Abstract
Clustered DNA damage, when multiple lesions are generated in close proximity, has various biological consequences, including cell death, chromosome aberrations, and mutations. It is generally perceived as a hallmark of ionizing radiation. The enhanced mutagenic potential of lesions within a cluster has been suggested to result, at least in part, from the selection of the strand with the mutagenic lesion as the preferred template strand, and that this process is relevant to the tolerance of persistent single-strand breaks generated during an attempted repair. Using a plasmid-based assay in Escherichia coli, we examined how the strand bias is affected in mutant strains deficient in different DNA polymerase I activities. Our study revealed that the strand-displacement and 5′-flap endonuclease activities are required for this process, while 3′-to-5′ exonuclease activity is not. We also found the strand template that the mutagenic lesion was located on, whether lagging or leading, had no effect on this strand bias. Our results imply that an unknown pathway operates to repair/tolerate the single-strand break generated at a bi-stranded clustered damage site, and that there exist different backup pathways, depending on which DNA polymerase I activity is compromised.
Similar content being viewed by others
Introduction
Under normal aerobic growth conditions, DNA is continuously attacked by reactive oxygen species, which can cause various types of DNA damage. Most of these DNA lesions are chemically indistinguishable from those caused by ionizing radiation1. However, the clustering of DNA damage is a distinctive feature of the effect of ionizing radiation on DNA2,3. Clustered DNA damage is defined as two or more DNA lesions within one to two helical DNA turns, and comprises distinct types of lesions, such as base damage, apurinic/apyrimidinic (AP) sites, and strand breaks. DNA damage clustering is thought to be highly relevant to the biological consequences of ionizing radiation2,3. Experiments have demonstrated that, after irradiation with low-density ionization, the yield of clustered DNA damage is at least four times greater than that of double-strand breaks (DSBs)4,5,6, although the overall yield decreases with increasing ionization density7. Based on extensive computer simulation studies, the complexity of clustered DNA damage, in terms of the number and types of damage, is predicted to increase with increasing ionizing radiation density8,9,10,11. Recently, atomic force microscopy was used to experimentally confirm that the complexity of clustered DNA damage depends on the ionization density of the radiation12,13.
Most data regarding the processing and biological consequences of clustered DNA damage have been obtained by studying synthetic DNA lesions located close to each other. Extensive in vitro studies have revealed that the repair of lesions within a cluster is often retarded, depending on the type, position, and number of lesions within a cluster (for review, see14,15,16). Detailed in vivo studies of the biological consequences of clustered DNA damage have shown that clustered DNA damage is often detrimental to cells (for review see16,17,18). The processing of bi-stranded clustered DNA damage, consisting of AP sites and single-strand breaks (SSB), likely leads to a DSB; therefore, the replication of DNA containing such clusters is strongly retarded. In contrast, DNA containing clusters of base damage generally replicates efficiently, accompanied by the enhanced mutagenic potential of the base damage.
Using a plasmid-based assay, we and others found that the increase in mutation frequency (MF) at the mutagenic base lesion in a bi-stranded clustered damage site likely results from the persistence of an SSB, which is generated as a repair intermediate of a lesion within a cluster. Taking evidence from in vitro experiments and previous in vivo studies in combination suggests that an SSB impairs the repair of the resulting mutagenic base lesion, such as 8-oxo-7,8-dihydroguanine (8-oxoG), and that the base lesion persists until replication19,20,21,22,23,24,25,26. Interestingly, the retardation of mutagenic lesion repair does not seem to be the only cause of the enhanced MF at the clustered DNA damage site. For instance, even in the absence of the main DNA glycosylase for 8-oxoG excision in Escherichia coli, Fpg, the MF of a bi-stranded cluster containing an 8-oxoG was markedly greater than that of an isolated 8-oxoG20,21,24,26. This is inconsistent with the expectation that the MF of the 8-oxoG should be similar in the absence of Fpg, whether within a cluster or not. Additionally, the MF of 5-hydroxyuracil (hU) within a bi-stranded clustered site was greater than 80%, significantly greater than the expected 50% limit27. This high MF was not observed if the opposite non-mutagenic strand replicated to a similar extent as the hU strand. Based on these results, it was postulated that the strand harboring the mutagenic lesion is preferentially replicated, leading to the concomitant loss of the opposite strand. To further gain insights into the template-strand preference, we developed an assay in which a mismatch serves as a strand-specific marker to determine the fraction of each strand used as the template around a bi-stranded clustered damage site28,29. The assay results indicated that (1) the 8-oxoG strand is the preferred template around the clustered DNA damage site to tolerate the persisting SSB on the opposite strand, (2) the size of the template-strand preference is in the order of several tens of base pairs, and (3) the size is determined by DNA polymerase I. If unrepaired, a nick or an SSB will be converted to a DSB by the arrival of a replication fork30, leading to chromosome fragmentation31,32. Although replication-dependent DSBs are repaired by a process involving RecA33,34, the central enzyme for strand transfer that mediates homologous recombination (HR), it is currently unknown whether other independent pathways are operative. We found that the template-strand preference near a bi-stranded clustered damage site was not affected either by RecA or by UvrA or UvrC, which function in recognizing or excising, respectively, DNA with bulky damage during nucleotide excision repair (NER)29.
DNA polymerase I (Pol I) participates in the maturation of the Okazaki fragment on the lagging strand during replication. During this process, the polymerase displaces the RNA primer, polymerizes DNA, and removes the displaced 5′-flap structure to enable ligation35. Additionally, Pol I functions in the DNA synthesis step of various DNA repair pathways, such as base excision repair (BER), NER, and ribonucleotide excision repair36,37. In BER, Pol I is involved in both short- and long-patch repair; it incorporates a mononucleotide at the damaged site38 and inserts multiple nucleotides 3′ downstream of the damaged site38,39. To fulfill these roles in vivo, Pol I is multifunctional and is known to possess (1) DNA polymerase activity, (2) strand-displacement activity, (3) 5′-flap endonuclease activity, and (4) 3′-to-5′ proofreading exonuclease activity. Each activity is located on a separate domain of the protein. The polymerase activity, strand-displacement activity, and 3′-to-5′ exonuclease activity are on the large (known as the Klenow) fragment, while the 5′-flap endonuclease activity is on the N-terminal side40,41. The essential residues related to these activities are known42,43,44,45,46,47, and Pol I-deficient strains have been constructed with loss of strand-displacement activity (S769A and F771A double mutants)37, 5′-flap endonuclease deficiency (Y77C)48,49, and loss of 3′-to-5′ exonuclease activity (D424A)50. To better understand the underlying mechanism(s) of the template-strand bias around a clustered DNA damage site, we aimed to explore which Pol I activity is responsible for this process. Here, using polA strains each lacking one of the Pol I activities, we present evidence that strand-displacement activity and 5′-flap endonuclease activity, which are required for the completion of long-patch repair in BER, affect the template-strand preference. Moreover, we found that the lack of 5′-flap endonuclease activity gives rise to a specific mode of strand bias around a bi-stranded clustered damage site.
