Abstract
Osteosarcoma (OS) is a heterogeneous malignant spindle cell tumor that is aggressive and has a poor prognosis. Although combining surgery and chemotherapy has significantly improved patient outcomes, the prognosis for OS patients with metastatic or recurrent OS has remained unsatisfactory. Therefore, it is imperative to gain a fresh perspective on OS development mechanisms and treatment strategies. After studying single-cell RNA sequencing (scRNA-seq) data in public databases, we identified seven OS subclonal types based on intra-tumor heterogeneity. Subsequently, we constructed a prognostic model based on pro-protein synthesis osteosarcoma (PPS-OS)-associated genes. Correlation analysis showed that the prognostic model performs extremely well in predicting OS patient prognosis. We also demonstrated that the independent risk factors for the prognosis of OS patients were tumor primary site, metastatic status, and risk score. Based on these factors, nomograms were constructed for predicting the 3- and 5-year survival rates. Afterward, the investigation of the tumor immune microenvironment (TIME) revealed the vital roles of γδ T-cell and B-cell activation. Drug sensitivity analysis and immune checkpoint analysis identified drugs that have potential application value in OS. Finally, the jumping translocation breakpoint (JTB) gene was selected for experimental validation. JTB silencing suppressed the proliferation, migration, and invasion of OS cells. Therefore, our research suggests that PPS-OS-related genes facilitate the malignant progression of OS and may be employed as prognostic indicators and therapeutic targets in OS.
Similar content being viewed by others
Introduction
Osteosarcoma (OS) is a heterogeneous malignant spindle cell tumor that most commonly occurs in children and adolescents1,2. It can arise in any bone but usually occurs in the metaphysis of long bones3. Surgery and chemotherapy, the standard treatment for OS established in the 1980s, has enabled long-term survival in 60% of patients with nonmetastatic tumors4. However, there are disadvantages to both chemotherapy and surgery. Excision by surgery frequently has a negative impact on athletic ability. Current chemotherapeutic drugs have low specificity, have serious adverse reactions, and are prone to drug resistance5. Tumors are composed of subpopulations (subclones) of cells of different phenotype due to the presence of ITH, which is also a powerful aid to cancer progression and therapeutic failure6. Over the past decade, the establishment of well-annotated tissue banks and the development of comprehensive molecular analysis techniques and preclinical models have enhance our understanding of the molecular mechanisms and biological heterogeneity of OS at the pathophysiological level7,8,9. However, progress in identifying new therapies has been slow, and treatment options are especially limited for patients with advanced or metastatic disease10. Thus, it is extremely valuable to study the molecular mechanisms and intra-tumor heterogeneity (ITH) related to the genesis and progression of OS, which may help us to filter out essential molecules or biomarkers for early diagnosis and targeted treatment.
Following an in-depth review of the scRNA-seq data, the current study identified seven unique subsets of OS cells based on ITH, each of which was annotated subjectively to facilitate further investigation. Next, the differentially expressed genes (DEGs) of each subtype were obtained, and a prognostic model was established based on the cell subtype with the worst survival indications. This cell subtype was annotated as pro-protein synthesis osteosarcoma (PPS-OS). Moreover, a nomogram was established by merging DEGs with clinical factors. Furthermore, the correlations of the model score with drug sensitivity and the TIME were assessed, thereby expanding the prognostic value of the gene signature for OS patients. In the final step, we conducted experiments in vitro to verify the effect of silencing the JTB gene. Our research suggests that PPS-OS-related genes play a vital role in the development and progression of OS and that JTB may be a novel target in the treatment of OS.
Methods
Data collection
The scRNA-seq data were obtained from the GSE152048 dataset in the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/, accessed on 15 January 2023). Eighty-five clinical case samples, including the survival information of OS patients, were obtained from the Therapeutically Applicable Research to Generate Effective Treatment (TARGET) database (https://ocg.cancer.gov/programs/target, accessed on 16 January 2023) and serve as the training cohort. Ninety clinical case samples from the GEO database were also gathered and served as the testing cohort. The samples (n = 396) for normal tissue were downloaded from the GTEx (https://gtexportal.org, accessed on 16 January 2023).
Processing of the scRNA-seq data and annotation of cell clusters
The data were processed statistically by the Seurat package in R 3.6.311. Firstly, the quality control of OS scRNA-seq data obtained from the database was performed, including correcting batch differences by using the Seurat3 software package12. The “LogNormalize” algorithm was employed to normalize the data before unsupervised clustering of cells, and dimensionality reduction and t-SNE were employed to visualize the data13. The SingleR package was used to annotate every cluster’s cell type14. The “FindAllMarkers” function in Seurat was employed to screen out differentially expressed genes (DEGs)15, and the criteria were set as follows: absolute log2-fold change (FC) ≥ 1, false discovery rate (FDR) < 0.05, adjusted P value < 0.05 (derived by Bonferroni’s multiple test correction). The cell clusters were annotated after subjective interpretation of marker genes and the outcomes of GO functional enrichment analysis.
GO and KEGG analysis
The clusterProfiler R package was employed to implement GO16 functional enrichment analysis and KEGG17 pathway enrichment analysis. The GO results include multiple biological processes (BPs), cellular components (CCs), and molecular functions (MFs). The threshold for significant enrichment was set as P < 0.05.
Development of a prognostic model based on PPS-OS-related genes
Analysis of gene differential expression and univariate cox regression analysis were implemented to filter genes linked to prognosis from the PPS-OS gene set using a p < 0.05 threshold. We used the LASSO regression algorithm in the “glmnet” R package to select the best genes and prevent overfitting. Simultaneously, the risk score for each OS patient was calculated as follows18:
The symbols exp (x) and coef (x) represent the expression level and gene coefficient of gene X, respectively. The median risk score served as the division criterion, and samples of patients were categorized into low- and high-risk subgroups. The overall survival difference between the two risk groups was assessed by Kaplan–Meier (K‒M) survival analysis with the log-rank test19. A receiver operating characteristic (ROC) curve was employed to evaluate the sensitivity and specificity of the PPS-OS signature20.
