Abstract
The cGAS-STING pathway appears to contribute to dysregulated inflammation during coronavirus disease 2019 (COVID-19); however, inflammatory factors related to long COVID are still being investigated. In the present study, we evaluated the association of cGAS and STING gene expression levels and plasma IFN-α, TNF-α and IL-6 levels with COVID-19 severity in acute infection and long COVID, based on analysis of blood samples from 148 individuals, 87 with acute COVID-19 and 61 in the post-COVID-19 period. Quantification of gene expression was performed by real-time PCR, and cytokine levels were quantified by ELISA and flow cytometry. In acute COVID-19, cGAS, STING, IFN-α, TNF-α, and IL-6 levels were higher in patients with severe disease than in those with nonsevere manifestations (p < 0.05). Long COVID was associated with elevated cGAS, STING and IFN-α levels (p < 0.05). Activation of the cGAS-STING pathway may contribute to an intense systemic inflammatory state in severe COVID-19 and, after infection resolution, induce an autoinflammatory disease in some tissues, resulting in long COVID.
Similar content being viewed by others
Introduction
Coronavirus disease 2019 (COVID-19) involves different clinical manifestations, ranging from mild and moderate forms to the most severe form of the disease, which is characterized by severe acute respiratory syndrome (SARS), with a high risk of death1,2.
Several factors may favor the development of severe COVID-19, including advanced age, sex, the presence of comorbidities and altered activation of immune mechanisms, which may contribute to increased replication of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and trigger systemic immunopathological reactions2,3,4.
Another condition resulting from SARS-CoV-2 infection is long COVID, which is defined by the presence of a variety of symptoms that persist after resolution of the acute infection and are not explained by other causes; the symptoms include cognitive and mental impairments, pain in the chest and joints, palpitations, myalgia, smell and taste dysfunctions, cough, headache, and gastrointestinal and heart problems5,6,7. Individuals with long COVID may develop myalgic encephalo-myelitis/chronic fatigue syndrome (ME/CFS)-related postexertional malaise, responsible for causing a severe inability to perform common activities that healthy people realize with ease8. Long COVID appears to be associated with a change in the inflammatory response, which persists in some individuals, regardless of the severity of the acute COVID-19 episode9,10.
The innate immune system is the body's first line of defense against invading pathogens, and it is mobilized by a variety of cell receptors that detect molecular patterns composing the viral structure. However, several strategies are exploited by SARS-CoV-2 to disrupt antiviral innate immune responses11, including the production of viral proteins that inhibit the translocation of stimulator of interferon genes (STING), attenuating the innate antiviral response12.
Canonically, the cyclic GMP-AMP synthase (cGAS)-STING pathway is activated when cGAS binds to double-stranded DNA (dsDNA) or DNA:RNA hybrids in the cell cytoplasm. This event leads to the production of cyclic GMP-AMP (cGAMP), which interacts with STING and promotes its dimerization, followed by interaction with TANK-binding kinase 1 (TBK1) and the transcription factor interferon regulatory factor 3 (IRF3). STING promotes the transphosphorylation of IRF3, leading to pIRF3 formation, which then translocates to the nucleus and helps to induce IFN-I transcription13,14,15.
Although the major IFN expression-inducing pattern recognition receptors (PRRs) that detect viral RNAs are members of the retinoic acid-inducible gene I (RIG-I) and Toll-like receptor (TLR) protein families, STING also plays an important role in RNA virus infection16 and has been associated with infections by vesicular stomatitis virus, dengue virus, coronavirus, and influenza virus. STING may play a relevant role in these infectious processes because, similar to SARS-CoV-2, these viruses have developed strategies to antagonize cGAS-STING signaling12,17,18,19,20. In addition, the cGAS-STING pathway can respond to viral signatures or mediate the inflammatory process via detection of cellular DNA released from mitochondria or the nucleus resulting from stress caused by the infection21.
To determine the role of STING in RNA virus infection, Franz et al.16 evaluated a panel of viruses from different families in the presence and absence of STING and observed that all viral infections were more productive in its absence, indicating that STING is required for controlling the replication of several RNA viruses. In SARS-CoV-2 infection, activation of the cGAS-STING pathway appears to be intensified as a result of a host collateral response to tissue damage22.
The intensity of the immune response is a critical factor during COVID-19, and evaluation of the components that activate innate immunity mechanisms may improve understanding of how the antiviral and inflammatory responses of the host influence the development of the severe form of acute or long COVID9,23. Thus, the present study aimed to investigate the association between the severe form of acute COVID-19 and long COVID with the gene expression of STING and cGAS and the plasma levels of IFN-α, TNF-α and IL-6.
