Abstract
The number of genome-level resources for non-model species continues to rapidly expand. However, frog species remain underrepresented, with up to 90% of frog genera having no genomic or transcriptomic data. Here, we assemble the first genomic and transcriptomic resources for the recently described southern stuttering frog (Mixophyes australis). The southern stuttering frog is ground-dwelling, inhabiting naturally vegetated riverbanks in south-eastern Australia. Using PacBio HiFi long-read sequencing and Hi-C scaffolding, we generated a high-quality genome assembly, with a scaffold N50 of 369.3 Mb and 95.1% of the genome contained in twelve scaffolds. Using this assembly, we identified the mitochondrial genome, and assembled six tissue-specific transcriptomes. We also bioinformatically characterised novel sequences of two families of antimicrobial peptides (AMPs) in the southern stuttering frog, the cathelicidins and β-defensins. While traditional peptidomic approaches to peptide discovery have typically identified one or two AMPs in a frog species from skin secretions, our bioinformatic approach discovered 12 cathelicidins and two β-defensins that were expressed in a range of tissues. We investigated the novelty of the peptides and found diverse predicted activities. Our bioinformatic approach highlights the benefits of multi-omics resources in peptide discovery and contributes valuable genomic resources in an under-represented taxon.
Similar content being viewed by others
Introduction
Since the completion of the Human Genome Project in 2003, there has been a proliferation of high quality genomic and transcriptomic resources for non-model species1,2,3. The ability to generate these ‘multi-omics’ resources in diverse species has clear comparative4, conservation5,6, and clinical benefits7. As of July 2023, over 5800 non-model animal genomes and 4600 non-model animal transcriptomes are available in the largest global genome repository, the National Center for Biotechnology Information (NCBI)8. Despite the acceleration of genomic resources, only 46 genera within the Order Anura (10% of all frog genera) have genomic or transcriptomic resources available on NCBI8,9. In Australia, despite 92% of frogs being endemic, only 4 of the 229 known endemic frog species currently have ‘omics level resources available9,10,11,12,13.
The global disparity in genetic resources across taxa has been driven in part by the diverse structures of tetrapod genomes and associated difficulties with sequencing and assembly12. While frogs have relatively stable chromosomal structures compared to other tetrapod groups14, they have highly variable genome sizes10,15,16; variable proportions of repetitive elements11,17; and can have ploidy variation18. As a result, the model Xenopus laevis genome was one of the only amphibian chromosome-scale assemblies prior to long-read sequencing19. More recent advances in long read sequencing technology20, alongside chromosome conformation capture techniques21, have since facilitated more contiguous assemblies of these challenging genomes10,22.
An emerging application of genomic resources is the bioinformatic discovery of novel antimicrobial peptides (AMPs). AMPs are small peptides found across all classes of life that form an important part of the innate immune response23,24,25. The comprehensive characterisation of AMPs is valuable in understanding how different species respond to pathogens in their environment and their pleiotropic immune effects have also been explored as potential therapeutics, with several currently undergoing clinical trials26,27. Frogs are known to express AMPs from the exocrine glands of their skin in response to their exposure to diverse, microbial communities in their amphibious, freshwater habitats28,29,30. Over 2500 novel AMPs have been characterised from only 167 frog species on the Database of Anuran Defence Peptides v1.6 (DADP)31. To date, frog AMPs have typically been characterised with a peptidomic approach; the iterative purification of AMPs showing antimicrobial activity from frog skin secretions using high-performance liquid chromatography (HPLC)32. However, this method may not capture the suite of AMPs expressed in other tissues33, nor AMPs with immune functions other than antimicrobial activity34. In response, there has been increased interest in ‘mining’ genomic and transcriptomic resources for the genes that encode for AMPs35. These methods are primarily homology-driven, using known sequences of large, well-characterised AMP families like cathelicidins and β-defensins to search for homologs36. A genomics-driven approach has been applied to discover novel AMPs from a range of non-model species35,37,38,39. However, despite the evolutionary divergence of Australian frog families (Limnodynastidae and Myobatrachidae diverged from South American frogs an estimated 80–100 million years ago)40,41, none have yet been investigated for AMPs using a bioinformatic approach.
The southern stuttering frog (Mixophyes australis)42 is a small, ground-dwelling frog inhabiting naturally vegetated riverbanks in south-eastern Australia, and is part of the Myobatrachidae family of endemic Australian ground frogs43,44. The northern (Mixophyes balbus) and southern (Mixophyes australis) species of stuttering frog were once considered a single species, but were recently split42. Although yet to be officially assessed, application of the International Union for the Conservation of Nature (IUCN) threat assessment methods for the southern stuttering frog warrants a listing of ‘Endangered’ for this newly defined species42. As a ground-dwelling frog, southern stuttering frogs spend most of their life cycle exposed to large bodies of water and decomposing detritus, which may carry pathogens such as chytrid fungus (Batrachochytrium dendrobatidis) and Ranaviruses42,45. We hypothesise that the selective pressures of the southern stuttering frog’s environment make it a good candidate for the discovery of novel and diverse AMPs. Preliminary studies on other Mixophyes species demonstrated some resistance against chytrid fungus in skin secretions46,47. While these secretions have not been sequenced, these studies suggest possible AMP activity in the Mixophyes genus.
In this study, we use a combination of PacBio HiFi long-read, short-read, and Hi-C sequencing data to generate the first genomic and transcriptomic resources for the southern stuttering frog. Using these high-quality resources, we use bioinformatics to characterise and analyse the AMP families cathelicidins and β-defensins. By generating the first ‘omics level resources for the Mixophyes genus, this study provides a unique opportunity to investigate an otherwise largely understudied, Australian taxon.
Results
Multi-omics resources for the southern stuttering frog
Using a combination of PacBio HiFi long-read and Hi-C sequencing data, we generated a high-quality genome assembly for the southern stuttering frog (Fig. 1A). The genome is 3.13 Gbp in length, has a read coverage of 26× and a scaffold N50 of 369 Mbp (Table 1). Repeat-masking the genome revealed a high proportion of repetitive elements (53.7%; Supplementary Table S1), within the broad expected range for Anuran genomes (32.0–77.1%;8). The GC content of the genome, 40.3%, was also within the range of expected Anuran GC values (26.9–44.5%;8). From the assembly, we identified a scaffold representing the mitochondrial genome, which contained 36 genes: 21 tRNAs, 2 rRNAs and 13 protein-coding genes (Fig. 1B).
