparisons indicate recent transfer of wRi-like Wolbachia between sister species Drosophila suzukii and D. subpulchrella. Ecol. Evol. 7, 9391–9404 (2017).
Gerth, M. & Bleidorn, C. Comparative genomics provides a timeframe for Wolbachia evolution and exposes a recent biotin synthesis operon transfer. Nat. Microbiol. 2, 16241 (2017).
Turelli, M. et al. Rapid global spread of wRi-like Wolbachia across multiple Drosophila. Curr. Biol. 28, 963–971.e8 (2018).
Cooper, B. S., Vanderpool, D., Conner, W. R., Matute, D. R. & Turelli, M. Wolbachia acquisition by Drosophila yakuba-clade hosts and transfer of incompatibility loci between distantly related Wolbachia. Genetics 212, 1399–1419 (2019).
Shropshire, D. et al. Rapid turnover of pathogen-blocking Wolbachia and their incompatibility loci. Preprint at bioRxiv https://doi.org/10.1101/2023.12.04.569981 (2023).
Vancaester, E. & Blaxter, M. Phylogenomic analysis of Wolbachia genomes from the Darwin Tree of Life biodiversity genomics project. PLoS Biol. 21, e3001972 (2023).
Caspari, E. & Watson, G. On the evolutionary importance of cytoplasmic sterility in mosquitoes. Evolution 13, 568–570 (1959).
Turelli, M., Katznelson, A. & Ginsberg, P. S. Why Wolbachia-induced cytoplasmic incompatibility is so common. Proc. Natl Acad. Sci. USA 119, e2211637119 (2022).
Turelli, M. & Hoffmann, A. A. Rapid spread of an inherited incompatibility factor in California Drosophila. Nature 353, 440–442 (1991).
Kriesner, P., Conner, W. R., Weeks, A. R., Turelli, M. & Hoffmann, A. A. Persistence of a Wolbachia infection frequency cline in Drosophila melanogaster and the possible role of reproductive dormancy. Evolution 70, 979–997 (2016).
Hadfield, S. J. & Axton, J. M. Germ cells colonized by endosymbiotic bacteria. Nature 402, 482 (1999).
Veneti, Z., Clark, M. E., Karr, T. L., Savakis, C. & Bourtzis, K. Heads or tails: host-parasite interactions in the Drosophila-Wolbachia system. Appl. Environ. Microbiol. 70, 5366–5372 (2004).
Ferree, P. M. et al. Wolbachia utilizes host microtubules and Dynein for anterior localization in the Drosophila oocyte. PLoS Pathog. 1, e14 (2005).
Serbus, L. R. & Sullivan, W. A cellular basis for Wolbachia recruitment to the host germline. PLoS Pathog. 3, e190 (2007).
Russell, S. L., Lemseffer, N. & Sullivan, W. T. Wolbachia and host germline components compete for kinesin-mediated transport to the posterior pole of the Drosophila oocyte. PLoS Pathog. 14, e1007216 (2018).
Prout, T. Some evolutionary possibilities for a microbe that causes incompatibility in its host. Evolution 48, 909–911 (1994).
Turelli, M. Evolution of incompatibility-inducing microbes and their hosts. Evolution 48, 1500–1513 (1994).
Hague, M. T., Caldwell, C. N. & Cooper, B. S. Pervasive effects of Wolbachia on host temperature preference. mBio 11, e01768–20 (2020).
Hague, M. T. J., Woods, H. A. & Cooper, B. S. Pervasive effects of Wolbachia on host activity. Biol. Lett. 17, 20210052 (2021).
Pagel, M. Inferring the historical patterns of biological evolution. Nature 401, 877–884 (1999).
Freckleton, R. P., Harvey, P. H. & Pagel, M. Phylogenetic analysis and comparative data: a test and review of evidence. Am. Nat. 160, 712–726 (2002).
Rajpurohit, S. & Schmidt, P. S. Measuring thermal behavior in smaller insects: a case study in Drosophila melanogaster demonstrates effects of sex, geographic origin, and rearing temperature on adult behavior. Fly 10, 149–161 (2016).
Christensen, S. et al. Quantitative methods for assessing local and bodywide contributions to Wolbachia titer in maternal germline cells of Drosophila. BMC Microbiol. 19, 1–17 (2019).
