Abstract
The COVID-19 influenza emerged and proved to be fatal, causing millions of deaths worldwide. Vaccines were eventually discovered, effectively preventing the severe symptoms caused by the disease. However, some of the population (elderly and patients with comorbidities) are still vulnerable to severe symptoms such as breathlessness and chest pain. Identifying these patients in advance is imperative to prevent a bad prognosis. Hence, machine learning and deep learning algorithms have been used for early COVID-19 severity prediction using clinical and laboratory markers. The COVID-19 data was collected from two Manipal hospitals after obtaining ethical clearance. Multiple nature-inspired feature selection algorithms are used to choose the most crucial markers. A maximum testing accuracy of 95% was achieved by the classifiers. The predictions obtained by the classifiers have been demystified using five explainable artificial intelligence techniques (XAI). According to XAI, the most important markers are c-reactive protein, basophils, lymphocytes, albumin, D-Dimer and neutrophils. The models could be deployed in various healthcare facilities to predict COVID-19 severity in advance so that appropriate treatments could be provided to mitigate a severe prognosis. The computer aided diagnostic method can also aid the healthcare professionals and ease the burden on already suffering healthcare infrastructure.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
The COVID-19 began in late 2019 and caused a significant uproar worldwide1. Most patients experienced mild-moderate symptoms such as cough, cold, myalgia, sore throat, muscle pain, nausea, loss of taste/smell and headaches. However, people also developed severe symptoms such as accurate respiratory disorder syndrome (ARDS), severe hypoxia and multi-organ failure and succumbed to this deadly disease2. As of today, the virus is still spreading, and new mutations are being created. Cytokine storm manifests in COVID-19 patients, distinguished by an enormous release of cytokines such as IL-6 and IL-1. This condition has led to the immune system attacking itself and has caused deaths in many Sars-Cov-2 patients3.
The severe symptoms of COVID-19 have decreased after the introduction of vaccines4. However, some COVID-19 patients are still vulnerable to severe prognoses3. Older patients and people with comorbidities such as hypertension, diabetes, cancer etc., are still at risk. It is crucial to identify these patients early so that appropriate medications and treatments can be provided to them to avoid unnecessary casualties. A few drugs have been created and shown to prevent the onset of severe COVID-19 symptoms3. These medicines must be administered during the illness's initial stages to be effective.
Artificial Intelligence (AI) applications have been extensively utilized in the healthcare sector5,6,7. Diagnostic and prognostic models, decision support systems and predictive modelling are being developed to assist healthcare professionals using machine learning (ML). The above technologies are also being used in the fight against COVID-198,9,10. Explainable artificial intelligence (XAI) makes the models more transparent and understandable. The reasoning behind a patient prediction can be visually represented using XAI. It has also been utilized in various domains such as finance, engineering, pharmacy, medicine and commerce.
A few markers such as c-reactive protein (CRP), D-Dimer, lactate dehydrogenase (LDH), neutrophil to lymphocyte ratio (NLR) and Ferritin are known to change excessively before the actual onset of the severe symptoms2. Machine learning models can be deployed using the markers to predict COVID-19 severity in advance. The early detection of patients with poor prognoses and the development of reliable forecasting techniques that are simple to use in routine clinical practice are crucial for ensuring the highest level of treatment in clinics.
Several researchers have utilized machine learning to predict COVID-19 severity using hematological parameters. Huyut11 developed an automatic decision support system to predict mild and severe coronavirus patients. The dataset consisted of 194 severe and 4010 mild patients. Twenty-nine markers were considered, and the local weighted algorithm obtained a maximum accuracy of 97.86%. Wendland et al.12 used classifiers to predict severe COVID-19 cases. They were able to predict the severity status with an AUC of 0.918. The most important markers in this study were CRP and blood sugar levels. A COVID-19 severity prediction model was developed by Nguyen et al.13. Two hundred sixty-one patients from Vietnam were considered for this research. The random forest obtained a accuracy of 97%. The best prognostic markers were CRP, IL-6, dyspnea, D-Dimer and ferritin. A nature-inspired was developed to predict COVID-19 severity14. The study used details of 65,000 patients, which consisted of twenty-six features. A variant of the artificial bee colony algorithm was used for feature selection. Among all the algorithms, the support vector machine obtained an accuracy of 96%. The model categorized the patients into mild, moderate and severe. Laatifi et al.15 used two explainable AI techniques to predict COVID-19 severity. Eighty-seven patients were considered in this study. Shapley additive values (SHAP) and local interpretable model-agnostic explanations (LIME) were used to make the models understandable. The most critical cytokine markers are VEGF-A and IL-7.
COVID-19 vaccines have been successful in preventing severe COVID-19 in most patients. However, a small part of the population still experiences severe symptoms. It is of utmost importance to prevent the onset of severe prognosis in these patients. The machine learning models can be beneficial in predicting the same.
The above studies show that COVID-19 severity could be predicted effectively using clinical and laboratory markers. The main objective of this research is to forecast the severity of a COVID-19 patient. The other contributions are given below:
-
Descriptive statistical analysis of the data has been conducted to understand various trends and patterns in the data.
-
Fourteen feature selection methods including nature-inspired algorithms have been used to choose the most important markers.
-
Machine learning models including bagging, boosting, voting and stacking have been used to predict COVID-19 severity. The classifiers have been further compared to with the state-of-the-art deep learning models such as deep neural network (DNN), one-dimensional convolutional neural network (1D-CNN) and Long short-term memory (LSTM).