The southern stuttering frog (Mixophyes australis) genome assembly. (A) The 12 longest southern stuttering frog genome scaffolds, ordered by length. Red dots indicate the presence of telomeric sequences at the end of scaffolds. Horizontal, black lines indicate contig joins informed by the Hi-C sequencing data. (B) Circos plot of the southern stuttering frog mitochondrial genome generated using Proksee48. Outer ring contains genes on the forward strand, while the inner ring contains genes on the reverse strand.
Over 95% of the genome was assembled in twelve scaffolds (Fig. 1A; Supplementary Fig. S1), which likely represent the chromosomes based on previous karyotypic analysis of other Mixophyes species (2n = 24), including M. hihihorlo50, M. fasciolatus and M. schevilli51. Six scaffolds were flanked at each end by long, telomere-like repeats, while four scaffolds had telomere-like repeats on a single end (Fig. 1A). Additionally, a substantial drop in scaffold length between the 12th and 13th scaffold (94.13 Mbp vs. 4.75 Mbp) was noted, further suggesting 12 chromosome-level scaffolds.
To compare the contiguity and completeness of the genome to other frog genomes, we calculated several genomic statistics, including benchmarking universal single-copy orthologs (BUSCO)49 using the vertebrata_odb10 lineage. Genome statistics were calculated for two model frog species and the three publicly available scaffolded Australian frog genomes (Table 1). Our BUSCO analysis revealed that our assembly contained 91.8% of complete BUSCOs, 2.9% were present but fragmented, while 5.3% were missing. The proportion of complete BUSCOs in the southern stuttering frog is slightly lower, but still comparable to the model reference X. laevis (95.9%). An unbiased assessment of whole genome completeness with merqury52 showed that our genome assembly was highly complete (93.5%) and accurate (Q60+).
Using Illumina short-read RNA sequencing, we generated six reference-guided, tissue-specific transcriptome assemblies (dorsal skin, ventral skin, liver, spleen, brain, and gonads) that were aligned into a global transcriptome. All tissues had high mapping rates to the genome (78.62–89.37%), with both the tissue-specific and global transcriptomes yielding high complete BUSCO scores (84.0–96.1%; Supplementary Table S2). A total of 36,540 genes were annotated using FGENESH++53. Of these, 16,355 were annotated using evidence from the coding regions of the global transcriptome, similar to the 16,279 predicted coding regions provided as input. A further 12,892 genes were annotated via homology to non-redundant, metazoan, proteins from NCBI, and 7293 were annotated ab initio. Proteins from these annotated genes yielded a complete BUSCO score of 86.0%, comparable to the genome annotation of Limnodynastes dumerilii, and lower than the Xenopus spp. annotations (Supplementary Table S2). However, a large proportion of the stuttering frog annotated genes were fragmented (8.5%), similar to L. dumerilii (8.0%).
Characterisation of cathelicidins and β-defensins
We used the genome and transcriptome assemblies of the southern stuttering frog to characterise two families of AMPs: cathelicidins and β-defensins. These families were selected due to their conservation across vertebrate species54,55, highly conserved structures and motifs56,57,58 and demonstrated antimicrobial potency57,59. We identified 12 cathelicidin (MA-CATH1-12) and two β-defensin (MA-BD1 and MA-BD2) genes in the genome using homology-based search strategies. Full-length transcripts of all cathelicidins and β-defensins were identified in the global transcriptome. All putative AMPs contained the characteristic features of each family, including expected exon number (four for cathelicidins and two for β-defensins), conserved amino acid residues, motifs, and domain structure (Supplementary Figs. S2 and S3). As expected, each AMP family was encoded in clusters within the genome; all cathelicidins were located on scaffold 5 and all β-defensins on scaffold 2 (Supplementary Fig. S4). Cathelicidin and β-defensin genes were named in the order in which they are encoded within the genome (Supplementary Table S3; Fig. S4). Amino acid sequences of all AMPs identified in this study, with predicted signal peptide and mature peptide domains are provided in Supplementary Figs. S2 and S3.
Properties, evolutionary relationships and expression patterns of cathelicidins and β-defensins
The mature peptide domains of cathelicidins and β-defensins are the bioactive, antimicrobial portion of the peptide. In the southern stuttering frog, the mature peptides of the cathelicidins ranged between 10 and 161 amino acids in length (Supplementary Fig. S2), while the two β-defensins were both 47 amino acids (Supplementary Fig. S3). The mature peptides of AMPs generally have a high positive charge which facilitates electrostatic interaction and attachment to microbial cell membranes24. Stuttering frog cathelicidins MA-CATH1, 2 and 3 had a high cationic charge of 8.9, 4.9 and 27.3 respectively at pH 7 (Supplementary Table S4). However, several AMPs identified in this study were weakly cationic (charge 0–3), and MA-CATH8 and MA-CATH2 were anionic. Similarly, while a large proportion (10/14) of the characterised AMPs had > 30% hydrophobic residues, another general trend of AMPs60, some peptides like MA-CATH3 had as few as 12.42% hydrophobic residues (Supplementary Table S4). Amphipathicity is often seen in AMPs to permeabilise microbial membranes60. Through visual inspection of Kyte and Dolittle hydropathicity plots, almost all the southern stuttering frog AMPs exhibited amphipathicity, with the N-terminus of the peptides being generally more hydrophilic than the C-terminus (Supplementary Fig. S5). MA-CATH3 exhibited some regions of hydrophobicity but was largely hydrophilic (Supplementary Fig. S5).
As the first AMPs characterised in the Mixophyes genus, we explored their sequence diversity and relationship to other known frog AMPs. All stuttering frog AMPs displayed low percent identity to other known AMPs by BLAST, with MA-BD2 having the highest percent identity of all AMPs in this study to a Chinese spiny frog (Quasipaa spinosa) β-defensin (64.15%) (Supplementary Table S4). As expected, maximum likelihood phylogenetic trees generated using all known frog cathelicidins and β-defensins reflected this diversity, as indicated by the long branch lengths (Fig. 2). MA-CATH1-3 and MA-CATH4-8 formed two species-specific clades within the cathelicidin tree, some of which were strongly supported with > 95% ultrafast bootstrap support (Fig. 2). MA-CATH9-12 clustered within clades containing AMPs from multiple Asian and African frog species, albeit with low bootstrap support (Fig. 2). While Fig. 3 suggests that the southern stuttering frog β-defensins are more closely related than the other β-defensins, this relationship was not well supported.