Materials and methods
Participant characteristics and sample collection
In the present study, blood samples from 148 individuals with COVID-19, were analyzed. Of all the patients, 87 had COVID-19 at the time of sample collection, that is, acute COVID-19, and were classified as having severe (n = 44) or nonsevere (n = 43) clinical manifestations according to the criteria established by the World Health Organization1. This study also included 61 individuals who had no active infection at the time of sample collection (the post-COVID-19 period); of these subjects, 30 had long and 31 had already recovered from COVID-19 and did not have any symptoms related to long COVID syndrome (these individuals were followed up for 6 months after infection resolution)24. All individuals in the post-COVID period experience mild clinical manifestations during COVID-19.
The participants included individuals of both sexes, aged 18 years or over, who were untreated, had not yet been vaccinated against SARS-CoV-2 and were treated at the COVID-19 outpatient clinic of the State University of Pará, in the Belém Adventist Hospital and at the Evandro Chagas Institute from July 2020 to May 2021. The diagnostic criteria for long-COVID consisted of: (I) Having had an acute COVID-19 diagnosis with symptoms (severe, moderate or severe) with confirmation of SARS-CoV-2 infection by real-time polymerase chain reaction amplification; (II) Have presented prolonged clinical manifestation of at least one symptom of COVID-19, such as fatigue, dyspnea, cough, chest pain, muscle pain or weakness, headache, insomnia, visual disturbances, tremor, loss of balance, edema lower limb pain, arthralgia, palate and/or smell disorders; (III) Symptoms persist for at least 3 months after resolution of the acute infection; (IV) Post-COVID-19 symptoms cannot be attributed to any other possible cause. The prevalence of symptoms related to long COVID in the investigated group is described in Table 1. These patients were treated at the outpatient clinic for long COVID at the State University of Pará25.
Individuals with an established diagnosis of other diseases (including genetic, infectious, autoimmune diseases, cancer and physical trauma) were excluded from the study.
Blood samples (10 mL) were collected by intravenous puncture using a vacuum collection system containing ethylenediaminetetraacetic acid (EDTA) as an anticoagulant. The samples were transported to the Virology Laboratory of the Federal University of Pará, where they were processed for separation of plasma and leukocytes. Leukocyte samples were used for RNA extraction, and plasma samples were used for plasma cytokine assessment24.
RNA extraction and reverse transcription
Total RNA was extracted from peripheral blood leukocytes using a TRIzol™ Plus RNA Purification Kit (Thermo Fisher Scientific, Waltham, Massachusetts, USA), and all steps followed the protocol recommended by the manufacturer. The concentration of the extracted RNA was determined using a BioDrop™ (Bio-Rad, Hercules, California, USA) according to the manufacturer's instructions. The concentrations of all total RNA samples for synthesis of complementary DNA (cDNA) were equal to 50 ng/µL25.
The extracted RNA was converted to cDNA using a High Capacity cDNA Reverse Transcription® with RNAse Inhibitor Kit (Applied Biosystems, Foster City, CA, USA). For the cDNA reaction, a mixture was prepared with a final volume of 20.0 µL containing 2 µL of 10X RT Buffer, 0.8 µL of 25X dNTP Mix (100 nM), 2 µL of random primer, 1 µL of MultiScribe™ Reverse Transcriptase, 1 µL of RNaseOUT™ and 3.2 µL of ultra-pure water provided by the kit as well as 10.0 µL of extracted RNA. The mixture was subjected to cycles of 25 °C for 10 min, 37 °C for 120 min and 85 °C for 5 min using a Mastercycler Personal thermocycler (Eppendorf, Hamburg, Germany)25.
Quantification of gene expression
Gene expression was determined by mRNA quantification using real-time PCR. Initially, standardization of qPCRs with cDNAs and probes (endogenous and target genes) was carried out to calculate the efficiency of the amplification reactions in which different concentrations of cDNA were tested (neat and in 4 dilutions of factor 2 - − 1:2, 1:4, 1:8 and 1:16). All reactions were performed in microtiter plates and in triplicate while simultaneously analyzing the same cDNA (at different dilutions) with different probes to construct an efficiency curve to validate the 2-ΔΔCT analysis method. All assays showed good efficiency, as expected (100% ± 10)26.
Relative quantification of gene expression consisted of amplification of the target gene with the endogenous gene (normalizer) using TaqMan™ assays (Applied Biosystems, Foster City, CA, USA) and the StepOnePLUS™ Real-Time PCR System (Thermo Fisher Scientific, Waltham, MA, USA). The reactions were performed in the singleplex format according to the manufacturer's protocol. Hs00736955_g1 was used for STING and Hs00403553_m1 for cGAS; glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as an endogenous control (Hs02786624_g1). All assay kits were obtained commercially (Thermo Fisher Scientific, Waltham, MA, USA). For the reaction, 15 µL of 2X TaqMan® Universal PCR Master Mix, 1.5 µL of 20 × TaqMan Gene Expression Assay, 3 µL of cDNA and 10.5 µL of RNase-free water were used, with the following thermocycling conditions: 2 min at 50 °C, followed by 10 min at 95 °C and 1 min at 60 °C25.