-
Five XAI techniques have been used to interpret the predictions such as SHAP, LIME, Eli5, QLattice and Anchor.
-
Further discussion about crucial COVID-19 prognostic markers from a medical perspective.
The reminder of the paper is structured as follows. Materials and methods are described in “Methods” section. Extensive explanation of the results is made in “Results” section. The discussion of the results obtained is made in “Discussion” section. The article concludes in “Conclusion” section.
Methods
Description of the dataset
The COVID-19 datasets were obtained from two Hospitals in India: Dr TMA Pai Hospital and Kasturba Medical College. The Manipal Academy of Higher Education has provided ethical clearance to conduct this research (IEC:613/2021). The patients have been completely anonymized in this study. COVID-19 patients who were tested between September 2021 and December 2021 have been considered in this study. Only patients above eighteen years of age have been included. Records of 899 patients have been utilized to train the machine learning models. The dataset included 599 non-severe patients and 300 severe patients. All patients whose condition deteriorated and required admission to the intensive care unit (ICU) and if the respiratory rate > 30/minute or SpO2 < 90% (World Health Organization standards) were grouped as severe cases16. Thirty-two clinical parameters were considered in this study (31—continuous and one categorical). The clinical markers chosen are tabulated in Table 1.
Data pre-processing
Pre-processing of the dataset is critical in machine learning. Missing values are imputed, categorical attributes are encoded, continuous values are scaled, data balancing is performed, and unnecessary attributes are dropped. In order to make sure that there are as few missing values as possible, we chose patients who completed the most clinical tests when gathering data. A few missing values in the dataset were replaced by their respective median. The “gender” attribute (categorical) had no missing values. Descriptive statistical analysis was conducted using the open-source statistical software Jamovi. Some statistical parameters utilized are described in Table 2.
Violin plots were used to find interesting patterns in the dataset, as shown in Fig. 1. From the figure, it can be seen that the median age was elevated in the severe COVID-19 cohort. Further, markers such as Neutrophils, HbA1c and CRP were elevated in severe patients. The lymphocytes and monocytes count decreased in the severe COVID-19 cohort.
The frequency of the “gender” attribute for severe/non-severe COVID-19 patients is described using a bar plot in Fig. 2. There were 347 male and 252 female patients in the non-severe cohort. There were 204 male patients and 96 female patients in the severe cohort.
In machine learning analysis, categorical values must be encoded since the classifiers do not handle text values. Several encoding techniques exist in machine learning17. In this study, we used the one-hot encoding technique to encode the “Gender” attribute18. This encoding mechanism solves the problem of ordinality, which can happen in categorical variables. Data scaling was performed using the standardization method19. When there is a considerable discrepancy in data points, the accuracy decreases. The classifiers also favour parameters with higher values, regardless of the units considered. Normalization and standardization are the two approaches utilized to scale the datasets in machine learning. Standardization was chosen in this study since they are better with outliers. The dataset was then split into training and testing in the ratio (80:20). There was a significant imbalance in the dataset. The number of severe COVID-19 cases was almost half compared to non-COVID-19 cases. The results obtained for the unbalanced data are completely biased since the models favour the majority classes. Hence, we used the oversampling technique called Borderline Synthetic Minority Oversampling Technique (SMOTE) to balance the training dataset20. This algorithm generates new synthetic samples using the K-nearest algorithm. The borderline cases are also handled well using the above technique. Under-sampling was not preferred in this study since we did not want to lose interesting trends and patterns. Further, the testing data was not balanced to protect data integrity.
Fourteen feature selection methods were used to choose the most important markers. Several metaheuristic nature-inspired algorithms have been utilized in this study. Feature selection is essential in machine learning since the classifiers perform better when removing redundant features. In this article, we have chosen multiple nature inspired algorithms. They have several advantages over traditional feature selection techniques. They are known for their global optimization, robustness, scalability, parallelism, adaptability, simplicity and stochasticity. Table 3 describes the features chosen by each algorithm. Among all the algorithms, the salp swarm optimization chose the maximum number of features (18). The whale optimization algorithm, flower pollination algorithm and mutual information chose 15 features. The sine cosine algorithm chose the minimum number of features (3). The Harris Hawk’s optimization and particle swarm optimization chose six features each. The markers chosen by the feature selection techniques are also described in Fig. 3. CRP was the most chosen feature since thirteen algorithms have included it. This was followed by neutrophils, NLR and AST, which were chosen 10, 9 and 8 times, respectively. The marker platelets were not chosen by any algorithm.
Machine learning concepts
Machine learning is a form of artificial intelligence that enables software programs to forecast predictive outcomes using past information as input. Several ML classifiers have been used in this study, such as random forest, decision tree, logistic regression, K nearest neighbors, catboost, adaboost, xgboost, lightgbm, stacking and voting algorithms. Stacking combines the result of multiple baseline models35. The stacking architecture consists of a classifier incorporating the initial model’s predictions. Aggregation of the models are performed based on their weights, improving the model’s accuracy. The meta-learner becomes a crucial factor in stacking. Logistic regression was the meta-learner used in this research. The stacking architecture is described in Fig. 4.
A voting classifier gathers training data from a large ensemble of classifiers, and predictions are made according to the class with the highest probability. It uses the concept of majority voting36. The voting algorithm is of two types: Hard-voting and soft-voting. The maximum number of votes is considered in hard-voting irrespective of the weights37. “Average probability” predicts the outcome of soft-voting38. The voting architecture is described in Fig. 5.