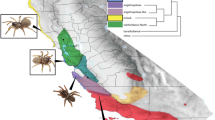
Phylogenetic relationships amongst frog cathelicidins. Tree was generated using the maximum likelihood method, and rooted using four fish cathelicidins as an outgroup. Branches are coloured by ultrafast bootstrap values, with values > 95% in red, and the remaining branches coloured in black. Stuttering frog cathelicidins are coloured green. Cathelicidins are labelled by geographic region; Oceania is green; Asia is blue; North America is pink; Africa is yellow; and Asia and Europe combined is purple. The tree was annotated using MEGA11. For frog cathelicidin sequences used, see Supplementary Table S5.
Phylogenetic relationships amongst frog defensins. Tree was generated using the maximum likelihood method and is unrooted. Branches are coloured by ultrafast bootstrap values, with values > 95% in red, and the remaining branches coloured in black. Stuttering frog defensins are coloured green. Defensins are labelled by geographic region; Oceania is green and Asia is blue. The tree was annotated using MEGA11. For frog defensin sequences used, see Supplementary Table S5.
Our comprehensive bioinformatic AMP discovery approach allowed us to explore the expression pattern of AMPs across a range of tissues beyond skin, which is typically the target of peptidomic approaches to AMP discovery. As expected, we observed high AMP gene expression in the dorsal and ventral skin in our specimen. However, we also observed AMP gene expression in other internal organs (Fig. 4). AMP gene expression was found in the liver, spleen, and gonads, with MA-CATH6 and 8 showing the highest expression. MA-CATH10 and MA-BD2 exhibited higher expression in the skin than other AMPs, while some peptides like MA-CATH12 were lowly expressed across all the tissues when compared to other AMPs (Fig. 4).
Discussion
Here, we used a combination of PacBio HiFi and Hi-C data to generate the first contiguous, annotated, reference genome for the southern stuttering frog. In addition, we assembled the mitochondrial genome and six tissue-specific transcriptomes that were merged into a global transcriptome. The level of resources generated for this single species is comparable with model Xenopus species, as few frogs have a publicly available nuclear genome, mitochondrial genome, as well as transcriptome. By comparing genome and transcriptome quality metrics between other Australian and model assemblies, we have also determined that these resources are comparable in contiguity and completeness to the model Xenopus assemblies. Additionally, they are the first ‘omics level resources in the Mixophyes genus. As the Myobatrachidae family is one of the oldest, most diverse frog families in Australia40, these resources add to our understanding of Australian fauna.
To demonstrate the insights that can be gained from these genomic and transcriptomic resources, we bioinformatically characterised 12 cathelicidins and two β-defensins, the first Australian frog AMPs to be discovered using this approach. While other peptidomics-based studies describing cathelicidins in frogs have revealed at most two per species61,62, 12 cathelicidins were discovered in this study. Similarly, while one β-defensin has been characterised per species in frogs57,58, two were found in this study. In our preliminary gene expression analysis, some of these AMPs, such as MA-CATH1, MA-CATH10 and MA-BD2, were primarily expressed in the dorsal and ventral skin. However, several others had little to no expression in the skin but were primarily expressed in the liver (MA-CATH6), spleen (MA-CATH7-8), and low levels in the brain (MA-BD1). Due to the endangered threat status of the southern stuttering frog, tissue-specific transcriptomes from multiple specimens were not available to determine if these expression patterns are consistent between individuals. However, the number and preliminary expression patterns of the discovered AMPs implies that the peptides that were not expressed in the skin may not have been identified if the more conventional, peptidomics approach of screening skin secretions was used. Our results reveal the benefits of a genomic-based AMP discovery approach and the need for more amphibian genomic resources to characterise such peptides.
It is possible that the evolutionary isolation of Australian amphibians compared to other characterised species has resulted in the diverse suite of cathelicidins and β-defensins observed. Alternatively, the range of cathelicidins and β-defensins characterised here may suggest that the southern stuttering frog is under stronger microbial exposure than other frogs. A previous study on Diptera (fly) species has shown that AMP diversity and gene duplication is positively correlated with microbial exposure25. This is also likely the case in frogs as they adapt to different global aquatic and terrestrial environments. It is more likely, however, that other frog species and genera may also exhibit a similar number and diversity of cathelicidins and β-defensins which are uncharacterised. The lack of bioinformatic AMP discovery for most frog species makes comparative analysis difficult. Future investigations into frog AMP diversity that incorporate a genomic-based discovery platform will facilitate direct comparisons between species.
Our phylogenetic analysis revealed that some southern stuttering frog cathelicidins formed species-specific sister clades to those containing cathelicidins from frogs of Europe, North America, and Asia (Fig. 2). This may indicate that these particular cathelicidins were the result of gene duplication events that occurred after the southern stuttering frogs diverged from the other frog species within the phylogenetic tree. Indeed, several southern stuttering frog cathelicidins have the same signal and cathelin region, but a different mature peptide. Other southern stuttering frog cathelicidins clustered with cathelicidins from European and African frog species (Fig. 2). This suggests these cathelicidins may be the result of gene duplication events in a more distant common ancestor. However, due in part to the great variability in AMP sequences, many of the relationships identified were not strongly supported, particularly for β-defensins, limiting the validity of insights drawn from these trees. It also remains difficult to ascertain whether the species-specific clusters of southern stuttering frog cathelicidins are truly unique to M. australis or the Mixophyes genus more broadly, as there are currently no other characterised cathelicidins from Mixophyes frogs. The number of known cathelicidins and β-defensins across the Anuran order is limited, with only one cathelicidin characterised in a European frog, and no known cathelicidins or β-defensins from South American frog species. As the Myobatrachidae family shares a distant common ancestor with South American frogs40, evolutionary relationships within AMP families across these geographical regions are likely not captured. As more frog cathelicidins and β-defensins are characterised, in particular from Australia and South America, future investigations may better identify the evolutionary patterns of AMP diversity across Mixophyes and other frogs.
The characterised and predicted properties of the stuttering frog AMPs suggest that they may play diverse immunological roles. AMPs are generally cationic and amphipathic due to their electrostatic interactions with anionic glycolipids on prokaryotic membranes, which facilitate membrane permeability and cell lysis63,64. However, AMPs can also exhibit diverse activities beyond antimicrobial activity and may serve other immune functions. For example, an anionic cathelicidin from a salamander species (TK-CATH) had no tested antimicrobial activity, but instead inhibited pro-inflammatory cytokine gene expression when added to mammalian macrophage cell lines65. Two of our AMPs were anionic (MA-CATH8 and MA-BD2; Supplementary Table S4), suggesting that they may have other immune functions. Future investigations will need to validate these in silico findings, such as by synthesising these AMPs and investigating their effects on immune gene expression in a range of cell lines.
