Abstract
The COVID-19 pandemic has altered the infection landscape for many pathogens. This retrospective study aimed to compare Haemophilus influenzae (H. influenzae) infections in pediatric CAP patients hospitalized before (2018–2019) and during (2020–2022) the COVID-19 pandemic. We analyzed the clinical epidemiology and antimicrobial resistance (AMR) patterns of H. influenzae from a tertiary hospital in southwest China. A total of 986 pediatric CAP patients with H. influenzae-associated infections were included. Compared to 2018, the positivity rate increased in 2019 but dropped significantly in 2020. Although it rose in the following 2 years, the rate in 2022 remained significantly lower than in 2019. Patients’ age during the pandemic was significantly higher than in 2018 and 2019, while gender composition remained similar across both periods. Notably, there were significant changes in co-infections with several respiratory pathogens during the pandemic. Resistance rates of H. influenzae isolates to antibiotics varied, with the highest resistance observed for ampicillin (85.9%) and the lowest for cefotaxime (0.0%). Resistance profiles to various antibiotics underwent dramatic changes during the COVID-19 pandemic. Resistance to amoxicillin-clavulanate, cefaclor, cefuroxime, trimethoprim-sulfamethoxazole, and the proportion of multi-drug resistant (MDR) isolates significantly decreased. Additionally, MDR isolates, alongside isolates resistant to specific drugs, were notably prevalent in ampicillin-resistant and β-lactamase-positive isolates. The number of pediatric CAP patients, H. influenzae infections, and isolates resistant to certain antibiotics exhibited seasonal patterns, peaking in the winter of 2018 and 2019. During the COVID-19 pandemic, sharp decreases were observed in February 2020, and there was no resurgence in December 2022. These findings indicate that the COVID-19 pandemic has significantly altered the infection spectrum of H. influenzae in pediatric CAP patients, as evidenced by shifts in positivity rate, demographic characteristics, respiratory co-infections, AMR patterns, and seasonal trends.
Similar content being viewed by others
Introduction
Globally, community acquired pneumonia (CAP) remains a profound public health challenge, being a primary cause for hospitalizations and constituting a significant fraction of antimicrobial prescriptions1. Children under 5 years old and adults over 65 years old are particularly susceptible2. Hospitalized CAP patients, especially those in intensive care settings, exhibit high mortality rate3,4,5. Pathogens with antimicrobial resistance (AMR) pose a heightened threat, escalating the risk of mortality in children infected with multi-drug resistant (MDR) isolates6,7. Yet, pediatric CAP treatments often hinge on empirical methods, as timely etiological identification and drug susceptibility testing aren't always feasible.
Haemophilus influenzae (H. influenzae) frequently instigates CAP in pediatric populations8. An understanding of local resistance trends is paramount for effective empiric therapy. As such, monitoring the bacterial epidemiology and AMR patterns of H. influenzae remains a priority. With the advent of H. influenzae type b (Hib) vaccination, incidences of H. influenzae infections, especially invasive diseases caused by Hib, have seen considerable reductions in numerous countries9. Nevertheless, H. influenzae continues to place a significant isolate on healthcare systems. For a long duration, ampicillin stood as the preferred drug against H. influenzae infections. However, there's a growing prevalence of ampicillin-resistant isolates10,11,12. For instance, in China, the resistance rate for H. influenzae isolates to ampicillin surged from 19.9% in 2009–2011 to 32.4% in 2013–201413. Alarmingly, child-isolated H. influenzae isolates now exhibit resistance to advanced antibiotics like fluoroquinolones, carbapenems, and third-generation cephalosporins14. This escalating resistance complicates the management of pediatric CAP triggered by H. influenzae.
COVID-19 was initially reported in December 2019 in Wuhan city, a densely populated area of Hubei province, China. The Chinese government classified COVID-19 as a Class B infectious disease and implemented preventive and containment measures typically reserved for Class A infectious diseases on January 20, 202015. A range of non-pharmaceutical interventions, including cordon sanitaire, social distancing, universal symptom surveys, quarantine strategies, and transport restrictions, were then enforced nationwide16. These measures proved effective in mitigating the spread of COVID-1917. Nonetheless, the epidemiological landscape of numerous pathogens experienced significant transformations during the COVID-19 pandemic. Notably, the seasonal distributions of H. influenzae infection changed in this period18,19. Meanwhile, the incidence of invasive bacterial diseases caused by Streptococcus pneumoniae (S. pneumoniae), H. influenzae, and Neisseria meningitidis significantly dropped20,21. Significant reduction was also shown in H. influenzae isolates from the respiratory tract of children in 2020 and 202122,23. However, comprehensive studies exploring the AMR patterns of H. influenzae from pediatric CAP patients in the context of the COVID-19 pandemic are sparse24. Nearly no study has reported the epidemiology and AMR of H. influenzae in 2022 (the third year of COVID-19 pandemic).
In light of this, our study endeavors to gauge the repercussions of the COVID-19 pandemic on the clinical epidemiology and AMR patterns of H. influenzae in pediatric CAP patients. Recognizing the public health significance of H. influenzae infections in children, this study encompasses data from 2018 to 2022, providing insights into AMR patterns concerning ten antibiotics.