Further, the data was subjected to a fivefold cross validation technique. Here, various subsets of data are trained to validate the model efficiency. The input data is divided into five equal groups. Four groups are used for training, while the fifth group is used for testing using various permutations and combinations in cross-validation. Hyperparameter tuning was performed to choose the best parameters using the grid search method. The performance of a classifier depends upon the hyperparameters chosen. Grid search automates the hyperparameter tuning and provides the best values as output.
We have chosen several classification and loss metrics to evaluate the models in this study. These include precision, recall, accuracy, F1-score, area under curve (AUC), average precision (AP), Mathew’s correlation, log loss, Jaccard score and hamming loss. Emphasis has been given to precision and recall since they focus on false-positive and false-negative cases.
In this research, three state-of-the-art deep learning models have been tested. They are DNN, 1D-CNN and LSTM. A DNN consists of multiple input, hidden and output layers39. The essential function of a deep neural network is to take input, process them through more sophisticated computations, and predict results. CNNs are primarily used for image classification. However, 1D-CNN models have also been highly influential in classifying tabular data40. LSTMs are highly used in sequence prediction problems41. Three types of gates are considered in LSTM: input gate, output gate, and forget gate. LSTMs have proven to be highly efficient in handling time series data.
After training and testing the ML and DL models, five XAI techniques have been used to demystify the predictions. The results obtianed by the XAI techniques are in the form of graphs and tables, which can be easily understood by the ML users. The entire process-flow of this study is described in Fig. 6.
Ethical approval
Ethical clearance has been obtained to collect patient data from Manipal Academy of Higher Education ethics committee with id IEC: 613/2021. The need for informed consent was waived by the ethics committee/Institutional Review Board of Manipal Academy of Higher Education, because of the retrospective nature of the study. All methods were carried out in accordance with relevant guidelines and regulations.
Results
Model testing
In this research, multiple machine learning and deep learning classifiers have been trained and tested to predict COVID-19 severity. The precision obtained by the models for various feature selection techniques is tabulated in Table 4. We emphasized the stacking and voting classifiers since they combine multiple models. From the table, it can be seen that the stacked model obtained the maximum precision of 94% after using mutual information. The soft-voting and hard-voting obtained a precision of 94% each. The bat algorithm performed well too. The stack, hard-voting and soft-voting classifier obtained a precision of 91%, 91% and 90%, respectively. The flower pollination algorithm was also efficient. The stack, hard-voting and soft-voting obtained a precision of 87%, 86% and 84%, respectively. The precision obtained for the stack, hard-voting and soft-voting after using the Jaya algorithm was 87%, 90% and 89%, respectively.
The recall obtained by the models for all the feature selection techniques is described in Table 5. Mutual information was the best feature selection method. The recall obtained by the stack, hard-voting and soft-voting algorithms were 93%, 95% and 94%, respectively. The bat algorithm was the next best-performing model. The recall obtained by the stack, hard-voting and soft-voting models were 90%, 93% and 91%, respectively. The flower pollination algorithm performed well too. The recall obtained by the stack, hard-voting and soft-voting models were 86%, 90% and 90%, respectively. The recall obtained by the stack, hard-voting and soft-voting classifiers after using the Jaya algorithm was 87%, 91% and 90%, respectively. For further analysis, the best four feature selection techniques were considered. They are mutual information, bat algorithm, flower pollination algorithm and Jaya algorithm.
The classification and the loss metrics are tabulated in Table 6. Mutual information performed the best among the four methods. The accuracy obtained for the stack, hard-voting and soft-voting classifiers were 90%, 95% and 94%, respectively. The bat algorithm was able to obtain excellent results too. The accuracies obtained by the stacking, hard-voting and soft-voting classifiers were 92%, 95% and 91%. The flower pollination algorithm performed relatively well. The accuracy obtained by the stacking, hard-voting and soft-voting classifiers were 87%, 85% and 86%. The accuracies obtained by the stack, hard-voting and soft-voting for the Jaya algorithm were 89%, 89% and 89%, respectively.
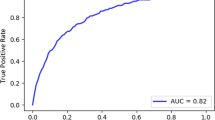
The ROC curves for the stacked model for the four feature selection methods are depicted in Fig. 7. The AUC was maximum for the mutual information algorithm with 0.96. The precision-recall curves for the stacked classifiers for the four feature selection methods are described in Fig. 8. The stacked model obtained a maximum average precision of 0,98 after being trained on features chosen by mutual information.
Further, the results obtained by the machine learning models were compared with the deep learning models. DNN, 1D-CNN and LSTM were the classifiers used in this study. The model architecture of the deep neural networks is described in Fig. 9. For the DNN, five layers were considered. The number of neurons used was 30, 11, 7, 4 and 1. “Relu” was the activation function used for the input and hidden layers. The “sigmoid” activation function was used for the output layer. “Adam” was the optimizer, and “binary cross entropy” was the loss function used. A learning rate of 0.0001 was utilized, and the batch size was set to 10. The neural network was run for 750 epochs to establish reliable results.
For the 1D-CNN, we included layers such as conv1D, max pooling, drop out and flatten layers. The loss function was “binary cross entropy”, and “Adam” was the optimizer. The number of epochs and batch size were set to 10 each. A learning rate of 0.001 was utilized, and “leaky relu” was the activation function used for the input and hidden layers. “sigmoid” was the activation function used for the output layer.