While our bioinformatic characterisation of novel AMPs has demonstrated one application of the newly generated genomic and transcriptomic resources, there are numerous other potential applications. For instance, custom DNA metabarcoding markers generated from the mitochondrial genome can now be developed for the southern stuttering frog, contributing to applied conservation outcomes in this threatened species. Metabarcoding has been extensively used to characterise the biodiversity of different environments66. Mitochondrial metagenomics (mtMG) has been previously used to distinguish between closely related species of nematodes67. As the southern stuttering frog is a recently defined species, closely related to the northern population of stuttering frogs (M. balbus)42, the mitochondrial genome generated in this study may be a useful monitoring tool, particularly in defining their range and overlap if applicable. Highly contiguous genomes and transcriptomes can also be used to characterise genomic regions that have high repeat content, such as the major histocompatibility complex (MHC)68. In frogs, the upregulation of MHC class I and II genes in the Montane brown frog (Rana ornativentris) has a functional role in tadpole development69. MHC heterozygosity is also a significant predictor of chytrid fungus resistance in the Lithobates genus70,71. Finally, these resources may be used comparatively with other genomes to investigate conserved and specialised traits across taxa. For example, emerging consortia like the Zoonomia Project have made significant progress in advancing our understanding of mammalian adaptations and evolutionary history72. Synteny analyses have previously been conducted in amphibians, but the variation in sequencing quality and gene annotation methods across the limited existing genomes has made deriving insights difficult73. Comparative studies in Anurans using high-quality genomic resources from representative genera would advance our understanding of a wealth of unique traits; some frog species can survive in extreme temperatures and environments74,75, produce a myriad of toxins76, and regenerate lost appendages as tadpoles77. The incorporation of the southern stuttering frog genome in future studies into amphibians will facilitate a better representation of evolutionarily unique Australian biodiversity in these investigations. In short, generating high-quality multi-omics resources facilitates a plethora of investigations into the southern stuttering frog and amphibians at large.
Methods
Sampling, extractions and sequencing
A wild caught, adult, male stuttering frog was medically euthanised in September 2021 (32° 59′ 52.4″ S 151° 24′ 27.3″ E). Heart and kidney tissue was flash frozen in liquid nitrogen for DNA extraction. Gonads, brain, dorsal skin, ventral skin, liver, and spleen tissue were stored in RNAlater at − 80 °C until RNA extraction. Lethal sampling was conducted under the University of Newcastle Animal Care and Ethics Committee (ACEC Number A-2013-339) and NSW scientific licence (SL190). All methods were performed in accordance with relevant guidelines and regulations; animal research was conducted in compliance with the ARRIVE guidelines.
DNA was extracted from heart and kidney tissue using a Nanobind Tissue Big DNA Kit (Circulomics). Total extracted DNA was verified to be > 20 μg through Qubit fluorometric quantification (ThermoFisher Scientific). DNA was pooled and sequenced at the Australian Genome Research Facility (Brisbane, Queensland, Australia) using a SMRTbell® prep kit 3.0 (PacBio), and circular consensus sequencing (CCS) was performed using three SMRT cells on a PacBio Sequel II system. For Hi-C sequencing, heart and kidney tissue were washed twice for 5 min with 1 × PBS using a rotator wheel at room temperature. Tissues were sequenced at the Biomolecular Resource Facility (Canberra, ACT, Australia), using the Arima Hi-C kit and sequenced as 150-bp paired-end (PE) reads on an Illumina NovaSeq 6000.
RNA was extracted from the six tissues using a Qiagen RNeasy Mini Kit. Concentrations of each sample were confirmed to be ≥ 25 ng/µl using a Nanodrop spectrophotometer (ThermoFisher Scientific) and the RNA integrity number (RIN) measured using the standard Agilent RNA 6000 Nano Kit Protocol and BioAnalyzer (Agilent Technologies). Extractions < 7 RIN were not sequenced. Extracted RNA was prepared at the Ramaciotti Centre for Genomics (Sydney, NSW, Australia), using the Illumina Stranded mRNA Prep Protocol and sequenced as 100-bp PE reads on an Illumina NovaSeq 6000 S1 flowcell.
De novo genome assembly
Over 15 million raw HiFi reads were generated from three SMRT cells. To prevent low quality reads introducing errors to the assembly, reads with Phred (Q) quality score < 20 were filtered out using bamtools v2.4.178. Reads containing adapter sequence were removed by HiFiAdapterFilt v2.0.079 with default parameters. Further details on the computational requirements and estimated run times for all bioinformatic analyses are provided in Supplementary Table S6. Hifiasm80 assembled the remaining reads, alongside paired Hi-C reads, into contigs. To merge the contigs into a scaffolded assembly, we first mapped the Hi-C reads to the unscaffolded genome following the Arima Hi-C mapping pipeline (A160156 v02; https://github.com/ArimaGenomics/mapping_pipeline). YaHS81 was used to merge contigs containing complementary pairs of reads, and the contact map visualised with Juicebox82. One misassembly in the first scaffold was manually corrected. We used MitoHiFi v3.2 to identify and annotate the mitochondrial genome using a closely-related mitogenome as input (Lechriodus melanopyga; NC_019999.1)83,84.
Genome statistics (e.g., N50 and L50 values), were calculated with bbmap v38.86 (https://sourceforge.net/projects/bbmap/). We identified regions matching canonical telomere hexamer repeats (TTAGGG/CCCTAA) using FindTelomeres (https://github.com/JanaSperschneider/FindTelomeres). Benchmarking Universal Single-Copy Orthologs (BUSCO) v5.3.249 analysis was performed on Galaxy Australia85, using the ‘genome’ mode, applying the ‘augustus’ gene-finding setting, and with the vertebrata_odb10 lineage. To compare the gene completeness of the stuttering frog with other genomes, BUSCO analysis was also performed with the same settings on two model frog genomes, the African clawed frog (Xenopus laevis; GCA_017654675.1) and the western clawed frog (Xenopus tropicalis; GCA_000004195.4), as well as the three publicly available scaffolded Australian frog genomes, the southern banjo frog (Limnodynastes dumerilii; GCA_011038615.1), ornate burrowing frog (Platyplectrum ornatum; GCA_016617825.1), and corroboree frog (Pseudophryne corroboree; GCA_028390025.1). An alternative assessment of genome completeness, inclusive of non-coding and repetitive regions, was performed for the stuttering frog and P. ornatum genomes using Merqury v1.352. RepeatModeler v2.0.186 was used to generate a de novo database of the repetitive regions in the stuttering frog genome. We characterised repeats and masked the genome with RepeatMasker v4.0.687. The repeat-masked genome was indexed with hisat2 v2.1.088.