Relative quantification (RQ) of target gene expression was determined using the comparative CT method (∆∆Ct) with the 2-ΔΔCT formula, where ∆∆Ct = ∆Ct sample–∆Ct reference (Life Technologies, Carlsbad, CA, USA)25.
Plasma concentration of cytokines
Quantification of TNF-α and IL-6 levels was performed using flow cytometry with a Cytometric Bead Array (CBA) Human Th1/Th2/Th17 Kit (BD Biosciences, San Diego, CA, USA) and BD FACS Canto II equipment. All procedures followed the manufacturer's guidelines. The methodology used is based on beads conjugated with a capture antibody, in which specific populations of beads with known fluorescence intensities and conjugated to a specific capture antibody are mixed to form the CBA, and fluorescence is then measured in the FL-3 channel of the flow cytometer24. IFN-α levels in plasma samples were quantified using an ELISA-type immunoenzymatic assay, the Elabbscience Human IFN alpha ELISA Kit (Houston, TX, USA), according to the manufacturer's recommendations25.
Statistical analysis
The information obtained was entered into a database in Microsoft Office Excel 2013 software. Normality analysis of the distribution of cytokine levels was performed using the Shapiro‒Wilk test. Based on the results of the normality test, evaluation of the plasma levels of these markers between the groups with acute, severe and nonsevere COVID-19 was performed using the nonparametric Mann‒Whitney test. A correlogram was generated to evaluate the correlation between the expression levels of cGAS and STING and the plasma levels of cytokines using the Spearman correlation test. The ROC curve was used to assess whether the levels of the investigated markers are capable of differentiating groups in acute COVID-19 (severe and non-severe) and in the post-COVID period (long COVID and non-long COVID). Tests were performed using GraphPad Prism 5.0 and RStudio 4.0.1 programs, with p < 0.05 considered to indicate significance.
Heatmap graphics were inferred using the R studio 4.2.1 program with the gplots, RColorBrewer and preprocessCore plotting packages; we utilized the ward d2 method as the distance type and the canberra method as the aggregation type27.
Ethics declaration
The study was approved by the Brazilian National Research Ethics Committee (CAEE: 33470020.1001.0018), protocolo nº 2.190.330, according to the guidelines of CNS Resolution nº 466/2012. Research involving human participants was carried out in accordance with the Declaration of Helsinki. All individuals were informed about the research objectives, and those who agreed to participate in the study signed a free and informed consent form and responded to a clinical-epidemiological questionnaire.
Results
Characterization of the investigated groups
The patients with acute COVID-19 had a mean age of 52 years, and the majority were male (n = 48; 55.17%). The individuals evaluated in the post-COVID-19 period had a mean age of 40.5 years, and 35 were female (57.37%). The characteristics of each analyzed group are described in Table 2.
Gene expression of cGAS and STING
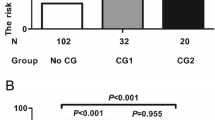
Evaluation of cGAS and STING gene expression levels showed that the group with the severe form of acute COVID-19 had higher levels of cGAS (p = 0.0009; Fig. 1A) and STING (p = 0.0269; Fig. 1C). Regarding the groups evaluated in the post-COVID-19 period, the group with long COVID had higher levels of cGAS (p < 0.0001; Fig. 1B) and STING (p = 0.0011; Fig. 1D).
Plasma levels of cytokines
In the analysis of plasma cytokine levels, it was observed that patients with the severe form of COVID-19 had higher levels of IFN-α (p = 0.0006; Fig. 2A), TNF-α (p = 0.0137; Fig. 2C) and IL-6 (p = 0.0071; Fig. 2E) than those with the nonsevere form. Furthermore, individuals with long COVID had higher levels of IFN-α (p = 0.0001; Fig. 2B) than those without long COVID symptoms, but there was no significant difference in TNF-α and IL-6 levels between these groups (Fig. 2D and F).
Correlation of the evaluated inflammatory markers
In assessing the correlation between the gene expression levels of cGAS, STING and cytokines in acute COVID-19, a positive correlation was observed between the levels of all markers in the group with the severe form of the disease. A positive correlation with almost all markers was also detected in the group with the nonsevere form, except for STING with TNF-α, STING with IL-6, IFN-α with TNF-α and IFN-α with IL-6 (Fig. 3).
Correlogram of the expression levels of cGAS and STING and plasma levels of the cytokines IFN-α, TNF-α and IL-6 in patients with acute COVID-19. Scatter plots show the correlation between markers, and density-adjusted curve plots show the distribution of individuals in relation to marker concentration.
In the group of individuals in the post-COVID-19 period, there was a positive correlation only between cGAS and STING and cGAS and IFN-α in the long COVID and without long COVID groups and between STING and IFN-α in the long COVID group (Fig. 4).