Materials and methods
Study population
A retrospective analysis was conducted at Yongchuan Hospital of Chongqing Medical University, a tertiary hospital in western Chongqing, covering the period from January 2018 to December 2022. Pediatric patients diagnosed with CAP were identified using a comprehensive approach involving clinical symptoms, chest imaging, and laboratory tests. Expert radiologists assessed the chest imaging to identify pneumonia-related findings such as infiltrates, consolidation, and other relevant abnormalities. The study encompassed patients aged 1 month to 18 years. Exclusion criteria comprised: (1) Pneumonia onset ≥ 48 h post-admission. (2) Chest radiography revealing interstitial infiltrate, alveolar infiltrate, lobar pneumonia, or pleural effusion > 72 h post-admission. (3) Lung infiltrate or interstitial changes suggestive of pulmonary tuberculosis, pulmonary edema, or atelectasis. (4) Patients with incomplete medical records or absent bacterial culture results. All methods were carried out in accordance with relevant guidelines and regulations. All experimental protocols were approved by the ethics committee of the Yongchuan Hospital of Chongqing Medical University (No. 2023-KeLunShen-76). As a retrospective study, the need for informed consent was waived by the ethics committee of the Yongchuan Hospital of Chongqing Medical University.
Identification and antimicrobial susceptibility testing of H. influenzae isolates
Sputum samples were collected upon admission and submitted for microbiological testing following established clinical protocols. For patients unable to expectorate sputum, samples were obtained either from the nasopharynx or via deep suction under negative pressure. Adequate sputum quality was defined as containing ≥ 25 leukocytes and ≤ 10 epithelial cells under low magnification. The samples underwent cultivation on MacConkey, blood, and chocolate agar plates, followed by an incubation for 18–24 h at 37℃ in a 5% CO2 environment. H. influenzae was identified using the Vitek-2 Compact automatic system (BioMérieux, France). The minimum inhibitory concentrations (MICs) of H. influenzae isolates were determined using ATB identification cards. The tested antibiotics included ampicillin, amoxicillin and clavulanate, cefaclor, cefuroxime, cefotaxime, rifampicin, ofloxacin, tetracycline, chloramphenicol, and trimethoprim-sulfamethoxazole. All tests aligned with the guidelines set by the Clinical and Laboratory Standards Institute (CLSI), which also provided the criteria for susceptibility classification. The nitrocefin-based method detected H. influenzae's β-lactamase, with a color shift to red indicating a positive result. This study exclusively included unique H. influenzae isolates, excluding any repeated isolates from the same patient during the same hospitalization episode. H. influenzae ATCC49247 served as the quality-control strain. MDR isolate was defined as the resistance to three or more antibiotic classes.
Identification of respiratory co-infections
Our study investigated respiratory co-infections involving bacteria, Mycoplasma pneumoniae (M. pneumoniae), and viruses among children with H. influenzae-associated CAP. We assessed bacterial co-infections for three specific isolates: S. pneumoniae, Staphylococcus aureus (S. aureus), and Moraxella catarrhalis (M. catarrhalis). Co-infections were confirmed when sputum specimens tested positive for both H. influenzae and another bacterial species. To detect M. pneumoniae co-infections, venous blood samples were collected to isolate serum. The presence of M. pneumoniae was determined by detecting Immunoglobulin M (IgM) antibodies in the serum, utilizing either an indirect immunofluorescence assay (IFA) or a passive particle agglutination test (Fujirebio, Japan), following the manufacturer’s instructions. For the passive agglutination test, an antibody titer of ≥ 1:160 was considered indicative of a M. pneumoniae infection.
Viral co-infections were detected by collecting venous blood or nasopharyngeal swab samples from patients upon admission. Our viral testing panel comprised five primary respiratory viruses: influenza virus A (IVA), influenza virus B (IVB), parainfluenza virus (PIV), respiratory syncytial virus (RSV), and adenovirus (ADV). Serum IgM antibodies targeting these viruses were quantified using IFA for venous blood samples. Simultaneously, nasopharyngeal swab samples underwent analysis using a multiplex direct immunofluorescence assay kit (Diagnostic Hybrids, Athens, Ohio, USA), following established protocols. Identification of viral co-infections relied on positive findings from either serum or nasopharyngeal swab samples.
Statistical analysis
The normality of quantitative data was assessed using the Kolmogorov–Smirnov test. Data conforming to a normal distribution were expressed as mean ± standard deviation (SD), and group comparisons were conducted using Student’s t-test. Non-normally distributed data were represented as medians and interquartile ranges, and group comparisons were carried out using the Mann–Whitney U test. According to the actual frequency and theoretical frequency, categorical variables were compared by two-tailed chi-square test, Fisher's exact test, or Yates’ continuity corrected chi-square test. Co-infections were analyzed after excluding patients without corresponding pathogenic results. All statistical evaluations were conducted utilizing the GraphPad Prism 9.0 Software (GraphPad Software, Inc., San Diego, CA, USA). P < 0.05 was considered to be statistically significant.
