Abstract
Most cancer-related deaths are due to the spread of tumour cells throughout the body—a process known as metastasis. While in the vasculature, these cells are referred to as circulating tumour cells (CTCs) and can be found as either single cells or clusters of cells (often including platelets), with the latter having the highest metastatic potential. However, the biology of CTC clusters is poorly understood, and there are no therapies that specifically target them. We previously developed an in vitro model system for CTC clusters and proposed a new extravasation model that involves cluster dissociation, adherence, and single-cell invasion in response to TGF-β1 released by platelets. Here, we investigated TGF-β1-induced gene expression changes in this model, focusing on genes for which targeted drugs are available. In addition to the upregulation of the TGF-β1 signalling pathway, we found that (i) genes in the EGF/EGFR pathway, including those coding for EGFR and several EGFR ligands, were also induced, and (ii) Erlotinib and Osimertinib, two therapeutic EGFR/tyrosine kinase inhibitors, decreased the TGF-β1-induced adherence and invasion of the CTC cluster-like line despite the line expressing wild-type EGFR. Overall, we suggest that EGFR inhibitors have the potential to decrease the dispersal of CTC clusters that respond to TGF-β1 and overexpress EGFR (irrespective of its status) and thus could improve patient survival.
Similar content being viewed by others
Introduction
Despite increasing research efforts and improved survival rates, cancer remains a major health concern with devastating consequences (e.g.,1). Most cancer-related deaths are due to the spread of tumour cells throughout the body (e.g.,2). During this process—known as metastasis, cancer cells break off the primary tumour, enter the vascular system, and disperse to new tissues that they subsequently colonize to form secondary metastatic tumours3. While in the vasculature, these cells are referred to as circulating tumour cells (CTCs). While CTCs primarily manifest as single cells4, clusters of circulating tumour cells (CTC clusters) are often detected5, and their presence is generally associated with poor prognosis6,7,8.
CTC clusters typically consist of 2–100 cells arranged in grape-like assemblages9. They are highly heterogeneous, being composed of both tumour cells expressing various genetic markers (e.g., epithelial, mesenchymal, or both)10 as well as several non-tumour cell types such as platelets, leukocytes, fibroblasts, and pericytes11,12. Although CTC clusters are found at a lower frequency than single CTCs, they have a significantly higher metastatic potential4. Several factors have been proposed to account for their increased metastatic abilities. For instance, the cluster phenotype provides cells with resistance to several factors, such as shear stress during circulation13, anoikis (i.e., cell death associated with the loss of cell–cell connections)14, immune attacks15, and drug therapies16. Additionally, the clustering of cells has been shown to promote the activation of DNA methylation patterns17 and metabolic switches18 that may increase their metastatic efficiency.
In spite of the significant contribution of CTC clusters to cancer dissemination and disease progression, their biology is not well understood and there are no therapies that specifically target them (for instance, see19,20). This is partly due to their low frequency in circulation4 and the challenges associated with their isolation and cultivation in the lab (e.g.,21). To circumvent these difficulties, we previously developed an in vitro model system for CTC clusters22,23. Specifically, we used a metastatic non-small cell lung cancer (NSCLC) line (NCI-H2122) that grows as a mixture of adherent and non-adherent cells and experimentally evolved a cell line that grows exclusively as clusters of cells in suspension (referred to as the “Suspension-Selected” or SS line). We showed that the SS line shares several characteristics with isolated CTC clusters, including similar morphology, cell–cell connections, cell heterogeneity (with respect to expression of epithelial and mesenchymal markers), and cell plasticity22,23.
While CTC clusters can be more successful and efficient at initiating metastatic tumours than single CTCs4, the cellular and molecular mechanisms involved in their ability to enter the vasculature (intravasate), exit the vasculature (extravasate), and disseminate in new tissues are not well understood (e.g.,20). Several studies provide conflicting explanations for the presence of CTC clusters in circulation. Certain lines of evidence suggest that CTC clusters can arise via the aggregation of single cells in or near the vasculature24, whereas other studies suggest that hypoxic environments cause clusters to break off the tumour and collectively enter the bloodstream25. Furthermore, several conflicting mechanisms have been proposed to explain how CTC clusters exit circulation and colonize new tissues. For instance, clusters are thought to become trapped in capillaries and then proliferate into new tumours at the site of arrest26. Contrary to this view, it has been argued that CTC clusters can, in fact, travel through capillaries by rearranging into single-file chain-like structures27. Alternatively, it has been shown that CTC clusters can cause extensive remodelling of the vasculature wall to promote their extravasation as clusters into new tissues28.
More recently, based on the extensive association between platelets and CTC clusters in the blood8, we proposed a new mechanism for the extravasation of CTC clusters involving TGF-β1 signalling29. Specifically, because single CTCs have been shown to activate platelets and cause them to release TGF-β1 that can facilitate the extravasation of single CTCs30, we hypothesized that platelets associated with CTC clusters might also contribute to the extravasation of CTC clusters that express TGF-β1 receptors, via TGF-β1-mediated paracrine signalling. To test this possibility, we examined the effect of TGF-β1 on the dispersal abilities of our evolved CTC cluster-like cell line29. We found that exogenous TGF-β1 induces the adherence and dissociation of cell clusters into single cells with increased migration and invasion abilities. Furthermore, these TGF-β1-induced adherent cells secrete their own TGF-β1, allowing them to migrate even in the absence of exogenous TGF-β1.
To gain insight into the potential mechanisms involved in the TGF-β1-mediated extravasation of CTC clusters, in the current study, we investigated changes in gene expression patterns in our experimentally evolved CTC cluster-like cell line in response to TGF-β1, focusing on pathways for which targeted drugs are available. TGF-β1 has been widely implicated in coordinating the development of invasive properties of tumour cells via the activation of the epithelial-mesenchymal transition (EMT) program31, alone or in conjunction with other signalling pathways, including the epidermal growth factor (EGF) pathway (e.g.,32,33,34). Several drugs have been developed and successfully used to target the mutant/overactive/overexpressed forms of the EGF receptor (EGFR) in tumours (e.g.,35). For instance, tyrosine kinase inhibitors (TKIs) such as Erlotinib and Osimertinib can be effective in reducing tumour progression (through inhibiting cell proliferation and/or inducing cell death) and improving the prognosis of NSCLC patients with activating (i.e., ligand-independent) EGFR mutations36,37. However, more recently, Erlotinib was also found to decrease the migration and invasion of several adherent cell lines expressing either mutant or wild-type EGFR32,38,39,40. Nevertheless, the mechanisms are not fully understood, and the potential impact of this drug on delaying metastasis in vivo has not been addressed. To this end, we used transcriptomics data to (i) address whether the TGF-β1-induced adherent H2122 SS cells overexpress EGFR, and (ii) test the effect of Erlotinib and Osimertinib on the viability, adherence, and invasion of the H2122 SS clusters exposed to TGF-β1. A better understanding of the pathways activated during the dispersal of CTC clusters can provide new therapeutic strategies to slow down the metastatic process by inhibiting their adherence and invasive abilities.
Materials and methods
Cell lines and culture conditions
The cancer cell line used in this study was experimentally evolved from a non-small cell lung cancer cell line obtained from the American Type Culture Collection—ATCC (Manassas, VA, USA). This original cell line (NCI-H2122) was established from the pleural effusion of a 46-year-old female with stage-4 adenocarcinoma and grows as a mixture of two distinct phenotypes: clusters of cells growing in suspension and adherent cells that grow as a monolayer. By selectively passaging only the adherent or suspension cell populations, two cell lines were experimentally evolved: one that grows as cell clusters in suspension (referred to as the H2122 Suspension-Selected or SS line) and one that grows as adherent cells (referred to as the H2122 Adherent-Selected or AS line)23. The SS line is an established model system for CTC clusters22,23,29. Cells were grown under 5% CO2, at 37°C, in RPMI-1640 media (MP Biomedicals) supplemented with 10% FBS and 1% Penicillin/Streptomycin/Amphotericin B mix (R&D Systems). The concentrations of glucose and glutamine in the media were adjusted to 5 mM and 0.5 mM, respectively, to mimic physiological levels29.
Reagents
Human recombinant TGF-β1 (R&D Systems; 7754-BH-005) was reconstituted in 4 mM HCl at 10 mg/ml and added directly to the culture medium at a final concentration of 10 ng/ml. Erlotinib hydrochloride (Sigma; SML2156) was dissolved (at 50°C, to increase solubility) in cell-culture-suitable DMSO to 10 mM and was added to the culture medium at final concentrations of 2.5, 5, 10, or 15 µM. Osimertinib mesylate (RayBiotech; 1421373-66-1) was dissolved in DMSO to a stock concentration of 5 mM and diluted to 5 µM in the culture medium.
Assessing the effect of Erlotinib and Osimertinib on the phenotype and population growth of the H2122 SS line exposed to TGF-β1
To assess the effect of Erlotinib or Osimertinib on the growth and TGF-β1-induced adherence of the SS line, cells were seeded (in triplicates) at 1 × 105 cells/ml in 12-well plates in the absence (control) or presence of Erlotinib (2.5, 5, 10, or 15 µM) or Osimertinib (5 µM), alone or in combination with TGF-β1 (10 ng/ml). The suspension and adherent cell fractions were then harvested separately, stained with Syto 9 and Propidium Iodide in PBS (at final concentrations of 1 µM and 6 µM, respectively) and live and dead cells were counted with the Countess II FL Automated Cell Counter (Thermo Fisher Scientific)29.