The LSTM used four layers consisting of 150, 75, 50 and 1 neurons, respectively. The loss function used was “binary cross-entropy, and the optimizer was “Adam”. The batch size was set to 32.
All three models were split into training and testing in the ratio of 80:20. The results obtained by the deep learning models are described in Table 7. Among the three, DNN performed the best, with an accuracy of 89%. 1D-CNN and LSTM obtained accuracies of 85% and 83%, respectively. The accuracy and loss curves for the models are depicted in Fig. 10. From the figure, the results obtained by the models are reliable and not overfitting.
Explainable artificial intelligence
In this study, five XAI methods: SHAP, LIME, QLattice, Eli5 and Anchor have been used to make the models more interpretable. We chose the stacked model for interpretation since they obtained good results and are generally reliable. Deep learning classifiers were not considered since many explainers do not support deep learning algorithms today. Further, machine learning algorithms performed better than deep learning models in this study. This is normal in artificial intelligence applications since deep learning models perform better only with comprehensive data.
SHAP is a widely used XAI technique that makes global and local interpretations42. SHAP uses game and probability theory to understand the impact of each attribute. The global interpretation of the models is explained using beeswarm plots as described in Fig. 11. A hyperplane separates the non-severe (left) and severe classes (right). Red indicates a higher value, and blue indicates a lower value. The markers are also arranged based on their importance (The best feature remains at the top). The figure shows that the most important markers are basophils, CRP, LDH, lymphocytes, albumin, protein and ferritin. CRP, LDH and Ferritin levels increased in severe COVID-19 patients. Basophils, lymphocytes, albumin and protein levels decreased in severe COVID-19 patients.
Local interpretations can be explained using the SHAP force plot, as shown in Fig. 12. Figure 12a,c indicate a non-severe prognosis. It can be seen that markers such as lymphocytes, SPO2, basophils and CRP are pushing the predictions towards a non-severe prognosis. Figure 12b,d indicate a severe COVID-19 prognosis. Markers such as CRP, AST, basophils and lymphocytes push the predictions towards severe COVID-19.
LIME is another explainer used to make local interpretations43. It uses a model-agnostic approach (It works for most ML models). It uses a ridge regression model and kernels such as Gaussian and RBF to explain the predictions. The LIME interpretations are depicted pictorially in Fig. 13. Figure 13a,b predict a severe prognosis, and Fig. 13c,d indicate a non-severe prognosis. The attributes are also arranged based on the descending order of their importance. The figure shows that the most important markers are albumin, D-Dimer, LDH, CRP, basophils, protein, AST, SPO2 and lymphocytes.
Eli5 is yet another method to demystify predictions44. It is a python package and is highly used with tree-based classifiers. Figure 14 depicts Eli5 predictions, and according to it, the most essential attributes are albumin, urea, lymphocytes, CRP, NLR, and basophils count. This explainer considers the “bias” (error rate).
Abzu developed the QLattice explainer45. It uses quantum computing and symbolic regression to explain the predictions. QLattice trains the models to understand the variation in data. The input attributes are called registers. A collection of registers is termed a QGraph. Every QGraph has a set of nodes (registers) and activation functions. Activation functions such as add, multiply, log, sine, tanh and Gaussian are generally used. The QGraphs are described in Fig. 15. It can be seen that the most important markers are lymphocytes, CRP and D-Dimer.
Anchor is an XAI technique that uses rules and conditions46. The strength of an anchor is measured using its precision and coverage. Precision defines the accuracy of the anchor. Coverage determines how many instances utilize the same conditions. The anchors for non-severe and severe cases are described in Table 8. The most important markers are basophils, albumin, lymphocytes, CRP, D-Dimer, neutrophils, protein and NLR.
Five XAI techniques have been utilized and their findings are similar. The most important markers that can predict a patient's severity are CRP, lymphocytes, basophils, albumin, D-Dimer, NLR, and neutrophils.
Discussion
This research used multiple machine learning algorithms to predict severe COVID-19 cases in advance so that appropriate treatments could be provided for vulnerable patients. To demystify the predictions, five heterogenous XAI techniques were used. Doctors and medical professionals can easily understand the variation in the markers provided by the explainers. This decision support system can be setup in various medical facilities to aid healthcare workers. In developing countries, this application can be used to make judicious use of essential medical assets such as ICU beds, ventilators and medicines. The models can also be utilized to present a second opinion to the doctors.
Fourteen feature selection methods were utilized and we chose the best four for further analysis. They are mutual information, bat algorithm, flower pollination algorithm and Jaya algorithm. A maximum accuracy of 95% was obtained by the mutual information algorithm. The F1-score, AUC and AP were 94%, 0.98 and 0.99. When the bat algorithm was utilized, a 93% accuracy was obtained. The F1-score, AUC and AP were 92%, 0.97 and 0.94. When the flower pollination algorithm was used, an accuracy of 89% was obtained. The F1-score, AUC and AP were 88%, 0.95 and 0.97. When the Jaya algorithm was utilized, a 90% accuracy was obtained. The F1-score, AUC and AP were 88%, 0.95 and 0.97. Most machine learning models performed relatively well.