Results
Experimental outline
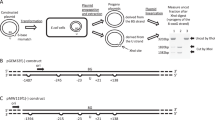
The plasmids shown in Fig. 1A were constructed to determine which function of DNA polymerase I is responsible for determining the template-strand preference around a clustered DNA damage site. The plasmids were constructed as previously described28,29 and contained a bi-stranded clustered damage site composed of a uracil and an 8-oxo-7,8-dihydroguanine (uracil + 8-oxoG), as well as a six-nucleotide XhoI recognition sequence inserted as a strand-specific marker. A synthetic 8-oxoG lesion was inserted into the strand complementary to the uracil (Fig. 1A(a)), and plasmids with the uracil and 8-oxoG in exchanged positions were also constructed (Fig. 1A(b)). After introduction of the plasmids into E. coli, the template-strand usage was measured based on the fraction of XhoI-uncut plasmid progeny (Fig. 1B). The strand-specific XhoI sequence was on the leading strand template, and so the XhoI-uncut fraction represents the fraction where the lagging strand was used as the template. In E. coli, a six-nucleotide mismatch is refractory to mismatch repair29,51, and thus strand synthesis past the mismatch site either eliminates or retains the XhoI recognition sequence, depending on which strand is used as a template. Notably, a uracil in a clustered DNA damage site is rapidly converted into an AP site and subsequently processed to generate an SSB in E. coli cells16,52. Hereafter, the Pol I-deleted strain (CC104mutY∆polA), the 3′-to-5′ exonuclease-deficient strain (NS2001), the strand displacement-deficient strain (NS2002), and the 5′-flap endonuclease-deficient strain (NS2003) (Table 1) are referred to as ∆polA, polAexo, polA_SD−, and polA107, respectively.
Schematic representations of the plasmids used in the study and the experiment to determine the level of strand bias near a clustered DNA damage site. (A) Constructed plasmids. (a) A bi-stranded clustered damage site comprised of uracil and 8-oxo-7,8-dihydroguanine (8-oxoG) was created by placing 8-oxoG on the lagging strand template (blue line) and uracil on the leading strand template (red line). The plasmid pMW119f1(−)NAEPBNG was used for construction. The black triangle represents the mismatch, which was generated by inserting a 6-nucleotide recognition sequence of XhoI and was used as a strand-specific marker. The numbers show the separation (in nucleotides) from 8-oxoG within the bi-stranded cluster, with + and − indicating the 5′ and 3′ directions on the leading strand, respectively. (b) A bi-stranded clustered damage site (uracil + 8-oxoG) was created by placing 8-oxoG on the leading strand template (red line) and uracil on the lagging strand template (blue line). pMW119f1(+)NAEPBNG, which synthesizes the complementary single strand DNA of that synthesized with pMW119f1(−)NAEPBNG, was used for construction. Uracil is placed at the position of 8-oxoG in (a), and the mismatches are placed as in (a). The black arrow indicates the direction of the movement of the replication fork. (B) Outline of the experiment. The constructed plasmids were introduced into E. coli, and the propagated plasmids were extracted, followed by linearization with ScaI. The progeny plasmids propagated from the strand with the mismatch were cut via XhoI treatment. The level of strand preference was measured by determining the fraction of the intensity of the XhoI-uncut plasmids to the intensity of all fragments after gel electrophoresis (see the text for further details).
Template-strand preference near a bi-stranded clustered damage site
When the 8-oxoG within the cluster (uracil + 8-oxoG) was on the lagging strand template in the wild-type (WT) Pol I strain, the fraction using the lagging strand template (8-oxoG strand) was greater at both +21 and −23 (greatest at −23) than the respective fraction observed with undamaged template (Fig. 2A,B), suggesting that a strand bias is generated near a bi-stranded clustered damage site by repair and/or replication29. The use of the 8-oxoG strand as the template in uracil + 8-oxoG was greatly enhanced in the ∆polA strain, as shown in our previous study29. In polAexo, the level of strand preference was comparable to that of WT. The extent of the increase at both +21 and −23 with uracil + 8-oxoG in the polA_SD− strain was markedly enhanced, and the level was only slightly less than that in the ∆polA strain. Interestingly, in polA107, the strand bias was completely different from WT. The extent of using the 8-oxoG strand as a template in uracil + 8-oxoG was markedly reduced compared with the undamaged construct at +21, but it was comparable to the undamaged construct at −23.
The fraction of the lagging strand template on which 8-oxo-7,8-dihydroguanine (8-oxoG) was placed when plasmids with bi-stranded cluster comprised of uracil and 8-oxoG (uracil + 8-oxoG) were propagated in Pol I-deficient strains. The relative positions of each damage and mismatch are schematically shown at the top of the figure. (A) The fraction of the lagging strand template (8-oxoG strand) at position +21. (B) The fraction of the lagging strand template (8-oxoG strand) at position −23. Black bars: no damage, Green bars: uracil + 8-oxoG bi-stranded cluster. polA genotypes of the strains are shown along the horizontal axis. ∆polA: strain lacking total Pol I activity (CC104mutY∆polA), polAexo: strain lacking 3′ to 5′ exonuclease activity (NS2001), polA_SD−: strain lacking strand displacement activity (NS2002), polA107: strain lacking 5′-flap endonuclease activity (NS2003). Each data represents the mean ± standard error of at least 3 independent experiments. Statistically significant differences between the fractions with and without a bi-stranded clustered damage site are indicated (**p < 0.01).
The strand preference of single lesions likely reflects the outcome of BER. Distinct levels of the use of lagging strand template were observed between single lesions (Fig. S1) and uracil + 8-oxoG (Fig. 2) at both +21 and −23 in the WT, implying that most, if not all, of the bi-stranded clustered damage site is specifically processed by an as yet unknown pathway29. The strand bias of single lesions in polAexo was quite similar to WT, while in ∆polA, polA_SD−, and polA107, different modes of strand bias were observed compared with WT. Furthermore, the strand bias around a single uracil was slightly different from that around a single 8-oxoG in each strain. We observed that the levels of strand preference around a single uracil in both polA_SD− and polA107 were less than ∆polA (Fig. 1S). This suggests that some of the single uracil bases are removed by short-patch repair, as only ∆polA lacks the ability to complete short-patch repair. Less frequent usage of the lagging strand template was observed with a single 8-oxoG at + 21 in ∆polA and polA107 than in WT, whereas the usage was comparable between polA_SD− and WT (Fig. 1S). These results suggest that the impairment of 5′-flap endonuclease activity strongly compromises short-patch repair of 8-oxoG, and that a backup pathway for 5′-flap removal is present.
Mutation frequencies of bi-stranded clustered damage sites in polA mutants
As the extent of template-strand preference around a bi-stranded clustered damage site depends on the activity of Pol I (Fig. 2), we next assessed the MF of uracil + 8-oxoG in polA mutants. As a mutation in the recognition sequence of a restriction enzyme leads to the inability to cut the sequence by the enzyme, the MF was estimated by the fraction of uncut Alw26I recognition sequence within which uracil and 8-oxoG were placed (Table S1)28,29. The 8-oxoG was placed on the lagging strand template. The MFs of the ∆polA and polA_SD− strains were significantly greater than WT (p < 0.01), while the MF of the polAexo strain was comparable to WT (Fig. 3). In polA107, the MF of uracil + 8-oxoG was quite low and comparable to that of the undamaged construct, supporting the finding from Fig. 2 that the processing mode of uracil + 8-oxoG in polA107 is quite different from that in WT or other polA mutants. We found that the level of MF in each strain was, to some extent, correlated with the level of the 8-oxoG strand fraction at + 21 (Fig. 2A).