Assessing the effect of Erlotinib on the invasion of the H2122 SS line exposed to TGF-β1
To assess the effect of Erlotinib on the invasion potential of H2122 SS, cells (in triplicates) were treated with TGF-β1 alone or in combination with Erlotinib (5 and 10 µM). Specifically, 1 × 105 cells were placed onto Transwell inserts with 8 µm pores (Corning) containing a layer of 1.5 mg/ml Matrigel (Corning), and the number of cells that invaded was assessed after 72 hours29. Invasive cells (i.e., attached to the underside of the insert) were fixed with 70% ethanol and stained with 0.2% crystal violet41. Four random fields of view from each stained insert were photographed using brightfield microscopy with a 20 × objective; cells were counted for each field of view and averaged for each triplicate29.
Statistical analyses
Statistical analyses were performed using GraphPad Prism 8 (https://www.graphpad.com/). The specific tests and thresholds used are indicated in figure legends. At least three independent experiments (with three replicates each) were performed; data were expressed as mean ± SE. Additional information and data regarding statistical analyses for each dataset are included in the Supplementary Material.
RNA extraction and sequencing
For RNA extraction, SS cells were seeded in triplicates in tissue-treated 12-well plates at a density of 1 × 105 cells/ml in the absence (control cultures) or presence (treatment cultures) of human recombinant TGF-β1 (at 10 ng/ml). After 24 h, total RNA from control and treatment cultures was extracted using the Qiagen RNeasy mini kit, following the manufacturer’s instructions. For the treatment cultures, unattached cells were removed before the adherent cells were trypsinized and harvested for RNA extraction. RNA quality control, library construction, and sequencing were performed by Novogene (https://en.novogene.com/).
RNAseq and DEG analyses
Initial RNAseq analyses were performed by Novogene. Specifically, raw reads were processed with in-house perl scripts designed to remove reads containing adapters, reads with uncertain nucleotides that represented more than 10% of either read, and low-quality reads (quality control information for RNA sequence data is included in Table S1). An index for the human reference genome (GRCh38/hg38) was built, and paired-end clean reads were aligned to the reference genome using Hisat2 v2.0.542. Quantification of gene expression levels was done first by using featureCounts v1.5.0-p3 to count the read numbers mapped to each gene and then calculating FPKM (Fragments Per Kilobase of transcript per Million) mapped reads42. The threshold for gene expression used was FPKM > 1. Principal component analysis (PCA) on the gene expression value (FPKM) of all samples was performed using Galaxy43,44,45 to evaluate intergroup differences and intragroup sample replication.
Differential expression analysis of control vs treatment (SS vs TGF-β1-induced adherent cells) cultures was performed using the DESeq2 R package (1.20.0)46. First, the raw readcount was normalized to correct for sequencing depth; then, a statistical model (Negative Binomial Distribution) was used to calculate P-values. The resulting P-values were adjusted using Benjamini and Hochberg’s approach for controlling the false discovery rate (FDR)47. Genes with an adjusted P-value < 0.05 and Log2FoldChange greater than 1 were considered differentially expressed. Gene Ontology (GO) enrichment analyses of differentially expressed genes were performed using ShinyGO 0.76.248 with an FDR cutoff of 0.05 and pathway size minimum of 2 and maximum of 2,000.
Results
TGF-β1 induces widespread gene expression changes in H2122 SS adherent cells
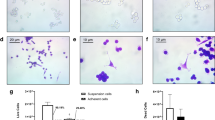
We have previously reported that when the H2122 SS line (growing as clusters of cells in suspension) is subjected to TGF-β1 (10 ng/ml) for 48 h, most clusters dissociate, and cells attach and acquire a mesenchymal-like phenotype29 (Fig. 1). To investigate early gene expression changes induced in the SS line in response to TGF-β1, we treated the cells for 24 h and then extracted total RNA from the control population (i.e., cell clusters in suspension) and the TGF-β1-induced adherent cell fraction (i.e., adherent cells). The transcriptomic profiles of the two phenotypes (naïve suspension and TGF-β1-induced adherent) revealed that, compared to the suspension phenotype, the adherent phenotype involves the expression of a slightly larger number of genes (11,791 vs 11,696); of these, 10,873 expressed genes are shared between the two groups, while 918 and 823 genes are uniquely expressed in the adherent and suspension phenotypes, respectively (Fig. 2a).
Micrographs of H2122 SS cells in their native state (control; clusters in suspension)—panel (a), and their TGF-β1-induced adherent state (treatment)—panels (b) and (c). Treated cells were subjected to TGF-β1 (10 ng/ml) for 48 h and stained with 0.2% crystal violet41; arrows in panel c (high magnification) indicate mesenchymal-like extensions.
Gene expression differences between the suspension and the TGF-β1-induced adherent phenotypes of the H2122 SS cell line. (a) Venn diagram showing the number of expressed genes that are common/unique to suspension cells and TGF-β1-induced adherent cells. (b) Volcano plot highlighting the upregulated (red) and downregulated (green) genes in adherent cells relative to suspension cells (Log2foldChange > 1; padj < 0.05). (c,d) Gene Ontology (biological processes) are enrichment summaries for upregulated (c) and downregulated (d) genes in adherent cells (fold enrichment, number of genes for each category and the corresponding -log10(FDR) values are indicated).
Overall, the comparison between the transcriptomic profiles of the suspension and adherent phenotypes revealed 2598 differentially expressed genes (DEGs); of these, 1390 DEGs were upregulated and 1208 DEGs were downregulated in the adherent phenotype relative to the suspension phenotype (Fig. 2b). The 15 most upregulated DEGs in the adherent cells included genes coding for a member of the Wnt pathway (Wnt7a; log2fold change = 11.5), three transcription factors (SNAI2, ZBED2, and MAF), two cytokines (IL32 and CCL20), and several factors associated with increased aggressivity in various cancers (e.g., COL8A1; STC1, ANGPTL4, and LAPTM5) (Supplementary Table S2). On the other hand, among the highest downregulated genes, we found genes associated with DNA and cell replication (e.g., E2F2—log2fold change = − 7.5; ESCO2—log2fold change = − 6; and MCM10—log2fold change = − 5.9) (Supplementary Table S3).
Differentially expressed genes in TGF-β1-induced adherent cells are enriched in adherence, migration, and cell proliferation processes
As expected from the drastic differences in phenotype, with SS cell clusters dissociating, adhering (Fig. 1), and being able to migrate when exposed to TGF-β129, Gene Ontology enrichment analyses of DEGs showed that the upregulated genes in TGF-β1-induced adherent cells are primarily involved in processes associated with cell adhesion and migration (Fig. 2c). Similarly, consistent with the TGF-β1-inhibitory effect on the proliferation of adherent cells29, the downregulated gene set was enriched in DNA replication, mitotic spindle formation, and cell cycle processes (Fig. 2d).
Based on the phenotypic changes observed and the results of enrichment analyses, we further analyzed the expression patterns of genes associated with specific processes and pathways, including cell adhesion and EMT. Consistent with the induction of adherence in the SS cells treated with TGF-β1, many genes involved in adherence to substrate were upregulated (Supplementary Table S4), including genes coding for nine integrin subunits, seven semaphorins, and various proteins that support integrin function such as TGM2, FERMT1, FERMT2, and NEDD9. Additionally, the transcription of genes coding for various ECM components was drastically upregulated after TGF-β1 treatment (e.g., three laminin and six collagen subunits; Supplementary Table S4), which is consistent with the role of TGF-β1 in inducing ECM deposition during fibrosis49.
Because TGF-β1 is widely recognized as an EMT initiating factor, we searched for components of the EMT program that were transcriptionally altered. Various factors that have been reported to initiate the EMT process are upregulated in TGF-β1-induced adherent cells relative to suspension cells (Supplementary Table S5). These include members of the BMP family—BMP1, BMP2, and BMP4, Wnt ligands—Wnt7a and Wnt7b, as well as VEGFA and HB-EGF. Two transcription factors with roles in regulating the EMT program—SNAI2 and SOX4, were also found to be upregulated in SS adherent cells (Supplementary Table S6; see Discussion).
Also as expected, we found an upregulation of the expression of several TGF-β1 signalling components (discussed later), including the TGF-β1 ligand itself, the two TGF-β1 receptors (TGFβRI and TGFβRII), the TGF-β induced (TGFβI) and TGF-β1 homeobox (TGIF1) proteins as well as SMAD3, SMAD7, and two SMAD specific ubiquitin ligases (SMURF 1 and 2) (Table 1). To address whether the canonical pathway is activated in the adherent cells, we specifically looked for upregulated target genes of the TGF-β1/Smad signalling pathway. In addition to SMAD7 and TGF-β1, upregulated genes included SERPINE1, LTBP2, and SNAI2 (Table 2).