Nomogram construction and validation
Univariate and multivariate Cox regression analyses were performed to determine clinical parameters associated with prognosis and to derive their hazard ratio (HR). These parameters included gender, age, tumor site, metastatic status, and risk scores. Based on these clinical parameters, “rms” R software was used to plot the clinical nomogram21. In addition, calibration curves were used to assess how well the predicted and actual survival rates agreed with one another.
Evaluation of the TIME
The stromal score, ESTIMATE score, immune score and tumor purity level of the entire sample were calculated through the “ESTIMATE” algorithm22. The degrees of infiltration of different immune cells in the two risk subgroups were obtained using the ssGSEA algorithm. The immunophenoscore (IPS) of OS patients derived from The Cancer-Immune Group Atlas (TCIA) (https://tcia.at/home) was evaluated using ggpubr R software. Effector cell (EC) score, immunosuppressive cell (SC) score, MHC molecule (MHC) score, and immune checkpoint (CP) score are the four categories that make up the IPS23. The IPS (range 0–10) was calculated using the gene expression in the corresponding cell type, and the score was proportional to the immunogenicity.
Prediction of immunotherapy and chemotherapy
The tumor immune dysfunction and exclusion (TIDE) score and subclass mapping were used to estimate the clinical immune checkpoint effect on CTLA4 and PD-1 to reflect the tumor response to immune checkpoint blockade (ICB). The limma, ggpubr, and pRRophetic packages of R were used to screen potential chemotherapy drugs for OS.
Cell culture and transfection
The human OS cell lines HOS, MG-63, 143B, R-1059D, U-2OS, and SAOS-2 were obtained from the American Type Culture Collection (US). MEM or DMEM containing FBS (10%, Gibco, USA) and penicillin‒streptomycin solution (100 µg/ml, Beijing Solarbio Science & Technology Co., Ltd.) was used as the culture medium. We discovered that research on JTB (jumping translocation breakpoint) in OS is scarce and has not been experimentally validated. Therefore, to target and knockdown JTB, we employed siRNA synthesized by General Biol (Anhui, China). When the cells were cultured to an optimal density in a 6-well plate, the culture medium was replaced, and siRNA was transfected using jetPrime reagent (Poly plus-transfection®). The sequences for the two JTB siRNAs were as follows: JTB-1-forward GGAAGAGUUUGUGGUAGCATT, JTB-1-forward UGCUACCACAAACUCUUCCTT, JTB-2-forward CAGCGACAAUUGGACAGAATT, JTB-2-forward UUCUGUCCAAUUGUCGCUGTT.
RNA extraction and RT-qPCR
Total cellular RNA was isolated using TRIzol reagent (Invitrogen, Thermo Fisher Scientific), followed by detection of the quality and concentration of RNA with a Nanodrop spectrophotometer (IMPLEN GmbH). cDNAs were obtained by reverse transcription. Subsequently, RT-qPCR experiments were carried out with SYBR Green mix (TaKaRa Biotechnology, China). Expression levels of target genes were reported by the 2-ddCt method. The primer sequences were as follows: GAPDH forward: 5ʹ-CGCTCTCTGCTCCTCCTGT-3ʹ; reverse: 5ʹ-ATCCGTTGACTCCGACCTA-3ʹ. JTB forward: 5ʹ-AATAGGCAACTCCGGCCTTC-3ʹ; reverse: 5ʹ-AGAGGGACCTACTCCACAGG-3ʹ.
Cell viability assay
The transfected cells (3 × 103) were seeded in 96-well plates and incubated at 37 °C. After incubation for 24, 48, and 72 h, CCK-8 reagent (Kumamoto, Japan) was injected and incubated again for 4 h at 37 °C. The absorbance at 450 nm was detected by a microplate reader (Thermo Fisher Scientific).
Migration and invasion assay
The assays were conducted in a 24-well plate utilizing a chamber insert with a pore size of 8 μm (3422, Corning, USA). For the migration assays, 2 × 104 cells in serum-free medium were transferred to the upper chamber. The lower chamber was filled with culture media (600 μl) containing 30% FBS.
The invasion assay was performed the same as the migration assay, with a few exceptions: 1 × 105 cells were seeded into the upper chamber, which had been precoated with Matrigel (356234; BD Biocoat). Later, they were all incubated for 24 h (migration assay) or 48 h (invasion assay) at 37 °C and 5% CO2. After incubation, cells were fixed and stained with 4% paraformaldehyde and 0.5% crystal violet. After the cells on the upper surface of the chamber were wiped away, the cells were photographed and counted under an inverted microscope.
Western blotting
RIPA lysis buffer was used to extract total protein from 143B and HOS cells. The BCA Protein Assay Kit and SDS-PAGE were employed for protein content detection and protein separation. The proteins were transferred onto a PVDF membrane (Millipore Corp, USA), and the blots were blocked with TBST with 5% skim milk for 3 h and then incubated with the primary antibody overnight at 4 °C. Afterward, membranes were washed with TBST once per minute for a total of 3 times, and TBST was further diluted to a ratio of 1:10,000, followed by the addition of a second antibody and shaking for 1 h at room temperature. Afterward, immune complexes were detected using ECL reagent. The antibodies used in this experiment included anti-β-actin (Abcam, ab8226), anti-ZEB1 and anti-PCNA (Proteintech, China).