Several markers showed variation between the two cohorts. Among all, CRP was chosen by all the XAI techniques. CRP levels increased in severe COVID-19 patients in this study47. Lymphocyte levels decreased in severe COVID-19 patients. Lymphopenia has been commonly recorded in patients when their conditions deteriorate48. This research also observed Basopenia in severe COVID-19 patients49. Low serum albumin has been associated with severe COVID-1950. This marker variation was also observed in this study. D-Dimer has already been an important marker in predicting COVID-1951. Elevated D-Dimer levels were found in the severe COVID-19 cohort. NLR is a vital marker which has already been utilized to diagnose and predict severity in patients. NLR levels elevates in severe COVID-19 patients52. The same trend has been observed in this research. Spo2 levels decreased rapidly in the severe COVID-19 cohort. Lower oxygen levels can seriously threaten COVID-19 patients since it causes hypoxia53. Age was also observed to be an important factor in predicting COVID-19 severity. Older patients were more vulnerable to experiencing severe symptoms54. The above markers can be monitored carefully to prevent a fatal prognosis.
Various machine researches have been conducted to predict the severity of COVID-19. Raman et al.55 used machine learning to predict COVID-19 severity during hospital admission. Patient data was collected from the University of Texas. The random forest obtained a sensitivity of 72% and a specificity of 78%. Their model could predict the severity within six hours of hospital admission. Ershadi et al.56 used image and clinical data to predict COVID-19 severity. A fuzzy-based classifier was developed to forecast severe cases. Two datasets were used, and the accuracies obtained for them were 92% and 90%, respectively. Chest X-ray images and clinical data were used to predict COVID-19 severity in another research57. 930 COVID-19 patients from Italy were considered for this research, and the stacking classifier achieved an accuracy of 89.03%. The most important markers were LDH, CRP, age, WBC and SpO2. Bello et al.58 used clinical markers and omics to predict COVID-19 severity. The model obtained an accuracy of 91.6%. The most important markers were LDH, albumin, creatinine, lymphocytes, neutrophils and potassium.
However, no articles that use five different XAI techniques to predict COVID-19 severity exist. Explainers such as Anchor, QLattice and Eli5 have been rarely used in medical machine learning. There are some limitations in our study too. The data collected was from a single geographical territory (India). Multiple datasets from different sources must be considered to make the classifiers more reliable. This research made exclusive use of supervised learning. Unsupervised learning and reinforcement learning algorithms were not considered. Graphical processing units (GPU) increase computational speed while training. However, they were not used in this study. We had divided the dataset into training and testing and had performed cross validation. However, we could not test the data on real-time patients as our study was retrospective. Prospective study can be conducted in the future to test real patients’ prognosis. Our machine learning models could also be used for other diseases and public health issues59,60,61,62,63,64,65.
Conclusions
XAI is a part of machine learning, generally used to demystify the predictions made by the classifiers. In this study, we used several supervised learning algorithms and XAI techniques to predict the COVID-19 severity in advance. The patients vulnerable to severe COVID-19 symptoms can be identified early, and appropriate treatments can be provided to save them. Various patterns and trends in the clinical markers were observed using descriptive statistics in the initial part of this research. Multiple feature selection techniques, including nature-inspired algorithms, were utilized to select the most crucial parameters. Several algorithms, such as bagging, boosting, stacking, voting and state-of-art deep learning, were used to make accurate predictions. The mutual information algorithm proved to be the most efficient feature selection technique obtaining a maximum accuracy of 95%. Five heterogeneous XAI algorithms such as, SHAP, LIME, QLattice, Eli5 and Anchor, have been used to understand the classification predictions. According to them, the most essential marker was CRP. Other markers such as D-Dimer, lymphocytes, neutrophils, albumin and basophils were also crucial. The classifiers can be utilized as a decision support system in various hospitals for prediction. The models can be used to predict the COVID-19 severity in advance. It can also aid the medical professionals and can offer them a second opinion. The algorithms can also be used for a rapid diagnosis too.
In the future, cloud-based models can be deployed. They can easily store both the data and code more efficiently. High-end GPUs can be utilized to train deep learning algorithms. Other diagnostic methods, such as rapid antigen tests, chest X-rays and genome sequencing, can be combined suitably. Prognosis can be predicted for various COVID-19 variants. Electronic health records from multiple hospitals across various countries can be combined before training the models. Other deep learning techniques such as fuzzy ensembling techniques could be utilized.
Data availability
Data will be made available by Mr. Krishnaraj Chadaga after obtaining prior permission from Manipal Academy of Higher Education.
References
Yang, L. et al. COVID-19: Immunopathogenesis and Immunotherapeutics. Signal Transduct. Target. Ther. 5(1), 128. https://doi.org/10.1038/s41392-020-00243-2 (2020).
Velavan, T. P. & Meyer, C. G. Mild versus severe COVID-19: Laboratory markers. Int. J. Infect. Dis. 1(95), 304–307. https://doi.org/10.1016/j.ijid.2020.04.061 (2020).
Hu, B., Huang, S. & Yin, L. The cytokine storm and COVID-19. J. Med. Virol. 93(1), 250–256. https://doi.org/10.1002/jmv.26232 (2021).
Ndwandwe, D. & Wiysonge, C. S. COVID-19 vaccines. Curr. Opin. Immunol. 1(71), 111–116. https://doi.org/10.1016/j.coi.2021.07.003 (2021).