TGF-β1-induced adherent H2122 SS cells overexpress EGFR
Finally, we investigated the expression of other growth factors and receptors known to be upregulated in cancer. Among growth factors and receptors, genes coding for two subunits of the Platelet-Derived Growth Factor (PDGFB and PDGFC), a Vascular Endothelial Growth Factor (VEGFA), the Fibroblast Growth Factor Receptor 1 (FGFR1), and several FGFR ligands (e.g., FGF1 and 2) were upregulated in the adherent cells relative to the control suspension cultures (Table 3). Interestingly, although we did not find an upregulation of the Epidermal Growth Factor (EGF), we found increased expression of the Epidermal Growth Factor Receptor (EGFR) and several of its other ligands, including TGF-\(\alpha\), a heparin-binding EGF-like growth factor, amphiregulin, epiregulin, and neuregulin (50; Table 3). Consistent with their induction, we also observed the upregulation of several proteases from the ADAM and ADAMT families, which are known to be involved in the proteolytic cleavage of these ligands that are synthesized as transmembrane proteins51,52,53.
Both Erlotinib and Osimertinib inhibit the TGF-β1-induced adherence of H2122 SS cells
To address whether the TGF-β1-induced upregulation of EGFR is specifically involved in the expression of the adherent phenotype (and thus could be a target to reduce TGF-β1-induced adherence), we used two EGFR inhibitors—Erlotinib and Osimertinib. Both drugs act by inhibiting the tyrosine kinase portion of the EGF receptor, but Osimertinib is a more effective, third-generation drug that irreversibly binds the receptor and is more specific to both sensitizing and TKI-resistance mutations (e.g.,37,54). We evaluated their effects (at pharmacologically relevant concentrations; see Discussion) on the proliferation, viability, and adherence of SS cells alone or in the presence of TGF-β1.
First, we observed that following a 72 h-treatment, both Erlotinib and Osimertinib alone had a negative effect on population growth (ca. 50% decrease); this was primarily due to inhibiting cell proliferation (as the number of dead cells was only slightly affected) (Fig. 3a). We also noted that, as previously observed29, exposure to 10 ng/ml TGF-β1 induced the adherence of SS cells, concomitant with a strong inhibition of cell proliferation. Specifically, up to 88% of the cells adhered, while the overall population only reached ca. 50% of the control population size (Fig. 3a).
The effect of Erlotinib and Osimertinib on the TGF-β1-induced adherence (top row) and viability (bottom row) of H2122 SS cultures exposed to TGF-β1. (a) H2122 SS cultures were seeded at a density of 1 × 105 live cells/ml (indicated by the dotted line) in triplicates and treated with 5 µM Erlotinib, 5 µM Osimertinib, 10 ng/ml TGF-β1, 10 ng/ml TGF-β1 with 5 µM Erlotinib and 10 ng/ml TGF-β1 with 5 µM Osimertinib for 72 h. ns indicates not significant (p ≥ 0.05), while *, **, and **** denote p < 0.05, p < 0.01, and p < 0.0001, respectively (One-way Brown-Forsythe and Welch ANOVA tests); significance levels above individual graph bars are relative to controls. (b,c) Same as (a), but using 2.5 µM (b) or 15 µM (c) Erlotinib. (d) H2122 SS cultures were first seeded at a density of 1 × 105 live cells/ml (in triplicates) in the presence of 10 ng/ml TGF-β1 and allowed to adhere for 72 h. Then, the old medium and unattached cells were removed and replaced with fresh medium without or with 10 µM Erlotinib for 72 h. ns indicates not significant (p ≥ 0.05), while * denotes p < 0.05 (Welch’s t-tests). Data shown are from one representative independent experiment (of at least 3) with three replicates; graphs showing each data point are included in Supplementary Material (to avoid overcrowding in stacked bars).
However, we found that in cultures exposed to TGF-β1 in the presence of 5 µM Erlotinib or 5 µM Osimertinib, the number of adherent cells was drastically reduced (while the number of cells in suspension increased), resulting in a significant change (relative to TGF-β1 alone) in the proportion of adherent cells in SS cells exposed to TGF-β1, from 88 to 15% and 18%, respectively (Fig. 3a). Yet, the addition of Erlotinib or Osimertinib to TGF-β1-treated cultures did not decrease the number of live cells relative to TGF-β1-treated cultures (both were maintained around the initial seeding density); nevertheless, Erlotinib slightly increased the number of dead cells (Fig. 3a).
Next, we wanted to address whether the observed effects are dose dependent. Since H2122 is not EGFR-mutated and Osimertinib is less effective against wild-type EGFR (see Discussion), the subsequent experiments were performed with Erlotinib only. We found that in the presence of TGF-β1, a lower Erlotinib concentration (2.5 µM) still affected the TGF-β1-induced adherence (only 32% of cells adhered) but had a smaller effect on cell proliferation (Fig. 3b). On the other hand, a higher Erlotinib concentration (15 µM) affected both the proportion of adherent cells (down to 18%) and the total number of live cells (a ca. 30% reduction; from 1.2 × 105 to 8.6 × 104 cells/ml) (Fig. 3c).
Because most TGF-β1-induced adherent SS cells remain adherent even in the absence of TGF-β129, we also addressed whether the overexpression of EGFR is implicated in the stability of the adherent phenotype. SS cultures were treated with 10 ng/ml TGF-β1 and allowed to adhere for 72 h. Then, the medium and the unattached cells were removed and replaced with fresh medium without (control) or with 10 µM Erlotinib. After 72 h, the number of live and dead cells was assessed in both the adherent and suspension fractions (Fig. 3d). We found that in the presence of Erlotinib, there is a decreased proportion of live adherent cells relative to control cultures (36.5% vs. 75.5%), indicating that Erlotinib facilitates the detachment of cells that adhered in response to TGF-β1 (note that the numbers of dead cells are similar between control and Erlotinib cultures, suggesting that the detachment is not associated with cell death).
Erlotinib inhibits the invasion of TGF-β1-induced adherent H2122 SS cells
Lastly, because we previously found that TGF-β1 also promotes the invasiveness of H2122 SS cells29, we addressed whether EGFR is involved in the observed TGF-β1-induced invasion and whether EGFR inhibitors can decrease the invasive potential of SS cells. To do this, we assessed the number of cells that were able to invade (over 72 h) in response to TGF-β1 in the presence or absence of 5 or 10 µM Erlotinib (Fig. 4). We observed that, as expected29, exposure to TGF-β1 drastically increased the invasive potential of SS cells. Furthermore, we found that the addition of Erlotinib decreased the number of invaded cells (by 23% and 40% for 5 and 10 µM Erlotinib, respectively), suggesting that in addition to adherence, EGFR also contributes to the TGF-β1-induced invasion of SS cells.
The effect of Erlotinib at 5 µM (a) and 10 µM (b) on the TGF-β1-induced invasion of SS cells. Untreated H2122 SS cells (control) and cells in the presence of either Erlotinib (5 µM and 10 µM), TGF-β1 (10 ng/ml), or TGF-β1 and Erlotinib were allowed to invade for 72 h, and the number of invaded cells was assessed. ns indicates not significant (p ≥ 0.05), while *, **, and *** denote p < 0.05, p < 0.01, and p < 0.001, respectively (One-way ANOVA: Brown-Forsythe and Welch ANOVA tests). Data shown are from one representative independent experiment (of at least 3) with three replicates.
Discussion
Recently, based on the extensive association between platelets and CTC clusters in the blood and a series of in vitro experiments using the H2122 SS cell line, we proposed a new mechanism for the extravasation of CTC clusters that express TGF-β1 receptors29. Specifically, to mimic the association between CTC clusters and platelets that has been observed in vivo8, we exposed H2122 SS cells to exogenous TGF-β1 (at a physiologically relevant concentration) and found that TGF-β1 caused a distinct phenotypic switch from clusters of cells to single adherent cells with invasive capabilities. In addition, we also observed that once adherent, the H2122 SS cells released their own TGF-β1, which is required for their ability to migrate in the absence of exogenous TGF-β129. In the current study, we set out to investigate the signalling pathways involved in the observed TGF-β1-induced phenotypic changes with the goal of identifying druggable targets to decrease the metastatic potential of TGF-β1-responsive CTC clusters.
TGF-β1-induced adherence of H2122 SS involves the TGF-β/Smad and Wnt/β-catenin signalling pathways
TGF-β1 is known to have various, sometimes contradictory effects highly dependent on the cellular context. This context-dependent response is partly due to extensive crosstalk between the pathways it activates55 as well as with other signalling pathways such as the Wnt, BMP, and TNF pathways56. TGF-β1 ligands act as dimers and initiate intracellular signals when each monomer binds to a set of receptor tyrosine kinases—TGFβRI and TGFβRII57. Upon receptor binding, the cytoplasmic kinase portion of the receptors triggers a set of phosphorylation cascades in the cell, activating various intracellular proteins. The canonical pathway, known as the SMAD signalling pathway, involves direct phosphorylation of SMAD2/3 proteins by the kinase portion of TGFβRI58. Phosphorylated SMAD 2 and 3 then complex with SMAD4 and collectively enter the nucleus, where they associate with various cofactors and bind the promoter region of target genes (e.g.,59) (Fig. 5a). Various non-canonical signalling pathways also exist in which TGF-β1 receptors activate non-SMAD proteins such as MAPK/ERK, PI3K/AKT/mTOR, and JNK/p38/NF-κB (e.g.,55).
Our proposed model depicting (a) the crosstalk between TGF-β1 and EGFR signalling pathways resulting in increased adherence and invasion of CTC clusters, and (b) the effect of Erlotinib on the dispersal potential of CTC clusters. TGF-β1 released by platelets induces the adherence and invasion of CTCs through both the activation of the TGF-β1 canonical pathway and the overexpression of EGFR and EGFR ligands that can activate the EGF pathway. The activation of the EGF receptors by EGFR ligands can be inhibited by EGFR inhibitors, which can result in a decrease in the adherence and extravasation of CTC clusters.