Analysis of data
For all statistical testing and analysis during this investigation, GraphPad Prism 8 and R software were used. Continuous variables were compared using the Wilcoxon test. For correlation analysis, the Spearman correlation test was employed. The findings are presented in terms of the mean ± standard deviation (SD). The t test was employed to analyze intergroup differences. “NS” indicates P > 0.05, “*” indicates P < 0.05, “**” indicates P < 0.01, and “***” indicates P < 0.001. All experiments were repeated three times.
Results
Identification of 7 cell clusters
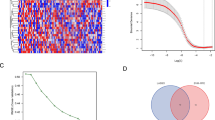
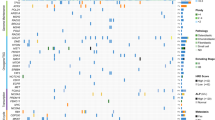
A schematic diagram for the research is displayed in Fig. 1A. After removing batch effects and performing an initial quality control assessment, the scRNA-seq data from public databases were used for further analysis. Unbiased clustering of the cells identified 16 main clusters based on t-SNE analyses (Fig. 1B, Supplementary Table 1). Figure 1C displays the average expression of DEGs of 16 clusters. The bubble plots compare the proportions and relative expression levels of specific markers in 16 clusters (Fig. 1D). The OS cells were extracted according to the marker genes (COL1A1, COL3A1, RUNX2, etc.) of malignant osteoblastic cells24,25 and divided into seven subtypes (Supplementary Table 2). Cluster 0 contained 412 genes; Cluster 1 contained 594 genes; Cluster 2 contained 370 genes; Cluster 3 contained 351 genes; Cluster 4 contained 259 genes; Cluster 5 contained 602 genes; and Cluster 6 contained 381 genes.
Flowchart and the single-cell transcriptomic analysis of OS tissues. (A) Flowchart of this study. (B) Unbiased clustering of the cells identified 16 main clusters in parallel according to their gene profiles and canonical markers. (C) Heatmap showing the DEGs of 16 cell subtypes. (D) The proportion of specific markers in 16 clusters and their scaled relative expression levels. (E) GO analysis of one cluster of PPS-OS. (F) The annotation of seven distinct cell subtypes of OS.
Functional enrichment analysis and cell subtype annotation
The GO functional enrichment analysis revealed that cluster 2-correlated genes were mainly enriched in crucial factors or steps in protein synthesis. Figure 1E shows the biological processes (BPs), including ‘cytoplasmic translation’ (GO:0002181) and ‘ribosomal small subunit assembly’ (GO:0000028). The significantly enriched cellular components (CCs) included ‘cytosolic ribosome’ (GO:0022626), ‘ribosomal subunit’ (GO:0044391), and ‘ribosome’ (GO:0005840). Among the molecular function (MF) terms, the genes were enriched for ‘structural constituent of ribosome’ (GO:0003735) and ‘protein self-association’ (GO:0043621). The results of GO analysis of the remaining subtypes were detailed in the (Supplementary Fig. 1A–F). To facilitate valuable researches on the biological significance of each cell subtype, the biological functions and interactions of GO terms were analyzed, and annotated the subtype function as highly invasive osteosarcoma (HI-OS), homeostatic type osteosarcoma (HST-OS), pro-protein synthesis osteosarcoma (PPS-OS), pro-angiogenic osteosarcoma (PA-OS), immunoreactive osteosarcoma (IR-OS), stress-related osteosarcoma (SR-OS), and extracellular matrix-enriched osteosarcoma (ECM-OS) (Fig. 1F).
Construction of a prognostic model containing six PPS-OS marker genes
The overall survival rates of different risk groups in every cell subtype were shown by K‒M curves (Fig. 2F, Supplementary Fig. 2A–F). The P values representing the significance of the survival differences are 0.0000015 for PPS-OS, 0.000097 for ECM-OS, 0.00014 for SR-OS, 0.00028 for HST-OS, 0.0035 for PA-OS, 0.0057 for HI-OS, 0.067 for IR-OS, respectively. The PPS-OS with the most significant survival differences (minimum P value) between the two risk groups was finally identified as the focus of this study.
Establishment and validation of the risk prognostic mode. (A) The heatmap shows expression levels of 23 PPS-OS related genes in normal and OS tissues. (B) Univariate Cox regression analysis presents the hazard ratios and P-value of 6 PPS-OS related genes. (C) Obtainment of the optimal λ value. (D) The LASSO Cox analysis identified 6 genes apparently affecting the model. (E) The 6 risk signature genes and the corresponding coefficients. (F,I) The survival analysis in the training (F) and testing set (I). (G,J) The distribution of risk score as well as survival time for the two risk subgroups in the training (G) and testing set (J). (H,K) The predictive effificiency of the risk score is presented by the ROC curves in both training (H) and testing set (K).
Subsequently, 23 PPS-OS-related genes that were significantly associated with prognosis were extracted from 370 genes. In the TARGET and GTEx merged cohort, differential expression analysis was applied to compare the DEGs between the tumor and normal sets (Fig. 2A). Afterward, six genes for modeling were identified by univariate cox regression analysis, and the HR and P values of each gene were also calculated (Fig. 2B). Figure 2C,D shows that with the decline of log λ, the corresponding coefficient of the genes likewise diminished to 0, and finally, 6 genes in cross-validation were within the partial likelihood estimation bias minimum value. These genes (CSAG1, RPS27, RPS28, CD320, JTB, and S100A13) are significantly associated with overall survival and are potential prognostic genes. The risk coefficients for these six genes are shown in Fig. 2E.