Correlogram of the expression levels of cGAS and STING and plasma levels of the cytokines IFN-α, TNF-α and IL-6 in individuals in the post-COVID-19 period. Scatter plots show the correlation between markers, and density-adjusted curve plots show the distribution of individuals in relation to marker concentration.
A heatmap was constructed for joint analysis of the gene expression levels of cGAS and STING and plasma levels of IFN-α, TNF-α and IL-6 in acute COVID-19 and in the post-COVID-19 period. In acute COVID-19, two groups were observed: one consisting mainly of patients with severe COVID-19, who showed greater levels of gene expression and the cytokines evaluated, and another including patients with nonsevere manifestations of the disease (Fig. 5A). In evaluation of the groups in the post-COVID-19 period, the heatmap illustrated two groups, with the group formed by patients with long COVID having higher gene expression of cGAS and STING and cytokine IFN-α levels (Fig. 5B).
The ability of the levels of the investigated markers to differentiate groups in acute COVID-19 and in the post-COVID period was evaluated by the ROC curve. In acute COVID-19, TNF-α, IFN-α, STING and cGAS showed a performance that varied from fair to good in differentiating severe and non-severe cases, with area under the curve (AUC) ranging from 0.7051 to 0.8145 (p < 0.05; Fig. 6A). In the post-COVID period, the IFN-α, STING and cGAS markers show good ability to differentiate individuals with and without long COVID, with AUC that ranged from 0.8195 to 0.8904 (p < 0.005; Fig. 6B).
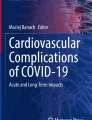
Inflammatory markers evaluated in relation to risk factors for COVID-19 and long COVID
The gene expression levels of cGAS, STING and the plasma levels of the cytokines IFN-α, TNF-α and IL-6 in patients with acute COVID-19 were evaluated according to factors considered risk factors for the severity of the disease, namely, sex, age and comorbidities. However, no association was observed between the levels of any of the markers and male sex, advanced age (> 60 years) or the presence of comorbidities (Table 3).
Analysis of the long COVID group showed that there was no statistical difference in the expression levels of cGAS and STING and the cytokines IFN-α, TNF-α and IL-6 between individuals with different amounts of symptoms and those with the presence or absence of comorbidities (Table 4).
Discussion
Based on increased susceptibility in STING-knockout mice, initial studies on the role of STING in RNA virus infections described the importance of this adapter in controlling different infections28. In some infections, proteins produced by RNA viruses promote STING degradation, reducing IFN-I levels and facilitating viral replication17,18,19,20. STING is important for host defense during RNA virus infection, as its activity can also limit the initial steps of viral mRNA translation16.
In the present study, higher levels of STING and cGAS gene expression and plasma levels of IFN-α, IL-6, and TNF-α were identified in patients with the severe form of acute COVID-19 than in patients with the nonsevere form. A positive correlation between the levels of all evaluated markers was also found for severe COVID-19 patients, a relationship also demonstrated by a heatmap.
In viral infections, IFN-I is important for restricting replication by inducing the expression of IFN-stimulated genes (ISGs) and enhancing the immune response of myeloid cells, NK cells, B cells and T lymphocytes29. IFN-I, induced by cGAS-STING, binds to the heterodimeric transmembrane receptor composed of the subunits IFNAR1 and IFNAR2, which activates two proteins, Janus receptor-associated tyrosine kinase (JAK1) and tyrosine kinase e (TYK2), which induce dimerization and nuclear translocation of transcription activator 1 (STAT1) and 2 (STAT2) and binding to IRF9. This complex associates with the IFN-stimulated response elements (ISREs), inducing the production of several ISGs. Some ISGs act in the antiviral response such as MX1, OAS and TRIM and others as positive regulators of IFN signaling, including STAT1 and 2, IRF3, 7 and 9 and cGAS, contributing to the maintenance of INF-I production through the cGAS-STING pathway30,31.
TNF-α promotes intercellular communication and regulates cell survival, apoptosis and necroptosis32. The synthesis of IL-6 at the injury site leads to a change in homeostasis induced by the production of acute-phase proteins and platelets, which promotes changes in vascular permeability and thrombocytopenia33. The actions induced by high levels of IL-6, IFN-α, TNF-α have been associated with persistence and dysfunction of severe COVID-19 and with a greater probability of death34,35,36.
Although STING has been shown to participate in the control of RNA virus infections, including SARS-CoV-2 infection, elevated expression of cGAS and STING in patients with severe acute COVID-19 appears to be a consequence of the intense damage caused to infected cells rather than an attempt to limit virus infection. Domizio et al.22 demonstrated that the main cells involved in cGAS, STING and IFN-I dysfunction in COVID-19 are endothelial cells and phagocytes. Infection of endothelial cells by SARS-CoV-2 results in damage and the release of mitochondrial DNA into the cytoplasm, inducing cGAS-STING pathway signaling, IFN-I and inflammatory cytokine production and cell death; in phagocytes, enhanced IFN-I production occurs upon phagocytosis of dying endothelial cells and recognition of the DNA from these cells by cGAS. In cells infected with SARS-CoV-2, cGAS-STING is mainly responsible for the inflammatory cytokine production mediated through NF-κβ activation21.