As expected, in response to TGF-β1 we found that SS cells upregulated the expression of several TGF-β1 signalling components, including the TGF-β1 ligand itself, the two TGF-β1 receptors (TGFβRI and TGFβRII), the TGF-β induced (TGFβI) protein, and the TGF-β1 homeobox (TGIF1) (Table 1). TGF-β1 has been shown to upregulate its own transcription60 as well as that of its type II receptors61. The observed upregulation of TGF-β1 itself (Table 1) is consistent with our previous finding that once adhered, the SS cells release their own TGF-β1 to which they continue to respond29. Autocrine signalling (autonomy; self-sufficiency) involving growth factors has been recently proposed to be specifically induced in CTCs; and, consistent with their increased metastatic potential, this “autonomy signature” was more significant in CTC clusters than in single CTCs62. Interestingly, the expression of the TGFβI protein was also reported in a highly malignant circulating tumour cell subline from pancreatic cancer63.
To address whether the canonical pathway is activated in the TGF-β1-induced adherent cells, we looked for specific components and target genes. In addition to SMAD3, SMAD7, and two SMAD-specific ubiquitin ligases (SMURF 1 and 2), we found that several target genes of the TGF-β1/Smad signalling pathway were upregulated, including SERPINE1, LTBP2, and SNAI264,65 (Table 2). Overall, these findings suggest that the SMAD2/3 pathway plays a role in the induction of the adherent phenotype. Notably, LTBP2 is associated with poor prognosis and an aggressive phenotype in several cancers66. For example, in lung cancer, increased expression of LTBP2 is associated with increased migration67.
We also found that the most upregulated gene in response to TGF-β1 codes for the Wnt ligand Wnt7a (Log2FoldChange = 11.49). In the canonical Wnt signalling pathway, Wnt ligands bind the extracellular domain of Frizzled family receptors, which disrupts the degradation of β-catenin causing its accumulation in the cytoplasm68. Then, β-catenin translocates into the nucleus where it interacts with the TCF/Lef1 complex and induces gene transcription of target genes68. Besides Wnt7a, we also found a significant upregulation of the genes encoding Wnt9a and Wnt7b (Tables 3, S2, S5). Similarly, we observed an increase in the transcription of β-catenin and several Wnt/β-catenin target genes such as JUN, CD24, and CD4469,70. Overall, these findings suggest that Wnt/β-catenin signalling also plays a role alongside the Smad2/3 signalling pathway to induce the adherent phenotype and/or prime cells for migration. Indeed, the TGF-β and Wnt signalling pathways are known to overlap in a variety of cancer types71,72,73. In lung cancer, crosstalk between these pathways has been shown to promote metastasis by controlling the development of stem cell properties and promoting EMT74.
TGF-β1 and EGFR signalling pathways work cooperatively to increase the dispersal potential of H2122 SS clusters
In addition to activating the Smad and Wnt signalling pathways, TGF-β1 induced other genes known to be overexpressed in cancer (Table 3). Unexpectedly, among them, we noticed the upregulation of the EGFR gene. The EGF/EGFR signalling pathway regulates various cellular processes (e.g.,75) and is involved in triggering EMT in several cell lines (e.g.,76,77,78). Generally, a specific balance between the cell survival and proliferative effects of the EGF/EGFR pathway and the antiproliferative and anti-apoptotic effects of TGF-β signalling is necessary to maintain healthy tissue homeostasis. However, in advanced carcinogenesis, the two pathways can act synergistically to induce a malignant phenotype through cooperative interactions. For instance, crosstalk between TGF-β1 and EGFR signalling has been shown to be involved in the acquisition of mesenchymal characteristics and increased migration/invasion in various cancer cell lines; though the exact mechanisms (including the specific downstream pathways activated by EGFR) remain to be fully elucidated32,33,34,79,80.
Interestingly, using two breast cancer lines, Zhao et al.32 showed that TGF-β1 elicited a significant increase in EGFR transcription and that EGFR was essential for the TGF-β1-induced increase in migration and invasion. Specifically, both the knockdown of EGFR and the inhibition of EGFR using Erlotinib decreased the TGF-β1-induced cell migration and invasion32. Based on their data, the authors proposed a novel model for how TGF-β1 and EGFR pathways act synergistically to increase the migration and invasion abilities of breast cancer cells through the upregulation of EGFR (via Smad3 and ERK/Sp1). Notably, although Erlotinib is generally considered a targeted drug against EGFR with activating mutations, the two breast cancer cell lines whose migration and invasion were reduced by Erlotinib express wild-type EGFR.
Consistent with these findings, we also found that TGF-β1 induced the overexpression of EGFR in the lung cancer H2122 SS line. Furthermore, Erlotinib also negatively affected the TGF-β1-induced cell adherence and invasion of SS cells, indicating that EGFR contributes to the TGF-β1-induced adherence and invasion of the SS cells despite the absence of an activating EGFR mutation in H212281. However, our finding that TGF-β1 also induces the overexpression of TGF-α and other EGFR ligands suggests that cells exposed to TGF-β1 and overexpressing EGFR can actually become Erlotinib-sensitive due to ligand-dependent activation of EGFR.
Thus, based on our in vitro data, we propose a possible model for (i) the TGF-β1-induced adherence and invasion of CTCs involving the transactivation of the EGFR pathway through the TGF-β1-induced transcription of genes coding for both EGFR and EGFR ligands (Fig. 5a) and (ii) the inhibitory effect of Erlotinib on the dispersal potential of CTCs (Fig. 5b). Although the exact mechanism remains to be investigated, our results are consistent with previous studies reporting that Erlotinib can reduce the migration and invasion of several adherent cell lines with wild-type EGFR genes that are overexpressed or are upregulated by TGF-β1 (e.g., breast, lung, colon, gastric, and ovarian32,38,39,40).
Notably, in addition to the upregulation of the EGF/EGFR pathway, we observed the induction of the FGF/FGFR pathway. FGFR1 is also a tyrosine kinase receptor whose overexpression (in FGFR1-amplified lung cancer) was associated with EMT and metastasis through the activation of the FGFR1-ERK1/2-SOX2 axis82. Interestingly, FGFR1 was found to be overexpressed in CTC clusters isolated from a patient with stage IV adenocarcinoma83. Also, the TGF-β1-induction of FGF1 and FGF2 that we found suggests that, in addition to the involvement of the EGF/EGFR pathway (Fig. 5), the FGF/FGFR pathway might also contribute to the adherence and invasion of SS cells. This possibility is consistent with the report of FGFR1 upregulation interfering with the response to TKIs in EGFR-mutated NSCLC84.
Clinical significance
We have previously argued that the H2122 SS line can be used as an in vitro model system to understand the biology of CTC clusters and to develop strategies to decrease the metastatic potential of TGF-β1-responsive CTC clusters22,23. Here, we show that the TGF-β1-induced adherence in the H2122 SS line is associated with the upregulation of several genes that have been previously implicated in the ability of real CTC clusters to metastasize. For example, consistent with our findings, a study that investigated the molecular characteristics of breast CTC clusters coated with platelets found their transcriptome to be enriched in TGF-β and Wnt signalling components as well as extracellular matrix (ECM) production and ECM-related receptors (including integrin receptors)85. Additionally, the most abundant transcript was SERPINE1 (a target of the canonical TGF-β pathway—known to promote invasiveness86), which was also induced by TGF-β1 in the adherent H2122 SS cells (Table 1). Similarly, consistent with our transcriptomic data, EGFR was often found to be expressed (and even phosphorylated) in CTCs and thought to contribute to their intravasation and survival87,88,89,90. In fact, one study used the expression of EGFR to detect CTCs in lung cancer patients with brain metastases (note that 52.3% and 62.7% of primary tumours and brain metastases, respectively, were positive for EGFR)91.
Furthermore, autocrine signalling involving the EGFR pathway has recently been proposed to be a signature of CTCs and provide them with the autonomy required for maintaining their proliferative and metastatic potential while in circulation and at new sites62. The TGF-β1-induced expression of both TGF-β1 as well as EGFR and EGF ligands that we observed in our system provides such autocrine signalling and autonomy that could allow CTCs to extravasate in the absence of additional sources of growth factors. Furthermore, our gene expression analyses identified three DEGs associated with the clinically validated lung metastasis gene signature for breast cancer involved in vascular remodelling during intravasation and extravasation92. They code for an EGFR ligand (Epiregulin), the prostaglandin-synthesizing enzyme cyclooxygenase 2 (COX2/PTGS2), and a metalloproteinase (MMP1) (Supplementary Table 4). Overall, our data indicate that this in vitro model system could be used to further explore (and test in vivo) therapeutic strategies directed against CTC clusters that overexpress EGFR (see discussion below), either constitutively or induced by TGF-β1.