The MF of a bi-stranded clustered DNA damage site in strains lacking Pol I activity. 8-oxo-7,8-dihydroguanine (8-oxoG) was placed on the lagging strand template. Black bars: no damage, green bars: uracil + 8-oxoG bi-stranded cluster. polA genotypes of the strains are shown along the horizontal axis. ∆polA: strain lacking total Pol I activity (CC104mutY∆polA), polAexo: strain lacking 3′ to 5′ exonuclease activity (NS2001), polA_SD−: strain lacking strand displacement activity (NS2002), polA107: strain lacking 5′-flap endonuclease activity (NS2003). Each data represents the mean ± standard error from at least 3 independent experiments. Statistically significant differences between the fractions with and without a bi-stranded clustered damage site are indicated (**p < 0.01).
Strand annealing by RecT or RecO is not involved in the process leading to strand preference around a bi-stranded clustered damage site
We found previously29 that the key protein in HR (RecA) and the proteins required for NER (UvrA and UvrC) do not take part in determining the template-strand preference around a clustered DNA damage site. Here, we examined mutants that are deficient in strand-annealing activity to determine whether this activity plays any role in strand preference. RecT and RecO are known to possess strand-annealing activity53,54 and to function in the RecE and RecF recombination pathways, respectively55. Although our experimental strain derived from CC10456 does not harbor a sbcA mutation activating the RecE recombination pathway57, we aimed to determine whether RecE recombination plays a role in the biased template-strand usage by any spontaneous activation of sbcA in our strains. Our results with the recET-disrupted strain revealed that the strand-annealing activity of RecT is unlikely to play a significant role in determining strand preference, as the template-strand usage fractions around the uracil + 8-oxoG cluster (+21 and −23) in each mutant did not significantly differ from WT (Fig. 4). Similarly, RecO was not strongly involved in determining strand bias around the uracil + 8-oxoG cluster (Fig. 4).
The fraction of the lagging strand template on which 8-oxo-7,8-dihydroguanine (8-oxoG) was placed when plasmids were propagated in strains lacking RecET or RecO. (A) The fraction of the lagging strand template (8-oxoG strand) at +21. (B) The fraction of the lagging strand template (8-oxoG strand) at − 23. Black bars: no damage, Green bars: uracil + 8-oxoG bi-stranded cluster. Strains deficient in strand annealing activity are shown along the horizontal axis. recET: strain that lacks RecET activity (NS2004). recO: strain that lacks RecO activity (NS2005). Each data represents the mean ± standard error from at least 3 independent experiments. Statistically significant differences between the fractions with and without a bi-stranded clustered damage site are indicated (**p < 0.01).
Template-strand preference around clusters with reversed lesion positions relative to the direction of replication fork movement
Strand bias may emerge at or after replication, and thus, the different modes of replication for the leading and lagging strands may affect the outcome. Therefore, we next determined the strand preference when the location of the lesions was reversed relative to the direction of replication fork movement. In this reversed setup, the 8-oxoG within the cluster was on the leading strand template (Fig. 1A(b)). In the WT, the fraction of the lagging strand template used, representing the level of use of the uracil strand, was lower at either position than the respective fractions without any damage (+21 or −23, although much less at +21 with uracil + 8-oxoG) (Fig. 5). The fractions in the Pol I mutants, except for polA107, also showed less usage of the uracil strand. Comparing these results with the results in Fig. 2 indicates that the 8-oxoG strand is preferentially used as the template in both configurations of the lesions, and there is little strand specificity. Even in polA107, where the usage of the uracil strand at position −23 was increased in the presence of bi-stranded clustered damage (Fig. 5, p < 0.01), strand specificity was not observed. These results indicate that the template-strand preference around a bi-stranded clustered damage site is not strongly affected by the direction of replication fork movement.
The fraction of the lagging strand template, on which uracil was placed, when plasmids with uracil + 8-oxoG bi-stranded cluster were propagated in Pol I-deficient strains. (A) The fraction of the lagging strand template (uracil strand) at +21. (B) The fraction of the lagging strand template (uracil strand) at −23. Black bars: no damage, Green bars: uracil + 8-oxoG bi-stranded cluster. polA genotypes of the strains are shown along the horizontal axis. ∆polA: strain lacking total Pol I activity (CC104mutY∆polA), polAexo: strain lacking 3′ to 5′ exonuclease activity (NS2001), polA_SD−: strain lacking strand displacement activity (NS2002), polA107: strain lacking 5′-flap endonuclease activity (NS2003). Each data represents the mean ± standard error of at least 3 independent experiments. Statistically significant differences between the fractions with and without a bi-stranded clustered damage site are indicated (**p < 0.01).
With single lesions, a strand-specific mode of template preference was observed, to some extent. The strand preference was unaffected at the 5′ side of a single 8-oxoG (−23) when it was located on the lagging strand in each strain, except for ∆polA (Fig. S1), whereas when the 8-oxoG was on the leading strand, the 5′ side of the 8-oxoG (+21) gave rise to a slight strand bias in every strain (Fig. S2). This difference suggests that the repair of a single 8-oxoG is, to some extent, strand specific.
Template-strand usage in wild-type or polA107 clones
To better understand how this strand bias was caused, we next assessed the distribution of the template-strand usage level for each transformed clone (Table 2). Notably, the average template-strand usage in each clone was comparable to the overall template-strand usage analyzed in Figs. 2 and 5. In the WT, when the uracil + 8-oxoG contained the 8-oxoG on the lagging strand template, the distribution markedly changed from that without damage (Table 2, p < 0.01 vs. no damage at both the +21 and −23 sites): most clones used only one of the strands at +21 (79.6%) or −23 (87.8%). Similarly, when the 8-oxoG of the uracil + 8-oxoG was on the leading strand, most clones at +21 (69.4%) or −23 (78.4%) used only one of the strands as a template (Table 2, p < 0.01 vs. no damage at both the +21 and −23 sites). These results were expected, because the 8-oxoG strand is the preferred template in the WT (Figs. 2 and 5); thus, the majority of clones used only one strand as a template near a bi-stranded clustered damage site.
The characteristic strand bias in polA107 (Figs. 2 and 5) prompted us to investigate the distribution of template usage in polA107. One of the template strands was predominantly used at +21 (77.1%) or −23 (60.0%) when the 8-oxoG of the uracil + 8-oxoG was on the lagging or leading strand, respectively. The respective fractions were much lower in plasmids without damage (44.9% and 37.5%) (Table 2, p < 0.01). However, the presence of uracil + 8-oxoG did not alter the mode of distribution of the template-strand usage (Table 2) at −23 or +21 when the 8-oxoG was on the lagging or leading strand, respectively.
Discussion
In a previous study29, we showed that Pol I is required to define the level and size of the preferential use of a template strand harboring a mutagenic lesion in a cluster. Herein, we have gained further insights into the preferential use of a mutagenic template by studying how the Pol I activity, and whether the relative direction of the movement of the replication fork, affect the template-strand usage.