Although the role of cGAS, STING and cytokines in the severity of COVID-19 has been described in other populations, our study includes initial information on the relationship among these components in the population of Brazil, more specifically, in the population of the Brazilian Amazon, a mixed-race population (consisting of whites, blacks and indigenous people)37 that presents genetic and cultural differences compared to populations evaluated in other studies. The results of the present study reinforce that, regardless of characteristics specific to the population, cGAS-STING pathway contributes significantly to the severity of COVID-19, as these patients have higher gene expression levels of cGAS and STING as well as the cytokines IFN-α, TNF-α and IL-6 in peripheral blood. This pathway promotes maintenance of a systemic inflammatory state, which induces thromboembolic changes and multiple organ failure, the main causes of death in patients with severe disease38.
Patients with long COVID had higher cGAS, STING and IFN-α levels than individuals who did not manifest any symptoms in the post-COVID-19 period. Although the levels of several inflammatory markers have been associated with the severity of acute COVID-19, the development of long COVID is still poorly understood since the persistence of inflammation is associated with only certain conditions, such as neurological manifestations and insulin resistance39,40. In a more general assessment of the disease, patients with long COVID had higher levels of IL-17 and lower levels of the anti-inflammatory cytokines IL-10 and IL-4 (compared to individuals who did not develop the syndrome), which suggests a possible dysregulation of the immune-inflammatory balance10.
Since in long COVID there is deregulation of the functions of various systems of the human body41 and, possibly, high levels of cGAS-STING can contribute to the development of low-grade inflammatory diseases in various organs such as the heart (cardiomyopathy and myocardial infarction), liver (fat accumulation), kidneys (chronic kidney disease), pancreas (type 1 diabetes mellitus) and brain (ischemic stroke)43, the elevated levels of cGAS and STING observed in this study may suggest an important role for these markers in the development and maintenance of long COVID. Although STING is essential to promote host defense against viral infections by mediating IFN-I production, chronic activation of STING must be inhibited to prevent the development of inflammatory and autoimmune diseases42,43,44,45.
Several researchers have struggled to understand the causes of long COVID, which can be disabling. They are developing studies with three main foci, which may explain the possible mechanisms of disease pathogenesis46, including changes in blood flow, which can lead to the formation of microclots47,48, the persistence of the virus in certain body tissues, mainly in the intestinal mucosa49 and the unregulated activation of cells of the immune system50,51,52. Although these mechanisms differ, they converge to a common point, the activation of the inflammatory response.
As the processes in the possible causes of long COVID that are being investigated may be related to the persistence of the virus's genetic material or to cell damage, these components may induce overactivation of innate immune pathways, including activation of cGAS-STING. One of the main causes of inflammation is the activation of STING in phagocytes, mediated by the ingestion of damaged cells, since the DNA of these cells can intensify STING activity and induce greater inflammatory cytokine and antibody production and leukocyte activation against the antigens themselves, leading to the development of autoinflammatory disease. In the case of residual virus, this can directly cause the activation of STING in infected cells or favor the phagocytosis of these cells, which further aggravates the activation of STING in phagocytes22,53,54,55,56. The presence of pro-inflammatory factors induces the activation of pathways that promote neurodegenerative processes and mitochondrial damage, such as the kynurenine pathway (KP)57 which can lead to the activation of cGAS and STING and the maintenance of high levels of IFN-I, contributing to the development of some long COVID symptoms, mainly related to neurodegenerative disorders. KP activation was more prolonged and associated with high levels of IFN-β in individuals who have long COVID with neurological changes58.
The results of the present study are relevant to understanding that all possible causes of long COVID that can promote chronic inflammatory activation in certain tissues, which can be mainly mediated by activation of the cGAS-STING pathway and IFN-I production. È possivel que outros mecanismos possam contribuir.
The study provides an initial understanding of the role of the cGAS-STING pathway, as one of the possible mechanisms that may be contributing to the development of morbidities associated with long COVID, nevertheless the presence of a higher frequency of female individuals, of an older age (40 to 60 years old) and with comorbidities in the group with long COVID constituted a limitation of the study, since these characteristics are considered risk factors for the severity of COVID-19 and can influence the development of symptoms in long COVID. However, we emphasize that all individuals in the post-COVID period (with and without long COVID) presented mild manifestations during the COVID-19 phase, suggesting the need for a better assessment of the contribution of these risk factors in the establishment of long COVID.