Ethical approval
All procedures performed in these studies involving human participants were approved by the ethics review committee of the Yongchuan Hospital of Chongqing Medical University, Chongqing, China (No. 2023-KeLunShen-76).
Consent to participate
As a retrospective study, the need for informed consent was waived.
Consent for publication
As a retrospective study, the need for informed consent was waived.
Results
Impact of the COVID-19 pandemic on the positivity rates, demographic characteristics, and respiratory co-infections of H. influenzae-associated CAP patients
This retrospective study involved 6115 pediatric patients hospitalized with CAP from January 2018 to December 2022. Among the 5941 non-duplicate respiratory specimens sent for bacterial culture upon admission, 986 H. influenzae isolates were isolated from sputum samples for the AMR data analysis. Despite a significant increase in 2019, the positivity rate of H. influenzae notably decreased in 2020 compared to 2018 and 2019. The rate rebounded in the subsequent 2 years, but the level in 2022 was still significantly lower than that in 2019. The H. influenzae-associated CAP patients had a median age of 14.5 months (interquartile range: 6–36 months) and included 58.8% males. The distribution of H. influenzae infections across various age groups was as follows: 41.6% (410 cases) were children under 1 year, 33.0% (325 cases) were children aged 1 to less than 3 years, 22.8% (225 cases) were children aged 3 to less than 6 years, 2.5% (25 cases) were children aged 6 to less than 12 years, and 0.1% (1 case) were children aged 12 years and older. During the COVID-19 pandemic from 2020 to 2022, the median age was notably higher than that in 2018 and 2019, but no significant changes were observed in the gender composition, as shown in Table 1.
The predominant admission symptoms among patients were cough (98.1%) and fever (45.2%), with wheezing present in 26.6% of cases. In bacterial co-infections, the rate of H. influenzae-associated CAP patients co-infected with S. pneumoniae, S. aureus, and M. catarrhalis was 12.2%, 3.0%, and 5.2%, respectively. Comparing to pre-pandemic levels, significant changes were observed in 2021, with more S. pneumoniae and M. catarrhalis co-infections, but fewer S. aureus co-infections (P < 0.05). Concerning other pathogens, 31.1% of patients were co-infected with M. pneumoniae, and 158 patients had viral co-infections, with RSV being the predominant virus. In comparison to 2018 and 2019, M. pneumoniae co-infections notably decreased in 2022. Additionally, the rate of IVA co-infections in 2021 was significantly lower than that in 2019 (P < 0.05). However, the co-infections of other viruses did not significantly change during the COVID-19 pandemic, as detailed in Table 1.
Impact of the COVID-19 pandemic on the antibiotic resistance rates of H. influenzae isolates from pediatric CAP patients
As shown in Table 2, H. influenzae isolates exhibited relatively high resistance rates to ampicillin, trimethoprim-sulfamethoxazole, cefaclor, and cefuroxime. However, all isolates displayed sensitivity to cefotaxime. MDR isolates accounted for 38.4% of the cases. The most prevalent MDR patterns were resistance to aminopenicillins/ cephalosporins/ sulfonamides/ aminopenicillin with β-lactamase inhibitor (n = 136/379, 35.9%) and aminopenicillins/ cephalosporins/ sulfonamides (n = 135/379, 35.6%). In comparison to 2018, there was a significant decrease in the resistance rate to amoxicillin-clavulanate, cefaclor, cefuroxime, trimethoprim-sulfamethoxazole, and the proportion of MDR isolates in 2019. Notable changes were also observed during the COVID-19 pandemic. Specifically, the resistance rates to amoxicillin-clavulanate, cefaclor, and cefuroxime in 2020 were lower than pre-pandemic levels. Furthermore, the resistance rate to trimethoprim-sulfamethoxazole and the proportion of MDR isolates decreased steadily from 2020 to 2022 (P < 0.05). However, the resistance rates to other antibiotics showed no significant changes during this period.
Comparison of antibiotic resistance between ampicillin-resistant and sensitive isolates, and β-lactamase positive and negative isolates
Considering the high resistance of H. influenzae to ampicillin, a comparative AMR analysis between ampicillin-resistant and ampicillin-sensitive isolates was essential. The resistance rates to amoxicillin-clavulanate, cefaclor, cefuroxime, tetracycline, chloramphenicol, trimethoprim-sulfamethoxazole, and the proportion of MDR isolates were significantly higher in ampicillin-resistant group (P < 0.05). The resistance rates to other antibiotics were comparable between two groups (Table 3).
Remarkably, 83.5% (823/986) of the H. influenzae isolates were positive for β-lactamase. The resistance rates to ampicillin, cefaclor, cefuroxime, tetracycline, chloramphenicol, trimethoprim-sulfamethoxazole, and the proportion of MDR isolates were significantly higher among β-lactamase-positive isolates when compared to β-lactamase-negative isolates (P < 0.05) (Table 4).