Therapeutic implications
Dysregulated EGF/EGFR signalling has been shown in many cancer types (e.g., breast, colon, lung, head and neck, pancreas, liver, and bladder). It involves multiple mechanisms, including EGFR gene amplifications, EGFR overexpression, EGFR activating mutations (resulting in constitutive/ligand-independent activation), mutations that specifically trigger the expression of EGFR and its ligands, as well as the EGFR transactivation by other receptor tyrosine kinases, including TGF-β1 (e.g.33,93,94,95,96,97). For instance, EGFR is amplified or overexpressed in most head and neck cancers98 and a significant proportion of pancreatic cancers99. Consequently, many drugs have been developed to inhibit proliferation or activate apoptosis of cells with specific EGFR amplifications or activating mutations (e.g.,96,100,101).
Erlotinib is a tyrosine kinase inhibitor that reversibly binds the ATP-binding site and affects the kinase activity of EGFR. Because of its greater binding affinity for mutant EGFR proteins compared to wild-type counterparts, Erlotinib has been used effectively against tumours with activating EGFR mutations (e.g.,100) to inhibit cell proliferation and/or induce apoptosis. However, this drug has also been shown to affect tumour progression and overall survival in patients with wild-type EGFR102,103,104, though its use is not approved anymore in these cases105. The exact mechanisms for the efficacy of Erlotinib in such cases are not well understood but might reflect the presence of mutations in other pathways that result in the overexpression and/or activation/transactivation of EGFR (e.g.,96,106). The former suggestion is consistent with reports of Erlotinib improving outcome in NSCLC patients with wild-type EGFR but high levels of expression104. Since EGFR is not mutated or amplified in H212281, our results support the possibility that TGF-β1-induced EGFR overexpression and activation can render cancer cells sensitive to EGFR inhibitors in the absence of amplifying or activating EGFR mutations81.
In addition, consistent with other reports, our data argue that TKIs can be effective against cells that express wild-type EGFR cells not only in terms of decreasing proliferation and viability but also as far as inhibiting cell invasion. Furthermore, since EGFR can be overexpressed in CTC clusters or could be induced and activated in response to exogenous TGF-β1 released by platelets, we suggest that the use of TKIs has the potential to reduce the dispersal of CTCs by affecting their extravasation potential.
Notably, the lowest concentration of Erlotinib that we found to affect the adherence and invasion of the H2122 SS cells (i.e., 2.5 µM) is in the lower range reported for plasma Erlotinib levels (i.e., between 1.3 and 9.5 µM) in patients taking standard doses of Erlotinib (150 mg/day)107,108,109,110,111,112,113,114,115. This suggests that Erlotinib might be effective at low doses, which could decrease its toxicity and allow for long-term use. Moreover, since Erlotinib (at concentrations as low as 1–5 µM) was shown to decrease the migration and invasion of several adherent cell lines with wild-type EGFR (e.g., breast, lung, colon, gastric, and ovarian32,38,39,40), its use could contribute to improving the prognosis of patients with advanced cancers by affecting both the potential of cells to invade nearby tissues (and enter the circulatory system) along with their ability to extravasate and disseminate, irrespective of their EGFR status.
The fact that both Osimertinib and Erlotinib had similar outcomes indicates that the observed inhibition of adherence was not due to off-target effects and suggests that similar responses can be induced by other TKIs. However, the concentration of Osimertinib we found to be effective in reducing the TGF-β1-induced adherence of SS cells (5 µM) was higher than the plasma levels reported in patients with mutant EGFR taking daily doses of 80 or 160 mg (between 0.62 and 1.26 µM116). The increased levels of Osimertinib required to inhibit adherence in our experiments likely reflect its much lower affinity for wild-type EGFR54; specifically, Osimertinib shows a ca. 200-fold greater potency against mutant EGFR117. Thus, lower doses of Osimertinib might be effective against the TGF-β1-induced extravasation of CTCs expressing EGFR with sensitizing or TKI-resistant mutations32,38,39,40. Notably, a novel EGFR inhibitor that targets both wild-type and mutant EGFR forms as well as an agent that potentiates Erlotinib to wild-type EGFR have recently been reported118,119.
Interestingly, Cetuximab—an approved drug that targets the extracellular portion of the EGFR (and is effective against cancer cells with amplified EGFR120), in combination with a COX2 inhibitor (Celecoxib—a non-steroidal anti-inflammatory drug), inhibited the formation of lung metastases by interfering with the extravasation of CTCs92. In addition, breast cancer cells that preferentially infiltrate the lungs and brain were found to overexpress EGFR ligands, and the addition of Cetuximab inhibited their in vitro migration and increased brain metastasis-free survival in mice121. Cetuximab has also been shown to inhibit EGFR-mediated invasion in a head and neck squamous cell carcinoma line122, suggesting that it might affect the dispersal of CTC clusters with amplified EGFR. Notably, one of the differentially expressed genes during EGFR-mediated EMT in this line (whose expression was reduced by Cetuximab) was ITGB4 (integrin beta 4), which is a gene that was up-regulated in the SS line in response to TGF-β1.
Overall, we argue that the impact of EGFR inhibitors, alone or in combination with other TKIs (e.g.38,40), on the metastatic potential of CTC clusters should be further investigated. Furthermore, consistent with other reports, our findings indicate that EGFR inhibitors have the potential to be effective in improving the prognosis of cancer patients without EGFR mutations (which represent, for instance, ca. 85–90% of Caucasian lung adenocarcinoma patients and up to 60% of patients of Asian descent123) but with overexpressed and/or ligand-activated EGFR including through non-cell-autonomous mechanisms (such as TGF-β1 released by other cells).
Conclusion
This study (i) investigated the molecular basis of our recently proposed model for the extravasation of CTC clusters involving TGF-β1 released by platelets, (ii) revealed potential pathways involved in the TGF-β1-induced adherence of CTC clusters and (iii) identified TGF-β1-induced overexpression and activation of EGFR as a potential therapeutic target to reduce the metastatic potential of CTC clusters that are responsive to TGF-β1 and/or overexpress EGFR. Based on our findings, we suggest that EGFR inhibitors might be effective drugs to slow cancer progression by affecting the dispersal potential of CTC clusters even in the absence of specific EGFR mutations. Our proof-of-principle study underscores the need for a better understanding of the molecular mechanisms involved in the extravasation and dissemination of CTC clusters to develop new therapeutic strategies that can specifically affect this important step in the metastatic cascade and improve clinical outcomes. Furthermore, our findings highlight an under-explored target for the development of drugs—namely, induced changes in gene expression associated with specific cellular states and stages during cancer progression.
Data availability
All relevant data can be found within the article and its supplementary information. Raw sequence reads were submitted to NCBI under Accession PRJNA1093148.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 Countries. CA Cancer J. Clin. 71(3), 209–249. https://doi.org/10.3322/caac.21660 (2021).
Dillekås, H., Rogers, M. S. & Straume, O. Are 90% of deaths from cancer caused by metastases?. Cancer Med. 8(12), 5574–5576. https://doi.org/10.1002/cam4.2474 (2019).
Massagué, J. & Obenauf, A. C. Metastatic colonization by circulating tumour cells. Nature 529(7586), 298–306. https://doi.org/10.1038/nature17038 (2016).
Aceto, N. et al. Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell 158(5), 1110–1122. https://doi.org/10.1016/j.cell.2014.07.013 (2014).
Lambert, A. W., Pattabiraman, D. R. & Weinberg, R. A. Emerging biological principles of metastasis. Cell 168(4), 670–691. https://doi.org/10.1016/j.cell.2016.11.037 (2017).
Zhang, D. et al. Circulating tumor microemboli (CTM) and vimentin+ circulating tumor cells (CTCs) detected by a size-based platform predict worse prognosis in advanced colorectal cancer patients during chemotherapy. Cancer Cell Int. 17(1), 6. https://doi.org/10.1186/s12935-016-0373-7 (2017).
Jansson, S., Bendahl, P. O., Larsson, A. M., Aaltonen, K. E. & Rydén, L. Prognostic impact of circulating tumor cell apoptosis and clusters in serial blood samples from patients with metastatic breast cancer in a prospective observational cohort. BMC Cancer 16(1), 433. https://doi.org/10.1186/s12885-016-2406-y (2016).
Lim, M. et al. Circulating tumor cell clusters are cloaked with platelets and correlate with poor prognosis in unresectable pancreatic cancer. Cancers 13(21), 5272. https://doi.org/10.3390/cancers13215272 (2021).
Au, S. H. et al. Microfluidic isolation of circulating tumor cell clusters by size and asymmetry. Sci. Rep. 7(1), 2433. https://doi.org/10.1038/s41598-017-01150-3 (2017).
Hou, J. M. et al. Circulating tumor cells as a window on metastasis biology in lung cancer. Am. J. Pathol. 178(3), 989–996. https://doi.org/10.1016/j.ajpath.2010.12.003 (2011).
Umer, M., Vaidyanathan, R., Nguyen, N. T. & Shiddiky, M. J. A. Circulating tumor microemboli: Progress in molecular understanding and enrichment technologies. Biotechnol. Adv. 36(4), 1367–1389. https://doi.org/10.1016/j.biotechadv.2018.05.002 (2018).
Pereira-Veiga, T., Schneegans, S., Pantel, K. & Wikman, H. Circulating tumor cell-blood cell crosstalk: Biology and clinical relevance. Cell Rep. https://doi.org/10.1016/J.CELREP.2022.111298 (2022).
Maeshiro, M. et al. Colonization of distant organs by tumor cells generating circulating homotypic clusters adaptive to fluid shear stress. Sci. Rep. 11(1), 6150. https://doi.org/10.1038/s41598-021-85743-z (2021).