Reference-aligned global transcriptome assembly
Over 900 million PE reads were generated across the six tissue transcriptomes. Low quality sequence calls, adapter and primer sequences were trimmed using Trimmomatic v0.3989 with ILLUMINACLIP:TruSeq3-PE.fa:2:30:10, SLIDINGWINDOW:4:5, LEADING:5, TRAILING:5, and MINLEN:25 settings. The reads from each tissue sample were aligned to the genome with hisat2 v2.1.088. StringTie v2.1.690 was used to merge aligned reads into tissue-specific transcriptomes. We used Transcriptome Annotation by Modular Algorithms (TAMA) v1.091 to combine the tissue-specific transcriptomes into a global transcriptome with adjustments to minimise duplicate transcripts. Briefly, the ‘-d merge_dup’ flag was applied to merge identical transcripts, and the ‘-z 500’ flag was applied to facilitate transcripts with variable 3’ ends (differences of up to 500 bp) to be merged. Transcripts with weak evidence were removed, including transcripts that were found in only one tissue and were lowly expressed (fragments per kilobase of transcript per million fragments mapped [FPKM] < 0.1). CPC2 v2019-11-1992 was used to predict coding regions; non-coding regions were removed. Open reading frames were predicted by TransDecoder v2.0.193. BUSCO v5.3.249 analysis was performed using ‘transcriptome’ mode for both tissue-specific transcriptomes and the global transcriptome, as well as the X. tropicalis transcriptome for comparison.
Genome annotation
The genome was annotated with FGENESH++ v7.2.253 informed by the global transcriptome. Non-mammalian settings were applied throughout the pipeline, and parameters were optimised for Anuran gene discovery by providing the Xenopus gene-finding matrix (Softberry). BUSCO v5.3.249 analysis was performed on the annotation using ‘protein’ mode, as well as for the X. laevis, X. tropicalis and L. dumerilii annotations. The number of genes, exons and introns from the stuttering frog annotated assembly was calculated using the ‘genestats’ script (https://github.com/darencard/GenomeAnnotation/blob/master/genestats).
AMP characterisation
To ensure a comprehensive search of genomic regions that could encode cathelicidins or β-defensins, the annotated genome and global transcriptome were queried using Basic Local Alignment Search Tool (BLAST) v2.2.30+94,95 and HMMER v3.396. A dual program approach has previously been applied in other bioinformatic searches for cathelicidins and β-defensins97,98,99. Further explanation of the approaches used is provided in the Supplementary Extended Methods.
AMP phylogeny
An unrooted phylogenetic tree was generated for the cathelicidin and β-defensin gene families. We first aligned amino acid sequences for each gene family through ClustalW alignments. Both alignments included the full prepropeptide sequences from the stuttering frog and other available frogs (Supplementary Table S5). The cathelicidin alignment also included four fish cathelicidins as an outgroup (Supplementary Table S5). The best-fitting substitution models for each alignment were selected using the ModelFinder option in IQ-TREE v2.2.0100 according to the Bayesian information criterion101. For the cathelicidin alignment, the Jones Taylor Thornton (JTT) substitution model102 was optimal, incorporating Invariant Sites (+ I) and four components of gamma rate heterogeneity (+ G4). For the β-defensin alignment, the Dayhoff model was optimal103. Maximum likelihood analysis was conducted in IQ-TREE v2.2.0100 with node support estimated by ultrafast bootstrap approximation with 1000 replicates104. The results were visualised and annotated using MEGA11105.
Characterisation and prediction of AMP structure, properties and expression
Cathelicidins are made up of a signal peptide, a conserved cathelin pro-region, as well as the mature, bioactive peptide56,106. To predict the signal peptide from the full prepropeptide sequence, the SignalP v6.0 webserver was used107 with the ‘Eukarya’ organism setting and the ‘Slow’ model mode. While the enzyme used to cleave the mature peptide from the cathelin pro-region is not known for amphibians, there is experimental evidence to support proprotein convertases or trypsin-like proteases that cleave at dibasic residues like lysine (K) and arginine (R)61,108,109. Therefore, a two-tiered approach predicted the mature peptide of the novel cathelicidins. Proprotein convertase cleavage sites were first predicted from the last exon of the cathelicidins using ProP v1.0110. If no cleavage was identified, trypsin cleavage sites were predicted using ExPASy’s peptide cutter tool111. For β-defensins, which generally consist of a signal peptide and a mature, bioactive peptide112,113,114, the signal peptide was predicted using SignalP, and the remaining peptide was annotated as the mature peptide.
The molecular weight and charge at pH 7 for each putative stuttering frog cathelicidin and β-defensin was calculated using Protein Calculator v3.4 (https://protcalc.sourceforge.net/). The percentage of hydrophobic residues in the AMPs was calculated on Peptide v2.0 (https://www.peptide2.com/) using the ‘Peptide Hydrophobicity/Hydrophilicity Analysis’ tool. The amphipathicity of the AMPs was determined by generating Kyte and Doolittle hydropathicity plots on ExPASy using Protscale111, with a window size of 5. These plots were inspected for the presence of ‘peaks’ and ‘troughs’, which indicate sections in the AMP of high and low hydropathicity115. For the expression analysis, the Transcripts Per Million (TPM) values for each AMP was generated from the tissue-specific transcriptomes using StringTie v2.1.690.
Data availability
The genome assembly, mitochondrial genome, and raw transcriptome sequencing data generated in this study have been submitted to the NCBI BioProject database (https://www.ncbi.nlm.nih.gov/bioproject/) under accession number PRJNA991157. Raw genome sequencing data is publicly available through the Bioplatforms Australia Threatened Species Initiative (https://data.bioplatforms.com/organization/threatened-species). The global transcriptome and annotation generated in this study are available on Amazon Web Services Australasian Genomes Open Data Store (https://awgg-lab.github.io/australasiangenomes/genomes.html).
References
Lewin, H. A. et al. Earth BioGenome Project: Sequencing life for the future of life. Proc. Natl. Acad. Sci. 115(17), 4325–4333 (2018).
Lewin, H. A. et al. The Earth BioGenome Project 2020: Starting the clock. Proc. Natl. Acad. Sci. U S A 119(4), e2115635118 (2022).