The preference for the 8-oxoG strand and the significance of long-patch repair in the Pol I-proficient strain
In the WT, the preferred usage of the 8-oxoG strand within the uracil + 8-oxoG is in line with the observed enhanced mutagenic potential of the mutagenic lesion in a bi-stranded clustered damage site (Fig. 3)16,17,18. We found that the 8-oxoG strand was used preferentially, irrespective of which strand (the leading or lagging strand template) the 8-oxoG was located on within the cluster (Figs. 2 and 5). These results are consistent with the interpretation that the uracil in the uracil + 8-oxoG is excised first, retarding the subsequent repair of the 8-oxoG, and that the template-strand preference in the WT is not affected by which strand is used as the template for replication, but by the relative location of the lesions (Figs. 2 and 5). The usage of the 8-oxoG strand independent of the direction of the replication fork is in agreement with our earlier findings that reversing the direction of (and thus inverting the damage-containing strand relative to) the movement of the replication fork did not affect the level of MF of a bi-stranded clustered damage site21,58. Further, the lack of strand specificity suggests that any RecA-independent recombination or template switching mechanism associated with crossovers, which are proposed to proceed via annealing of nascent strands across a blocked replication fork, are unlikely to be involved, as these events requires discontinuity on the lagging strand, and thus, exhibits lagging strand specificity59. In addition, no strand bias during the preferential use of the 8-oxoG strand suggests that the process is unlikely to be related to the process with increased fidelity of lagging strand replication, which is proposed to result from the increased dissociability of lagging strand DNA polymerase60.
During BER long-patch repair, Pol I synthesizes DNA by incorporating multiple nucleotides while displacing the downstream strand, and subsequently excises the 5′-flap generated38,39. The last step, SSB ligation, is most likely inhibited in the WT, because a study using mammalian cell extracts demonstrated that, with a bi-stranded cluster, long-patch repair is impeded at ligation, not at the strand-displacement or 5′-flap removal steps61. The observation that cells display a greater fraction of the 8-oxoG strand at the 3′ end of the opposing lesion (uracil) irrespective of the direction of the fork (Figs. 2 and 5) supports the notion that incomplete long-patch repair initiates an SSB tolerance pathway and leads to template-strand preference in the WT. Since the tract size for long-patch repair in E. coli is in the range of 9 to 27 nucleotides62, we consider it likely that Pol I displaces the majority of the strands 3′ of the uracil. In contrast to the significance of the strand-displacement and 5′-flap endonuclease activities, the 3′-to-5′ proofreading exonuclease activity of Pol I had a minor, if any, effect on strand usage (Figs. 2 and 5). This suggests that the 3′-to-5′ exonuclease activity does not affect the 5′ side of the uracil, and that this activity is not essential for the strand bias surrounding a bi-stranded clustered DNA damage site.
Template-strand preference in strains deficient in different Pol I activities
The preferential use of the 8-oxoG strand as the template in the presence of uracil + 8-oxoG was further promoted when the strand-displacement activity was impaired (Figs. 2 and 5). This enhanced use of the 8-oxoG strand is consistent with the elevated MF in the polA_SD− strain (Fig. 3). The deficiency of this activity resulted in only a slight decrease in the level of strand bias compared with that seen with total loss of Pol I activity (∆polA)29, implying that a bi-stranded clustered damage site is not mainly processed by short-patch repair and that a similar, if not the same, unknown backup pathway operates in both polA_SD− and ∆polA. With single damage sites (Figs. S1 and S2), we found that the strand bias was less pronounced in polA_SD− than in ∆polA. We believe that some of the single lesions are repaired by short-patch repair in the polA_SD− strain.
Interestingly, the lack of 5′-flap endonuclease activity in polA107 led to a completely different mode of strand usage around a bi-stranded clustered damage site from that seen in the WT, or in the ∆polA and polA_SD− strains (Figs. 2 and 5). In polA107, the strand bias at the 3′ side of the U (−23 and +21 when 8-oxoG is on the lagging and leading strand templates, respectively) in a bi-stranded clustered damage site was quite similar, if not the same, to the strand bias observed for the undamaged construct at the respective sites (Figs. 2 and 5, Table 2). These results indicate that the uracil strand was most likely not displaced, which is consistent with an in vitro study demonstrating that Pol I deficient in 5′-flap endonuclease activity does not displace the downstream strand and inhibits ligation63. Interestingly, a certain fraction of the 3′ strand of a single uracil (not within a cluster) was displaced in polA107 (Figs. S1 and S2), indicating that the uracil processing pathway depends on the presence of a nearby lesion. In contrast to the strand bias at the 3′ side of the U, the majority of the 5′ side of the uracil (which is the 3′ side of the opposing 8-oxoG) in a bi-stranded clustered damage site used the uracil strand as a template, irrespective of on which strand (leading or lagging strand template) the 8-oxoG was placed. Together with the markedly reduced MF in polA107, we believe that 8-oxoG is excised at a bi-stranded clustered damage site and that repair synthesis occurs downstream (3′) of 8-oxoG in polA107. This strand bias at the 3′ side of 8-oxoG was also observed with a single 8-oxoG in polA107.
These findings in polA_SD− and polA107 suggest that an SSB in a bi-stranded clustered damage site is tolerated even in the absence of fully functional Pol I, and that different pathways operate to tolerate the SSB, based on the residual activities of Pol I. Notably, we observed no influence of the direction of replication fork movement on these SSB toleration mechanisms, indicating that they are not strand specific.
The process leading to strand preference
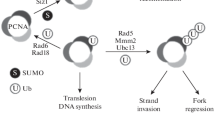
The findings—(1) that a lesion opposed to an 8-oxoG within a bi-stranded cluster is converted to an SSB whose repair is strongly retarded16,17,18, and (2) that the strand preference around a bi-stranded clustered damage site is affected very differently depending on which activity of Pol I is disrupted (Figs. 2 and 5, Table 1)—led us to suggest that the template-strand preference is associated with SSB tolerance pathways in which Pol I plays a crucial role. These potential SSB tolerance pathways are all unaffected by the relative position of the lesions with regard to the direction of the movement of the replication fork (Figs. 2 and 5), and are unrelated to the activity of RecA29 and the strand-annealing activities of RecT and RecO (Fig. 4). We, therefore, suggest that the mechanisms likely do not involve recombination between sister chromatids, but operate before replication. Our study proposes a model in which at least three different pathways give rise to strand bias (Fig. 6). The uracil in the uracil + 8-oxoG is likely processed first, by uracil DNA glycosylase generating an AP site. The major enzyme that cleaves AP site from uracil is most likely exonuclease III in E. coli, as mutants in dUTPase (dut) that incorporate large amounts of uracil into DNA are inviable in the presence of additional mutations in exonuclease III64. With its 3′ phosphatase activity, exonuclease III cleavage of the 5′ side of the AP site leaves 3′OH and 5′-deoxyribose-phosphate (dRP) at the break65. In the Pol I-proficient strain (Fig. 6, Pathway A), SSB is tolerated by a mechanism that leads to the formation of a gap with a size of several tens of base pairs, which is subsequently filled in most probably by Pol I using the mutagenic strand as a template29. The potential 3′ to 5′ exonuclease(s) involved in generating the gap 5′ upstream of the SSB is currently unknown. The pathway in the Pol I-proficient strain (pathway A) may resemble the long-patch repair subpathway identified by Woodrick et al.66, in which gap filling occurs not only downstream (3′) of the SSB but also upstream (5′). In this context, gap filling gives rise to strand bias and thus elevates the MF, as the 8-oxoG:adenine pair is likely to be generated prior to replication. In strains that are deficient in strand displacement (∆polA and polA_SD−) (Fig. 6, Pathway B), the 5′-flap is not generated, and the SSB likely persists, as the short patch repair of an SSB within a cluster is strongly compromised15,16,17. The persisting SSB is likely tolerated by an unknown pathway, which forms a gap by both 5′ and 3′ digestion from the SSB. As this digestion on both sides of the break generates a gap with a relatively large size of several hundred base pairs29, we assume that the 5′ and 3′ nucleases involved in pathway B are different from the nucleases in pathway A. One potential candidate for the 5′ nuclease is RecJ, as RecJ possesses a dRPase activity which cleaves the 5′-dRP at the break67. The difference in the MF between the Pol I-proficient and strand displacement-deficient strains (Fig. 3) can be explained by the involvement of distinct DNA polymerases with different frequencies of adenine insertion opposite the 8-oxoG. Interestingly, the strand bias in cells deficient in 5′-flap endonuclease activity indicates that the 5′-flap generated 3′ of the SSB derived from the uracil remains unexcised (Figs. 2 and 5, Table 2). We assume that the 5′-flap is eventually reannealed to the original template (Fig. 6, Pathway C). The strand that is synthesized downstream of the initial SSB is unwound and removed by 3′ nuclease(s), followed by gap filling and ligation. The use of the 5′ side of the SSB as a template (Figs. 2 and 5, Table 2), together with the low MF observed in polA107 (Fig. 3), suggests that the 8-oxoG is subsequently excised and that, at least, some of the 3′ side of the 8-oxoG is displaced with repair synthesis. The precise mechanisms underlying the excision or digestion of the single-stranded DNA in the 5′-flap endonuclease-deficient polA107 strain remain unknown. Taken together, our study suggests that (1) different SSB tolerance/repair mechanisms are provoked, depending on how the SSB is processed by WT or mutated Pol I, and (2) the different tolerance/repair mechanisms lead to a specific template-strand bias and determine the mutagenic potential of the bi-stranded clustered damage site.