Zhao, Q. et al. Interaction between circulating galectin-3 and cancer-associated MUC1 enhances tumour cell homotypic aggregation and prevents anoikis. Mol. Cancer 9(24), 1937–1947. https://doi.org/10.1186/1476-4598-9-154 (2010).
Lo, H. C. et al. Circulating tumor cell clusters exhibit enhanced immune evasion from natural killer cells. J. Immunol. 204(1 Supplement), 8818 (2020).
Bithi, S. S. & Vanapalli, S. A. Microfluidic cell isolation technology for drug testing of single tumor cells and their clusters. Sci. Rep. 7, 41707. https://doi.org/10.1038/srep41707 (2017).
Gkountela, S. et al. Circulating tumor cell clustering shapes DNA methylation to enable metastasis seeding. Cell 176(1–2), 98-112.e14. https://doi.org/10.1016/j.cell.2018.11.046 (2019).
Labuschagne, C. F., Cheung, E. C., Blagih, J., Domart, M. C. & Vousden, K. H. Cell clustering promotes a metabolic switch that supports metastatic colonization. Cell Metabol. 30(4), 720-734.e5. https://doi.org/10.1016/j.cmet.2019.07.014 (2019).
Wrenn, E., Huang, Y. & Cheung, K. Collective metastasis: Coordinating the multicellular voyage. Clin. Exp. Metastasis 38(4), 373–399. https://doi.org/10.1007/S10585-021-10111-0 (2021).
Nasr, M. M. & Lynch, C. C. How circulating tumor cluster biology contributes to the metastatic cascade: From invasion to dissemination and dormancy. Cancer Metastasis Rev. 42(4), 1133–1146. https://doi.org/10.1007/S10555-023-10124-Z (2023).
Sharma, S. et al. Circulating tumor cell isolation, culture, and downstream molecular analysis. Biotechnol. Adv. 36(4), 1063–1078. https://doi.org/10.1016/J.BIOTECHADV.2018.03.007 (2018).
May, A. N., Crawford, B. D. & Nedelcu, A. M. In vitro model-systems to understand the biology and clinical significance of circulating tumor cell clusters. Front. Oncol. https://doi.org/10.3389/fonc.2018.00063 (2018).
Jong, E. D., Chan, I. C. W. & Nedelcu, A. M. A model-system to address the impact of phenotypic heterogeneity and plasticity on the development of cancer therapies. Front. Oncol. 9, 842. https://doi.org/10.3389/fonc.2019.00842 (2019).
Liu, X. et al. Homophilic CD44 interactions mediate tumor cell aggregation and polyclonal metastasis in patient-derived breast cancer models. Cancer Discov. 9(1), 96–113. https://doi.org/10.1158/2159-8290.CD-18-0065 (2019).
Donato, C. et al. Hypoxia triggers the intravasation of clustered circulating tumor cells. Cell Rep. https://doi.org/10.1016/j.celrep.2020.108105 (2020).
Giuliano, M. et al. Perspective on circulating tumor cell clusters: Why it takes a village to metastasize. Cancer Res. 78(4), 845–852. https://doi.org/10.1158/0008-5472.CAN-17-2748 (2018).
Au, S. H. et al. Clusters of circulating tumor cells traverse capillary-sized vessels. Proc. Natl. Acad. Sci. USA 113(18), 4947–4952. https://doi.org/10.1073/pnas.1524448113 (2016).
Allen, T. A. et al. A zebrafish model of metastatic colonization pinpoints cellular mechanisms of circulating tumor cell extravasation. Front. Oncol. 11, 641187. https://doi.org/10.3389/fonc.2021.641187 (2021).
Hapeman, J. D., Carneiro, C. S. & Nedelcu, A. M. A model for the dissemination of circulating tumour cell clusters involving platelet recruitment and a plastic switch between cooperative and individual behaviours. BMC Ecol. Evolut. https://doi.org/10.1186/s12862-023-02147-5 (2023).
Labelle, M., Begum, S. & Hynes, R. O. Direct signaling between platelets and cancer cells induces an epithelial-mesenchymal-like transition and promotes metastasis. Cancer Cell 20(5), 576–590. https://doi.org/10.1016/j.ccr.2011.09.009 (2011).
Colak, S. & ten Dijke, P. Targeting TGF-β Signaling in Cancer. Trends Cancer 3(1), 56–71. https://doi.org/10.1016/j.trecan.2016.11.008 (2017).
Zhao, Y. et al. TGF-β transactivates EGFR and facilitates breast cancer migration and invasion through canonical Smad3 and ERK/Sp1 signaling pathways. Mol. Oncol. 12(3), 305–321. https://doi.org/10.1002/1878-0261.12162 (2018).
Kang, M. et al. Cross-talk between TGFβ1 and EGFR signalling pathways induces TM4SF5 expression and epithelial-mesenchymal transition. Biochem. J. 443(3), 691–700. https://doi.org/10.1042/BJ20111584 (2012).
Li, L. et al. Transforming growth factor-β1 induces EMT by the transactivation of epidermal growth factor signaling through HA/CD44 in lung and breast cancer cells. Int. J. Mol. Med. 36(1), 113–122. https://doi.org/10.3892/ijmm.2015.2222 (2015).
Xu, M. J., Johnson, D. E. & Grandis, J. R. EGFR-targeted therapies in the post-genomic era. Cancer Metastasis Rev. 36(3), 463–473. https://doi.org/10.1007/s10555-017-9687-8 (2017).
Shepherd, F. A. et al. Erlotinib in previously treated non-small-cell lung cancer. N. Engl. J. Med. 353(2), 123–132. https://doi.org/10.1056/NEJMoa050753 (2005).
Yuan, Z. et al. Instability mechanism of osimertinib in plasma and a solving strategy in the pharmacokinetics study. Front. Pharmacol. 13, 928983. https://doi.org/10.3389/FPHAR.2022.928983 (2022).
Glaser, F. et al. KCa channel blockers increase effectiveness of the EGF receptor TK inhibitor erlotinib in non-small cell lung cancer cells (A549). Sci. Rep. https://doi.org/10.1038/s41598-021-97406-0 (2021).
Shen, T. et al. Erlotinib inhibits colon cancer metastasis through inactivation of TrkB-dependent ERK signaling pathway. J. Cell. Biochem. 120(7), 11248–11255. https://doi.org/10.1002/jcb.28400 (2019).
Hu, X., Wu, L.-W., Weng, X., Lin, N.-M. & Zhang, C. Synergistic antitumor activity of aspirin and erlotinib: Inhibition of p38 enhanced aspirin plus erlotinib-induced suppression of metastasis and promoted cancer cell apoptosis. Oncol. Lett. 16(2), 2715–2724. https://doi.org/10.3892/ol.2018.8956 (2018).
Justus, C. R., Leffler, N., Ruiz-Echevarria, M. & Yang, L. V. In vitro cell migration and invasion assays. J. Vis. Exp. https://doi.org/10.3791/51046 (2014).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5(7), 621–628. https://doi.org/10.1038/nmeth.1226 (2008).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46(W1), W537–W544. https://doi.org/10.1093/nar/gky379 (2018).
Tang, Y., Horikoshi, M. & Li, W. ggfortify: Unified interface to visualize statistical results of popular R packages. R J. 8(2), 478–489 (2016).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. https://doi.org/10.1186/gb-2010-11-10-r106 (2010).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: Accounting for selection bias. Genome Biol. 11(2), 1–12. https://doi.org/10.1186/GB-2010-11-2-R14/TABLES/4 (2010).
Xijin Ge, S., Jung, D. & Yao, R. ShinyGO: A graphical gene-set enrichment tool for animals and plants. Bioinformatics 36(8), 2628–2629. https://doi.org/10.5281/zenodo.1451847 (2019).
Walton, K. L., Johnson, K. E. & Harrison, C. A. Targeting TGF-β mediated SMAD signaling for the prevention of fibrosis. Front. Pharmacol. 8, 277037. https://doi.org/10.3389/FPHAR.2017.00461/BIBTEX (2017).
Wilson, K. J., Gilmore, J. L., Foley, J., Lemmon, M. A. & Riese, D. J. Functional selectivity of EGF family peptide growth factors: Implications for cancer. Pharmacol. Ther. 122(1), 1–8. https://doi.org/10.1016/j.pharmthera.2008.11.008 (2009).
Kenny, P. A. & Bissell, M. J. Targeting TACE-dependent EGFR ligand shedding in breast cancer. J. Clin. Investig. 117(2), 337–345. https://doi.org/10.1172/JCI29518 (2007).
Gschwind, A., Hart, S., Fischer, O. M. & Ullrich, A. TACE cleavage of proamphiregulin regulates GPCR-induced proliferation and motility of cancer cells. EMBO J. 22(10), 2411–2421. https://doi.org/10.1093/EMBOJ/CDG231 (2003).
Lu, X. et al. ADAMTS1 and MMP1 proteolytically engage EGF-like ligands in an osteolytic signaling cascade for bone metastasis. Genes Dev. 23(16), 1882–1894. https://doi.org/10.1101/GAD.1824809 (2009).
Jiang, T. et al. A consensus on the role of osimertinib in non-small cell lung cancer from the AME Lung Cancer Collaborative Group. J. Thoracic Dis. 10(7), 3909–3921. https://doi.org/10.2103/jtd.2018.07.61 (2018).