The median risk value (9.92) was used as the dividing point of the calculated risk score, and 85 patients in TARGET were categorized into low- and high-risk groups. Figure 2F shows a significant difference (P value = 0.0000015) between the two risk groups and a negative correlation between risk scores and patient survival time. The two-dimensional distribution of survival status as well as risk scores for the two risk groups are displayed in Fig. 2G and reflect the poorer survival rate in the high-risk group. Moreover, the areas under the curves (AUCs) for 1, 3, and 5 years were 0.779, 0.847, and 0.881, respectively (Fig. 2H), reflecting that the model has accurate and meaningful predictive capabilities. In the testing set, the survival curve (Fig. 2I), risk score and distribution of patient survival status (Fig. 2J), and AUC values (Fig. 2K) were similar to those in the training set. These results indicated that the prognosis model containing six PPS-OS marker genes can predict disease outcomes in OS patients. Figure 3A–F shows the expression of these six genes in the tumor and normal groups as well as the overall survival differences between the high- and low-risk groups.
Assessment of the independent prognostic value of clinical factors
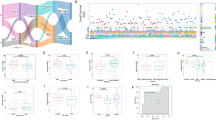
In analyzing the relationship between clinical variable information and prognostic value, we discovered significant differences in survival status, primary tumor site, tumor metastasis status and risk score between risk groups. The high-risk group had a shorter survival time, more diverse primary tumor sites (including the limbs and pelvis), and a higher rate of tumor metastasis (Fig. 4A). Figure 4B shows that the risk score (P < 0.001) and metastasis (P < 0.001) were independent factors affecting the prognosis of patients with OS, and multivariate Cox regression analysis generated comparable results (Fig. 4C). Figure 4D–K shows the relationship between various clinical factors in a more intuitive and accurate manner.
Relationship between the clinical variables in the TARGET dataset. (A) The heatmap displays the differential distribution of clinical variables and 6 PPS-OS related genes expression levels in low- and high-risk. (B,C) Univariate (B) and multivariate (C) cox regression analyses including age, gender, metastatic, primary tunmor site and risk score. (D–K) The various clinical variables were precisely compared one by one between the high- and low-risk groups. (L) A prognostic nomogram for predicting the overall survival of OS patients for the 3 and 5 years. (M) The calibration curves of the nomogram for the 3 and 5 years.
The nomogram we built is a highly reliable mathematical model. As shown in the nomogram, the most common age for OS patients was approximately 15 years old. Another concern was the tumor metastasis status. Distant metastatic tumors were clearly much more likely to cause death than localized tumors. The nomogram also predicted the survival rate of OS patients at 3 and 5 years in a systematic manner (Fig. 4L). Figure 4M shows that the calibration curve is relatively close to the 45° reference line, suggesting that the nomogram-predicted overall survival was consistent with actual overall survival.
Comprehensive analysis of the TIME
Analysis of the training (Fig. 5A–D) and validation sets (Fig. 5E–H) revealed that the high-risk subgroup had a lower immune score, stromal score, and ESTIMATE score and higher tumor purity. Figure 5I–J indicates that the infiltration of B cells and gamma delta T cells (Tgd, γδ T cells) was higher in the high-risk group. Figure 5K–N shows the differences between the four components of the IPS. We found that IPS was higher in both risk groups, and it represents OS as a more immunogenic tumor (Fig. 5O). Together, these results indicated that PPS-OS-related genes have a substantial link with immune infiltration, regulate multiple aspects of tumor immunity, and may play an essential role in the progression of OS.
Analysis of tumor immune microenvironment. (A–H) The score of Stromal, Immune, ESTIMATE, and Tumor Purity between the two risk subgroups in the training (A–D) and testing set (E–H). (I) Heatmap showing the infiltration of 24 immune cells in the high- and low-risk groups. (J) Violin plots showing the difference in immune cell infiltration between the high- and low-risk groups. (K–O) The immunophenoscore analysis of two risk groups.
Prediction for immunotherapy and drug sensitivity analysis
We found several molecules that are significant in tumor initiation and treatment by immune checkpoint analysis, such as lymphocyte-activation gene 3 (LAG3), PTPRC, HAVCR2, B2M, LDHA, and LDHB, which may be used as a reference for tumor immunotherapy (Fig. 6A). According to the analysis, the predicted treatment effects of anti-CTLA4 and anti-PD-1 therapy for the high- and low-risk groups are presented in Fig. 6B,C. Five drugs were demonstrated to be effective in the high-risk group in the drug sensitivity analysis: CGP.082996, elesclomol, pictilisib, MK.2206, and thapsigargin (Fig. 6D–H).
Prediction for immunotherapy and drug sensitivity analysis. (A) Boxplots depict the degree of expression of the 38 immune checkpoint molecules in the two risk subgroups. (B) TIDE value and the effect of immunotherapy of patients with OS. (C) According to submap analysis, the high-risk subgroup may benefit more from anti-PD-1 therapies. (D–H) Estimated IC50 indicated that 5 drugs were sensitive to patients in the high-risk group.
In vitro JTB knockdown experiments
Figure 3A shows that OS tissue had a much higher level of JTB expression than normal tissue. Between the low and high JTB expression groups, there was a notable difference in the overall survival rate. Additionally, no research or reports on OS and JTB were discovered in a search of published literature. Thus, we decided to focus our experiments on PPS-OS-related genes specifically on JTB.
RT-qPCR revealed that HOS and 143B cells had higher JTB expression levels than the other cell lines (Fig. 7A). JTB was then knocked down in 143B and HOS cells using si-JTB-1 and -2, and the results were noticeable (Fig. 7B). The CCK-8 assay showed that the survivability of HOS and 143B cells after JTB knockdown was decreased compared with that of negative control cells (Fig. 7C,D). Then, the cell transwell experiment demonstrated that the number of 143B and HOS cells penetrating the micropore membrane was significantly decreased after silencing JTB, indicating that OS cell capacity for migration and invasion was impaired when JTB was silenced (Fig. 7E,F). Finally, the results of Western blotting demonstrated that si-JTB-1 and si-JTB-2 can effectively decrease JTB protein expression in 143B and HOS cells (Fig. 7I,J).