Zhou, L. et al. The SNHG1-centered ceRNA network regulates cell cycle and is a potential prognostic biomarker for hepatocellular carcinoma. Tohoku J. Exp. Med. 258(4), 265–276. https://doi.org/10.1620/tjem.2022.J083 (2022).
Zhu, Y. et al. Deep learning-based predictive identification of neural stem cell differentiation. Nat. Commun. 12(1), 2614. https://doi.org/10.1038/s41467-021-22758-0 (2021).
Lu, S. et al. Soft tissue feature tracking based on DeepMatching network. CMES Comput. Model. Eng. Sci. https://doi.org/10.32604/cmes.2023.025217 (2023).
Akter, S., Shamrat, F. J., Chakraborty, S., Karim, A. & Azam, S. COVID-19 detection using deep learning algorithm on chest X-ray images. Biology. 10(11), 1174 (2021).
Shamrat, F. J. et al. LungNet22: a fine-tuned model for multiclass classification and prediction of lung disease using X-ray images. J. Personal. Med. 12(5), 680. https://doi.org/10.3390/jpm12050680 (2022).
Shamrat, F. J. et al. High-precision multiclass classification of lung disease through customized MobileNetV2 from chest X-ray images. Comput. Biol. Med. 1(155), 106646. https://doi.org/10.1016/j.compbiomed.2023.106646 (2023).
Huyut, M. T. Automatic detection of severely and mildly infected COVID-19 patients with supervised machine learning models. IRBM. 44(1), 100725. https://doi.org/10.1016/j.irbm.2022.05.006 (2023).
Wendland, P. et al. Machine learning models for predicting severe COVID-19 outcomes in hospitals. Inf. Med. Unlock. 1(37), 101188. https://doi.org/10.1016/j.imu.2023.101188 (2023).
Nguyen, H. T. et al. Outcome prediction model and prognostic biomarkers for COVID-19 patients in Vietnam. ERJ Open Res. https://doi.org/10.1183/23120541.00481-2022 (2023).
Suma, L. S., Anand, H. S. & Vinod Chandra, S. S. Nature inspired optimization model for classification and severity prediction in COVID-19 clinical dataset. J. Ambient Intell. Human. Comput. 31, 1–3. https://doi.org/10.1007/s12652-021-03389-1 (2021).
Laatifi, M. et al. Explanatory predictive model for COVID-19 severity risk employing machine learning, shapley addition, and LIME. Sci. Rep. 13(1), 5481. https://doi.org/10.1038/s41598-023-31542-7 (2023).
World Health Organization. Clinical management of COVID-19: interim guidance, 27 May 2020. World Health Organization; 2020.
Cerda, P., Varoquaux, G. & Kégl, B. Similarity encoding for learning with dirty categorical variables. Mach. Learn. 107(8–10), 1477–1494. https://doi.org/10.1007/s10994-018-5724-2 (2018).
Rodríguez, P., Bautista, M. A., Gonzalez, J. & Escalera, S. Beyond one-hot encoding: lower dimensional target embedding. Image Vision Comput. 1(75), 21–31. https://doi.org/10.1016/j.imavis.2018.04.004 (2018).
Ali, P. J., Faraj, R. H., Koya, E., Ali, P. J. & Faraj, R. H. Data normalization and standardization: A technical report. Mach. Learn Tech. Rep. 1(1), 1–6 (2014).
Han, H., Wang, W. Y., Mao, B. H. Borderline-SMOTE: A new over-sampling method in imbalanced data sets learning. In Advances in Intelligent Computing: International Conference on Intelligent Computing, ICIC 2005, Hefei, China, August 23–26, 2005, Proceedings, Part I, 878–887 (Springer Berlin Heidelberg, 2005).
Mirjalili, S. & Lewis, A. The whale optimization algorithm. Adv. Eng. Softw. 1(95), 51–67. https://doi.org/10.1016/j.advengsoft.2016.01.008 (2016).
Abualigah, L. & Diabat, A. Advances in sine cosine algorithm: A comprehensive survey. Artif. Intell. Rev. 54(4), 2567–2608. https://doi.org/10.1007/s10462-020-09909-3 (2021).
Castelli, M., Manzoni, L., Mariot, L., Nobile, M. S. & Tangherloni, A. Salp swarm optimization: A critical review. Exp. Syst. Appl. 1(189), 116029. https://doi.org/10.1016/j.eswa.2021.116029 (2022).
Wang, D., Tan, D. & Liu, L. Particle swarm optimization algorithm: An overview. Soft Comput. 22, 387–408. https://doi.org/10.1007/s00500-016-2474-6 (2018).
Zitar, R. A., Al-Betar, M. A., Awadallah, M. A., Doush, I. A. & Assaleh, K. An intensive and comprehensive overview of JAYA algorithm, its versions and applications. Arch. Comput. Methods Eng. 29(2), 763–792. https://doi.org/10.1007/s11831-021-09585-8 (2022).
Heidari, A. A. et al. Harris hawks optimization: Algorithm and applications. Future Gener. Comput. Syst. 1(97), 849–872. https://doi.org/10.1016/j.future.2019.02.028 (2019Aug).
Mirjalili, S., Mirjalili, S. M. & Lewis, A. Grey wolf optimizer. Adv. Eng. Softw. 1(69), 46–61. https://doi.org/10.1016/j.advengsoft.2013.12.007 (2014).
Mirjalili, S. & Mirjalili, S. Genetic algorithm. Evol. Algor. Neural Netw. Theory Appl. https://doi.org/10.1007/978-3-319-93025-1_4 (2019).