Impact of the COVID-19 pandemic on the seasonal patterns of H. influenzae infections and drug resistant isolates
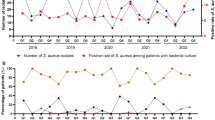
The number of pediatric CAP patients and H. influenzae isolates exhibited seasonal distribution patterns in 2018 and 2019, with increases observed during winter and peaks occurring in December or January. However, the seasonal distributions, particularly those of isolates, underwent shifts during the COVID-19 pandemic. In 2020, there was a significant decrease in February, with isolates absent in March and September, maintaining low levels between April and August. In 2021, H. influenzae infections rebounded in April, reaching the highest level for that year. In 2022, isolates were absent in September and then remained at low levels without resurgence (Fig. 1A).
Seasonal trends were also observed in the prevalence of H. influenzae isolates resistant to ampicillin, amoxicillin-clavulanate, cefaclor, cefuroxime, and trimethoprim-sulfamethoxazole. Generally, resistance increased during winter months, peaking in December or January, but exhibited lower levels from August to October in both 2018 and 2019. However, these seasonal patterns shifted during the COVID-19 pandemic. Notably, there was a sharp decrease in resistant isolates in February 2020. Moreover, resistant isolates were either absent or maintained at low levels between March and September in 2020, as well as from September to December in 2022. Due to the low resistance rate, the seasonal distribution of H. influenzae isolates resistant to other antibiotics remained inconspicuous throughout the five-year period (Fig. 1B).
Discussion
CAP persists as a dominant contributor to pediatric pneumonia morbidity25. The prompt initiation of effective antibiotics remains pivotal in treating pediatric CAP. Continual surveillance of pathogen epidemiology and AMR trends is paramount for informed clinical decision-making, especially with H. influenzae being a frequent CAP culprit in children26. Our study revealed an 85.9% resistance rate for H. influenzae to ampicillin, considerably higher than prior studies14,26. The primary mechanism of resistance to ampicillin involves the production of β-lactamase, which hydrolyzes β-lactam antibiotics, rendering them ineffective against bacterial cell wall synthesis27. Another mechanism is the mutation of penicillin binding protein 3 (PBP3), which decreases the susceptibility of H. influenzae to ampicillin and other β-lactam antibiotics28. The β-lactam resistance phenotype mediated by mutation in PBP3 is named β-lactamase-negative ampicillin resistant (BLNAR)10,26,29. In our cohort, 83.5% of the H. influenzae isolates were β-lactamase positive. But there were 25 BLNAR isolates, revealing a higher positive rate of β-lactamase but lower proportion of BLNAR than previous studies26,30. Every BLNAR isolate was resistant to amoxicillin-clavulanate, contrasting with the resistance rate of 20.0% in β-lactamase-positive ampicillin resistant (BLPAR) isolates. This aligns with expectations, given that BLNAR isolates are typically resistant to amoxicillin-clavulanate and ampicillin-sulbactam26. While BLPAR isolates are usually sensitive to these β-lactamase-inhibiting compounds11,13,26.
Consistent with prior studies14,26,30,31, our study noted minimal resistance to rifamycins, fluoroquinolones, and third-generation cephalosporins. H. influenzae isolates resistant to fluoroquinolones are generally uncommon in children and have been mainly isolated from older people with chronic lung diseases exposed to frequent quinolone treatments32,33,34. BLNAR isolates with PBP3 mutations or BLPAR isolates may have elevated MIC values of third generation cephalosporins, but the level did not transgress the resistance breakpoint in most isolates29,35,36. While rifamycins and fluoroquinolones are infrequently administered to pediatric CAP patients, third-generation cephalosporins emerge as potential treatments for MDR isolates. Nonetheless, vigilant monitoring remains crucial because isolates resistant to third generation cephalosporins have been reported37,38.
MDR H. influenzae isolates, with their enhanced AMR profiles, pose a growing challenge to pediatric healthcare. In our study, 38.4% of isolates were identified as MDR. The proportion of MDR isolates, and the resistance rates to cefaclor, cefuroxime, tetracycline, chloramphenicol, and trimethoprim-sulfamethoxazole were significantly higher in ampicillin-resistant and β-lactamase-positive isolates. Additionally, we observed prevalence of amoxicillin-clavulanate resistance in ampicillin-resistant isolates and ampicillin resistance in β-lactamase-positive isolates. Ampicillin-resistant isolates from another population has demonstrated same trend of cefuroxime resistance39. Our analysis of AMR patterns between β-lactamase-positive and negative isolates parallels the findings of a multicenter study conducted in China30. However, disparities in sample sources and study durations have led to divergent results in other populations. For instance, while one study observed a significant prevalence of cefotaxime resistance in β-lactamase-positive isolates14, another study reported substantially lower resistance to amoxicillin-clavulanate in this group40.