Serbus, L. R. et al. A feedback loop between Wolbachia and the Drosophila gurken mRNP complex influences Wolbachia titer. J. Cell Sci. 124, 4299–4308 (2011).
Clark, M. E., Veneti, Z., Bourtzis, K. & Karr, T. L. Wolbachia distribution and cytoplasmic incompatibility during sperm development: the cyst as the basic cellular unit of CI expression. Mech. Dev. 120, 185–198 (2003).
Chrostek, E. et al. Wolbachia variants induce differential protection to viruses in Drosophila melanogaster: a phenotypic and phylogenomic analysis. PLoS Genet 9, e1003896 (2013).
Noda, H., Koizumi, Y., Zhang, Q. & Deng, K. Infection density of Wolbachia and incompatibility level in two planthopper species, Laodelphax striatellus and Sogatella furcifera. Insect Biochem. Mol. Biol. 31, 727–737 (2001).
Duron, O., Fort, P. & Weill, M. Influence of aging on cytoplasmic incompatibility, sperm modification and Wolbachia density in Culex pipiens mosquitoes. Heredity 98, 368–374 (2007).
Kaur, R., Martinez, J., Rota-Stabelli, O., Jiggins, F. M. & Miller, W. J. Age, tissue, genotype and virus infection regulate Wolbachia levels in Drosophila. Mol. Ecol. 29, 2063–2079 (2020).
Tortosa, P. et al. Wolbachia age-sex-specific density in Aedes albopictus: a host evolutionary response to cytoplasmic incompatibility? PLoS ONE 5, e9700 (2010).
Unckless, R. L., Boelio, L. M., Herren, J. K. & Jaenike, J. Wolbachia as populations within individual insects: causes and consequences of density variation in natural populations. Proc. R. Soc. B Biol. Sci. 276, 2805–2811 (2009).
Shropshire, J. D., Hamant, E. & Cooper, B. S. Male age and Wolbachia dynamics: investigating how fast and why bacterial densities and cytoplasmic incompatibility strengths vary. mBio 12, e02998–21 (2021).
Layton, E. M., On, J., Perlmutter, J. I., Bordenstein, S. R. & Shropshire, J. D. Paternal grandmother age affects the strength of Wolbachia-induced cytoplasmic incompatibility in Drosophila melanogaster. mBio 10, 10–1128 (2019).
Frydman, H. M., Li, J. M., Robson, D. N. & Wieschaus, E. Somatic stem cell niche tropism in Wolbachia. Nature 441, 509 (2006).
White, P. M. et al. Mechanisms of horizontal cell-to-cell transfer of Wolbachia spp. in Drosophila melanogaster. Appl. Environ. Microbiol. 83, e03425–16 (2017).
Richardson, M. F. et al. Population genomics of the Wolbachia endosymbiont in Drosophila melanogaster. PLoS Genet 8, e1003129 (2012).
Riegler, M., Sidhu, M., Miller, W. J. & O’Neill, S. L. Evidence for a global Wolbachia replacement in Drosophila melanogaster. Curr. Biol. 15, 1428–1433 (2005).
Ilinsky, Y. Coevolution of Drosophila melanogaster mtDNA and Wolbachia genotypes. PLoS ONE 8, e54373 (2013).
Early, A. M. & Clark, A. G. Monophyly of Wolbachia pipientis genomes within Drosophila melanogaster: geographic structuring, titre variation and host effects across five populations. Mol. Ecol. 22, 5765–5778 (2013).
Strunov, A. et al. Historic museum samples provide evidence for a recent replacement of Wolbachia types in European Drosophila melanogaster. Mol. Biol. Evol. 40, msad258 (2023).
Poinsot, D., Bourtzis, K., Markakis, G., Savakis, C. & Merçot, H. Wolbachia transfer from Drosophila melanogaster into D. simulans: Host effect and cytoplasmic incompatibility relationships. Genetics 150, 227–237 (1998).
Strunov, A., Lerch, S., Blanckenhorn, W. U., Miller, W. J. & Kapun, M. Complex effects of environment and Wolbachia infections on the life history of Drosophila melanogaster hosts. J. Evol. Biol. 35, 788–802 (2022).
