Abstract
Germline TP53 pathogenic variants can lead to a cancer susceptibility syndrome known as Li–Fraumeni (LFS). Variants affecting its activity can drive tumorigenesis altering p53 pathways and their identification is crucial for assessing individual risk. This study explored the functional impact of TP53 missense variants on its transcription factor activity. We selected seven TP53 missense variants (c.129G > C, c.320A > G, c.417G > T, c.460G > A, c,522G > T, c.589G > A and c.997C > T) identified in Brazilian families at-risk for LFS. Variants were created through site-directed mutagenesis and transfected into SK-OV-3 cells to assess their transcription activation capabilities. Variants K139N and V197M displayed significantly reduced transactivation activity in a TP53-dependent luciferase reporter assay. Additionally, K139N negatively impacted CDKN1A and MDM2 expression and had a limited effect on GADD45A and PMAIP1 upon irradiation-induced DNA damage. Variant V197M demonstrated functional impact in all target genes evaluated and loss of Ser15 phosphorylation. K139N and V197M variants presented a reduction of p21 levels after irradiation. Our data show that K139N and V197M negatively impact p53 functions, supporting their classification as pathogenic variants. This underscores the significance of conducting functional studies on germline TP53 missense variants classified as variants of uncertain significance to ensure proper management of LFS-related cancer risks.
Similar content being viewed by others
Introduction
Pathogenic germline variants in TP53 are associated with a complex inherited cancer-predisposing disorder known as Li–Fraumeni Syndrome (LFS)1. Patients with LFS are prone to develop a wide range of early-onset cancers, such as breast cancer, glioblastomas, osteosarcomas, and adrenocortical carcinomas, among others2,3,4,5. Thus, identifying individuals at risk for LFS is particularly important to optimize surveillance aimed at downstaging and ultimately improved survival.
Li–Fraumeni families face a significantly increased risk for several cancer types, nearly 24 times higher than the general population (standardized incidence ratio 23.9; 95% CI 21.9–26.0) for any cancer6. While guidelines exist for managing and preventing carriers of TP53 pathogenic/likely pathogenic (P/LP) variants, as well as those with other syndromes related to TP53 germline variants4,7,8,9, establishing appropriate management for carriers of variants of uncertain significance (VUS) remains challenging, relying solely on family history10. Missense variants account for nearly 74% of all germline TP53 alterations reported, and many of them remain unclassified11,12. Variants classified as VUS yield inconclusive genetic information, often with limited data on their functional impact on tumorigenesis. This limitation hampers proper patient management and follow-up.
The TP53 gene encodes for a tumor suppressor protein that preserves genome integrity and prevents tumorigenesis by regulating DNA repair, cell growth, senescence, and cell death13,14. p53 acts as a transcription factor, promoting or repressing transcription, and interacts with DNA as a tetramer15,16,17. Secondary and tertiary structural features of p53 determine its affinity and specificity during DNA sequence recognition and binding18. Theoretically, deleterious alterations in highly conserved amino acid residues can disrupt the conformation of structural domains, thereby influencing p53 function.
Functional assays provide key information for variant classification. Particularly in the analysis of VUS with low population frequency. These assays unveil the impact of the VUS on protein function, when combined with available genetic and epidemiologic information, can contribute to cancer causality prediction algorithms19,20,21,22. In this study, we used a TP53-mutated human ovarian cancer cell line that lacks p53 expression at both protein and mRNA levels23. This model was utilized to assess the functional analysis of seven TP53 missense variants identified in Brazilian families at-risk for LFS.
Results
Personal and family history of cancer, in silico predicted impact of TP53 variants, and previously reported functional analysis
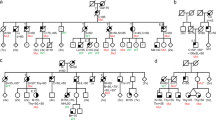
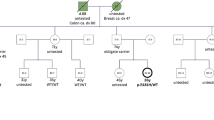
The personal and family history of patients carrying the selected TP53 variants are described in Table 1 and were previously published by Bittar et al.24.
Regarding the in silico predictions and the previously reported functional assays, REVEL predominantly classified most variants as pathogenic or having a medium impact and no variant showed an impact on splice sites (Table 1). The L43F and R333C variants were considered functional, and the others were considered non-functional or partially functional by Kato et al.20. Other in silico prediction tools were also accessed so that, together with our results from functional assays, we could propose a new classification for the variants evaluated. All results are described in Supplementary Table 1.
Expression and transcription transactivation activity of p53 variants
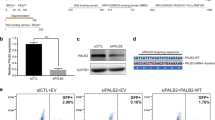
To verify the ability of our TP53 constructs, with the selected variants (Fig. 1A), to restore p53 expression in the p53-null SK-OV-3 cells (characterized by the absence of functional p53 protein), we transfected the cell and evaluated mRNA and protein expression. Figure 1B shows the presence of p53 mRNA in all conditions (pcDNA3-TP53 WT and variants), except for the empty vector (pcDNA3). These results were corroborated by protein analyses (Supplementary Fig. 1).