Possible mechanisms for the occurrence of strand bias around a bi-stranded cluster containing uracil and 8-oxo-7,8-dihydroguanine (8-oxoG). The uracil in the uracil + 8-oxoG cluster is excised first and generates an SSB. Pathway (A): When wild-type Pol I is present, a gap is generated from 5′-flap cleavage after strand displacement and from the digestion of 3′ single-stranded (ss) DNA. Filling in the gap leads to a relatively short patch size of several tens of nucleotides. Pathway (B): When strand displacement activity (or total Pol I activity) is absent, the persistent SSB generated at a bi-stranded clustered damaged site is processed by an unknown mechanism that requires a long patch size of several hundred nucleotides. Since the preferential use and the mutagenic potential of the 8-oxoG strand is more enhanced in the strand displacement-deficient strain, the polymerase(s) involved in pathways A and B could be different. Pathway (C): In the absence of 5′-flap endonuclease activity, the 5′-flap is not excised and reanneals to the original template. The resulting 3′ ssDNA is digested to generate a strand break. Before reannealing the 5′-flap, 8-oxoG could be excised and processed similarly to a single 8-oxoG. Subsequently, the strand breaks generated after reannealing of the 5′-flap and generated downstream of 8-oxoG are both sealed.
The findings of the present study further deepen our understanding of the essential role played by Pol I activity in determining the extent of the preferential use of the mutagenic strand as a template, and that different backup pathways likely rescue the persistent SSB depending on which activity of Pol I (strand-displacement activity or 5′-flap endonuclease activity) is compromised. Pol I has long been known to play a significant role in BER, but despite extensive studies on the function of Pol I, its importance in preventing detrimental consequences after irradiation remains unclear. As SSBs are regularly generated in the genome by direct scission or after the excision of diverse types of damage, further studies are needed to fully reveal the role of Pol I in the tolerance/repair of SSBs in cells.
Materials and methods
Strains
The strains used in the study are listed in Table 2. The CC104mutY::tet strain was provided by Prof. Q.M. Akiyama (Kyoto University). The MG1655 polA_SD− strain harboring polA F769A/F771A::cat allele was a kind gift from Prof. R. Woodgate (National Institutes of Health), and the KA796 polAD424A::cat strain was obtained from Prof. I. J. Fijalkowska (Institute of Biochemistry and Biophysics). Strain KMBL1789 was obtained from the Coli Genetic Stock Center (CGSC). Strains with the CC104mutY::tet background that lacked one of the activities of Pol I were constructed through P1 transduction, which uses P1 phage for moving portions of the E. coli genome from one genetic variant to another, or by disrupting the gene via the method developed by Datzenko and Wanner68, which utilizes λ Red recombinase to integrate PCR amplified fragments into the E. coli genome (Table 2). For NS2003 construction, firstly, a DNA fragment containing the chloramphenicol acetyltransferase (cat) gene was PCR amplified from KA796 polAD424A::cat using the primers 5′-ATCAGGCGCACTAAGATTCG-3′ and 5′-GACTGGGGCGGCTAAAATA-3′. Then, the amplified cat gene was inserted at the 3′ side of the polA107 allele in the strain KMBL1789 using the method of Datzenko and Wanner to construct NS0001. The polA107::cat fragment in NS0001 was further transferred by P1 transduction to CC104mutY::tet to produce NS2003. The plasmids used for gene disruption, pKD46, pKD13, and pCP20, were from CGSC. The strains used to analyze strand bias were MutY deficient. As MutY is responsible for removing adenine residues misincorporated in 8-oxoG:adenine mispairs, and thus, avoiding G:C to T:A mutations, this deficiency enabled monitoring of the reparability of 8-oxoG due to its mutagenic potential21,24. Bacterial cultures were grown at 37 °C in Luria–Bertani (LB) medium supplemented with appropriate antibiotics.
Oligonucleotides and plasmids
The oligonucleotides, whose sequences are shown in Table S1, were purchased from Tsukuba Oligo Service (Tsukuba, Japan). U-1EP and 8G-2EP are identical to NAEP except for the presence of uracil in U-1EP and 8-oxoG in 8G-2EP. Both U-1EPX + 21 and 8G-2EPX + 21 contain the 6-nucleotide XhoI recognition sequence (5′-CTCGAG-3′) and are identical to NAEPX + 21 except for the presence of uracil in U-1EPX + 21 and 8-oxoG in 8G-2EPX + 21. Similarly, U-1EPX-23 and 8G-2EPX-23 both contain the XhoI sequence and are identical to NAEPX-23 except for the presence of uracil in U-1EPX-23 and 8-oxoG in 8G-2EPX-23. Ura and 8OXOG are identical to BNG except for uracil and 8-oxoG in Ura and 8OXOG, respectively. Uracil and 8-oxoG were located within the recognition sequence of Alw26I. NAEP and BNG are complementary sequences, and pMW119f1(−)NAEPBNG and pMW119f1(+)NAEPBNG were constructed by replacing the HindIII–NdeI fragment of pMW119 with the HindIII–NdeI fragment of pGEM3Zf(−)NAEPBNG and pGEM3Zf(+)NAEPBNG, respectively, as described previously29. pMW119f1(+)NAEPBNG is identical to pMW119f1(−)NAEPBNG except for the direction of the origin of the f1 phage. Thus, pMW119f1(+)NAEPBNG generates single-stranded DNA (ssDNA) complementary to that from pMW119f1(−)NAEPBNG upon the addition of the helper phage. The number (e.g., +21 or −23) in the oligonucleotide indicates the position of the mismatch in the plasmids pMW119f1(−)NAEPBNG (Fig. 1A(a)) and pMW119f1(+)NAEPBNG (Fig. 1A(b)).