Zhang, Y. E. Non-Smad signaling pathways of the TGF-β family. Cold Spring Harbor Persp. Biol. https://doi.org/10.1101/cshperspect.a022129 (2017).
Guo, X. & Wang, X. F. Signaling cross-talk between TGF-β/BMP and other pathways. Cell Res. 19(1), 71–88. https://doi.org/10.1038/cr.2008.302 (2009).
Huang, T. et al. TGF-β signalling is mediated by two autonomously functioning TβRI:TβRII pairs. EMBO J. 30(7), 1263–1276. https://doi.org/10.1038/emboj.2011.54 (2011).
vander Ark, A., Cao, J., & Li, X. (2018). TGF-β receptors: In and beyond TGF-β signaling. In Cellular Signalling (Vol. 52, pp. 112–120). Elsevier Inc. https://doi.org/10.1016/j.cellsig.2018.09.002
Massagué, J. TGFβ signalling in context. Nat. Rev. Mol. Cell Biol. 13(10), 616–630. https://doi.org/10.1038/nrm3434 (2012).
Zhang, M., Fraser, D. & Phillips, A. ERK, p38, and Smad signaling pathways differentially regulate transforming growth factor-β1 autoinduction in proximal tubular epithelial cells. Am. J. Pathol. 169(4), 1282–1293. https://doi.org/10.2353/ajpath.2006.050921 (2006).
Duan, D. & Derynck, R. Transforming growth factor-β (TGF-β)-induced up-regulation of TGF-β receptors at the cell surface amplifies the TGF-β response. J. Biol. Chem. 294(21), 8490–8504. https://doi.org/10.1074/jbc.RA118.005763 (2019).
Sinha, S. et al. Growth signaling autonomy in circulating tumor cells aids metastatic seeding. PNAS Nexus https://doi.org/10.1093/PNASNEXUS/PGAE014 (2024).
Sato, T., Muramatsu, T., Tanabe, M. & Inazawa, J. Identification and characterization of transforming growth factor beta-induced in circulating tumor cell subline from pancreatic cancer cell line. Cancer Sci. 109(11), 3623–3633. https://doi.org/10.1111/cas.13783 (2018).
Morita, T., Mayanagi, T. & Sobue, K. Dual roles of myocardin-related transcription factors in epithelial-mesenchymal transition via slug induction and actin remodeling. J. Cell Biol. 179(5), 1027–1042. https://doi.org/10.1083/jcb.200708174 (2007).
Zhang, Y. et al. High throughput determination of TGFβ1/SMAD3 targets in A549 lung epithelial cells. PLoS ONE https://doi.org/10.1371/journal.pone.0020319 (2011).
Zhao, J. et al. The prognostic significance of LTBP2 for malignant tumors: Evidence based on 11 observational studies. J. Bone Joint Surg. 101(17), E29207. https://doi.org/10.1097/MD.0000000000029207 (2022).
Li, Z., Xia, J., Fang, M. & Xu, Y. Epigenetic regulation of lung cancer cell proliferation and migration by the chromatin remodeling protein BRG1. Oncogenesis https://doi.org/10.1038/s41389-019-0174-7 (2019).
Pai, S. G. et al. Wnt/beta-catenin pathway: Modulating anticancer immune response. J. Hematol. Oncol. https://doi.org/10.1186/s13045-017-0471-6 (2017).
Mann, B. et al. Target genes of beta-catenin-T cell-factor/lymphoid-enhancer-factor signaling in human colorectal carcinomas. Med. Sci. 96, 1603–1608. https://doi.org/10.1073/pnas.96.4.160 (1999).
Stewart, D. J. (2014). Wnt signaling pathway in non-small cell lung cancer. In Journal of the National Cancer Institute (Vol. 106, Issue 1). Oxford University Press. https://doi.org/10.1093/jnci/djt356
Zhang, L. et al. Simultaneous overactivation of Wnt/β-catenin and TGFβ signalling by miR-128–3p confers chemoresistance-associated metastasis in NSCLC. Nat. Commun. https://doi.org/10.1038/ncomms15870 (2017).
Mishra, L., Shetty, K., Tang, Y., Stuart, A. & Byers, S. W. The role of TGF-β and Wnt signaling in gastrointestinal stem cells and cancer. Oncogene 24(37), 5775–5789. https://doi.org/10.1038/sj.onc.1208924 (2005).
Incassati, A., Pinderhughes, A., Eelkema, R. & Cowin, P. Links between transforming growth factor-β and canonical Wnt signaling yield new insights into breast cancer susceptibility, suppression and tumor heterogeneity. Breast Cancer Res. 11(3), 104. https://doi.org/10.1186/bcr2253 (2009).
Minoo, P. & Li, C. Cross-talk between transforming growth factor-β and Wingless/Int pathways in lung development and disease. Int. J. Biochem. Cell Biol. 42(6), 809–812. https://doi.org/10.1016/j.biocel.2010.02.011 (2010).
Wee, P., Shi, H., Jiang, J., Wang, Y. & Wang, Z. EGF stimulates the activation of EGF receptors and the selective activation of major signaling pathways during mitosis. Cell. Signal. 27(3), 638–651. https://doi.org/10.1016/j.cellsig.2014.11.030 (2015).
Davis, F. M. et al. Non-stimulated, agonist-stimulated and store-operated ca2+ influx in MDA-MB-468 breast cancer cells and the effect of EGF-induced EMT on calcium entry. PLoS ONE 7(5), e36923. https://doi.org/10.1371/journal.pone.0036923 (2012).
Kim, J., Kong, J., Chang, H., Kim, H. & Kim, A. EGF induces epithelial-mesenchymal transition through phospho-Smad2/3-Snail signaling pathway in breast cancer cells. Oncotarget 7(51), 85021–85032. https://doi.org/10.18632/ONCOTARGET.13116 (2016).
Lo, H. W. et al. Epidermal growth factor receptor cooperates with signal transducer and activator of transcription 3 to induce epithelial-mesenchymal transition in cancer cells via up-regulation of TWIST gene expression. Cancer Res. 67(19), 9066–9076. https://doi.org/10.1158/0008-5472.CAN-07-0575 (2007).
Fabregat, I., & Caballero-Díaz, D. (2018). Transforming growth factor-β-induced cell plasticity in liver fibrosis and hepatocarcinogenesis. In Frontiers in Oncology (Vol. 8, Issue SEP). Frontiers Media S.A. https://doi.org/10.3389/fonc.2018.00357
Tian, Y.-C. et al. Epidermal growth factor and transforming growth factor-β1 enhance HK-2 cell migration through a synergistic increase of matrix metalloproteinase and sustained activation of ERK signaling pathway. Exp. Cell Res. 313(11), 2367–2377. https://doi.org/10.1016/j.yexcr.2007.03.022 (2007).
Xiao, Z., Sperl, B., Ullrich, A. & Knyazev, P. Metformin and salinomycin as the best combination for the eradication of NSCLC monolayer cells and their alveospheres (cancer stem cells) irrespective of EGFR, KRAS, EML4/ALK and LKB1 status. Oncotarget 5(24), 12877–12890. https://doi.org/10.18632/oncotarget.2657 (2014).
Wang, K. et al. FGFR1-ERK1/2-SOX2 axis promotes cell proliferation, epithelial–mesenchymal transition, and metastasis in FGFR1-amplified lung cancer. Oncogene 39(42), 6619–6620. https://doi.org/10.1038/S41388-020-01441-6 (2020).
Kouhmareh, K. et al. Capture of circulating metastatic cancer cell clusters from a lung cancer patient can reveal a unique genomic profile and potential anti-metastatic molecular targets: A proof of concept study. BioRxiv https://doi.org/10.1101/2023.09.19.558270 (2023).
Gammelgaard, K. R. et al. Up-Regulated FGFR1 Expression as a Mediator of Intrinsic TKI Resistance in EGFR-Mutated NSCLC. Transl. Oncol. 12(3), 432. https://doi.org/10.1016/J.TRANON.2018.11.017 (2019).
Yu, M. et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339(6119), 580–584. https://doi.org/10.1126/science.1228522 (2013).
Kong, H. J. et al. Crosstalk between YAP and TGFβ regulates SERPINE1 expression in mesenchymal lung cancer cells. Int. J. Oncol. 58(1), 111–121. https://doi.org/10.3892/ijo.2020.5153 (2020).
Serrano, M. J. et al. Significance of EGFR expression in circulating tumor cells. Adv. Exp. Med. Biol. 994, 285–296. https://doi.org/10.1007/978-3-319-55947-6_16/COVER (2017).
Payne, R. I. et al. Measurements of EGFR expression on circulating tumor cells are reproducible over time in metastatic breast cancer patients. Pharmacogenomics 10(1), 51–57. https://doi.org/10.2217/14622416.10.1.51 (2009).
Turetta, M. et al. Assessment of the mutational status of NSCLC using hypermetabolic circulating tumor cells. Cancers https://doi.org/10.3390/CANCERS10080270 (2018).
Kallergi, G. et al. Phosphorylated EGFR and PI3K/Akt signaling kinases are expressed in circulating tumor cells of breast cancer patients. Breast Cancer Res BCR https://doi.org/10.1186/BCR2149 (2008).
Scharpenseel, H. et al. EGFR and HER3 expression in circulating tumor cells and tumor tissue from non-small cell lung cancer patients. Sci. Rep. 9(1), 7406. https://doi.org/10.1038/s41598-019-43678-6 (2019).