Overexpression of TP53 variants and wild-type (WT) constructs selected for functional analysis. (A)—Distribution of TP53 variants analyzed in this study. TAD: transactivation domain, PRD: proline-rich domain, DBD: DNA-binding domain, NLS: nuclear localization signal, TD: tetramerization domain, and BD: basic domain. (B)—TP53 mRNA expression levels for all transfected constructs, including empty vector (EV), p53 wild-type (WT) and p53 variants. (C)—Analysis of transcriptional activation (TA) capacity of TP53 variants. Variant activity is represented in relation to wild-type construction activity. The black bars represent the positive (WT) and negative controls (EV), and the gray bars represent the analyzed variants. p-values < 0.05 were considered statistically significant; (*) p < 0.05; (****) p < 0.0001.
To assess the functional effects of the seven TP53 VUS, we performed a luciferase reporter assay to quantitatively measure the capacity of each p53 variant to transactivate luciferase expression through the p53 DNA-binding site (derived from WAF1 promoter)19. Overall, four variants (L43F, Y107C, G154S and R174S) showed comparable activity to wild-type p53 (100%), ranging from 86.7 to 124.1% (Fig. 1C). The R333C variant significantly increased luciferase expression (198.7%), while the K139N variant showed residual activity with a statistically significant reduction in transactivation activity compared to the WT construct (62.9%). Notably, V197M showed negligible reporter activity (3.6%), similar to the absence of p53 condition (empty vector). No transcriptional activity was reported for V197M, even upon DNA damage induction (data not shown).
K139N and V197M variants impact p53 target genes expression
To confirm the functional impact observed in the transcription transactivation assay for all seven variants, we evaluated the ability of p53 variants to induce expression of key downstream target, including CDKN1A, MDM2, GADD45A and PMAIP1. Cells expressing wild-type p53 exhibited elevated mRNA levels for CDKN1A (Fig. 2A), MDM2 (Fig. 2B), GADD45A (Fig. 2C) and PMAIP1 (Fig. 2D) compared to cells transfected with empty vector. Upon DNA damage induction through ionizing irradiation, the expression of all tested target genes increased, except for GADD45A, which maintained its expression levels.
p53 overexpression impact in mRNA relative expression of p53’s target genes. The quantifications were normalized by endogenous housekeeping gene GUSB. The WT conditions were settled = 1 and the mRNA expression of CDKN1A (A), MDM2 (B), GADD45A (C) and PMAIP (D) genes were relativized by WT. The statistical comparisons between the variants were performed with their respective WT conditions (control or 10 Gy). p-values < 0.05 (*) were considered statistically significant; (**) p < 0.005; (***) p < 0.001; and (****) p < 0.0001. UT = untreated.
In general, the L43F, Y107C, G154S, R174S and R333C variants showed comparable or higher expression levels of CDKN1A, MDM2, GADD45A and PMAIP1 compared to WT condition. Cells expressing the K139N variant showed limited CDKN1A, MDM2 and GADD45A expression induction, with the exception of the PMAIP1 gene. While the K139N variant displayed higher levels of PMAIP1 expression than the WT control before irradiation, a substantial decrease was observed after DNA damage. The V197M variant substantially impacts mRNA levels for the CDKN1A gene. However, this variant shows a limited induction of mRNA expression for the other target genes (MDM2, GADD45A, and PMAIP1) in both, irradiated and non-irradiated, conditions. The observed results for target gene expression in cells carrying the K139N and V197M variants align with the functional impact observed during transcriptional transactivation with these p53 variants.
Loss of p53 Ser15 phosphorylation and impact on p21 protein levels
We also examined the impact of each variant on the phosphorylation of serine residue 15 in p53 (phospho-p53 Ser15) and p21 protein expression (Fig. 3). All variants, except for K139N and V197M, led to an increase in p21 protein levels after transfection (Fig. 3A–C). Following DNA damage, L43F and Y107C induced higher p21 protein levels than those induced by wild-type p53, while the other three variants showed similar p21 induction.
Analysis of p21 and p-p53 Ser15 protein levels upon DNA damage. (A)—Western blot of p53 (53KDa), p-p53 Ser15 (53KDa), p21 (21KDa) and β-actin (42KDa), in a control or irradiated (10 Gy) conditions. Cellular protein levels of p-p53Ser15 (B) and p21 (C) in SK-OV-3 cell line overexpressing exogenous p53 (WT or variants) were quantified in control and 4 h after ionizing irradiation (10 Gy) conditions. p-values < 0.05 (*) were considered statistically significant. The analyzes were performed from two independent experiments.
The phospho-p53 Ser15 levels significantly increased with wild-type p53 and the variants, except for V197M (Fig. 3A, B). Notably, V197M demonstrated a significant reduction in phospho-p53 Ser15 levels, equivalent to the control (empty vector), even after DNA damage.
Presence of K139N and V197M variants affects p53 nuclear stabilization
Phosphorylation of serine residue 15 is crucial for stabilization of p53 in the nucleus after DNA damage8. Consequently, we compared the capacity of p53 WT, K139N and V197M variants to translocate to the nucleus upon DNA damage. The evaluation of p53 WT, K139N and V197M variants localization after irradiation was performed through immunofluorescence assay. As expected, the presence of the K139N variant significantly impairs the nuclear accumulation of p53 compared to p53 WT condition (p < 0.005). In contrast, the presence of V197M variant does not abolish p53 translocation nor does it affect p53’s nuclear levels compared to p53 WT (Fig. 4).