Discussion
Over the past 30 years, the 5-year survival rate of patients with OS has improved with a combination of radiotherapy, chemotherapy and surgery26. However, metastatic or drugresistant OS continues to pose a challenge27. For refractory OS, new therapeutic targets are urgently needed for clinical application.
Verification of the silencing effect. (A) Relative expression of JTB in six OS cell lines. (B) The expression levels of JTB in HOS and 143B were detected by RT-qPCR after knocking down with siRNA. (C,D) Cellular viability (%) of CCK-8 experiment in HOS (C) cells and 143B (D) cells. (E,F) Representative imaging of migration assays (E) and invasion assays (F) after silencing with JTB-1 and JTB-2 in HOS and 143B cells. (G,H) Cell counts for migration assays (G) and invasion assays (H) after silencing with JTB-1 and JTB-2 in HOS and 143B cells. (I,J) 143B (I) and HOS (J) cells were transfected with si-NC/si-JTB followed by western blot analysis.
Previous evidence has shown that the heterogeneity of tumor cells reflects differences in the biological behavior of tumor cells. To date, studies on cancer, especially studies focused on identifying the interplay between ITH and TME in different subclones, have benefited greatly from scRNA-seq technology28. Therefore, further understanding of the heterogeneity between OS cells and its mechanism of action based on scRNA-seq data may provide clues for developing novel therapies. We conducted a functional enrichment analysis of genes related to PPS-OS in this study, and the results were mainly related to the critical step and molecular machine of protein synthesis. A transgenic mouse model of B-cell Burkitt lymphoma driven by Eµ-Myc revealed a possible role for hyperactive ribosomal biogenesis in the progression of cancer29. Both the increased efficiency of protein synthesis and the decreased fidelity of translation are associated with aberrant ribosome biogenesis, which may lead to tumorigenesis30,31,32. Therefore, the identification of genes associated with PPS-OS may aid in the study of the heterogeneity and underlying mechanisms of OS cells, which may provide a foundation for improving OS diagnosis and treatment33.
First, we assessed scRNA-seq data from public databases and annotated seven cell subtypes. Among them, PPS-OS indicated the worst prognosis of patients. Next, by LASSO regression analysis, we constructed a model based on 6 genes. The ROC analysis displayed the accuracy with which the model predicts prognosis. In the nomogram we constructed, metastasis status had the highest weighted score, followed by the risk score and tumor primary site. Nomograms, which are multivariable regression models that generate individual numerical probabilities of clinical events by integrating several prognostic and determinant variables34, have been widely used in various studies35. The calibration curves demonstrated the nomogram’s prognosis prediction efficiency in a straightforward manner, confirming the accuracy of our model.
The ratio of malignant cells among all cells in a tumor tissue specimen was recognized as the tumor purity, and it was directly correlated with a poor prognosis36. The ESTIMATE algorithm is a novel algorithm relying on gene expression data, and the infiltration of nontumor cells from cancer samples determines the stromal score and immune score. The transcriptional data were then used to determine tumor purity based on immune score and stromal score37. In accordance with our research, lower overall survival, stromal score, immune score, and ESTIMATE score, as well as higher tumor purity, were observed in the high-risk group. This partially explains why patients who were classified into the high-risk group had worse survival rates.
Tumor-infiltrating immune cells (TIICs) make up the majority of the complex mixture of cells in the tumor immune microenvironment (TIME)38. Numerous studies have demonstrated that TIICs play a crucial role in the occurrence, recurrence and metastasis of OS39,40. Understanding TIICs is crucial in treatment optimization and prognosis improvement in patients41. Our research indicates that B cells and gamma delta T cells (Tgd, γδ T cells) were higher in the high-risk group. B cells and γδ T cells are important tumor immune cells in vivo, especially in tumor immunity. They would, however, evolve into immunosuppressive cells to promote carcinogenesis42,43,44. After mitogen activation, γδ T cells can be stimulated by autologous B cells45. Subsequently, γδ T cells are able to influence B-cell function by suppressing the secretion of IgG46. Disruption of the immune balance maintained by these cells leads to inflammation and promotes tumor immune escape47. Studies have also shown that the exosomes released by OS have been found to contain an immunomodulatory substance that targets T cells, which reduces T cell activity and promote the regulatory phenotype T48. Experiments in mouse models have shown that B cells can inhibit the antitumor T cell response to promote tumorigenesis49,50. Therapies that boost the antitumor response mediated by innate immune cells, including T cells, are beneficial for OS patients51. Thus, this immunological profile may contribute to the malignant features of OS.
TIDE is an algorithm for predicting the immune checkpoint blockade (ICB) response in cancer by predicting tumor immune evasion activity based on specific expression signatures (T-cell dysfunction and T-cell exclusion)52. As such, its predictive value for the response to immune checkpoint inhibitors is evident53. Based on the two reliable targets CTLA-4 and PD-1, a variety of targeted agents have been approved for the treatment of different cancers54. The US Food and Drug Administration has authorized pembrolizumab (PD-1) and ipilimumab (CTLA-4) for the treatment of metastatic melanoma55. In recent years, CTLA-4 and PD-1/PD-L1 blockade have exhibited great application potential for OS immunotherapy56,57. The results we obtained imply that low-risk patients identified by our model may respond well to PD-1 and CTLA4 blockade.
For malignant tumors, chemotherapy is a common treatment, but because of its poor bioavailability and lack of targets, its efficacy still confronts severe clinical challenges. Previous studies have suggested that tumor-derived or tumor-associated exosomes are critical in modulating tumor drug resistance58. According to our analysis and prediction, five drugs were obtained: elesclomol, pictilisib, MK.2206, and thapsigargin. Among them, MK-2206, an AKT inhibitor, could suppress the observed decrease in sensitivity to chemotherapeutic agents induced by exosomes59. Based on the results of our study, a combined strategy utilizing immunotherapy and targeted drugs may serve as a new approach for the treatment of OS51.