Hotaling, S., Kelley, J. & Frandsen, P. Toward a genome sequence for every animal: Where are we now?. Proc. Natl. Acad. Sci. 118, e2109019118 (2021).
Horgan, R. & Kenny, L. ‘Omic’ technologies: Genomics, transcriptomics, proteomics and metabolomics. Obstet. Gynaecol. 13, 189 (2011).
Formenti, G. et al. The era of reference genomes in conservation genomics. Trends Ecol. Evol. 37, 197 (2022).
Paez, S. et al. Reference genomes for conservation. Science 377(6604), 364–366 (2022).
Wong, A. K. et al. Decoding disease: From genomes to networks to phenotypes. Nat. Rev. Genet. 22(12), 774–790 (2021).
Sayers, E. W. et al. Database resources of the national center for biotechnology information. Nucleic Acids Res. 50(D1), D20-d26 (2022).
AmphibiaWeb. Amphibian Species by the Numbers (University of California, 2022).
Lamichhaney, S. et al. A bird-like genome from a frog: Mechanisms of genome size reduction in the ornate burrowing frog, Platyplectrum ornatum. Proc. Natl. Acad. Sci. USA 118(11), e2011649118 (2021).
Li, Q. et al. A draft genome assembly of the eastern banjo frog Limnodynastes dumerilii dumerilii (Anura: Limnodynastidae). Gigabyte 2020, 1–13 (2020).
Rhie, A. et al. Towards complete and error-free genome assemblies of all vertebrate species. Nature 592(7856), 737–746 (2021).
Farquharson, K. et al. The genome sequence of the critically endangered Kroombit tinkerfrog (Taudactylus pleione) [version 1; peer review: 2 approved]. F1000Research 12, 845 (2023).
Bredeson, J. V. et al. Conserved chromatin and repetitive patterns reveal slow genome evolution in frogs. Nat. Commun. 15(1), 579 (2024).
Liedtke, H. C. et al. Macroevolutionary shift in the size of amphibian genomes and the role of life history and climate. Nat. Ecol. Evol. 2(11), 1792–1799 (2018).
Sun, Y.-B., Zhang, Y. & Wang, K. Perspectives on studying molecular adaptations of amphibians in the genomic era. Zool. Res. 41(4), 351 (2020).
Seidl, F. et al. Genome of Spea multiplicata, a rapidly developing, phenotypically plastic, and desert-adapted spadefoot toad. G3 Genes Genomes Genetics 9(12), 3909–3919 (2019).
Novikova, P. Y. et al. Polyploidy breaks speciation barriers in Australian burrowing frogs Neobatrachus. PLoS Genet. 16(5), e1008769 (2020).
Session, A. M. et al. Genome evolution in the allotetraploid frog Xenopus laevis. Nature 538(7625), 336–343 (2016).
Pollard, M. O. et al. Long reads: Their purpose and place. Hum. Mol. Genet. 27(R2), R234-r241 (2018).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326(5950), 289–293 (2009).
Streicher, J. W. The genome sequence of the common frog, Rana temporaria Linnaeus 1758. Wellcome Open Res. 6, 286 (2021).
Wang, G., Li, X. & Wang, Z. APD3: The antimicrobial peptide database as a tool for research and education. Nucleic Acids Res. 44(D1), D1087–D1093 (2016).
Huan, Y. C. et al. Antimicrobial peptides: Classification, design, application and research progress in multiple fields. Front. Microbiol. 11, 582779 (2020).
Hanson, M. A., Lemaitre, B. & Unckless, R. L. Dynamic evolution of antimicrobial peptides underscores trade-offs between immunity and ecological fitness. Front. Immunol. 10, 2620 (2019).
Mercer, D. K. et al. NP213 (Novexatin®): A unique therapy candidate for onychomycosis with a differentiated safety and efficacy profile. Med. Mycol. 58(8), 1064–1072 (2020).
Ridyard, K. E. & Overhage, J. The potential of human peptide LL-37 as an antimicrobial and anti-biofilm agent. Antibiotics 10(6), 650 (2021).
Wang, Q. et al. Diversity of antimicrobial peptides in three partially sympatric frog species in Northeast Asia and implications for evolution. Genes (Basel) 11(2), 158 (2020).
Varga, J. F. A., Bui-Marinos, M. P. & Katzenback, B. A. Frog skin innate immune defences: Sensing and surviving pathogens. Front. Immunol. 9, 3128 (2018).
Ladram, A. & Nicolas, P. Antimicrobial peptides from frog skin: Biodiversity and therapeutic promises. Front. Biosci. Landmark 21, 1341–1371 (2016).
Novković, M. et al. DADP: The database of anuran defense peptides. Bioinformatics 28(10), 1406–1407 (2012).
Yang, Y. et al. A non-bactericidal cathelicidin provides prophylactic efficacy against bacterial infection by driving phagocyte influx. eLife 11, e72849 (2022).
Hao, X. et al. Amphibian cathelicidin fills the evolutionary gap of cathelicidin in vertebrate. Amino Acids 43(2), 677–685 (2012).
He, X. et al. A frog-derived immunomodulatory peptide promotes cutaneous wound healing by regulating cellular response. Front. Immunol. 10, 2421 (2019).
Peel, E. et al. Cathelicidins in the Tasmanian devil (Sarcophilus harrisii). Sci. Rep. 6, 35019 (2016).
Dalla Valle, L. et al. Bioinformatic and molecular characterization of beta-defensins-like peptides isolated from the green lizard Anolis carolinensis. Dev. Comp. Immunol. 36(1), 222–229 (2012).
Wang, M. et al. Identification and characterization of antimicrobial peptides from butterflies: An integrated bioinformatics and experimental study. Front. Microbiol. 12, 720381 (2021).
Pérez de la Lastra, J. M. et al. Bioinformatic analysis of genome-predicted bat cathelicidins. Molecules 26(6), 1811 (2021).
Yoo, W. G. et al. Genome-wide identification of antimicrobial peptides in the liver fluke, Clonorchis sinensis. Bioinformation 11(1), 17–20 (2015).
Brennan, I. G. et al. Populating a continent: Phylogenomics reveal the timing of Australian frog diversification. Syst. Biol. https://doi.org/10.1093/sysbio/syad048 (2023).
Irisarri, I. et al. The origin of modern frogs (Neobatrachia) was accompanied by acceleration in mitochondrial and nuclear substitution rates. BMC Genomics 13(1), 626 (2012).