Immunofluorescence analysis of p53 protein levels in p53 WT and K139N or V197M variants after irradiation (10 Gy). (A)—Representative images showing nuclear localization of p53 in p53 WT and K139N or V197M variants; scale bar, 40 µm; (B)—The ratio between Integrated Density (IntDen) and cell number of each condition. The presence of variants decreased the nuclear p53 levels compared to p53 WT. p-values < 0.05 were considered statistically significant; (**) p < 0.005.
Discussion
The presence of pathogenic/likely pathogenic (P/LP) variants in the TP53 gene can lead to Li–Fraumeni Syndrome or Li–Fraumeni like syndrome. Interpreting the pathogenicity of variants identified in patients, particularly missense variants, can be challenging24,25, however adequate classification means appropriate management for carriers of variants in TP53 gene. The process of variant classification is complex and considers specificities of each gene, such as the minimum population frequency that takes into account the penetrance of the gene in question (PM2/BS1), personal and family history (PP4), variant segregation (PP1), case–control studies (PS4), variant type (PVS1), in silico prediction (PP3/BP4), and functional assays (PS3/BS3), among other criteria. From this perspective, we selected seven missense variants, identified in families at-risk for hereditary cancer and classified as VUS by ClinVar26, to conduct functional assays and provide additional evidence, as the ACMG guidelines recognize them among the most robust types of evidence for determining variant pathogenicity10.
Considering the results observed in the transactivation capability (TA assay) for K139N and V197M variants in addition to defective expression of selected p53 target genes (CDKN1A, MDM2, GADD45A, and PMAIP1) we can confirm the impact on transcriptional function for both variants. P53 is a well-characterized transcriptional activation factor for genes involved in cell-cycle arrest, apoptosis, and DNA repair15,27,28, consequently, the failure of K139N and V197M variants to induce the expression of these genes demonstrated a deleterious behavior, akin to other TP53 variants previously associated with cancer predisposition19,20,29.
The K139N variant showed a reduction in mRNA levels of CDKN1A and MDM2, while V197M showed a clear loss of expression for CDKN1A. The impacts in transcriptional activation for CDKN1A and MDM2 genes can be stronger because their expression is mainly regulated by p53, meanwhile, GADD45A and PMAIP1 mRNA levels are regulated by many other proteins, including p5330,31,32. For this reason, the impact on their expression levels is much more subtle. WT p53 induces a slight increase in GADD45A expression after irradiation which may be explained by the plethora of proteins regulating it. For MDM2, we also observed an increase in mRNA levels after irradiation, but surely, we expected a bigger difference. Here, we used SK-OV-3 cells, a p53 null cell line, that has been previously shown to induce an increase in MDM2 protein levels when expressing exogenous WT p5333,34. We corroborated this by observing our mRNA data comparing empty vector and WT p53 conditions without irradiation. Thus, we expected that, after irradiation, induction of MDM2 expression wouldn’t be substantial since its expression has already been influenced by p53 transfection.
The loss of phosphorylation of p53 at Ser15 in the presence of the V197M variant could be explained by the failure in the transcriptional activation. This phenomenon is critical for its tetramer stabilization, the expression of target genes, and cell cycle arrest35, corroborating that p21 protein levels were significantly lower than in the p53 WT condition. On the hand, the presence of the K139N variant maintained its Ser15 phosphorylation; however, also presented p21 protein levels significantly lower than WT condition. Regarding the R333C variant, the observed functional profile like WT, is consistent with Fisher et al. who demonstrated that the cells were still able to form a multimeric structure in an in vitro p53-null model36. The results also corroborated with Kato et al.20 and Giacomelli et al.37, which described this variant as functional and not-DNE/LOF, respectively. In addition, the classification by ClinVar26 has recently changed, after we select variants and performance of functional assays in this work, and now is considered likely benign. We can also highlight that even carriers of variants classified as B/LB should benefit from better management, demonstrating the importance of improving the variant classification system.
Concerning the reduction in the nuclear levels of p53 upon DNA damage in the presence of the K139N variant, this lysine residue (139), along with two other residues, serves as an MDM2 target for ubiquitination and is responsible for p53 nuclear export38. Therefore, the alteration of the MDM2 ubiquitination site could lead to the abolishment of p53 nuclear levels. In the case of the V197M variant, the loss of Ser15 phosphorylation could have resulted in a reduction of p53’s tetramerization and, consequently, a decrease in its nuclear levels.
All analyses were conducted to enhance the characterization of pathogenicity for seven missense variants carried by families at-risk for hereditary cancer, some of whom reported a classical personal or family history of Li–Fraumeni Syndrome2,3,4. The results obtained from functional assays corroborated the personal and family history reported by carriers of variants demonstrating an impact on p53 functionality, as they reported a significant family history of cancer. Typically, families carrying variants considered pathogenic exhibit this pattern of family history, characterized by multiple tumors, some of which may manifest at young ages. The functional characterization of variants facilitates improved management and screening of patients and families carrying the same variants, enabling early tumor diagnosis and the establishment of an effective cancer surveillance program39. Additionally, it has been previously demonstrated that the adoption of surveillance for patients with LFS by the Brazilian National Health Care System is cost-beneficial40.
