Abstract
Esophageal carcinoma (ESCA) is a leading cause of cancer-related death worldwide, and certain oral and intestinal pathogens have been associated with cancer development and progression. We asked if esophageal microbiomes had shared alterations that could provide novel biomarkers for ESCA risk. We extracted DNA from tumor and non-tumor tissue of 212 patients in the NCI-MD case control study and sequenced the 16S rRNA gene (V3-4), with TCGA ESCA RNA-seq (n = 172) and WGS (n = 123) non-human reads used as validation. We identified four taxa, Campylobacter, Prevotella, Streptococcus, and Fusobacterium as highly enriched in esophageal cancer across all cohorts. Using SparCC, we discovered that Fusobacterium and Prevotella were also co-enriched across all cohorts. We then analyzed immune cell infiltration to determine if these dysbiotic taxa were associated with immune signatures. Using xCell to obtain predicted immune infiltrates, we identified a depletion of megakaryocyte-erythroid progenitor (MEP) cells in tumors with presence of any of the four taxa, along with enrichment of platelets in tumors with Campylobactor or Fusobacterium. Taken together, our results suggest that intratumoral presence of these co-occurring bacterial genera may confer tumor promoting immune alterations that allow disease progression in esophageal cancer.
Similar content being viewed by others
Introduction
Esophageal carcinoma (ESCA) is a rapidly increasing malignancy, with global rates increasing nearly 50% from 2012 to 2019 (Surveillance, Epidemiology, and End Results (SEER), National Cancer Institute). ESCA is predominantly classified as adenocarcinoma (EAC) and squamous cell carcinoma (ESCC), which show striking disparities. EAC is more common in men, younger patients, and Western countries while ESCC is more common in women, older patients, and African and Asian countries1,2. Development of ESCA also varies by histology, with EAC linked to a pro-inflammatory condition called Barrett’s esophagus and ESCC linked to environmental factors, including obesity and smoking, and for both somatic mutations such as TP533,4,5,6.
These risk factors are also known to play a role in modulating the gastrointestinal microbiome7. Several studies have demonstrated community and taxonomic alterations of the esophageal microbiome in ESCA patients8,9,10, showing a transition from Gram-positive dominated to a Gram-negative dominated microbiota before development of EAC. In vivo studies indicate that microbiome changes occur during the development of ESCA that correlate with changes in gene expression in the esophageal epithelium, including multiple microbial sensing pathways (e.g. toll-like receptors) that influence immune signaling and immune cell recruitment patterns11,12. These data suggest that alterations within the microbiome, or dysbiosis, may result in chronic inflammation and contribute to the development of ESCA.
To better understand how microbial dysbiosis and its interplay with the immune system contributions to ESCA development, we analyzed the microbiome of three datasets from the National Cancer Institute-Maryland (NCI-MD) case control study, The Cancer Genome Atlas (TCGA) RNA sequencing (RNA-seq) and whole genome sequencing (WGS) datasets. We compared taxonomic abundance between non-tumor adjacent and tumor tissues and identified four genera, Campylobacter, Fusobacterium, Prevotella, and Streptococcus, co-enriched in each dataset. Further examination of clinicopathological factors such as gender, race, smoking status, and histology revealed no association with the above four taxa. However, taxa association with immune cell abundance suggested platelet differentiation was increased in tumors with high taxa abundance. These data suggest certain opportunistic pathogenic taxa may promote esophageal cancer development by altering the immune microenvironment.
Results
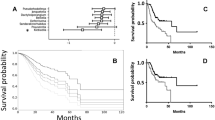
To better understand the microbial and immune system contributions to ESCA development, we comprehensively evaluated the esophageal tissue microbiome and inferred immune cell infiltration in patients with ESCA in two cohorts, NCI-MD and TCGA, with latter divided into WGS and RNA-seq, which results in three datasets. We extracted DNA from patients enrolled in the NCI-MD case control study from the Baltimore, Maryland area (118 non-tumor tissues, 94 tumors; 45 NT-T pairs) for 16S V3-4 sequencing as previously described13. Briefly, sequence reads were filtered for length (> 200bp) and max error rate (0.5%), and submitted for high-resolution taxonomic assignment (Resphera Insight) to assess taxa abundance. After QC, 154 samples were used for analysis. TCGA ESCA RNA-seq data (11 non-tumor tissue, 162 tumors; 11 NT-T pairs) and whole genome sequencing (WGS) (61 non-tumor tissue, 62 tumors; 61 NT-T pairs) data were downloaded (GDC Data Portal, NCI) as validation cohorts (Fig. 1A, Tables S1-S3). Stringent quality control measures were applied on both data sets (Methods).