Mahony, M. et al. A new species of barred frog, Mixophyes (Anura: Myobatrachidae) from south-eastern Australia identified by molecular genetic analyses. Zootaxa 5297, 301–336 (2023).
Barker, J., Grigg, G. & Tyler, M. J. A Field Guide to Australian frogs (Surrey Beatty and Sons, 1995).
Cogger, H. G. Reptiles and Amphibians of Australia 6th edn. (Reed New Holland, 2000).
Murray, B. R. & Hose, G. C. Life-history and ecological correlates of decline and extinction in the endemic Australian frog fauna. Austral Ecol. 30(5), 564–571 (2005).
Woodhams, D. C. et al. Resistance to chytridiomycosis varies among amphibian species and is correlated with skin peptide defenses. Anim. Conserv. 10(4), 409–417 (2007).
Hollanders, M. et al. Recovered frog populations coexist with endemic Batrachochytrium dendrobatidis despite load-dependent mortality. Ecol. Appl. 33(1), e2724 (2023).
Grant, J. R. et al. Proksee: In-depth characterization and visualization of bacterial genomes. Nucleic Acids Res. 51(W1), W484-w492 (2023).
Simão, F. A. et al. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31(19), 3210–3212 (2015).
Donnellan, S. C., Mahony, M. J. & Davies, M. A new species of mixophyes (Anura: Leptodactylidae) and first record of the genus in New Guinea. Herpetologica 46(3), 266–274 (1990).
Schmid, M. et al. Chromosome banding in Amphibia. XXV. Karyotype evolution and heterochromatin characterization in Australian Mixophyes (Anura, Myobatrachidae). Cytogenet. Genome Res. 97(3–4), 239–53 (2002).
Rhie, A. et al. Merqury: Reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21(1), 245 (2020).
Solovyev, V. et al. Automatic annotation of eukaryotic genes, pseudogenes and promoters. Genome Biol. 7(1), S10 (2006).
Kosciuczuk, E. M. et al. Cathelicidins: Family of antimicrobial peptides. A review. Mol. Biol. Rep. 39(12), 10957–10970 (2012).
Tu, J. et al. Molecular evolutionary analysis of β-defensin peptides in vertebrates. Evol. Bioinform. Online 11, 105–114 (2015).
Chen, J. et al. Molecular characterization of cathelicidin in tiger frog (Hoplobatrachus rugulosus): Antimicrobial activity and immunomodulatory activity. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 247, 109072 (2021).
Shen, W. et al. A novel defensin-like antimicrobial peptide from the skin secretions of the tree frog, Theloderma kwangsiensis. Gene 576(1, Part 1), 136–140 (2016).
Yu, S. S. et al. Antimicrobial and immunomodulatory activity of beta-defensin from the Chinese spiny frog (Quasipaa spinosa). Dev. Comp. Immunol. 126, 104264 (2022).
Wei, L. et al. The first anionic defensin from amphibians. Amino Acids 47(7), 1301–1308 (2015).
Chen, C. H. & Lu, T. K. Development and challenges of antimicrobial peptides for therapeutic applications. Antibiotics 9(1), 24 (2020).
Yu, H. et al. Identification and polymorphism discovery of the cathelicidins, Lf-CATHs in ranid amphibian (Limnonectes fragilis). FEBS J. 280(23), 6022–6032 (2013).
Ling, G. et al. Cathelicidins from the Bullfrog Rana catesbeiana provides novel template for peptide antibiotic design. PLoS ONE 9(3), e93216 (2014).
Hancock, R. E. W. & Rozek, A. Role of membranes in the activities of antimicrobial cationic peptides. FEMS Microbiol. Lett. 206(2), 143–149 (2002).
Rivas, L., Luque-Ortega, J. R. & Andreu, D. Amphibian antimicrobial peptides and Protozoa: Lessons from parasites. Biochim. Biophys. Acta Biomembranes 1788(8), 1570–1581 (2009).
Luo, X. J. et al. A novel anionic cathelicidin lacking direct antimicrobial activity but with potent anti-inflammatory and wound healing activities from the salamander Tylototriton kweichowensis. Biochimie 191, 37–50 (2021).
Ruppert, K. M., Kline, R. J. & Rahman, M. S. Past, present, and future perspectives of environmental DNA (eDNA) metabarcoding: A systematic review in methods, monitoring, and applications of global eDNA. Glob. Ecol. Conserv. 17, e00547 (2019).
Gendron, E. M. et al. Nematode mitochondrial metagenomics: A new tool for biodiversity analysis. Mol. Ecol. Resour. 23(5), 975–989 (2023).
Vekemans, X. et al. Whole-genome sequencing and genome regions of special interest: Lessons from major histocompatibility complex, sex determination, and plant self-incompatibility. Mol. Ecol. 30(23), 6072–6086 (2021).
Lau, Q. et al. Expression changes of MHC and other immune genes in frog skin during ontogeny. Animals 10(1), 91 (2020).
Savage, A. & Zamudio, K. MHC genotypes associate with resistance to a frog-killing fungus. Proc. Natl. Acad. Sci. USA 108, 16705–16710 (2011).
Trujillo, A. L. et al. Spatiotemporal adaptive evolution of an MHC immune gene in a frog-fungus disease system. Heredity 126(4), 640–655 (2021).
Christmas, M. J. et al. Evolutionary constraint and innovation across hundreds of placental mammals. Science 380(6643), eabn3943 (2023).
Kosch, T. A. et al. Comparative analysis of amphibian genomes: An emerging resource for basic and applied research. bioRxiv https://doi.org/10.1101/2023.02.27.530355 (2023).
Larson, D. J. et al. Wood frog adaptations to overwintering in Alaska: New limits to freezing tolerance. J. Exp. Biol. 217(12), 2193–2200 (2014).
van Beurden, E. K. Energy metabolism of dormant Australian water-holding frogs (Cyclorana platycephalus). Copeia 1980(4), 787–799 (1980).
Raaymakers, C. et al. Antimicrobial peptides in frog poisons constitute a molecular toxin delivery system against predators. Nat. Commun. 8(1), 1495 (2017).
Aztekin, C. et al. Identification of a regeneration-organizing cell in the Xenopus tail. Science 364(6441), 653–658 (2019).
Barnett, D. W. et al. BamTools: A C++ API and toolkit for analyzing and managing BAM files. Bioinformatics 27(12), 1691–1692 (2011).
Sim, S. B. et al. HiFiAdapterFilt, a memory efficient read processing pipeline, prevents occurrence of adapter sequence in PacBio HiFi reads and their negative impacts on genome assembly. BMC Genomics 23(1), 157 (2022).