It is worth mentioning that there are already specific guidelines for classifying variants in the TP53 gene41, recommending the use of, among several other criteria, an in silico tool and some specific functional assays20,37. In this work, we assessed the results of Kato et al.20 and Giacomelli et al.37, to help in our final suggestion of classification. Furthermore, we chose to evaluate specific functions of the protein, with assays used in other works in the literature42,43,44, including expression of other genes regulated by p53, also evaluated by Kato et al.20 and Kawaguchi et al.45. Our results corroborate Kato et al.20 observations for the K139N variant, while this variant was not classified by Giacomelli et al.37. Additionally, V197M variant was classified as DNE/LOF37 and partially functional20, in a similar way to our findings. We also point out that, recently, the classification of the V197M variant in ClinVar26 changed to conflicting interpretations of pathogenicity (one LP classification and three VUS), and this germline variant was previously identified in another Brazilian family suggestive of Li–Fraumeni46.
Finally, evidence from functional assays is among the highest levels of evidence in variant classification10. There is an increasing emergence of large-scale assays to evaluate the pathogenicity of variants, mainly aiming to reduce the number of variants classified as VUS and to improve the classification system for little-known variants. However, it is important to mention that it was observed in the analysis of variants present in genes associated with the Lynch Syndrome that, although these assays provide important data on mechanisms, they also have several challenges and offer less precision when compared to individual assessments47. Taken together, our results, along with the patients’ family history (another important parameter to be considered), demonstrate that the presence of K139N or V197M variants adversely affects the functionality of p53, as well as the pathways regulated by this protein. The use of robust functional assays can help in the classification of identified variants, in addition to validating data from in silico prediction computational tools. However, it is worth highlighting that the variant’s classification is a dynamic process, and other parameters are also considered for the classification of a variant, such as population frequency, family segregation studies and case–control studies10.
Accordingly, we propose that these data could be taken into consideration in the classification of the K139N and V197M variants, potentially warranting their categorization as likely pathogenic (Supplementary Table 1). Another possibility would be that like what already exists in variant curation expert panels, such as BRCA1/2, differentiated weights of PS3/BS3 could be assigned to functional assays so that they can assist in the classification of a given variant, even with a lower weight than the originally proposed criterion. Patients with variants classified as pathogenic or likely pathogenic could benefit from well-established management according to the National Comprehensive Cancer Network (NCCN)7, as well as from the possibility to perform cascade testing in the family, thus offering personalized prevention measures for those relatives.
Conclusions
In summary, using multiple independent functional assays, we demonstrated a deleterious functional impact on p53 caused by the K139N and V197M variants. Furthermore, the functional analyses conducted here, suggesting structural interference for both variants, may contribute to the annotation of these variants, enabling a more accurate assessment of cancer risk for carrier families and influencing their clinical outcomes.
Methods
Ethical considerations
All clinical data presented in this work was previous published by Bittar et al.24, we did not generate any datasets, use any human sample for de functional assays or human database for analysis. No additional clinical information was disclosed in this work.
Patients and variant selection
Genetic germline testing results, including analysis of the TP53 (NM_000546; GrCh37) gene of patients from cancer risk evaluation programs in Southern and Southeastern Brazil, were analyzed (Hospital de Clínicas de Porto Alegre and Barretos Cancer Hospital). Variants classified as VUS according to the American College of Genetics and Genomics (ACMG)10 in a clinical diagnostic setting24 were selected for functional analyses in the present study. Variants were selected based on their location within p53’s important motifs and domains that impact the correct functioning of the protein (Supplementary Fig. 1A), classification at ClinVar26 (VUS) and phenotype (personal and family history of cancer). We also considered the in silico predictions by REVEL and Splice AI, and the functional assays previously published by Kato et al.20 as recommended by Fortuno et al.41.
Screening and confirmation of TP53 variants
Variants selected for functional studies were confirmed by Sanger or NGS sequencing by the Molecular Diagnostic Laboratory from Barretos Cancer Hospital and/or Hospital das Clínicas de Porto Alegre. Genomic DNA was extracted from peripheral blood using the QIAmp DNA Blood Mini QIAcube Kit (QIAGEN, #51126) with the automated QIAcube (Qiagen) platform following the manufacturer’s instructions. Genomic DNA was amplified by PCR and purified with the Affymetrix™ExoSAP-IT™ (#15513687) enzyme. Sequencing was performed with the BigDye™ Terminator v3.1 Cycle Sequencing Kit (Thermo Fisher Scientific, #4337456) and purified with the X-terminator kit v3.1 (Thermo Fisher Scientific, #4376484). Finally, sequencing was performed on an automated sequencer model 3500 (Applied Biosystems).
Plasmid vector constructions
TP53 missense variants were generated using as template the pcDNA3:flag-p53 plasmid containing human wild type (WT) TP53 cDNA (Addgene, #10838). TP53 variants were generated by site-directed mutagenesis as described previously48. Briefly, PCR amplification conditions were designed using PrimeSTAR® Max Polymerase (Takara, #R045A) according to manufacturer’s instructions. Mutagenesis primers are listed in Supplementary Table 2. The PCR products were digested with DpnI (New England Biolabs, #R0176S) overnight and transformed into NEB® 5-alpha Competent Escherichia coli cells (New England Biolabs, #C2981I). Single colonies were selected and sequenced to confirm the insertion of each variant of interesting and no additional alterations.