Construction of plasmids with DNA damage and mismatch
Plasmids with DNA damage and mismatch were constructed according to methods described previously29. In brief, VCM13 helper phages were added to cultures of CJ236 harboring pMW119f1(−)NAEPBNG or pMW119f1(+)NAEPBNG grown in the presence of uridine (75 µg/mL), and phage particles were precipitated overnight in the presence of 5% polyethylene glycol (average MW of 6000) and 0.5 M NaCl at 4 °C. Uracil-containing circular ssDNA was extracted from the pelleted phage particles with lysis buffer (10 mM MOPS, pH 6.5, 1% Triton X-100, 500 mM guanidine-HCl) and a Qiagen HiSpeed Plasmid Midi kit (Qiagen, Tokyo, Japan). Fifty pmol of oligonucleotides (Table S1) were annealed with 20 pmol of Uracil-containing circular ssDNA. The first strand was synthesized from the annealed primer in the presence of 50 mM Tris–HCl, pH 7.5, 5 mM MgCl2, 1 mM DTT, 0.5 mM ATP, 0.185 mM dNTPs, 5% PEG (average MW of 8000), 0.2 units of pyrophosphatase (New England Biolabs, Tokyo, Japan), 50 units of native T7 DNA polymerase (New England Biolabs, Tokyo, Japan) and 15 units of T4 DNA ligase (Thermo Fisher Scientific, Tokyo, Japan) at 33 °C for 30 min, followed by heat inactivation of the enzymes at 75 °C for 20 min. After digestion of the uracil-containing template strand at 37 °C for 1 h with 3 units of UDG, 50 units of exonuclease I, and 50 units of exonuclease III (all from New England Biolabs, Tokyo, Japan), the resulting single stranded circular first strand was purified with a QIAquick Gel Extraction Kit (Qiagen, Tokyo, Japan). Twenty pmol of the 5′-phosphorylated primer was annealed to the first strand, and the second strand was synthesized. The 8-oxoG-containing primers were used for first strand synthesis, and the uracil-containing primers were used for second strand synthesis. The final product (circular dsDNA) was gel-purified with a QIAquick Gel Extraction Kit.
Transformation and analysis of the amplified plasmids
The constructed plasmids (6.6 ng) were electroporated into E. coli cells with a Bio-Rad E. coli pulser set at 1.8 kV, 200 Ω and 25 µF. Under our experimental conditions, each type of plasmid generated 2000–3000 transformants (Table 2S and 3S). Plasmids from the overnight culture of transformants were extracted with a QIAprep Spin Miniprep Kit (Qiagen, Tokyo, Japan). For the analysis of the plasmids from each clone, bacterial culture was grown from each colony, and the plasmids were subsequently extracted. To quantify the fraction of the strand used as a template, plasmids were first linearized with ScaI, and the linearized fragments were further digested with XhoI (Fig. 1B). To measure the MF, the propagated plasmids were digested with Alw26I. After digestion, the plasmids were electrophoresed in an agarose gel, and the relative intensities of the ethidium bromide-stained DNA fragments were measured with a gel imaging system (FluoroChem Imager, Proteinsimple, Tokyo, Japan). The fraction of the strand used as a template was estimated based on the intensity of the uncut fragment relative to all fragments (both cut and uncut fragments). Similarly, the fraction of the uncut Alw26I plasmid was quantified to estimate the MF. The transformation was repeated at least three times for each plasmid construct, and the statistical significance of the differences in the fractions of uncut fragments between the plasmids was determined with the two-tailed t-test. For the analysis of the distribution of template strand usage in each clone, the significant difference was verified by the χ-square test.
Data availability
The strains and plasmids used are available upon request. The authors affirm that all data necessary for confirming the conclusions of the article are present within the article, figures, and tables.
References
Ward, J. F. Complexity of damage produced by ionizing radiation. Cold Spring Harb. Symp. Quant. Biol. 65, 377–382. https://doi.org/10.1101/sqb.2000.65.377 (2000).
Ward, J. F. The complexity of DNA damage: Relevance to biological consequences. Int. J. Radiat. Biol. 66, 427–432 (1994).
Goodhead, D. T. Initial events in the cellular effects of ionizing radiations: Clustered damage in DNA. Int. J. Radiat. Biol. 65, 7–17 (1994).
Gulston, M., Fulford, J., Jenner, T., de Lara, C. & O’Neill, P. Clustered DNA damage induced by gamma radiation in human fibroblasts (HF19), hamster (V79–4) cells and plasmid DNA is revealed as Fpg and Nth sensitive sites. Nucleic Acids Res. 30, 3464–3472 (2002).
Sutherland, B. M., Bennett, P. V., Sidorkina, O. & Laval, J. Clustered DNA damages induced in isolated DNA and in human cells by low doses of ionizing radiation. Proc. Natl. Acad. Sci. USA 97, 103–108 (2000).
Sutherland, B. M., Bennett, P. V., Sutherland, J. C. & Laval, J. Clustered DNA damages induced by X rays in human cells. Radiat. Res. 157, 611–616 (2002).
Tsao, D. et al. Induction and processing of oxidative clustered DNA lesions in 56Fe-ion-irradiated human monocytes. Radiat. Res. 168, 87–97 (2007).
Watanabe, R., Rahmanian, S. & Nikjoo, H. Spectrum of radiation-induced clustered non-DSB damage—A Monte Carlo track structure modeling and calculations. Radiat. Res. 183, 525–540. https://doi.org/10.1667/RR13902.1 (2015).
Moeini, H., Mokari, M., Alamatsaz, M. H. & Taleei, R. Calculation of the initial DNA damage induced by alpha particles in comparison with protons and electrons using Geant4-DNA. Int. J. Radiat. Biol. 96, 767–778. https://doi.org/10.1080/09553002.2020.1730015 (2020).
Semenenko, V. A. & Stewart, R. D. A fast Monte Carlo algorithm to simulate the spectrum of DNA damages formed by ionizing radiation. Radiat. Res. 161, 451–457 (2004).
Nikjoo, H., O’Neill, P., Wilson, W. E. & Goodhead, D. T. Computational approach for determining the spectrum of DNA damage induced by ionizing radiation. Radiat. Res. 156, 577–583 (2001).
Nakano, T. et al. Formation of clustered DNA damage in vivo upon irradiation with ionizing radiation: Visualization and analysis with atomic force microscopy. Proc. Natl. Acad. Sci. USA 119, e2119132119. https://doi.org/10.1073/pnas.2119132119 (2022).
Xu, X. et al. Direct observation of damage clustering in irradiated DNA with atomic force microscopy. Nucleic Acids Res. 48, e18. https://doi.org/10.1093/nar/gkz1159 (2020).
Weinfeld, M., Rasouli-Nia, A., Chaudhry, M. A. & Britten, R. A. Response of base excision repair enzymes to complex DNA lesions. Radiat. Res. 156, 584–589 (2001).