Cheng, H. et al. Haplotype-resolved de novo assembly using phased assembly graphs with Hifiasm. Nat. Methods 18(2), 170–175 (2021).
Zhou, C., McCarthy, S. A. & Durbin, R. YaHS: Yet another Hi–C scaffolding tool. Bioinformatics 39(1), btac808 (2022).
Durand, N. C. et al. Juicebox provides a visualization system for Hi–C contact maps with unlimited zoom. Cell Syst. 3(1), 99–101 (2016).
Allio, R. et al. MitoFinder: Efficient automated large-scale extraction of mitogenomic data in target enrichment phylogenomics. Mol. Ecol. Resour. 20(4), 892–905 (2020).
Uliano-Silva, M. et al. MitoHiFi: A python pipeline for mitochondrial genome assembly from PacBio high fidelity reads. BMC Bioinform. 24(1), 288 (2023).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46(W1), W537–W544 (2018).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. 117(17), 9451–9457 (2020).
Smit, A., Hubley, R., Green, P. RepeatMasker Open-4.0. 2013–2015.
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 12(4), 357–360 (2015).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30(15), 2114–2120 (2014).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33(3), 290–295 (2015).
Kuo, R. I. et al. Illuminating the dark side of the human transcriptome with long read transcript sequencing. BMC Genomics 21(1), 751 (2020).
Kang, Y.-J. et al. CPC2: A fast and accurate coding potential calculator based on sequence intrinsic features. Nucleic Acids Res. 45(W1), W12–W16 (2017).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protocols 8(8), 1494–1512 (2013).
Altschul, S. F. et al. Basic local alignment search tool. J. Mol. Biol. 215(3), 403–410 (1990).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 25(17), 3389–3402 (1997).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7(10), e1002195 (2011).
Whittington, C. M. et al. Defensins and the convergent evolution of platypus and reptile venom genes. Genome Res. 18(6), 986–994 (2008).
Schutte, B. C. et al. Discovery of five conserved beta-defensin gene clusters using a computational search strategy. Proc. Natl. Acad. Sci. USA 99(4), 2129–2133 (2002).
Helbing, C. C. et al. Antimicrobial peptides from Rana [Lithobates] catesbeiana: Gene structure and bioinformatic identification of novel forms from tadpoles. Sci. Rep. 9(1), 1529 (2019).
Minh, B. Q. et al. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37(5), 1530–1534 (2020).
Kalyaanamoorthy, S. et al. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 14(6), 587–589 (2017).
Jones, D. T., Taylor, W. R. & Thornton, J. M. The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. 8(3), 275–282 (1992).
Dayhoff, M., Schwartz, R. & Orcutt, B. A model of evolutionary change in proteins. In Atlas of Protein Sequence and Structure (ed. Dayhoff, M.) 345–352 (National Biomedical Research Foundation, 1978).
Hoang, D. T. et al. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 35(2), 518–522 (2017).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38(7), 3022–3027 (2021).
Tomasinsig, L. & Zanetti, M. The cathelicidins—Structure, function and evolution. Curr. Protein Pept. Sci. 6(1), 23–34 (2005).
Teufel, F. et al. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat. Biotechnol. 40(7), 1023–1025 (2022).
Shi, Y. et al. Cathelicidin-DM is an antimicrobial peptide from Duttaphrynus melanostictus and has wound-healing therapeutic potential. ACS Omega 5(16), 9301–9310 (2020).
Mu, L. et al. The first identified cathelicidin from tree frogs possesses anti-inflammatory and partial LPS neutralization activities. Amino Acids 49(9), 1571–1585 (2017).
Duckert, P., Brunak, S. & Blom, N. Prediction of proprotein convertase cleavage sites. Protein Eng. Des. Sel. 17(1), 107–112 (2004).
Gasteiger, E. et al. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 31(13), 3784–3788 (2003).
Huttner, K. M. & Bevins, C. L. Antimicrobial peptides as mediators of epithelial host defense. Pediatr. Res. 45(6), 785–794 (1999).
Tang, Y. Q. & Selsted, M. E. Characterization of the disulfide motif in BNBD-12, an antimicrobial beta-defensin peptide from bovine neutrophils. J. Biol. Chem. 268(9), 6649–6653 (1993).
Semple, F. & Dorin, J. R. β-Defensins: Multifunctional modulators of infection, inflammation and more?. J. Innate Immunity 4(4), 337–348 (2012).
Peel, E. et al. Marsupial and monotreme cathelicidins display antimicrobial activity, including against methicillin-resistant Staphylococcus aureus. Microbiology 163(10), 1457–1465 (2017).
Acknowledgements
We acknowledge the traditional custodians of the land where the specimen was collected, the Awabakal people, and pay respects to their elders past and present, and their ongoing connection to country. Thank you to M. Mahony, University of Newcastle, who collected the specimen. Thanks also to Elspeth McLennan for assistance with the DNA and RNA extractions. We thank the two reviewers for comments that improved the manuscript. Computational resources were provided by Amazon Web Services and RONIN; the Australian FGENESH++ Service provided by the Australian Biocommons and the Pawsey Supercomputing Research Centre with funding from the Australian Government and the Government of Western Australia; Galaxy Australia, a service provided by the Australian Biocommons and its partners; and the University of Sydney’s High Performance Computing facility Artemis provided by the Sydney Informatics Hub. The authors wish to acknowledge the use of the services and facilities of the Australian Genome Research Facility; the Ramaciotti Centre for Genomics, UNSW; and the Biomolecular Resource Facility, Canberra. Funding was provided by the Threatened Species Initiative through a grant from the Australian Government Bushfire Recovery Scheme (ERF-WRR2-020), and the Australian Research Council Centre of Excellence for Innovations in Peptide and Protein Science (CE200100012).
Author information
Authors and Affiliations
Contributions
The project was conceived, and funding sought, by C.J.H. and K.B. S.T. undertook the genome and transcriptome assembly with assistance from K.A.F. S.T. characterised the AMPs with assistance from K.A.F. and E.P. S.T. drafted the manuscript with assistance from K.A.F. All authors provided review and feedback.
Corresponding author
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tang, S., Peel, E., Belov, K. et al. Multi-omics resources for the Australian southern stuttering frog (Mixophyes australis) reveal assorted antimicrobial peptides. Sci Rep 14, 3991 (2024). https://doi.org/10.1038/s41598-024-54522-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-54522-x
- Springer Nature Limited