Cell culture
The SK-OV-3 (HTB-77; ATCC) ovarian adenocarcinoma cell line was cultured in RPMI 1640 medium (GIBCO, #11530586) supplemented with 10% fetal bovine serum (GIBCO, #26140079), 100 U/mL penicillin, and 100 mg/mL streptomycin (Invitrogen). Cells were maintained in a 5% CO2 atmosphere at 37 °C. Cell line authenticity was confirmed by DNA short tandem repeat analysis49. Moreover, mycoplasma contamination was assay using MycoAlert® Mycoplasma Detection Assays (Lonza, #LT07-318).
RNA extraction, RT-PCR, and real-time PCR
To evaluate the expression of genes regulated by p53, total RNA was extracted from transfected SK-OV-3 cells after DNA damage induction or in a basal condition (untreated), and the RT-PCR was performed. For this, cells were plated at a density of 5 × 105 cells per well in a 6-well plate, allowed to adhere overnight, transfected with 4 µg of plasmid constructions, and incubated for 24 h. Subsequently, for DNA damage condition, cells were irradiated with 10 Gy using the Rad Source 2000 Irradiator (Rad Source Technologies), after that, they were incubated for 4 h and the RNA was extracted, using TRIzol™ Reagent (Thermo Fisher Scientific, #15596018). The same extraction step was performed for untreated cells. Then, 1 µg of RNA was used for cDNA synthesis with SuperScript™ III Reverse Transcriptase (Thermo Fisher Scientific, #18080093) following the manufacturer’s instructions. Quantitative real-time PCR was performed using TaqMan™ Fast Advanced Master Mix (Applied Biosystems, #4444557) and PrimeTime Standard® qPCR Assay (IDT) probes. The evaluated targets were CDKN1A (Hs.PT.58.40874346.g), MDM2 (Hs.PT.58.358457), GADD45A (Hs.PT.58.20049396), PMAIP1 (Hs.PT.58.21318159), and endogenous genes GUSB (Hs.PT.58v.27737538) and TP53 (Hs.PT.58.123122), were used as endogenous controls. Reactions were performed in triplicate using the QuantStudio™ 6 Flex Real Time PCR System (Applied Biosystems). Reference genes were normalized by geometric means, and relative quantification measurements of target genes were determined using the comparative Ct method (2-ΔΔCt).
Transcription activation assay
To assess TP53 missense variant functionality, we used the transcriptional activation (TA) assay37. Briefly, SK-OV-3 cells were seeded at a density of 3.5 × 104 cells per well in a 96-well plate. Co-transfection was performed using vector constructions containing p53 WT (pcDNA3:flag-p53) or the selected missense variants, the pG13-Luc (Addgene, #16442) and pLX313-Renilla Luciferase (Addgene, #118016) plasmids, with Lipofectamine™3000 (Invitrogen, #L3000008). Cells were harvested 24 h post-transfection and TA was measured using the Dual-Luciferase® Reporter Assay System (Promega, #E1910). All assays were conducted in triplicate and repeated at least three times.
Immunoblotting and antibodies
To compare protein expression among different p53 conditions, SK-OV-3 cells were plated at a density of 5 × 105 cells per well in a 6-well plate, allowed to adhere overnight, transfected with 4 µg of plasmid constructions, and incubated for 24 h. Subsequently, cells were irradiated with 10 Gy using the Rad Source 2000 Irradiator (Rad Source Technologies), incubated for 4 h and lysed in ice-cold RIPA buffer (0.01 M EDTA, 1 mM de PMSF, 1 mM DTT, 0.1 µg/mL aprotinin e 1 mM leupeptin) for 30 min. Protein quantification was performed by the Bradford method50. Sample proteins (25 µg) were separated by polyacrylamide gel electrophoresis and transferred to Hybond-C™ Extra—(Amersham Biosciences, #45004018) membranes, using the Trans-Blot® Turbo™ Transfer System (Bio-Rad Laboratories). Non-specific sites were blocked with 5% milk, and membranes were incubated overnight at 4 °C with primary antibodies, including mouse monoclonal β-actin (Abcam, #ab49900), mouse monoclonal p53 (Cell Signaling Technology, #2524), mouse monoclonal phospho-p53 Ser15 (Cell Signaling Technology, #9286), rabbit monoclonal p21 (Cell Signaling Technology, #2947). Membranes were further incubated for 1 h at room temperature with anti-rabbit IgG, HRP linked antibody (Cell Signaling, #7074) or anti-mouse IgG, HRP-linked Antibody (Cell Signaling, #7076). The membranes were exposed to the SignalFire™ ECL Reagent (Cell Signaling, #6883) and photo documented by ImageQuant LAS 4000 mini (Cytiva).