JTB is a gene located on human chromosome 1q2160. While JTB is widely expressed in normal cells, it has been discovered that cancer cells overexpress it61. Numerous investigations have demonstrated that JTB can promote the proliferation, invasion and metastasis of a variety of cancer cells62. Jayathirtha et al. explored the impact of downregulation of JTB expression in MCF-7 breast cancer cells, laying the foundation for its potential application as a biomarker in breast cancer63. Sanford et al. concluded that JTB in myeloid malignancies was associated with treatment resistance and poor survival64. However, its role in OS is unknown. In our study, the analysis of OS data and in vitro experimental results suggested that JTB knockdown may prevent OS cells from proliferating, migrating, and invading and may affect protein expression. These findings deepen our understanding of the biological basis of OS and indicate that JTB may become a novel therapeutic target for OS in humans.
Conclusion
In this study, the collected single-cell OS data were divided into seven subtypes, and the PPS-OS subtype was further investigated. Based on the six PPS-OS correlated genes, we constructed a model with the ability to predict the prognosis of OS patients. Moreover, the difference in the TIME features of OS patients in different risk groups OS were presented. Corresponding functional experiments demonstrated that the capacity of OS cells to migrate and invade was impaired when JTB was silenced. Although this study achieved ideal positive results, further studies are also warranted. To conclude, our findings may contribute to a strategy for predicting prognosis and facilitate the identification of novel therapeutic targets for OS patients.
Data availability
All data included in this study are available upon request by contact with the corresponding author.
References
Aran, V. et al. Osteosarcoma, chondrosarcoma and Ewing sarcoma: Clinical aspects, biomarker discovery and liquid biopsy. Crit. Rev. Oncol. Hematol. 162, 103340. https://doi.org/10.1016/j.critrevonc.2021.103340 (2021).
Guo, J. et al. Single-cell profiling of tumor microenvironment heterogeneity in osteosarcoma identifies a highly invasive subcluster for predicting prognosis. Front. Oncol. 12, 732862. https://doi.org/10.3389/fonc.2022.732862 (2022).
Ferguson, J. L. & Turner, S. P. Bone cancer: Diagnosis and treatment principles. Am. Fam. Phys. 98, 205–213 (2018).
Isakoff, M. S., Bielack, S. S., Meltzer, P. & Gorlick, R. Osteosarcoma: Current treatment and a collaborative pathway to success. J. Clin. Oncol. 33, 3029–3035. https://doi.org/10.1200/jco.2014.59.4895 (2015).
Jiang, Z. Y., Liu, J. B., Wang, X. F., Ma, Y. S. & Fu, D. Current status and prospects of clinical treatment of osteosarcoma. Technol. Cancer Res. Treat. 21, 15330338221124696. https://doi.org/10.1177/15330338221124696 (2022).
Turajlic, S., Sottoriva, A., Graham, T. & Swanton, C. Resolving genetic heterogeneity in cancer. Nat. Rev. Genet. 20, 404–416. https://doi.org/10.1038/s41576-019-0114-6 (2019).
Gill, J. & Gorlick, R. Advancing therapy for osteosarcoma. Nat. Rev. Clin. Oncol. 18, 609–624. https://doi.org/10.1038/s41571-021-00519-8 (2021).
Feleke, M. et al. Single-cell RNA sequencing reveals differential expression of EGFL7 and VEGF in giant-cell tumor of bone and osteosarcoma. Exp. Biol. Med. 247, 1214–1227. https://doi.org/10.1177/15353702221088238 (2022).
Xu, F., Yan, J., Peng, Z., Liu, J. & Li, Z. Comprehensive analysis of a glycolysis and cholesterol synthesis-related genes signature for predicting prognosis and immune landscape in osteosarcoma. Front. Immunol. 13, 1096009. https://doi.org/10.3389/fimmu.2022.1096009 (2022).
Somarelli, J. A. et al. A comparative oncology drug discovery pipeline to identify and validate new treatments for osteosarcoma. Cancers 12, 335. https://doi.org/10.3390/cancers12113335 (2020).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420. https://doi.org/10.1038/nbt.4096 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902. https://doi.org/10.1016/j.cell.2019.05.031 (2019).
Do, V. H. & Canzar, S. A generalization of t-SNE and UMAP to single-cell multimodal omics. Genome Biol. 22, 130. https://doi.org/10.1186/s13059-021-02356-5 (2021).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172. https://doi.org/10.1038/s41590-018-0276-y (2019).
Gianferante, D. M., Mirabello, L. & Savage, S. A. Germline and somatic genetics of osteosarcoma—Connecting aetiology, biology and therapy. Nat. Rev. Endocrinol. 13, 480–491. https://doi.org/10.1038/nrendo.2017.16 (2017).
Gene Ontology Consortium. Gene Ontology Consortium: Going forward. Nucleic Acids Res. 43, 1049–1056. https://doi.org/10.1093/nar/gku1179 (2015).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353-d361. https://doi.org/10.1093/nar/gkw1092 (2017).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
George, B., Seals, S. & Aban, I. Survival analysis and regression models. J. Nucl. Cardiol. 21, 686–694. https://doi.org/10.1007/s12350-014-9908-2 (2014).
Mandrekar, J. N. Receiver operating characteristic curve in diagnostic test assessment. J. Thorac. Oncol. 5, 1315–1316. https://doi.org/10.1097/JTO.0b013e3181ec173d (2010).
Iasonos, A., Schrag, D., Raj, G. V. & Panageas, K. S. How to build and interpret a nomogram for cancer prognosis. J. Clin. Oncol. 26, 1364–1370. https://doi.org/10.1200/jco.2007.12.9791 (2008).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612. https://doi.org/10.1038/ncomms3612 (2013).