Conclusion
The SARS-CoV-2 infection induces immune response activation and cell injury, which can vary in intensity. In acute infection, patients with severe COVID-19 had higher levels of cGAS, STING, IFN-α, TNF-α and IL-6 expression than patients with nonsevere manifestations of the disease, demonstrating that cGAS and STING activation, which is responsible for inducing IFN-I and proinflammatory cytokine production, contributes to the maintenance of an intense systemic inflammatory state characteristic of severe COVID-19. In long COVID, elevated levels of cGAS, STING and IFN-α after the resolution of SARS-CoV-2 infection may be a consequence of the development of a possible autoinflammatory disease in some tissues, maintained by cGAS-STING activation.
Data availability
The data that support the fndings of this study are available on request from the corresponding author.
Abbreviations
- AMP:
-
Adenosine monophosphate
- cGAMP:
-
Cyclic GMP-AMP
- cGAS:
-
Cyclic GMP-AMP synthase
- COVID-19:
-
Coronavirus disease 2019
- GMP:
-
Guanosine monophosphate
- IFNAR1:
-
Interferon alpha/beta receptor 1
- IFNAR2:
-
Interferon-alpha/beta receptor 2
- IFN-α:
-
Interferon alpha
- IFN-I:
-
Type I interferon
- IL-4:
-
Interleukin 4
- IL-6:
-
Interleukin 6
- IL-10:
-
Interleukin 10
- IRF3:
-
Interferon regulatory factor 3
- IRF9:
-
Transcription interferon regulatory factor 9
- ISGs:
-
IFN-stimulated genes
- ISREs:
-
IFN stimulated response elements
- JAK1:
-
Janus receptor-associated tyrosine kinase
- MX1:
-
MX dynamin like GTPase 1
- OAS:
-
2'-5'-Oligoadenylate synthetases
- PRRs:
-
Pattern recognition receptors
- RIG-I:
-
Retinoic acid-inducible gene I
- ROC:
-
Receiver operating characteristic curve
- SARS:
-
Severe acute respiratory syndrome
- SARS-CoV-2:
-
Severe acute respiratory syndrome coronavirus 2
- STAT1:
-
Transcription activator 1
- STAT2:
-
Transcription activator 2
- STING:
-
Stimulator of interferon genes
- TBK1:
-
TANK-binding kinase 1
- TLR:
-
Toll-like receptor
- TNF-α:
-
Tumor necrosis factor alpha
- TRIM:
-
Tripartite motif-containing proteins
- TYK2:
-
Tyrosine kinase 2
References
WHO. WHO Global Clinical Platform for the Clinical Characterization of COVID-19: Statistical Analysis Plan. https://www.who.int/publications/i/item/WHO-2019-nCoV-Clinical-Analytic-plan-2021.1 (2021).
Wang, F. et al. Epidemiological characteristics of patients with severe COVID-19 infection in Wuhan, China: Evidence from a retrospective observational study. Int. J. Epidemiol. 49(6), 1940–1950 (2021).
Nagant, C. et al. A score combining early detection of cytokines accurately predicts COVID-19 severity and intensive care unit transfer. Int. J. Infect. Dis. 101, 342–345 (2020).
da Silva Torres, M. K. et al. The complexity of SARS-CoV-2 infection and the COVID-19 pandemic. Front. Microbiol. 13, 789882 (2022).
Collins, F.S. NIH Launches New Initiative to Study “Long COVID”. National Institutes of Health; Bethesda, MD, USA. Available at: https://www.nih.gov/about-nih/who-we-are/nih-director/statements/nih-launches-new-initiative-study-long-covid.
Parums, D. V. Editorial: Long COVID, or post-COVID syndrome, and the global impact on health care. Med. Sci. Monit. 27, e933446 (2021).
Yong, S. J. Long COVID or post-COVID-19 syndrome: Putative pathophysiology, risk factors, and treatments. Infect. Dis. 53(10), 737–754 (2021).
Landhuis, E. W. How primary care physicians can recognize and treat long COVID. JAMA 329(20), 1727–1729 (2023).
Doykov, I. et al. “The long tail of Covid-19” - The detection of a prolonged inflammatory response after a SARS-CoV-2 infection in asymptomatic and mildly affected patients. F1000Res 9, 1349 (2020).
Queiroz, M. A. F. et al. Cytokine profiles associated with acute COVID-19 and long COVID-19 syndrome. Front. Cell. Infect. Microbiol. 12, 922422 (2022).
Zhang, Y. et al. An update on innate immune responses during SARS-CoV-2 infection. Viruses 13(10), 2060 (2021).
Han, L. et al. SARS-CoV-2 ORF10 antagonizes STING-dependent interferon activation and autophagy. J. Med. Virol. 94(11), 5174–5188 (2022).
Ishikawa, H., Ma, Z. & Barber, G. N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461(7265), 788–792 (2009).
Barber, G. N. STING-dependent cytosolic DNA sensing pathways. Trends Immunol. 35(2), 88–93 (2014).