Nuclear p53 levels by immunofluorescence
To assess the impairment of p53 nuclear translocation, we performed an immunofluorescence analysis. A density of of 2 × 104 cells was seeded per coverslip and incubated for 24 h. Constructs carrying p53 WT, K139N or V197M variants were transfected into the cells. After 24 h, cells were irradiated with 10 Gy and incubated for 30 minutes51. Cells were then fixed in a 4% paraformaldehyde solution and permeabilized with 0.5% Triton X-100. A blocking solution (5% BSA) was added for one hour, followed by incubation with anti-p53 (1C12) antibody (Cell Signaling Technology, #2524), diluted at 1:50, overnight at 4 °C. Subsequently, coverslips were incubated with donkey anti-mouse IgG (AlexaFluor® 488; Abcam, #ab150105), at a concentration of 1:1000 for one hour in the dark, at room temperature. Finally, ProLong™ Gold Antifade Mountant with DAPI (Invitrogen, #P36941) was added, and the coverslip was mounted onto the slide. Images were acquired using the Olympus FluoView FV10i (Olympus) fluorescent confocal microscope. Analysis was performed in the ImageJ software (National Institutes of Health), and the ratio between Integrated Density and cell number (according to DAPI) was calculated.
Statistical analysis
Data were expressed as mean ± standard deviation. Statistical analysis was performed using GraphPad Prism 9.0 software (California, USA). Comparisons of two variances were assessed by the One-way ANOVA test with Fisher’s Least Significant Difference test. The threshold for significance was established at p < 0.05.
Ethics declarations
All clinical data presented in this work was previous published by Bittar et al.25. No additional clinical information was disclosed in this work. Patient samples were not used for the functional assays of this work.
Consent to participate
We did not use patient’s sample, databases or generate datasets in this study.
Data availability
The datasets generated and/or analyzed during the current study are available in the PubMed repository, [Bittar et al.25; PMID: 31,321,604; DOI: https://doi.org/10.1007/s10689-019-00140-w]. We did not generate additional data.
References
Malkin, D. et al. Germ line p53 mutations in a familial syndrome of breast cancer, sarcomas, and other neoplasms. Science 250, 1233–1238. https://doi.org/10.1126/science.1978757 (1990).
Amadou, A., Achatz, M. I. W. & Hainaut, P. Revisiting tumor patterns and penetrance in germline TP53 mutation carriers: Temporal phases of Li–Fraumeni syndrome. Curr. Opin. Oncol. 30, 23–29. https://doi.org/10.1097/CCO.0000000000000423 (2018).
Fortuno, C. et al. Genotype-phenotype correlations among TP53 carriers: Literature review and analysis of probands undergoing multi-gene panel testing and single-gene testing. Cancer Genet. 248–249, 11–17. https://doi.org/10.1016/j.cancergen.2020.09.002 (2020).
Mai, P. L. et al. Risks of first and subsequent cancers among TP53 mutation carriers in the National Cancer Institute Li–Fraumeni syndrome cohort. Cancer 122, 3673–3681. https://doi.org/10.1002/cncr.30248 (2016).
Schneider, K., Zelley, K., Nichols, K. E. & Garber (1993) J. in GeneReviews (R) (eds.) M. P. Adam et al.
de Andrade, K. C. et al. Cancer incidence, patterns, and genotype-phenotype associations in individuals with pathogenic or likely pathogenic germline TP53 variants: An observational cohort study. Lancet Oncol. 22, 1787–1798. https://doi.org/10.1016/S1470-2045(21)00580-5 (2021).
Daly, M. B. et al. NCCN Guidelines(R) Insights: Genetic/Familial high-risk assessment: Breast, ovarian, and pancreatic, version 2.2024. J. Natl. Compr. Cancer Netw. 21, 1000–1010. https://doi.org/10.6004/jnccn.2023.0051 (2023).
Frebourg, T. et al. Guidelines for the Li–Fraumeni and heritable TP53-related cancer syndromes. Eur. J. Hum Genet. 28, 1379–1386. https://doi.org/10.1038/s41431-020-0638-4 (2020).
Kratz, C. P. et al. Analysis of the Li–Fraumeni spectrum based on an international germline TP53 variant data set: An international agency for research on cancer TP53 database analysis. JAMA Oncol. 7, 1800–1805. https://doi.org/10.1001/jamaoncol.2021.4398 (2021).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of medical genetics and genomics and the association for molecular pathology. Genet. Med. Offic. J. Am. College Med. Genet. 17, 405–424. https://doi.org/10.1038/gim.2015.30 (2015).
Guha, T. & Malkin, D. Inherited TP53 mutations and the Li–Fraumeni syndrome. Cold Spring Harb Perspect Med. 7, a026187. https://doi.org/10.1101/cshperspect.a026187 (2017).
Zhou, X., Hao, Q. & Lu, H. Mutant p53 in cancer therapy-the barrier or the path. J. Mol. Cell Biol. 11, 293–305. https://doi.org/10.1093/jmcb/mjy072 (2019).
Carlsen, L. & El-Deiry, W. S. Differential p53-mediated cellular responses to DNA-damaging therapeutic agents. Int. J. Mol. Sci. 22, 11828. https://doi.org/10.3390/ijms222111828 (2021).