Abdel-Basset, M. & Shawky, L. A. Flower pollination algorithm: a comprehensive review. Artif. Intell. Rev. 52, 2533–2557. https://doi.org/10.1007/s10462-018-9624-4 (2019).
Kumar, V. & Kumar, D. A systematic review on firefly algorithm: Past, present, and future. Archiv. Comput. Methods Eng. 28, 3269–3291. https://doi.org/10.1007/s11831-020-09498-y (2021).
Pant, M., Zaheer, H., Garcia-Hernandez, L. & Abraham, A. Differential evolution: A review of more than two decades of research. Eng. Appl. Artif. Intell. 1(90), 103479. https://doi.org/10.1016/j.engappai.2020.103479 (2020).
Guerrero-Luis, M., Valdez, F. & Castillo, O. A review on the cuckoo search algorithm. Fuzzy Logic Hybrid Extens. Neural Optim. Algor. Theory Appl. https://doi.org/10.1007/978-3-030-68776-2_7 (2021).
Agarwal, T. & Kumar, V. A systematic review on bat algorithm: Theoretical foundation, variants, and applications. Archiv. Comput. Methods Eng. https://doi.org/10.1007/s11831-021-09673-9 (2021).
Vergara, J. R. & Estévez, P. A. A review of feature selection methods based on mutual information. Neural Comput. Appl. 24, 175–186. https://doi.org/10.1007/s00521-008-0206-2 (2014).
Gollapalli, M. et al. A novel stacking ensemble for detecting three types of diabetes mellitus using a Saudi Arabian dataset: Pre-diabetes, T1DM, and T2DM. Comput. Biol. Med. 1(147), 105757. https://doi.org/10.1016/j.compbiomed.2022.105757 (2022).
Kumari, S., Kumar, D. & Mittal, M. An ensemble approach for classification and prediction of diabetes mellitus using soft-voting classifier. Int. J. Cogn. Comput. Eng. 1(2), 40–46. https://doi.org/10.1016/j.ijcce.2021.01.001 (2021).
Atallah, R., Al-Mousa, A. Heart disease detection using machine learning majority voting ensemble method. In 2019 2nd international conference on new trends in computing sciences (ictcs). 1–6. (IEEE, 2019).
Tasci, E., Uluturk, C. & Ugur, A. A voting-based ensemble deep learning method focusing on image augmentation and preprocessing variations for tuberculosis detection. Neural Comput. Appl.. 33(22), 15541–15555. https://doi.org/10.1007/s00521-021-06177-2 (2021).
Samek, W., Montavon, G., Lapuschkin, S., Anders, C. J. & Müller, K. R. Explaining deep neural networks and beyond: A review of methods and applications. Proc. IEEE. 109(3), 247–278. https://doi.org/10.1109/JPROC.2021.3060483 (2021).
Wang, X., Mao, D. & Li, X. Bearing fault diagnosis based on vibro-acoustic data fusion and 1D-CNN network. Measurement. 1(173), 108518. https://doi.org/10.1016/j.measurement.2020.108518 (2021).
Yu, Y., Si, X., Hu, C. & Zhang, J. A review of recurrent neural networks: LSTM cells and network architectures. Neural Comput. 31(7), 1235–1270 (2019).
Liu, Y., Liu, Z., Luo, X. & Zhao, H. Diagnosis of Parkinson’s disease based on SHAP value feature selection. Biocybernet. Biomed. Eng. 42(3), 856–869. https://doi.org/10.1016/j.bbe.2022.06.007 (2022).
Bhandari, M., Yogarajah, P., Kavitha, M. S. & Condell, J. Exploring the capabilities of a lightweight CNN model in accurately identifying renal abnormalities: Cysts, stones, and tumors, using LIME and SHAP. Appl. Sci. 13(5), 3125. https://doi.org/10.3390/app13053125 (2023).
Chadaga, K., Prabhu, S., Sampathila, N. & Chadaga, R. A machine learning and explainable artificial intelligence approach for predicting the efficacy of hematopoietic stem cell transplant in pediatric patients. Healthcare Anal. 1(3), 100170. https://doi.org/10.1016/j.health.2023.100170 (2023).
Khanna, V. V. et al. A distinctive explainable machine learning framework for detection of polycystic ovary syndrome. Appl. Syst. Innov. 6(2), 32. https://doi.org/10.3390/asi6020032 (2023).
Rahimi, S., et al. Explainable machine learning model to predict COVID-19 severity among older adults in the province of Quebec. https://doi.org/10.1370/afm.21.s1.3619.
Ahnach, M., Zbiri, S., Nejjari, S., Ousti, F. & Elkettani, C. C-reactive protein as an early predictor of COVID-19 severity. J. Med. Biochem. 39(4), 500. https://doi.org/10.5937/jomb0-27554 (2020).
Kosidło, J. W., Wolszczak-Biedrzycka, B., Matowicka-Karna, J., Dymicka-Piekarska, V. & Dorf, J. Clinical significance and diagnostic utility of NLR, LMR, PLR and SII in the course of COVID-19: A literature review. J. Inflamm. Res. 31, 539–562. https://doi.org/10.2147/JIR.S395331 (2023).
Alomair, B. M. et al. Mixed storm in SARS-CoV-2 infection: A narrative review and new term in the Covid-19 era. Immun. Inflamm. Dis. 11(4), e838. https://doi.org/10.1002/iid3.838 (2023).