Worldwide, the epidemiological landscape of numerous pathogens underwent discernible shifts due to the COVID-19 pandemic. As compared to pre-pandemic figures, there's a notable decline in invasive infections by S. pneumoniae, H. influenzae, and Neisseria meningitidis in 2020 and 202120,21,41,42,43,44,45,46. Our study analyzed respiratory infections caused by H. influenzae in pediatric CAP patients and revealed dramatic changes during the pandemic. Despite a notable increase in 2019, the positivity rate sharply decreased in 2020, dropping significantly below the levels of 2018 and 2019. Additionally, the rate in 2022 was significantly lower than that in 2019. Similarly, decreased H. influenzae infections during the COVID-19 pandemic have also been reported in other pediatric populations22,23. In our study, H. influenzae-associated CAP patients admitted between 2020 and 2022 were notably older compared to the pre-pandemic period. However, the proportion of males did not significantly change. Similar trends in the ages of children infected with H. influenzae during the COVID-19 pandemic have been reported23. By contrast, no changes in the distribution of age for most pathogens were observed in another study47. Respiratory co-infections can complicate effective treatment and impact prognosis. It has been shown that co-infection of H. influenzae and RSV is significantly associated with increased severity of CAP in children and adolescents48. In our study, M. pneumoniae was the most common pathogen in H. influenzae co-infections, affecting 31.1% of CAP patients. Additionally, S. pneumoniae and RSV emerged as the most prevalent bacterial and viral co-infections, respectively. This highlights the distinct co-infection patterns of H. influenzae among children in the community setting. During the COVID-19 pandemic, decreased viral and bacterial coinfections have been demonstrated in patients with acute respiratory infections49. In the H. influenzae-associated CAP patients of our study, the co-infections of S. aureus, M. pneumoniae, and IVA notably decreased. In contrast, we observed significant increases in S. pneumoniae and M. catarrhalis co-infections, revealing the differential effects of the COVID-19 pandemic on various pathogens.
Notably, the COVID-19 pandemic has also influenced the AMR patterns of H. influenzae in our study. A marked decrease in the proportion of MDR isolates and resistance rates to amoxicillin-clavulanate, cefaclor, cefuroxime, and trimethoprim-sulfamethoxazole was observed during this period. Similar trajectories of resistance to cefaclor, cefuroxime, and trimethoprim-sulfamethoxazole have been documented elsewhere24. Seasonal fluctuations in H. influenzae cases are not uncommon, often peaking during certain months24,50,51,52. Pre-pandemic of COVID-19, our findings pointed to pronounced seasonal distributions for H. influenzae infections and resistant isolates, with winter witnessing more cases. However, in line with the findings of two other investigations18,19, the seasonal pattern of H. influenzae in our study was disrupted following the onset of the COVID-19 pandemic. Few cases of H. influenzae-associated CAP and resistant isolates were recorded from April to August in 2020, and from November to December in 2022. Moreover, we observed a complete absence of H. influenzae isolates in March and September of 2020, as well as September of 2022. The implementation of non-pharmaceutical interventions against the COVID-19 pandemic likely impeded the respiratory transmission of H. influenzae among children.
This study has few limitations. Firstly, the decline in Hib infections due to Hib vaccine proliferation means that non-typeable unencapsulated isolate infections are on the rise53,54. These unencapsulated isolates exhibit higher β-lactamase positivity than encapsulated isolates10. Unfortunately, our study lacked data to assess changes in Hib vaccination status and H. influenzae serotype during the COVID-19 pandemic. Secondly, the absence of detailed clinical data concerning disease severity, treatment modalities, and patient responses limits our ability to conduct a thorough analysis of disease dynamics. Thirdly, drawing samples from a large tertiary hospital might have inadvertently introduced survivorship bias, potentially exaggerating community resistance rates of H. influenza. Fourthly, given its single-center and retrospective design, the findings might not be wholly reflective of broader regional patterns.
Conclusions
In conclusion, this study compared the clinical epidemiology and AMR patterns of H. influenzae in pediatric CAP patients during the periods before (2018–2019) and during (2020–2022) the COVID-19 pandemic. The positivity rate for H. influenzae notably surged in 2019 but then significantly declined in 2020. Although there was a resurgence in the following two years, the rate in 2022 was significantly lower than that in 2019. In contrast to the significant increases in age during the COVID-19 pandemic, the proportion of male patients remained comparable between the two periods. Significant changes were also observed in the prevalence of co-infections with certain other pathogens, the resistance rates to some antibiotics, and the proportion of MDR isolates. Additionally, the typical seasonal patterns in the number of pediatric CAP patients, H. influenzae-infected cases, and resistant isolates shifted to varying degrees. Therefore, a multicenter study involving more participants is essential for ongoing surveillance of H. influenzae infections in the post-COVID-19 era.
Data availability
The data set used in the study is available from the corresponding author on reasonable request.
References
Ho, J. & Ip, M. Antibiotic-resistant community-acquired bacterial pneumonia. Infect. Dis. Clin. North Am. 33, 1087–1103. https://doi.org/10.1016/j.idc.2019.07.002 (2019).
Prina, E., Ranzani, O. T. & Torres, A. Community-acquired pneumonia. Lancet (London, England) 386, 1097–1108. https://doi.org/10.1016/s0140-6736(15)60733-4 (2015).
Ewig, S. et al. New perspectives on community-acquired pneumonia in 388 406 patients. Results from a nationwide mandatory performance measurement programme in healthcare quality. Thorax 64, 1062–1069. https://doi.org/10.1136/thx.2008.109785 (2009).