Rizzotto, D., Englmaier, L. & Villunger, A. At a crossroads to cancer: How p53-induced cell fate decisions secure genome integrity. Int. J. Mol. Sci. 22, 10883. https://doi.org/10.3390/ijms221910883 (2021).
Gencel-Augusto, J. & Lozano, G. p53 tetramerization: at the center of the dominant-negative effect of mutant p53. Genes Dev. 34, 1128–1146. https://doi.org/10.1101/gad.340976.120 (2020).
Kitayner, M. et al. Structural basis of DNA recognition by p53 tetramers. Mol. Cell 22, 741–753. https://doi.org/10.1016/j.molcel.2006.05.015 (2006).
Ly, E., Kugel, J. F. & Goodrich, J. A. Single molecule studies reveal that p53 tetramers dynamically bind response elements containing one or two half sites. Sci. Rep. 10, 16176. https://doi.org/10.1038/s41598-020-73234-6 (2020).
Chillemi, G. et al. Structural evolution and dynamics of the p53 proteins. Cold Spring Harb Perspect Med. 7, a028308. https://doi.org/10.1101/cshperspect.a028308 (2017).
Doffe, F. et al. Identification and functional characterization of new missense SNPs in the coding region of the TP53 gene. Cell Death Differ. 28, 1477–1492. https://doi.org/10.1038/s41418-020-00672-0 (2021).
Kato, S. et al. Understanding the function-structure and function-mutation relationships of p53 tumor suppressor protein by high-resolution missense mutation analysis. Proc. Nat. Acad. Sci. USA. 100, 8424–8429. https://doi.org/10.1073/pnas.1431692100 (2003).
Quinn, E. A. et al. From uncertainty to pathogenicity: Clinical and functional interrogation of a rare TP53 in-frame deletion. Cold Spring Harb Mol Case Stud 5, a003921. https://doi.org/10.1101/mcs.a003921 (2019).
Soussi, T., Leroy, B. & Taschner, P. E. Recommendations for analyzing and reporting TP53 gene variants in the high-throughput sequencing era. Human Mutat. 35, 766–778. https://doi.org/10.1002/humu.22561 (2014).
Mullany, L. K. et al. Specific TP53 mutants overrepresented in ovarian cancer impact CNV, TP53 activity, responses to nutlin-3a, and cell survival. Neoplasia 17, 789–803. https://doi.org/10.1016/j.neo.2015.10.003 (2015).
Bittar, C. M. et al. TP53 variants of uncertain significance: Increasing challenges in variant interpretation and genetic counseling. Fam. Cancer 18, 451–456. https://doi.org/10.1007/s10689-019-00140-w (2019).
Butz, H. et al. Challenging interpretation of germline TP53 variants based on the experience of a national comprehensive cancer centre. Sci. Rep. 13, 14259. https://doi.org/10.1038/s41598-023-41481-y (2023).
Landrum, M. J. et al. ClinVar: Public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862-868. https://doi.org/10.1093/nar/gkv1222 (2016).
Fischer, M. Census and evaluation of p53 target genes. Oncogene 36, 3943–3956. https://doi.org/10.1038/onc.2016.502 (2017).
Raj, N. & Attardi, L. D. The transactivation domains of the p53 protein. Cold Spring Harb Perspect Med. 7, a026047. https://doi.org/10.1101/cshperspect.a026047 (2017).
Kharaziha, P. et al. Functional characterization of novel germline TP53 variants in Swedish families. Clin Genet. 96, 216–225. https://doi.org/10.1111/cge.13564 (2019).
Giono, L. E. & Manfredi, J. J. Mdm2 is required for inhibition of Cdk2 activity by p21, thereby contributing to p53-dependent cell cycle arrest. Mol. Cell Biol. 27, 4166–4178. https://doi.org/10.1128/MCB.01967-06 (2007).
Broude, E. V. et al. p21 (CDKN1A) is a negative regulator of p53 stability. Cell Cycle 6, 1468–1471 (2007).
Janus, P. et al. Pro-death signaling of cytoprotective heat shock factor 1: Upregulation of NOXA leading to apoptosis in heat-sensitive cells. Cell Death Differ. 27, 2280–2292. https://doi.org/10.1038/s41418-020-0501-8 (2020).
Ghosh, A., Chen, T. C. & Kapila, Y. L. Anoikis triggers Mdm2-dependent p53 degradation. Mol. Cell Biochem. 343, 201–209. https://doi.org/10.1007/s11010-010-0514-6 (2010).
Son, D. S., Kabir, S. M., Dong, Y. L., Lee, E. & Adunyah, S. E. Inhibitory effect of tumor suppressor p53 on proinflammatory chemokine expression in ovarian cancer cells by reducing proteasomal degradation of IkappaB. PLoS One 7, e51116. https://doi.org/10.1371/journal.pone.0051116 (2012).
Roos, W. P., Thomas, A. D. & Kaina, B. DNA damage and the balance between survival and death in cancer biology. Nat. Rev. Cancer 16, 20–33. https://doi.org/10.1038/nrc.2015.2 (2016).
Fischer, N. W., Prodeus, A., Tran, J., Malkin, D. & Gariepy, J. Association between the oligomeric status of p53 and clinical outcomes in Li–Fraumeni syndrome. J. Nat. Cancer Inst. 110, 1418–1421. https://doi.org/10.1093/jnci/djy114 (2018).