Yang, R. et al. Liver injury associated with the severity of COVID-19: A meta-analysis. Front. Public Health. 2(11), 1003352. https://doi.org/10.3389/fpubh.2023.1003352 (2023).
Saville, O., Elbatarny, M., Tera, Y., Deng, Y. & Othman, M. Sex differences in D-Dimer and critical illness in COVID-19 patients: A systematic review and meta-analysis. Res. Pract. Thromb. Haemost. 14, 100042. https://doi.org/10.1016/j.rpth.2023.100042 (2023).
Asperges, E. et al. Dynamic NLR and PLR in predicting Covid-19 severity: A retrospective cohort study. Infect. Dis. Therapy. 17, 1–6. https://doi.org/10.1007/s40121-023-00813-1 (2023).
Xiang, M., Wu, X., Jing, H., Novakovic, V. A. & Shi, J. The intersection of obesity and (long) COVID-19: Hypoxia, thrombotic inflammation, and vascular endothelial injury. Front. Cardiovasc. Med. 7(10), 1062491. https://doi.org/10.3389/fcvm.2023.1062491 (2023).
Zhang, J. J., Dong, X., Liu, G. H. & Gao, Y. D. Risk and protective factors for COVID-19 morbidity, severity, and mortality. Clin. Rev. Allergy Immunol. 64(1), 90–107. https://doi.org/10.1007/s12016-022-08921-5 (2023).
Raman, G. et al. Machine learning prediction for COVID-19 disease severity at hospital admission. BMC Med. Inf. Decis. Mak. 23(1), 1–6. https://doi.org/10.1186/s12911-023-02132-4 (2023).
Ershadi, M. M. & Rise, Z. R. Fusing clinical and image data for detecting the severity level of hospitalized symptomatic COVID-19 patients using hierarchical model. Res. Biomed. Eng. 39(1), 209–232. https://doi.org/10.1007/s42600-023-00268-w (2023).
Rahman, T. et al. BIO-CXRNET: A robust multimodal stacking machine learning technique for mortality risk prediction of COVID-19 patients using chest X-ray images and clinical data. Neural Comput. Appl. 4, 1–23. https://doi.org/10.1007/s00521-023-08606-w (2023).
Bello, B. et al. Integrating AI/ML models for patient stratification leveraging omics dataset and clinical biomarkers from COVID-19 patients: A promising approach to personalized medicine. Int. J. Mol. Sci. 24(7), 6250. https://doi.org/10.3390/ijms24076250 (2023).
Liu, M. et al. Three-dimensional modeling of heart soft tissue motion. Appl. Sci. 13(4), 2493. https://doi.org/10.3390/app13042493 (2023).
Rasheed, J. & Shubair, R. M. Screening lung diseases using cascaded feature generation and selection strategies. Healthcare 10, 1313. https://doi.org/10.3390/healthcare10071313 (2022).
Altameem, A., Mahanty, C., Poonia, R. C., Saudagar, A. K. & Kumar, R. Breast cancer detection in mammography images using deep convolutional neural networks and fuzzy ensemble modeling techniques. Diagnostics. 12(8), 1812. https://doi.org/10.3390/diagnostics12081812 (2022).
Mahanty, C. et al. Prediction of COVID-19 active cases using exponential and non-linear growth models. Exp. Syst. 39(3), e12648. https://doi.org/10.1111/exsy.12648 (2022).
Mahanty, C., Kumar, R. & Patro, S. G. Internet of medical things-based COVID-19 detection in CT images fused with fuzzy ensemble and transfer learning models. N. Gener. Comput. 40(4), 1125–1141. https://doi.org/10.1007/s00354-022-00176-0 (2022).
Mahanty, C., Kumar, R., Asteris, P. G. & Gandomi, A. H. COVID-19 patient detection based on fusion of transfer learning and fuzzy ensemble models using CXR images. Appl. Sci. 11(23), 11423. https://doi.org/10.3390/app112311423 (2021).
Mahanty, C., Kumar, R., Mishra, B. K. & Barna, C. COVID-19 detection with X-ray images by using transfer learning. J. Intell. Fuzzy Syst. 43(2), 1717–1726. https://doi.org/10.3233/JIFS-219273 (2022).
Acknowledgements
We would like to thank Manipal Academy of Higher Education for giving us a platform to conduct this study.
Funding
The article will be funded by Manipal Academy of Higher Education.
Author information
Authors and Affiliations
Contributions
K.C: Data Curation, Methodology, Software, Writing—Original Draft. S.P: Conceptualization, Supervision. N.S: Project administration, Funding acquisition. S.U: Validation, Resources. R.C: Visualization, Writing—Review and Editing. D.B: Review and Editing. S.K.G.S.: Funding acquisition, Writing—Review and Editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chadaga, K., Prabhu, S., Sampathila, N. et al. Explainable artificial intelligence approaches for COVID-19 prognosis prediction using clinical markers. Sci Rep 14, 1783 (2024). https://doi.org/10.1038/s41598-024-52428-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-52428-2
- Springer Nature Limited
This article is cited by
-
Prediction of 30-day mortality for ICU patients with Sepsis-3
BMC Medical Informatics and Decision Making (2024)
-
The predictive power of data: machine learning analysis for Covid-19 mortality based on personal, clinical, preclinical, and laboratory variables in a case–control study
BMC Infectious Diseases (2024)