Gupta, G. P. et al. Mediators of vascular remodelling co-opted for sequential steps in lung metastasis. Nature 446(7137), 765–770. https://doi.org/10.1038/nature05760 (2007).
Nishikawa, R. et al. A mutant epidermal growth factor receptor common in human glioma confers enhanced tumorigenicity (tumor progrsion/glioblastoma). Proc. Nati. Acad. Sci. USA 91(16), 7727–7731 (1994).
Grandis, J. R. et al. Levels of TGF-α and EGFR Protein in Head and Neck Squamous Cell Carcinoma and Neck Squamous Cell Carcinoma and Patient Survival. J. Natl. Cancer Inst. 90(11), 824–832 (1998).
Moscatello, D. K. et al. Frequent expression of a mutant epidermal growth factor receptor in multiple human tumors’. Cancer Res. 55(23), 5536–5539 (1995).
Cheng, L. et al. Molecular pathology of lung cancer: Key to personalized medicine. Mod. Pathol. 25(3), 347–369. https://doi.org/10.1038/modpathol.2011.215 (2012).
Al Moustafa, A. E., Achkhar, A. & Yasmeen, A. EGF-receptor signaling and epithelial-mesenchymal transition in human carcinomas. Front. Biosci. Scholar 4(2), 671–684. https://doi.org/10.2741/S292/PDF (2012).
Soulieres, D. et al. Multicenter phase II study of erlotinib, an oral epidermal growth factor receptor tyrosine kinase inhibitor, in patients with recurrent or metastatic squamous cell cancer of the head and neck. J. Clin. Oncol. 22(1), 77–85. https://doi.org/10.1200/JCO.2004.06.075 (2004).
Yamanaka, Y. et al. Coexpression of epidermal growth factor receptor and ligands in human pancreatic cancer is associated with enhanced tumor aggressiveness. Anticancer Res. 13(3), 565–569 (1993).
Kujtan, L. & Subramanian, J. Epidermal growth factor receptor tyrosine kinase inhibitors for the treatment of non-small cell lung cancer. Expert Rev. Anticancer Ther. 19(7), 547–559. https://doi.org/10.1080/14737140.2019.1596030 (2019).
Zubair, T. & Bandyopadhyay, D. Small molecule EGFR inhibitors as anti-cancer agents: Discovery, mechanisms of action, and opportunities. Int. J. Mol. Sci. 24, 2651. https://doi.org/10.3390/IJMS24032651 (2023).
Cheng, H. & Piperdi, B. Could erlotinib be used in EGF receptor wild-type non-small-cell lung cancer?. Lung Cancer Manag. 2(5), 333–336. https://doi.org/10.2217/LMT.13.39 (2013).
Jazieh, A. R. et al. Erlotinib in wild type epidermal growth factor receptor non-small cell lung cancer: A systematic review. Ann. Thoracic Med. 8(4), 204–208. https://doi.org/10.4103/1817-1737.118503 (2013).
Inno, A. et al. Erlotinib for patients with EGFR wild-type metastatic NSCLC: A retrospective biomarkers analysis. Pathol. Oncol. Res. 25(2), 513–520. https://doi.org/10.1007/s12253-018-0404-x (2019).
Itchins, M., Clarke, S. & Pavlakis, N. Do EGFR tyrosine kinase inhibitors (TKIs) still have a role in EGFR wild-type pre-treated advanced non-small cell lung cancer (NSCLC)?-The shifting paradigm of therapeutics. Transl. Lung Cancer Res. 7, S39–S45. https://doi.org/10.2103/TLCR.2018.01.06 (2018).
Piperdi, B. & Perez-Soler, R. Role of erlotinib in the treatment of non-small cell lung cancer: Clinical outcomes in wild-type epidermal growth factor receptor patients. Drugs 72(SUPPL. 1), 11–19. https://doi.org/10.2165/1163018-S0-000000000-00000 (2012).
Hao, J. et al. Paeoniflorin potentiates the inhibitory effects of erlotinib in pancreatic cancer cell lines by reducing ErbB3 phosphorylation. Sci. Rep. 6, 32809. https://doi.org/10.1038/srep32809 (2016).
Yamamoto, Y., Saita, T., Yamamoto, Y. & Shin, M. Quantitative determination of erlotinib in human serum using competitive enzyme-linked immunosorbent assay. J. Pharm. Anal. 8(2), 119–123. https://doi.org/10.1016/j.jpha.2017.10.002 (2018).
Lankheet, N. A. G. et al. Concentrations of erlotinib in tumor tissue and plasma in non-small-cell lung cancer patients after neoadjuvant therapy. Clin. Lung Cancer 16(4), 320–324. https://doi.org/10.1016/j.cllc.2014.12.012 (2015).
Hidalgo, M. et al. Phase I and pharmacologic study of OSI-774, an epidermal growth factor receptor tyrosine kinase inhibitor, in patients with advanced solid malignancies. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 19(13), 3267–3279. https://doi.org/10.1200/JCO.2001.19.13.3267 (2001).
Takeda, Y. et al. Phase I/II study of erlotinib to determine the optimal dose in patients with non-small cell lung cancer harboring only EGFR mutations. Clin. Transl. Sci. 13(6), 1150–1160. https://doi.org/10.1111/CTS.12796 (2020).
Petty, W. J. et al. Epidermal growth factor receptor tyrosine kinase inhibition represses cyclin D1 in aerodigestive tract cancers. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 10(22), 7547–7554. https://doi.org/10.1158/1078-0432.CCR-04-1169 (2004).
Tan, A. R. et al. Evaluation of biologic end points and pharmacokinetics in patients with metastatic breast cancer after treatment with erlotinib, an epidermal growth factor receptor tyrosine kinase inhibitor. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 22(15), 3080–3090. https://doi.org/10.1200/JCO.2004.08.189 (2004).
Frohna, P. et al. Evaluation of the absolute oral bioavailability and bioequivalence of erlotinib, an inhibitor of the epidermal growth factor receptor tyrosine kinase, in a randomized, crossover study in healthy subjects. J. Clin. Pharmacol. 46(3), 282–290. https://doi.org/10.1177/0091270005284193 (2006).
Gruber, A. et al. Monitoring of erlotinib in pancreatic cancer patients during long-time administration and comparison to a physiologically based pharmacokinetic model. Cancer Chemother. Pharmacol. 81(4), 763–771. https://doi.org/10.1007/S00280-018-3545-4/FIGURES/2 (2018).
Planchard, D. et al. Osimertinib Western and Asian clinical pharmacokinetics in patients and healthy volunteers: Implications for formulation, dose, and dosing frequency in pivotal clinical studies. Cancer Chemother. Pharmacol. 77(4), 767–776. https://doi.org/10.1007/S00280-016-2992-Z/METRICS (2016).
Cross, D. A. E., Ashton, S. E., Ghiorghiu, S. & Eberlein, C. AZD9291, an irreversible EGFR TKI, overcomes T790M-mediated resistance to EGFR inhibitors in lung cancer. Cancer Discov. 4(9), 1046–1061. https://doi.org/10.1158/2159-8290.CD-14-0337 (2014).
Wang, D. et al. A multifunctional nanotheranostic agent potentiates erlotinib to EGFR wild-type non-small cell lung cancer. Bioactive Mater. 13, 312–323. https://doi.org/10.1016/J.BIOACTMAT.2021.10.046 (2022).
Todsaporn, D. et al. Discovery of novel EGFR inhibitor targeting Wild-type and mutant forms of EGFR: In silico and in vitro. Molecules 28, 7. https://doi.org/10.3390/MOLECULES28073014/S1 (2023).
Blick, S. K. A. & Scott, L. J. Cetuximab: A review of its use in squamous cell carcinoma of the head and neck and metastatic colorectal cancer. Drugs 67(17), 2585–2607. https://doi.org/10.2165/00003495-200767170-00008 (2007).
Bos, P. D. et al. Genes that mediate breast cancer metastasis to the brain. Nature 459(7249), 1005–1009. https://doi.org/10.1038/NATURE08021 (2009).
Schinke, H. et al. A transcriptomic map of EGFR-induced epithelial-to-mesenchymal transition identifies prognostic and therapeutic targets for head and neck cancer. Mol. Cancer https://doi.org/10.1186/S12943-022-01646-1 (2022).
Tokumo, M. et al. The relationship between epidermal growth factor receptor mutations and clinicopathologic features in non-small cell lung cancers. Clin. Cancer Res. 11(3), 1167–1173 (2005).
Acknowledgements
This work was supported by the University of New Brunswick Research Fund (to AMN) and the Natural Sciences and Engineering Council of Canada (NSERC; to AMN). JH, RG and CSC were partly supported by the New Brunswick Innovation Foundation (NBIF).
Author information
Authors and Affiliations
Contributions
All authors contributed to (i) conception of the study and design of the experiments, (ii) data analyses, and (iii) manuscript writing and reviewing. J.P., C.S.C. and R.G. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hapeman, J.D., Galwa, R., Carneiro, C.S. et al. In vitro evidence for the potential of EGFR inhibitors to decrease the TGF-β1-induced dispersal of circulating tumour cell clusters mediated by EGFR overexpression. Sci Rep 14, 19980 (2024). https://doi.org/10.1038/s41598-024-70358-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-70358-x
- Springer Nature Limited