Woodhead, M., Welch, C. A., Harrison, D. A., Bellingan, G. & Ayres, J. G. Community-acquired pneumonia on the intensive care unit: secondary analysis of 17,869 cases in the ICNARC case mix programme database. Critical care (London, England) 10(Suppl 2), S1. https://doi.org/10.1186/cc4927 (2006).
Arnold, F. W., Wiemken, T. L., Peyrani, P., Ramirez, J. A. & Brock, G. N. Mortality differences among hospitalized patients with community-acquired pneumonia in three world regions: results from the community-acquired pneumonia organization (CAPO) international cohort study. Respir. Med. 107, 1101–1111. https://doi.org/10.1016/j.rmed.2013.04.003 (2013).
Folgori, L. et al. Epidemiology and clinical outcomes of multidrug-resistant, gram-negative bloodstream infections in a European tertiary pediatric hospital during a 12-month period. Pediatr. Infect. Dis. J. 33, 929–932. https://doi.org/10.1097/inf.0000000000000339 (2014).
Huemer, M., Mairpady Shambat, S., Brugger, S. D. & Zinkernagel, A. S. Antibiotic resistance and persistence-Implications for human health and treatment perspectives. EMBO Rep. 21, e51034. https://doi.org/10.15252/embr.202051034 (2020).
Butler, D. F. & Myers, A. L. Changing epidemiology of haemophilus influenzae in children. Infect. Dis. Clin. North Am. 32, 119–128. https://doi.org/10.1016/j.idc.2017.10.005 (2018).
Slack, M. P. E., Cripps, A. W., Grimwood, K., Mackenzie, G. A. & Ulanova, M. Invasive haemophilus influenzae infections after 3 decades of Hib protein conjugate vaccine use. Clin. Microbiol. Rev. 34, e0002821. https://doi.org/10.1128/cmr.00028-21 (2021).
Tsang, R. S. W. et al. Antibiotic susceptibility and molecular analysis of invasive haemophilus influenzae in Canada, 2007 to 2014. J. Antimicrob. Chemother. 72, 1314–1319. https://doi.org/10.1093/jac/dkw565 (2017).
Shiro, H., Sato, Y., Toyonaga, Y., Hanaki, H. & Sunakawa, K. Nationwide survey of the development of drug resistance in the pediatric field in 2000-2001, 2004, 2007, 2010 and 2012: evaluation of the changes in drug sensitivity of Haemophilus influenzae and patients background factors. J. Infect. Chemother. 21, 247–256. https://doi.org/10.1016/j.jiac.2014.11.012 (2015).
Torumkuney, D. et al. Results from the survey of antibiotic resistance (SOAR) 2012–14 in Thailand, India, South Korea and Singapore. J. Antimicrob. Chemother. 71(Suppl 1), i3-19. https://doi.org/10.1093/jac/dkw073 (2016).
Hu, F. et al. Results from the survey of antibiotic resistance (SOAR) 2009–11 and 2013–14 in China. J. Antimicrob. Chemother. 71(Suppl 1), i33-43. https://doi.org/10.1093/jac/dkw065 (2016).
Zhou, M. et al. Antimicrobial resistance of Haemophilus influenzae isolates from pediatric hospitals in Mainland China: report from the ISPED program, 2017–2019. Indian J. Med. Microbiol. 39, 434–438. https://doi.org/10.1016/j.ijmmb.2021.09.001 (2021).
Shi, Y. et al. An overview of COVID-19. J. Zhejiang Univ. Sci. B 21, 343–360. https://doi.org/10.1631/jzus.B2000083 (2020).
Hasnain, M., Pasha, M. F. & Ghani, I. Combined measures to control the COVID-19 pandemic in Wuhan, Hubei, China: a narrative review. J. Biosafety Biosecurity 2, 51–57. https://doi.org/10.1016/j.jobb.2020.10.001 (2020).
Lau, H. et al. The positive impact of lockdown in Wuhan on containing the COVID-19 outbreak in China. J. Travel Med. https://doi.org/10.1093/jtm/taaa037 (2020).
Deghmane, A. E. & Taha, M. K. Changes in invasive neisseria meningitidis and haemophilus influenzae infections in france during the COVID-19 pandemic. Microorganisms https://doi.org/10.3390/microorganisms10050907 (2022).
Meng, Q. et al. Comparison of the distribution and changes in the antibiotic resistance of clinical bacterial isolates from the lower respiratory tract of children in shenzhen before the epidemic, during the epidemic, and during the period of normalized prevention and control of COVID-19. Infect. Diseases Therapy 12, 563–575. https://doi.org/10.1007/s40121-022-00751-4 (2023).
Brueggemann, A. B. et al. Changes in the incidence of invasive disease due to streptococcus pneumoniae, haemophilus influenzae, and neisseria meningitidis during the COVID-19 pandemic in 26 countries and territories in the Invasive respiratory infection surveillance initiative: a prospective analysis of surveillance data. The Lancet Digital health 3, e360–e370. https://doi.org/10.1016/s2589-7500(21)00077-7 (2021).