Hart, B. L. Biological basis of the behavior of sick animals. Neurosci. Biobehav. Rev. 12, 123–137 (1988).
De Roode, J. C. & Lefèvre, T. Behavioral immunity in insects. Insects 3, 789–820 (2012).
Curtis, V. A. Infection-avoidance behaviour in humans and other animals. Trends Immunol. 35, 457–464 (2014).
Fedorka, K. M., Kutch, I. C., Collins, L. & Musto, E. Cold temperature preference in bacterially infected Drosophila melanogaster improves survival but is remarkably suboptimal. J. Insect Physiol. 93, 36–41 (2016).
White, P. M. et al. Reliance of Wolbachia on high rates of host proteolysis revealed by a genome-wide RNAi screen of Drosophila cells. Genetics 205, 1473–1488 (2017).
Grobler, Y. et al. Whole genome screen reveals a novel relationship between Wolbachia levels and Drosophila host translation. PLoS Pathog. 14, e1007445 (2018).
Chrostek, E. & Teixeira, L. Mutualism breakdown by amplification of Wolbachia genes. PLoS Biol. 13, e1002065 (2015).
Chrostek, E. & Teixeira, L. Within host selection for faster replicating bacterial symbionts. PLoS ONE 13, e0191530 (2018).
Duarte, E. H., Carvalho, A., López-Madrigal, S., Costa, J. & Teixeira, L. Forward genetics in Wolbachia: Regulation of Wolbachia proliferation by the amplification and deletion of an addictive genomic island. PLOS Genet 17, e1009612 (2021).
Woolfit, M. et al. Genomic evolution of the pathogenic Wolbachia strain, wMelPop. Genome Biol. Evol. 5, 2189–2204 (2013).
Werren, J. H., Windsor, D. & Guo, L. R. Distribution of Wolbachia among neotropical arthropods. Proc. R. Soc. Lond. B 262, 197–204 (1995).
Martinez, J., Klasson, L., Welch, J. J. & Jiggins, F. M. Life and death of selfish genes: comparative genomics reveals the dynamic evolution of cytoplasmic incompatibility. Mol. Biol. Evol. 38, 2–15 (2021).
Hoffmann, A. A., Hercus, M. & Dagher, H. Population dynamics of the Wolbachia infection causing cytoplasmic incompatibility in Drosophila melanogaster. Genetics 148, 221–231 (1998).
Reynolds, K. T. & Hoffmann, A. A. Male age, host effects and the weak expression or non-expression of cytoplasmic incompatibility in Drosophila strains infected by maternally transmitted Wolbachia. Genet. Res. 80, 79–87 (2002).
Giordano, R., O’Neill, S. L. & Robertson, H. M. Wolbachia infections and the expression of cytoplasmic incompatibility in Drosophila sechellia and D. mauritiana. Genetics 140, 1307–1317 (1995).
Hoffmann, A. & Turelli, M. Unidirectional incompatibility in Drosophila simulans: inheritance, geographic variation and fitness effects. Genetics 119, 435–444 (1988).
Rousset, F. & Solignac, M. Evolution of single and double Wolbachia symbioses during speciation in the Drosophila simulans complex. Proc. Natl Acad. Sci. USA 92, 6389–6393 (1995).
James, A. C. & Ballard, J. W. O. Expression of cytoplasmic incompatibility in Drosophila simulans and its impact on infection frequencies and distribution of Wolbachia pipientis. Evolution 54, 1661–1672 (2000).
Ballard, J. W. O. Sequential evolution of a symbiont inferred from the host: Wolbachia and Drosophila simulans. Mol. Biol. Evol. 21, 428–442 (2004).
Bourtzis, K., Nirgianaki, A., Markakis, G. & Savakis, C. Wolbachia infection and cytoplasmic incompatibility in Drosophila species. Genetics 144, 1063–1073 (1996).
Choi, J. Y., Bubnell, J. E. & Aquadro, C. F. Population genomics of infectious and integrated Wolbachia pipientis genomes in Drosophila ananassae. Genome Biol. Evol. 7, 2362–2382 (2015).
Osborne, S. E., Leong, Y. S., O’Neill, S. L. & Johnson, K. N. Variation in antiviral protection mediated by different Wolbachia strains in Drosophila simulans. PLOS Pathog. 5, e1000656 (2009).