Abe, T. & Barber, G. N. Cytosolic-DNA-mediated, STING-dependent proinflammatory gene induction necessitates canonical NF-κB activation through TBK1. J. Virol. 88(10), 5328–5341 (2014).
Franz, K. M., Neidermyer, W. J., Tan, Y. J., Whelan, S. P. J. & Kagan, J. C. STING-dependent translation inhibition restricts RNA virus replication. Proc Natl Acad Sci USA 115(9), E2058–E2067 (2018).
Sun, L. et al. Coronavirus papain-like proteases negatively regulate antiviral innate immune response through disruption of STING-mediated signaling. PLoS One 7(2), e30802 (2012).
Aguirre, S. et al. Fernandez-Sesma A. DENV inhibits type I IFN production in infected cells by cleaving human STING. PLoS Pathog. 8(10), e1002934 (2012).
Holm, C. K. et al. Influenza A virus targets a cGAS-independent STING pathway that controls enveloped RNA viruses. Nat. Commun. 7, 10680 (2016).
Stabell, A. C. et al. Dengue viruses cleave STING in humans but not in nonhuman primates, their presumed natural reservoir. Elife 7, e31919 (2018).
Neufeldt, C. J. et al. SARS-CoV-2 infection induces a pro-inflammatory cytokine response through cGAS-STING and NF-κB. Commun. Biol. 5(1), 45 (2022).
Domizio, J. D. et al. The cGAS-STING pathway drives type I IFN immunopathology in COVID-19. Nature 603(7899), 145–151 (2022).
Mortaz, E., Tabarsi, P., Varahram, M., Folkerts, G. & Adcock, I. M. The immune response and immunopathology of COVID-19. Front. Immunol. 11, 2037 (2020).
Queiroz, M. A. F. et al. Polymorphisms in the MBL2 gene are associated with the plasma levels of MBL and the cytokines IL-6 and TNF-α in severe COVID-19. Front. Immunol. 14, 1151058 (2023).
de Lima, L. L. P. et al. STING and cGAS gene expressions were downregulated among HIV-1-infected persons after antiretroviral therapy. Virol. J. 18(1), 78 (2021).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-∆∆CT method. Methods 25(4), 402–408 (2001).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing. https://www.R-project.org/ (2022).
Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455(7213), 674–678 (2008).
McNab, F., Mayer-Barber, K., Sher, A., Wack, A. & O’Garra, A. Type I interferons in infectious disease. Nat. Rev. Immunol. 15(2), 87–103 (2015).
Schneider, W. M., Chevillotte, M. D. & Rice, C. M. Interferon-stimulated genes: A complex web of host defenses. Annu. Rev. Immunol. 32, 513–545 (2014).
Ivashkiv, L. B. & Donlin, L. T. Regulation of type I interferon responses. Nat. Rev. Immunol. 14(1), 36–49 (2014).
Blaser, H., Dostert, C., Mak, T. W. & Brenner, D. TNF and ROS crosstalk in inflammation. Trends Cell Biol. 26(4), 249–261 (2016).
Tanaka, T., Narazaki, M., Masuda, K. & Kishimoto, T. Regulation of IL-6 in immunity and diseases. Adv. Exp. Med. Biol. 941, 79–88 (2016).
Zhou, X. et al. IL-6 drives T cell death to participate in lymphopenia in COVID-19. Int Immunopharmacol. 111, 109132 (2022).
Krämer, B. et al. Early IFN-α signatures and persistent dysfunction are distinguishing features of NK cells in severe COVID-19. Immunity 54(11), 2650-2669.e14 (2021).
Smail, S. W., Babaei, E., Amin, K. & Abdulahad, W. H. Serum IL-23, IL-10, and TNF-α predict in-hospital mortality in COVID-19 patients. Front. Immunol. 14, 1145840 (2023).
Santos, N. P. et al. Assessing individual interethnic admixture and population substructure using a 48-insertion-deletion (INSEL) ances-try-informative marker (AIM) panel. Hum. Mutat. 31, 84–90 (2010).
Lazzaroni, M. G. et al. Coagulation dysfunction in COVID-19: The interplay between inflammation, viral infection and the coagulation system. Blood Rev. 46, 100745 (2021).
Sun, B. et al. Characterization and biomarker analyses of post-COVID-19 complications and neurological manifestations. Cells 10(2), 386 (2021).
Montefusco, L. et al. Acute and long-term disruption of glycometabolic control after SARS-CoV-2 infection. Nat. Metab. 3(6), 774–785 (2021).
Desai, A. D., Lavelle, M., Boursiquot, B. C. & Wan, E. Y. Long-term complications of COVID-19. Am. J. Physiol. Cell Physiol. 322(1), C1–C11 (2022).
Bao, T., Liu, J., Leng, J. & Cai, L. The cGAS-STING pathway: More than fighting against viruses and cancer. Cell Biosci. 11(1), 209 (2021).