Giacomelli, A. O. et al. Mutational processes shape the landscape of TP53 mutations in human cancer. Nat. Genet. 50, 1381–1387. https://doi.org/10.1038/s41588-018-0204-y (2018).
Masuda, Y. et al. Stepwise multipolyubiquitination of p53 by the E6AP-E6 ubiquitin ligase complex. J. Biol. Chem. 294, 14860–14875. https://doi.org/10.1074/jbc.RA119.008374 (2019).
Mai, P. L. et al. Prevalence of cancer at baseline screening in the national cancer institute Li–Fraumeni syndrome cohort. JAMA Oncol. 3, 1640–1645. https://doi.org/10.1001/jamaoncol.2017.1350 (2017).
Frankenthal, I. A., Alves, M. C., Tak, C. & Achatz, M. I. Cancer surveillance for patients with Li–Fraumeni syndrome in Brazil: A cost-effectiveness analysis. Lancet Reg. Health Am. 12, 100265. https://doi.org/10.1016/j.lana.2022.100265 (2022).
Fortuno, C. et al. Specifications of the ACMG/AMP variant interpretation guidelines for germline TP53 variants. Human Mutat. 42, 223–236. https://doi.org/10.1002/humu.24152 (2021).
Berge, E. O., Huun, J., Lillehaug, J. R., Lonning, P. E. & Knappskog, S. Functional characterisation of p53 mutants identified in breast cancers with suboptimal responses to anthracyclines or mitomycin. Biochim. Biophys. Acta 830, 2790–2797. https://doi.org/10.1016/j.bbagen.2012.12.004 (1830).
Monti, P. et al. Dominant-negative features of mutant TP53 in germline carriers have limited impact on cancer outcomes. Mol. Cancer Res. 9, 271–279. https://doi.org/10.1158/1541-7786.MCR-10-0496 (2011).
Jordan, J. J. et al. Altered-function p53 missense mutations identified in breast cancers can have subtle effects on transactivation. Mol. Cancer Res. 8, 701–716. https://doi.org/10.1158/1541-7786.MCR-09-0442 (2010).
Kawaguchi, T. et al. The relationship among p53 oligomer formation, structure and transcriptional activity using a comprehensive missense mutation library. Oncogene 24, 6976–6981. https://doi.org/10.1038/sj.onc.1208839 (2005).
Achatz, M. I. et al. The TP53 mutation, R337H, is associated with Li–Fraumeni and Li–Fraumeni-like syndromes in Brazilian families. Cancer Lett. 245, 96–102. https://doi.org/10.1016/j.canlet.2005.12.039 (2007).
Abildgaard, A. B. et al. Lynch syndrome, molecular mechanisms and variant classification. Br. J. Cancer 128, 726–734. https://doi.org/10.1038/s41416-022-02059-z (2023).
Motohashi, K. A simple and efficient seamless DNA cloning method using SLiCE from Escherichia coli laboratory strains and its application to SLiP site-directed mutagenesis. BMC Biotechnol. 15, 47. https://doi.org/10.1186/s12896-015-0162-8 (2015).
Dirks, W. G., Faehnrich, S., Estella, I. A. & Drexler, H. G. Short tandem repeat DNA typing provides an international reference standard for authentication of human cell lines. Altex 22, 103–109 (2005).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254. https://doi.org/10.1006/abio.1976.9999 (1976).
Al Rashid, S. T. et al. Evidence for the direct binding of phosphorylated p53 to sites of DNA breaks in vivo. Cancer Res. 65, 10810–10821. https://doi.org/10.1158/0008-5472.CAN-05-0729 (2005).
Acknowledgements
We knowledge the Molecular Oncology Research Center from Barretos Cancer Hospital for the infrastructure support, the Hospital de Clínicas de Porto Alegre to contribute with data. We also thank the grant 2018/25118-8, São Paulo Research Foundation (FAPESP), and the Grant 25000.056766/2015-64, National Oncology Care Support Program (PRONON), from the Ministry of Health.
Funding
This work was supported by São Paulo Research Foundation (FAPESP, grant number 2018/25118-8) and National Oncology Care Support Program (PRONON, Grant Number 25000.056766/2015-64), from the Brazilian Ministry of Health.
Author information
Authors and Affiliations
Contributions
RBVA performed the data curation, investigation, formal analysis and validations, wrote original draft; ASP performed the investigation, formal analysis and validations, reviewed and edited the original draft; MNR performed the investigation, formal analysis and validations, reviewed and edited the original draft; PA-P was responsible for data curation, review and edit of the original draft; VAOS designed the project, acquired funding, reviewed and edited the original draft; MEM designed the project, reviewed and edited the original draft; EIP was responsible for data curation; supervision, conceptualization and administration of the project, acquired funding, reviewed and edited the original draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Abreu, R.B.V., Pereira, A.S., Rosa, M.N. et al. Functional evaluation of germline TP53 variants identified in Brazilian families at-risk for Li–Fraumeni syndrome. Sci Rep 14, 17187 (2024). https://doi.org/10.1038/s41598-024-67810-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-67810-3
- Springer Nature Limited