Martinez, J. et al. Symbionts commonly provide broad spectrum resistance to viruses in insects: a comparative analysis of Wolbachia strains. PLOS Pathog. 10, e1004369 (2014).
Gu, X. et al. A wMel Wolbachia variant in Aedes aegypti from field-collected Drosophila melanogaster with increased phenotypic stability under heat stress. Environ. Microbiol. 24, 2119–2135 (2022).
Ballard, J. & Melvin, R. Tetracycline treatment influences mitochondrial metabolism and mtDNA density two generations after treatment in Drosophila. Insect Mol. Biol. 16, 799–802 (2007).
Cooper, B. S., Ginsberg, P. S., Turelli, M. & Matute, D. R. Wolbachia in the Drosophila yakuba complex: Pervasive frequency variation and weak cytoplasmic incompatibility, but no apparent effect on reproductive isolation. Genetics 205, 333–351 (2017).
Canty, C. & Ripley, B. boot: Bootstrap R (S-Plus) Functions (2017).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2023).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
Singmann, H., Bolker, B., Westfall, J., Aust, F. & Ben-Shachar, M. S. Afex: Analysis of Factorial Experiments (2020).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res 29, e45–e45 (2001).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Klasson, L. et al. The mosaic genome structure of the Wolbachia wRi strain infecting Drosophila simulans. Proc. Natl Acad. Sci. USA 106, 5725–5730 (2009).
Salzberg, S. L. et al. Serendipitous discovery of Wolbachia genomes in multiple Drosophila species. Genome Biol. 6, R23 (2005).
Ellegaard, K. M., Klasson, L., Näslund, K., Bourtzis, K. & Andersson, S. G. Comparative genomics of Wolbachia and the bacterial species concept. PLoS Genet 9, e1003381 (2013).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Höhna, S. et al. RevBayes: Bayesian phylogenetic inference using graphical models and an interactive model-specification language. Syst. Biol. 65, 726–736 (2016).
Hoskins, R. A. et al. The Release 6 reference sequence of the Drosophila melanogaster genome. Genome Res. 25, 445–458 (2015).
Hu, T. T., Eisen, M. B., Thornton, K. R. & Andolfatto, P. A second-generation assembly of the Drosophila simulans genome provides new insights into patterns of lineage-specific divergence. Genome Res. 23, 89–98 (2013).
Conner, W. R. et al. A phylogeny for the Drosophila montium species group: a model clade for comparative analyses. Mol. Phylogenet. Evol. 158, 107061 (2021).
Clark, A. G. et al. Evolution of genes and genomes on the Drosophila phylogeny. Nature 450, 203–218 (2007).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Revell, L. J. phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 3, 217–223 (2012).
Boettiger, C., Coop, G. & Ralph, P. Is your phylogeny informative? Measuring the power of comparative methods. Evolution 66, 2240–2251 (2012).
Suvorov, A. et al. Widespread introgression across a phylogeny of 155 Drosophila genomes. Curr. Biol. 32, 111–123 (2022).
Acknowledgements
We are especially thankful to W. Sullivan for very useful discussions that improved the quality of this manuscript. We thank W. Conner for assistance with bioinformatic analyses. J. Statz provided thoughtful feedback on the manuscript. Support and instrumentation was provided by the Cellular Genetics Ecosystem in the Genomics Core, the Environmental Control for Organismal Research (ECOR) facility, and the Center for Biomolecular Structure and Dynamics at the University of Montana. Research reported in this publication was supported by a National Science Foundation CAREER (2145195) Award to B.S.C and by the National Institute of General Medical Sciences of the National Institutes of Health under award number R35GM124701.
Author information
Authors and Affiliations
Contributions
M.T.J.H. designed the project, collected data, performed analyses, and prepared the manuscript. T.B.W. collected data. B.S.C. acquired funding, designed the project, prepared the manuscript, and provided leadership. All authors reviewed and edited the manuscript prior to submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Hannes Schuler and Christina Karlsson Rosenthal.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hague, M.T.J., Wheeler, T.B. & Cooper, B.S. Comparative analysis of Wolbachia maternal transmission and localization in host ovaries. Commun Biol 7, 727 (2024). https://doi.org/10.1038/s42003-024-06431-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06431-y
- Springer Nature Limited