Konno, H., Konno, K. & Barber, G. N. Cyclic dinucleotides trigger ULK1 (ATG1) phosphorylation of STING to prevent sustained innate immune signaling. Cell 155(3), 688–698 (2013).
Jeremiah, N. et al. Inherited STING-activating mutation underlies a familial inflammatory syndrome with lupus-like manifestations. J. Clin. Invest. 124(12), 5516–5520 (2014).
Crow, M. K. Type I interferon in the pathogenesis of lupus. J. Immunol. 192(12), 5459–5468 (2014).
Couzin-Frankel, J. Clues to long COVID. Science 376(6599), 1261–1265 (2022).
Pretorius, E. et al. Persistent clotting protein pathology in long COVID/post-acute sequelae of COVID-19 (PASC) is accompanied by increased levels of antiplasmin. Cardiovasc. Diabetol. 20(1), 172 (2021).
Buonsenso, D. et al. Evidence of lung perfusion defects and ongoing inflammation in an adolescent with post-acute sequelae of SARS-CoV-2 infection. Lancet Child Adolesc. Health 5(9), 677–680 (2021).
Zollner, A. et al. Postacute COVID-19 is characterized by gut viral antigen persistence in inflammatory bowel diseases. Gastroenterology 163(2), 495-506.e8 (2022).
Peluso, M. J. et al. Markers of immune activation and inflammation in individuals with postacute sequelae of severe acute respiratory syndrome coronavirus 2 infection. J. Infect. Dis. 224(11), 1839–1848 (2021).
Phetsouphanh, C. et al. Immunological dysfunction persists for 8 months following initial mild-to-moderate SARS-CoV-2 infection. Nat. Immunol. 23(2), 210–216 (2022).
Son, K. et al. Circulating anti-nuclear autoantibodies in COVID-19 survivors predict long COVID symptoms. Eur. Respir. J. 61(1), 2200970 (2023).
Ahn, J., Gutman, D., Saijo, S. & Barber, G. N. STING manifests self DNA-dependent inflammatory disease. Proc. Natl. Acad. Sci. USA 109(47), 19386–19391 (2012).
Gall, A. et al. Autoimmunity initiates in nonhematopoietic cells and progresses via lymphocytes in an interferon-dependent autoimmune disease. Immunity 36(1), 120–131 (2012).
Ahn, J. et al. Extrinsic phagocyte-dependent STING signaling dictates the immunogenicity of dying cells. Cancer Cell 33(5), 862-873.e5 (2018).
Gao, K. M., Marshak-Rothstein, A. & Fitzgerald, K. A. Type-1 interferon-dependent and -independent mechanisms in cyclic GMP-AMP synthase-stimulator of interferon genes-driven auto-inflammation. Curr. Opin. Immunol. 80, 102280 (2023).
Mor, A., Tankiewicz-Kwedlo, A., Krupa, A. & Pawlak, D. Role of kynurenine pathway in oxidative stress during neurodegenerative disorders. Cells 10(7), 1603 (2021).
Cysique, L. A. et al. The kynurenine pathway relates to post-acute COVID-19 objective cognitive impairment and PASC. Ann. Clin. Transl. Neurol. 10(8), 1338–1352 (2023).
Acknowledgements
The authors thank all patients who agreed to voluntarily participate in this study.
Funding
This study was supported by the National Council for Scientific and Technological Development-CNPQ (n° 304835/2022-6, n° 401235/2020-3; n° 302935/2021-5), Instituto Nacional de Ciência e Tecnologia em Viroses Emergentes e Reemergentes (INCT-VER nº 406360/2022-7), Fundação Amazônia de Amparo a Estudos e Pesquisa do Pará-FAPESPA (n° 005/2020), Secretariat of Science, Technology and Higher, Professional and Technological Education-SECTET (n° 09/2021), Instituto Nacional de Ciência e Tecnologia em Viroses Emergentes e Reemergentes (INCT-VER #406360/2022-7) and Federal University of Para (PAPQ/2023).
Author information
Authors and Affiliations
Contributions
A.V., F.F. and E.S. conceived of the project. M.Q., A.V., I.B-C. and E.S. wrote and reviewed the manuscript. M.Q., L.P. and S.L. performed the statistical analyses. M.Q., W.B., K.P., E.A., S.L., E.Sa., F.C., K.S., M.C., M.B., A.S., M.L., M.V., F.R., R.S., G.V., T.C., A.Ve., M.C., D.H., C.S., J.N., I.C., I.C-V., I.B-C., J.Q. and F.F. collected the biological samples and performed the laboratory analyses. All authors reviewed and approved the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Queiroz, M.A.F., Brito, W.R.S., Pereira, K.A.S. et al. Severe COVID-19 and long COVID are associated with high expression of STING, cGAS and IFN-α. Sci Rep 14, 4974 (2024). https://doi.org/10.1038/s41598-024-55696-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-55696-0
- Springer Nature Limited