Xu, Q., Chen, S., Hu, Y. & Huang, W. Landscape of immune microenvironment under immune cell infiltration pattern in breast cancer. Front. Immunol. 12, 711433. https://doi.org/10.3389/fimmu.2021.711433 (2021).
Zhou, Y. et al. Single-cell RNA landscape of intratumoral heterogeneity and immunosuppressive microenvironment in advanced osteosarcoma. Nat. Commun. 11, 6322. https://doi.org/10.1038/s41467-020-20059-6 (2020).
Walsh, M. C., Takegahara, N., Kim, H. & Choi, Y. Updating osteoimmunology: Regulation of bone cells by innate and adaptive immunity. Nat. Rev. Rheumatol. 14, 146–156. https://doi.org/10.1038/nrrheum.2017.213 (2018).
Li, S. et al. CircDOCK1 promotes the tumorigenesis and cisplatin resistance of osteogenic sarcoma via the miR-339-3p/IGF1R axis. Mol. Cancer 20, 161. https://doi.org/10.1186/s12943-021-01453-0 (2021).
Kelley, L. M. et al. Pathological fracture and prognosis of high-grade osteosarcoma of the extremities: An analysis of 2847 consecutive cooperative osteosarcoma study group (COSS) patients. J. Clin. Oncol. 38, 823–833. https://doi.org/10.1200/jco.19.00827 (2020).
Wu, R. et al. Identification of cell subpopulations and interactive signaling pathways from a single-cell RNA sequencing dataset in osteosarcoma: A comprehensive bioinformatics analysis. Front. Oncol. 12, 853979. https://doi.org/10.3389/fonc.2022.853979 (2022).
Barna, M. et al. Suppression of Myc oncogenic activity by ribosomal protein haploinsufficiency. Nature 456, 971–975. https://doi.org/10.1038/nature07449 (2008).
Bhat, M. et al. Targeting the translation machinery in cancer. Nat. Rev. Drug Discov. 14, 261–278. https://doi.org/10.1038/nrd4505 (2015).
Harper, J. W. & Bennett, E. J. Proteome complexity and the forces that drive proteome imbalance. Nature 537, 328–338. https://doi.org/10.1038/nature19947 (2016).
Filbeck, S., Cerullo, F., Pfeffer, S. & Joazeiro, C. A. P. Ribosome-associated quality-control mechanisms from bacteria to humans. Mol. Cell 82, 1451–1466. https://doi.org/10.1016/j.molcel.2022.03.038 (2022).
Sun, Y., Zhang, C., Fang, Q., Zhang, W. & Liu, W. Abnormal signal pathways and tumor heterogeneity in osteosarcoma. J. Transl. Med. 21, 99. https://doi.org/10.1186/s12967-023-03961-7 (2023).
Balachandran, V. P., Gonen, M., Smith, J. J. & DeMatteo, R. P. Nomograms in oncology: More than meets the eye. Lancet Oncol. 16, e173–e180. https://doi.org/10.1016/s1470-2045(14)71116-7 (2015).
Harrell, F. E., Lee, K. L. & Mark, D. B. Multivariable prognostic models: Issues in developing models, evaluating assumptions and adequacy, and measuring and reducing errors. Stat. Med. 15, 361–387. https://doi.org/10.1002/(sici)1097-0258(19960229)15:4%3c361::Aid-sim168%3e3.0.Co;2-4 (1996).
Li, T. et al. TIMER: A web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res. 77, e108–e110. https://doi.org/10.1158/0008-5472.Can-17-0307 (2017).
Ren, N., Liang, B. & Li, Y. Identification of prognosis-related genes in the tumor microenvironment of stomach adenocarcinoma by TCGA and GEO datasets. Biosci. Rep. 40, 980. https://doi.org/10.1042/bsr20200980 (2020).
Hanahan, D. & Coussens, L. M. Accessories to the crime: Functions of cells recruited to the tumor microenvironment. Cancer Cell 21, 309–322. https://doi.org/10.1016/j.ccr.2012.02.022 (2012).
Binnewies, M. et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med. 24, 541–550. https://doi.org/10.1038/s41591-018-0014-x (2018).
Wu, T. & Dai, Y. Tumor microenvironment and therapeutic response. Cancer Lett. 387, 61–68. https://doi.org/10.1016/j.canlet.2016.01.043 (2017).
Pratt, H. G., Justin, E. M. & Lindsey, B. A. Applying osteosarcoma immunology to understand disease progression and assess immunotherapeutic response. Adv. Exp. Med. Biol. 1258, 91–109. https://doi.org/10.1007/978-3-030-43085-6_6 (2020).
Meng, X. et al. Exploiting Ca(2+) signaling in T cells to advance cancer immunotherapy. Semin. Immunol. 49, 101434. https://doi.org/10.1016/j.smim.2020.101434 (2020).
Lei, K., Kurum, A. & Tang, L. Mechanical immunoengineering of T cells for therapeutic applications. Acc. Chem. Res. 53, 2777–2790. https://doi.org/10.1021/acs.accounts.0c00486 (2020).
Bruno, T. C. et al. Antigen-presenting intratumoral B cells affect CD4(+) TIL phenotypes in non-small cell lung cancer patients. Cancer Immunol. Res. 5, 898–907. https://doi.org/10.1158/2326-6066.Cir-17-0075 (2017).
Horner, A. A., Jabara, H., Ramesh, N. & Geha, R. S. gamma/delta T lymphocytes express CD40 ligand and induce isotype switching in B lymphocytes. J. Exp. Med. 181, 1239–1244. https://doi.org/10.1084/jem.181.3.1239 (1995).