Shaw, D. et al. Trends in invasive bacterial diseases during the first 2 years of the COVID-19 pandemic: analyses of prospective surveillance data from 30 countries and territories in the IRIS consortium. The Lancet Digital health 5, e582–e593. https://doi.org/10.1016/s2589-7500(23)00108-5 (2023).
Fu, P. et al. Bacterial epidemiology and antimicrobial resistance profiles in children reported by the ISPED program in China, 2016 to 2020. Microbiol. Spectrum 9, e0028321. https://doi.org/10.1128/Spectrum.00283-21 (2021).
Zhou, J. et al. Changes of haemophilus influenzae infection in children before and after the COVID-19 pandemic, Henan. China. The Journal of infection 86, 66–117. https://doi.org/10.1016/j.jinf.2022.10.019 (2023).
Zhu, X. et al. Distribution and drug resistance of bacterial pathogens associated with lower respiratory tract infection in children and the effect of COVID-19 on the distribution of pathogens. Canadian J. Infect. Diseases Med. Microbiol. 29(2022), 1181283. https://doi.org/10.1155/2022/1181283 (2022).
Everard, M. L. Paediatric respiratory infections. Eur. Respiratory Review: Offic. J. Eur. Respiratory Society 25, 36–40. https://doi.org/10.1183/16000617.0084-2015 (2016).
Li, J. P. et al. Epidemiological features and antibiotic resistance patterns of haemophilus influenzae originating from respiratory tract and vaginal specimens in pediatric patients. J. Pediatr. Adolesc. Gynecol. 30, 626–631. https://doi.org/10.1016/j.jpag.2017.06.002 (2017).
Cherkaoui, A. et al. Ampicillin-resistant Haemophilus influenzae isolates in Geneva: serotype, antimicrobial susceptibility, and β-lactam resistance mechanisms. Eur. J. Clin. Microbiol. Infect. Dis.: Offic. Publ. Eur. Soc. Clin. Microbiol. 34, 1937–1945. https://doi.org/10.1007/s10096-015-2435-5 (2015).
Cherkaoui, A. et al. Imipenem heteroresistance in nontypeable Haemophilus influenzae is linked to a combination of altered PBP3, slow drug influx and direct efflux regulation. Clin. Microbiol. Infect.: Offic. Publ. Eur. Soc. Clin. Microbiol. Infect. Diseases 23(118), e119-118.e119. https://doi.org/10.1016/j.cmi.2016.10.009 (2017).
Lâm, T. T., Claus, H., Elias, J., Frosch, M. & Vogel, U. Ampicillin resistance of invasive Haemophilus influenzae isolates in Germany 2009–2012. Int. J. Med. Microbiol.: IJMM 305, 748–755. https://doi.org/10.1016/j.ijmm.2015.08.028 (2015).
Wang, H. J. et al. Antibiotic resistance profiles of haemophilus influenzae isolates from children in 2016: a multicenter study in China. Canadian J. Infect. Diseases Med. Microbiol. 14, 2019. https://doi.org/10.1155/2019/6456321(2019) (2019).
Zhao, C. et al. Antimicrobial resistance trends of the most common causative pathogens associated with community-acquired respiratory infections in China: 2009–2018. Infect. Drug Resistance 15, 5069–5083. https://doi.org/10.2147/idr.S374805 (2022).
Pérez-Vázquez, M., Román, F., Varela, M. C., Cantón, R. & Campos, J. Activities of 13 quinolones by three susceptibility testing methods against a collection of Haemophilus influenzae isolates with different levels of susceptibility to ciprofloxacin: evidence for cross-resistance. J. Antimicrob. Chemother. 51, 147–151. https://doi.org/10.1093/jac/dkg049 (2003).
Pérez-Vázquez, M., Román, F., García-Cobos, S. & Campos, J. Fluoroquinolone resistance in Haemophilus influenzae is associated with hypermutability. Antimicrob. Agents Chemother. 51, 1566–1569. https://doi.org/10.1128/aac.01437-06 (2007).
Kuo, S. C. et al. Levofloxacin-resistant haemophilus influenzae, Taiwan, 2004–2010. Emerg. Infect. Dis. 20, 1386–1390. https://doi.org/10.3201/eid2008.140341 (2014).
Kiedrowska, M. et al. β-Lactam resistance among Haemophilus influenzae isolates in Poland. Journal of Global Antimicrobial Resistance 11, 161–166. https://doi.org/10.1016/j.jgar.2017.08.005 (2017).
Søndergaard, A. & Nørskov-Lauritsen, N. Contribution of PBP3 Substitutions and TEM-1, TEM-15, and ROB-1 beta-lactamases to cefotaxime resistance in haemophilus influenzae and haemophilus parainfluenzae. Microbial. Drug Resistance (Larchmont, N.Y.) 22, 247–252. https://doi.org/10.1089/mdr.2015.0189 (2016).
Honda, H. et al. Multiclonal expansion and high prevalence of β-lactamase-negative haemophilus influenzae with high-level ampicillin resistance in japan and susceptibility to quinolones. Antimicrobial Agents Chemotherapy https://doi.org/10.1128/aac.00851-18 (2018).