Shikazono, N., Noguchi, M., Fujii, K., Urushibara, A. & Yokoya, A. The yield, processing, and biological consequences of clustered DNA damage induced by ionizing radiation. J. Radiat. Res. 50, 27–36 (2009).
Eccles, L. J., O’Neill, P. & Lomax, M. E. Delayed repair of radiation induced clustered DNA damage: Friend or foe?. Mutat. Res. 711, 134–141. https://doi.org/10.1016/j.mrfmmm.2010.11.003 (2011).
Sage, E. & Harrison, L. Clustered DNA lesion repair in eukaryotes: Relevance to mutagenesis and cell survival. Mutat. Res. 711, 123–133. https://doi.org/10.1016/j.mrfmmm.2010.12.010 (2011).
Sage, E. & Shikazono, N. Radiation-induced clustered DNA lesions: Repair and mutagenesis. Free Radic. Biol. Med. 107, 125–135. https://doi.org/10.1016/j.freeradbiomed.2016.12.008 (2017).
Malyarchuk, S., Brame, K. L., Youngblood, R., Shi, R. & Harrison, L. Two clustered 8-oxo-7,8-dihydroguanine (8-oxodG) lesions increase the point mutation frequency of 8-oxodG, but do not result in double strand breaks or deletions in Escherichia coli. Nucleic Acids Res. 32, 5721–5731 (2004).
Shikazono, N., Pearson, C., O’Neill, P. & Thacker, J. The roles of specific glycosylases in determining the mutagenic consequences of clustered DNA base damage. Nucleic Acids Res. 34, 3722–3730. https://doi.org/10.1093/nar/gkl503 (2006).
Shikazono, N. et al. Significance of DNA polymerase I in in vivo processing of clustered DNA damage. Mutat. Res. 749, 9–15. https://doi.org/10.1016/j.mrfmmm.2013.07.010 (2013).
Bellon, S., Shikazono, N., Cunniffe, S., Lomax, M. E. & O’Neill, P. Processing of thymine glycol in a clustered DNA damage site: Mutagenic or cytotoxic. Nucleic Acids Res. https://doi.org/10.1093/nar/gkp422 (2009).
Cunniffe, S., Walker, A., Stabler, R., O’Neill, P. & Lomax, M. E. Increased mutability and decreased repairability of a three-lesion clustered DNA-damaged site comprised of an AP site and bi-stranded 8-oxoG lesions. Int. J. Radiat. Biol. 90, 468–479. https://doi.org/10.3109/09553002.2014.899449 (2014).
Pearson, C. G., Shikazono, N., Thacker, J. & O’Neill, P. Enhanced mutagenic potential of 8-oxo-7,8-dihydroguanine when present within a clustered DNA damage site. Nucleic Acids Res. 32, 263–270 (2004).
Kozmin, S. G., Eot-Houllier, G., Reynaud-Angelin, A., Gasparutto, D. & Sage, E. Dissecting highly mutagenic processing of complex clustered DNA damage in yeast Saccharomyces cerevisiae. Cells https://doi.org/10.3390/cells10092309 (2021).
Noguchi, M., Urushibara, A., Yokoya, A., O’Neill, P. & Shikazono, N. The mutagenic potential of 8-oxoG/single strand break-containing clusters depends on their relative positions. Mutat. Res. 732, 34–42. https://doi.org/10.1016/j.mrfmmm.2011.12.009 (2012).
Sedletska, Y., Radicella, J. P. & Sage, E. Replication fork collapse is a major cause of the high mutation frequency at three-base lesion clusters. Nucleic Acids Res. 41, 9339–9348. https://doi.org/10.1093/nar/gkt731 (2013).
Takahashi, M., Akamatsu, K. & Shikazono, N. A polymerization-based method to construct a plasmid containing clustered DNA damage and a mismatch. Anal. Biochem. 510, 129–135. https://doi.org/10.1016/j.ab.2016.07.007 (2016).
Shikazono, N. & Akamatsu, K. Strand with mutagenic lesion is preferentially used as a template in the region of a bi-stranded clustered DNA damage site in Escherichia coli. Sci. Rep. 10, 9737. https://doi.org/10.1038/s41598-020-66651-0 (2020).
Kuzminov, A. Single-strand interruptions in replicating chromosomes cause double-strand breaks. Proc. Natl. Acad. Sci. USA 98, 8241–8246. https://doi.org/10.1073/pnas.131009198 (2001).
Kouzminova, E. A. & Kuzminov, A. Fragmentation of replicating chromosomes triggered by uracil in DNA. J. Mol. Biol. 355, 20–33. https://doi.org/10.1016/j.jmb.2005.10.044 (2006).
Mahaseth, T. & Kuzminov, A. Prompt repair of hydrogen peroxide-induced DNA lesions prevents catastrophic chromosomal fragmentation. DNA Repair (Amst) 41, 42–53. https://doi.org/10.1016/j.dnarep.2016.03.012 (2016).
Michel, B., Sinha, A. K. & Leach, D. R. F. Replication fork breakage and restart in Escherichia coli. Microbiol. Mol. Biol. Rev. https://doi.org/10.1128/MMBR.00013-18 (2018).
Kuzminov, A. Collapse and repair of replication forks in Escherichia coli. Mol. Microbiol. 16, 373–384. https://doi.org/10.1111/j.1365-2958.1995.tb02403.x (1995).
Kornberg, A. & Baker, T. A. DNA Replication 113–159 (University Science Books, 1992).
Friedberg, E. C. et al. DNA Repair and Mutagenesis 2nd edn. (ASM Press, 2006).
Vaisman, A. et al. Investigating the mechanisms of ribonucleotide excision repair in Escherichia coli. Mutat. Res. 761, 21–33. https://doi.org/10.1016/j.mrfmmm.2014.01.005 (2014).
Dianov, G. & Lindahl, T. Reconstitution of the DNA base excision-repair pathway. Curr. Biol. 4, 1069–1076. https://doi.org/10.1016/s0960-9822(00)00245-1 (1994).
Sung, J. S. & Mosbaugh, D. W. Escherichia coli uracil- and ethenocytosine-initiated base excision DNA repair: Rate-limiting step and patch size distribution. Biochemistry 42, 4613–4625. https://doi.org/10.1021/bi027115v (2003).
Setlow, P., Brutlag, D. & Kornberg, A. Deoxyribonucleic acid polymerase: Two distinct enzymes in one polypeptide. I. A proteolytic fragment containing the polymerase and 3′ leads to 5′ exonuclease functions. J. Biol. Chem. 247, 224–231 (1972).
Setlow, P. & Kornberg, A. Deoxyribonucleic acid polymerase: Two distinct enzymes in one polypeptide. II. A proteolytic fragment containing the 5′ leads to 3′ exonuclease function. Restoration of intact enzyme functions from the two proteolytic fragments. J. Biol. Chem. 247, 232–240 (1972).
Patel, P. H., Suzuki, M., Adman, E., Shinkai, A. & Loeb, L. A. Prokaryotic DNA polymerase I: Evolution, structure, and “base flipping” mechanism for nucleotide selection. J. Mol. Biol. 308, 823–837. https://doi.org/10.1006/jmbi.2001.4619 (2001).
Polesky, A. H., Steitz, T. A., Grindley, N. D. & Joyce, C. M. Identification of residues critical for the polymerase activity of the Klenow fragment of DNA polymerase I from Escherichia coli. J. Biol. Chem. 265, 14579–14591 (1990).