Li, Z. Potential of human γδ T cells for immunotherapy of osteosarcoma. Mol. Biol. Rep. 40, 427–437. https://doi.org/10.1007/s11033-012-2077-y (2013).
Le Menn, G., Jabłońska, A. & Chen, Z. The effects of post-translational modifications on Th17/Treg cell differentiation. Biochim. Biophys. Acta. Mol. Cell Res. 1869, 119223. https://doi.org/10.1016/j.bbamcr.2022.119223 (2022).
Li, S. The basic characteristics of extracellular vesicles and their potential application in bone sarcomas. J. Nanobiotechnol. 19, 277. https://doi.org/10.1186/s12951-021-01028-7 (2021).
Brodt, P. & Gordon, J. Anti-tumor immunity in B lymphocyte-deprived mice. I. Immunity to a chemically induced tumor. J. Immunol. 121, 359–362 (1978).
Qin, Z. et al. B cells inhibit induction of T cell-dependent tumor immunity. Nat. Med. 4, 627–630. https://doi.org/10.1038/nm0598-627 (1998).
Chen, C. et al. Immunotherapy for osteosarcoma: Fundamental mechanism, rationale, and recent breakthroughs. Cancer Lett. 500, 1–10. https://doi.org/10.1016/j.canlet.2020.12.024 (2021).
Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24, 1550–1558. https://doi.org/10.1038/s41591-018-0136-1 (2018).
Zheng, Y. et al. A novel defined endoplasmic reticulum stress-related lncRNA signature for prognosis prediction and immune therapy in glioma. Front. Oncol. 12, 930923. https://doi.org/10.3389/fonc.2022.930923 (2022).
Rotte, A. Combination of CTLA-4 and PD-1 blockers for treatment of cancer. J. Exp. Clin. Cancer Res. 38, 255. https://doi.org/10.1186/s13046-019-1259-z (2019).
Carlino, M. S., Larkin, J. & Long, G. V. Immune checkpoint inhibitors in melanoma. Lancet 398, 1002–1014. https://doi.org/10.1016/s0140-6736(21)01206-x (2021).
Wang, S. D. et al. The role of CTLA-4 and PD-1 in anti-tumor immune response and their potential efficacy against osteosarcoma. Int. Immunopharmacol. 38, 81–89. https://doi.org/10.1016/j.intimp.2016.05.016 (2016).
Sun, C. & Li, S. PTHR1 in osteosarcoma: Specific molecular mechanisms and comprehensive functional perspective. J. Cell. Mol. Med. 25, 3175–3181. https://doi.org/10.1111/jcmm.16420 (2021).
Guo, X., Gao, C., Yang, D. H. & Li, S. Exosomal circular RNAs: A chief culprit in cancer chemotherapy resistance. Drug Resist. Updates Rev. Comment. Antimicrob. Anticancer Chemother. 67, 100937. https://doi.org/10.1016/j.drup.2023.100937 (2023).
Tomita, R., Sasabe, E., Tomomura, A. & Yamamoto, T. Macrophage-derived exosomes attenuate the susceptibility of oral squamous cell carcinoma cells to chemotherapeutic drugs through the AKT/GSK-3β pathway. Oncol. Rep. 44, 1905–1916. https://doi.org/10.3892/or.2020.7748 (2020).
Kanome, T. et al. Characterization of jumping translocation breakpoint (JTB) gene product isolated as a TGF-beta1-inducible clone involved in regulation of mitochondrial function, cell growth and cell death. Oncogene 26, 5991–6001. https://doi.org/10.1038/sj.onc.1210423 (2007).
Hatakeyama, S., Osawa, M., Omine, M. & Ishikawa, F. JTB: a novel membrane protein gene at 1q21 rearranged in a jumping translocation. Oncogene 18, 2085–2090. https://doi.org/10.1038/sj.onc.1202510 (1999).
Jayathirtha, M. et al. Investigating the function of human jumping translocation breakpoint protein (hJTB) and its interacting partners through in-solution proteomics of MCF7 cells. Molecules 27, 301. https://doi.org/10.3390/molecules27238301 (2022).
Jayathirtha, M., Neagu, A. N., Whitham, D., Alwine, S. & Darie, C. C. Investigation of the effects of downregulation of jumping translocation breakpoint (JTB) protein expression in MCF7 cells for potential use as a biomarker in breast cancer. Am. J. Cancer Res. 12, 4373–4398 (2022).
Sanford, D. et al. Jumping translocations in myeloid malignancies associated with treatment resistance and poor survival. Clin. Lymphoma Myeloma Leukemia 15, 556–562. https://doi.org/10.1016/j.clml.2015.05.005 (2015).
Acknowledgements
The author thanks TARGET network, GEO network and TCIA database for their contributions.
Funding
This research was funded by Translational Medicine Research Foundation of the Second Affiliated Hospital of Anhui Medical University (2022ZHYJ13); Key Projects of Natural Science Research in Colleges and Universities in Anhui Province (2022AH040102); Research Foundation of Anhui Institute of Translational Medicine (No.2022zhyx-C49); Anhui Medical University Graduate Research and Practice Innovation Project (YJS20230197).
Author information
Authors and Affiliations
Contributions
D.T. and E.B. designed this study, guided the data analysis, funding acquisition and revised the manuscript. C.Y., J.L. and B.X. collected and processed data, performed the experiment, and drafted, edited and revised the manuscript. C.Y. and S.Z. deposited the data and constructed the database. D.G. and A.L. performed parts of the data analysis and experiment. All authors reviewed the manuscript and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yi, C., Liu, J., Zhao, S. et al. Identification of a pro-protein synthesis osteosarcoma subtype for predicting prognosis and treatment. Sci Rep 14, 16475 (2024). https://doi.org/10.1038/s41598-024-67547-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-67547-z
- Springer Nature Limited