Deghmane, A. E. et al. High diversity of invasive Haemophilus influenzae isolates in France and the emergence of resistance to third generation cephalosporins by alteration of ftsI gene. J. Infect. 79, 7–14. https://doi.org/10.1016/j.jinf.2019.05.007 (2019).
Wen, S. et al. Molecular epidemiology and antibiotic resistance analysis of non-typeable haemophilus influenzae (NTHi) in Guangzhou: a representative city of southern China. Antibiotics (Basel, Switzerland) https://doi.org/10.3390/antibiotics12040656 (2023).
Yuan, M. et al. Characterization of serotypes and molecular drug resistance patterns of haemophilus influenzae in kunming children. Pol. J. Microbiol. 72, 125–131. https://doi.org/10.33073/pjm-2023-006 (2023).
Prasad, N. et al. Changes in the Incidence of invasive bacterial disease during the COVID-19 pandemic in the United States, 2014–2020. J. Infect. Dis. 227, 907–916. https://doi.org/10.1093/infdis/jiad028 (2023).
Cheng, V. C. et al. Decreased antibiotic consumption coincided with reduction in bacteremia caused by bacterial species with respiratory transmission potential during the COVID-19 pandemic. Antibiotics (Basel, Switzerland) https://doi.org/10.3390/antibiotics11060746 (2022).
Tønnessen, R. et al. Molecular epidemiology and antibiotic resistance profiles of invasive Haemophilus influenzae from Norway 2017–2021. Front. Microbiol. 13, 973257. https://doi.org/10.3389/fmicb.2022.973257 (2022).
Steens, A. et al. Pathogen- and type-specific changes in invasive bacterial disease epidemiology during the first year of the COVID-19 pandemic in The Netherlands. Microorganisms https://doi.org/10.3390/microorganisms10050972 (2022).
Tang, H. J., Lai, C. C. & Chao, C. M. The collateral effect of COVID-19 on the epidemiology of airborne/droplet-transmitted notifiable infectious diseases in Taiwan. Antibiotics (Basel, Switzerland) https://doi.org/10.3390/antibiotics11040478 (2022).
Peradotto, M. et al. The impact of COVID-19 pandemic control on vaccine-preventable invasive bacterial diseases in Piedmont (Italy). Infection 50, 767–770. https://doi.org/10.1007/s15010-022-01770-6 (2022).
Nielsen, R. T. et al. COVID-19 preventive measures coincided with a marked decline in other infectious diseases in Denmark, spring 2020. Epidemiol. Infect. 150, e138. https://doi.org/10.1017/s0950268822001145 (2022).
Liu, Y. N. et al. Infection and co-infection patterns of community-acquired pneumonia in patients of different ages in China from 2009 to 2020: a national surveillance study. The Lancet Microbe 4, e330–e339. https://doi.org/10.1016/s2666-5247(23)00031-9 (2023).
Shi, T. et al. Immediate and long-term changes in the epidemiology, infection spectrum, and clinical characteristics of viral and bacterial respiratory infections in Western China after the COVID-19 outbreak: a modeling study. Arch. Virol. 168, 120. https://doi.org/10.1007/s00705-023-05752-3 (2023).
Li, D. et al. Molecular Epidemiology and clinical features of haemophilus influenzae among hospitalized children with community-acquired pneumonia in chengde, China. Biomed. Environ. Sci.: BES 33, 623–627. https://doi.org/10.3967/bes2020.082 (2020).
Li, X. et al. Prevalence and clinical significance of common respiratory pathogens in the upper respiratory tract of children with community-acquired pneumonia in Zunyi, China. Pediatric Pulmonol. 55, 2437–2443. https://doi.org/10.1002/ppul.24922 (2020).
Sun, Y. P. et al. Epidemiology of respiratory pathogens among children hospitalized for pneumonia in xiamen: a retrospective study. Infectious Dis. Therapy 10, 1567–1578. https://doi.org/10.1007/s40121-021-00472-0 (2021).
Langereis, J. D. & de Jonge, M. I. Invasive disease caused by nontypeable haemophilus influenzae. Emerging Infect. Dis. 21, 1711–1718. https://doi.org/10.3201/eid2110.150004 (2015).
Naito, S. et al. Clinical and bacteriologic analysis of nontypeable haemophilus influenzae strains isolated from children with invasive diseases in Japan from 2008 to 2015. J. Clin. Microbiol. 56(7), 10–128. https://doi.org/10.1128/jcm.00141-18 (2018).
Funding
This study was supported by the Project Fund of Yongchuan Science and Technology Bureau (No. 2020cc0202).
Author information
Authors and Affiliations
Contributions
Ling Ai, Liang Fang, Beizhong Liu, and Chanjuan Zhou: Investigation, Data curation, Writing-Original draft preparation, Conceptualization, Methodology, Software; Fang Gong: Visualization, Writing-Reviewing, Validation, Supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ai, L., Fang, L., Liu, B. et al. Impact of the COVID-19 pandemic on Haemophilus influenzae infections in pediatric patients hospitalized with community acquired pneumonia. Sci Rep 14, 12737 (2024). https://doi.org/10.1038/s41598-024-62728-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-62728-2
- Springer Nature Limited