Xu, Y. et al. Biochemical and mutational studies of the 5′–3′ exonuclease of DNA polymerase I of Escherichia coli. J. Mol. Biol. 268, 284–302. https://doi.org/10.1006/jmbi.1997.0967 (1997).
Xu, Y., Potapova, O., Leschziner, A. E., Grindley, N. D. & Joyce, C. M. Contacts between the 5′ nuclease of DNA polymerase I and its DNA substrate. J. Biol. Chem. 276, 30167–30177. https://doi.org/10.1074/jbc.M100985200 (2001).
Derbyshire, V. et al. Genetic and crystallographic studies of the 3′,5′-exonucleolytic site of DNA polymerase I. Science 240, 199–201. https://doi.org/10.1126/science.2832946 (1988).
Singh, K., Srivastava, A., Patel, S. S. & Modak, M. J. Participation of the fingers subdomain of Escherichia coli DNA polymerase I in the strand displacement synthesis of DNA. J. Biol. Chem. 282, 10594–10604. https://doi.org/10.1074/jbc.M611242200 (2007).
Joyce, C. M., Fujii, D. M., Laks, H. S., Hughes, C. M. & Grindley, N. D. Genetic mapping and DNA sequence analysis of mutations in the polA gene of Escherichia coli. J. Mol. Biol. 186, 283–293. https://doi.org/10.1016/0022-2836(85)90105-6 (1985).
Heijneker, H. L. et al. A mutant of Escherichia coli K12 deficient in the 5′–3′ exonucleolytic activity of DNA polymerase I. II. Purification and properties of the mutant enzyme. Mol. Gen. Genet. 124, 83–96. https://doi.org/10.1007/BF00267167 (1973).
Makiela-Dzbenska, K. et al. Role of Escherichia coli DNA polymerase I in chromosomal DNA replication fidelity. Mol. Microbiol. 74, 1114–1127. https://doi.org/10.1111/j.1365-2958.2009.06921.x (2009).
Parker, B. O. & Marinus, M. G. Repair of DNA heteroduplexes containing small heterologous sequences in Escherichia coli. Proc. Natl. Acad. Sci. USA 89, 1730–1734. https://doi.org/10.1073/pnas.89.5.1730 (1992).
Shikazono, N. & O’Neill, P. Biological consequences of potential repair intermediates of clustered base damage site in Escherichia coli. Mutat. Res. 669, 162–168. https://doi.org/10.1016/j.mrfmmm.2009.06.004 (2009).
Luisi-DeLuca, C. & Kolodner, R. Purification and characterization of the Escherichia coli RecO protein. Renaturation of complementary single-stranded DNA molecules catalyzed by the RecO protein. J. Mol. Biol. 236, 124–138. https://doi.org/10.1006/jmbi.1994.1123 (1994).
Hall, S. D. & Kolodner, R. D. Homologous pairing and strand exchange promoted by the Escherichia coli RecT protein. Proc. Natl. Acad. Sci. USA 91, 3205–3209. https://doi.org/10.1073/pnas.91.8.3205 (1994).
Kuzminov, A. Recombinational repair of DNA damage in Escherichia coli and bacteriophage lambda. Microbiol. Mol. Biol. Rev. 63, 751–813. https://doi.org/10.1128/MMBR.63.4.751-813.1999 (1999) (table of contents).
Cupples, C. G. & Miller, J. H. A set of lacZ mutations in Escherichia coli that allow rapid detection of each of the six base substitutions. Proc. Natl. Acad. Sci. USA 86, 5345–5349. https://doi.org/10.1073/pnas.86.14.5345 (1989).
Kolodner, R., Hall, S. D. & Luisi-DeLuca, C. Homologous pairing proteins encoded by the Escherichia coli recE and recT genes. Mol. Microbiol. 11, 23–30. https://doi.org/10.1111/j.1365-2958.1994.tb00286.x (1994).
Shikazono, N. & Akamatsu, K. Mutagenic potential of 8-oxo-7,8-dihydroguanine (8-oxoG) is influenced by nearby clustered lesions. Mutat. Res. 810, 6–12. https://doi.org/10.1016/j.mrfmmm.2018.05.001 (2018).
Lovett, S. T. Template-switching during replication fork repair in bacteria. DNA Repair (Amst) 56, 118–128. https://doi.org/10.1016/j.dnarep.2017.06.014 (2017).
Maslowska, K. H., Makiela-Dzbenska, K., Mo, J. Y., Fijalkowska, I. J. & Schaaper, R. M. High-accuracy lagging-strand DNA replication mediated by DNA polymerase dissociation. Proc. Natl. Acad. Sci. USA 115, 4212–4217. https://doi.org/10.1073/pnas.1720353115 (2018).
Mourgues, S., Lomax, M. E. & O’Neill, P. Base excision repair processing of abasic site/single-strand break lesions within clustered damage sites associated with XRCC1 deficiency. Nucleic Acids Res. 35, 7676–7687 (2007).
Radicella, J. P., Clark, E. A., Chen, S. & Fox, M. S. Patch length of localized repair events: Role of DNA polymerase I in mutY-dependent mismatch repair. J. Bacteriol. 175, 7732–7736. https://doi.org/10.1128/jb.175.23.7732-7736.1993 (1993).
Bhardwaj, A., Ghose, D., Thakur, K. G. & Dutta, D. Escherichia coli beta-clamp slows down DNA polymerase I dependent nick translation while accelerating ligation. PLoS One 13, e0199559. https://doi.org/10.1371/journal.pone.0199559 (2018).
Taylor, A. F. & Weiss, B. Role of exonuclease III in the base excision repair of uracil-containing DNA. J. Bacteriol. 151, 351–357. https://doi.org/10.1128/jb.151.1.351-357.1982 (1982).
Demple, B. & Harrison, L. Repair of oxidative damage to DNA: Enzymology and biology. Annu. Rev. Biochem. 63, 915–948. https://doi.org/10.1146/annurev.bi.63.070194.004411 (1994).
Woodrick, J. et al. A new sub-pathway of long-patch base excision repair involving 5′ gap formation. EMBO J. 36, 1605–1622. https://doi.org/10.15252/embj.201694920 (2017).
Dianov, G. et al. Release of 5′-terminal deoxyribose-phosphate residues from incised abasic sites in DNA by the Escherichia coli RecJ protein. Nucleic Acids Res. 22, 993–998. https://doi.org/10.1093/nar/22.6.993 (1994).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97, 6640–6645. https://doi.org/10.1073/pnas.120163297 (2000).
Acknowledgements
We thank Dr. T. Nakano for the helpful discussions. We would also like to thank C. Ishii and S. Kayamura for their technical assistance. We are grateful to Catherine Perfect, MA (Cantab), from Edanz, for editing a draft of this manuscript.
Funding
This work was supported, in part, by KAKENHI grant numbers 17K20052 and 21K18148.
Author information
Authors and Affiliations
Contributions
N. S. designed the study. N. S. and K. A. performed the experiments and analyzed the data. N. S. wrote the manuscript with input from K. A.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shikazono, N., Akamatsu, K. The role of DNA polymerase I in tolerating single-strand breaks generated at clustered DNA damage in Escherichia coli. Sci Rep 14, 19124 (2024). https://doi.org/10.1038/s41598-024-69823-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-69823-4
- Springer Nature Limited