Identification of microbial signatures in esophageal cancer. (A) Bacterial abundance within the NCI-MD case control study calculated from 16S V3-4 amplification. ESCA adj. indicates non-tumor adjacent tissue. (Panel) Total number of patients used in this study from three cohorts: NCI-MD case control study, TCGA RNA-seq, and TCGA whole genome seq (WGS). (B) Bacterial abundance within TCGA WGS, determined by quantification of non-human aligned reads. (C) Bacterial abundance within TCGA RNA-seq, determined by quantification of non-human aligned reads. A 1% cutoff was applied to all taxa as the minimal average (across samples) for plotting.
First, we sought to determine if any differences existed in the microbiome between ESCA tumor and non-tumor adjacent tissues. No differences in alpha or beta diversity were seen for the NCI-MD cohort, however alpha diversity decreased significantly in TCGA RNA-seq tumor samples but increased significantly in TCGA WGS tumor samples (Figure S1). Because tobacco smoking is a key risk-factor for developing esophageal cancer14, we asked if smoking status or other key ESCA risk factors were associated with alpha diversity. Interestingly, none of these clinicopathological factors (gender, histology, race, smoking status, or stage) showed significant difference in taxa abundance across all three cohorts (Figure S2).
Examination of the most abundant taxa in the NCI-MD cohort, independent of tissue type, identified Streptococcus, Pseudomonas, Prevotella, Veillonella, Lactobacillus, Stenotrophomonas, Fusobacterium, and Acinetobacter. One (Pseudomonas) and six (Streptococcus, Pseudomonas, Prevotella, Veillonella, Lactobacillus, and Fusobacterium) of these taxa were also highly abundant in TCGA RNA-seq and WGS cohorts, respectively (Fig. 1). We then performed a statistical concordance analysis (Methods, Table S4), which identified four common taxa across all cohorts: Campylobacter, Fusobacterium, Prevotella, and Streptococcus as enriched in ESCA (Fig. 2). All four taxa were enriched in tumors in at least two of three cohorts (Table S4). Additionally, given that TP53 is one of the most frequently mutated genes in ESCA4,6, we investigated TP53 mutation status in the TCGA cohort (WGS and RNA-seq) and found no relationship with abundance of these four taxa (Figure S3).
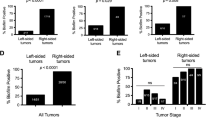
Four taxa are enriched in esophageal cancer across cohorts. Abundance of (A) Fusobacterium, (B) Campylobacter, (C) Prevotella, and (D) Streptococcus within NCI-MD case control study calculated from 16S V3-4 amplification. Abundance of the above four taxa within TCGA WGS and RNA-seq, determined by quantification of non-human aligned reads. Violin plots indicate relative abundance of each of the four taxa; % is the number of tumors with taxa present (ratio is number of samples with taxa present over number of total samples). Co-association was determined by statistical concordance analysis (Methods).*Statistical concordance analysis is located in Table S4.
Comparison of EAC versus ESCC revealed no significant differences in abundance of the four above taxa in any cohort (Figure S4) and expansion to include all taxa returned no differences after Benjamini–Hochberg correction. To determine if any relationship existed with microbial function, apart from community structure, we investigated the inferred metabolic profile of the ESCA microbiome using PICRUSt, but did not identify any associations with ESCA between tumor vs non-tumor tissue, overall or stratified by the four taxa (data not shown).
Having observed Fusobacterium was one of the most enriched taxa in ESCA tissue, we assessed whether this genus is co-abundant with specific taxa in ESCA. Specifically, Prevotella, and Streptococcus are often found in oral biofilms alongside Fusobacterium nucleatum where the two genera rely on F. nucleatum binding to salivary protein anchors (e.g. Statherin) and sharing nutrients to grow15. Furthermore, Fusobacterium and Prevotella have been previously described as co-enriched in ESCA9,16; therefore we asked if any of our four enriched taxa (Campylobacter, Fusobacterium, Prevotella, and Streptococcus) were co-enriched in the same tumors or were associated with other taxa. As most microbiome data is often sparse, correlation coefficients calculated by Pearson or Spearman methods are prone to spurious and false-positive relationships17 within such community networks18,19,20, so we used SparCC to compensate for the sparsity inherent to 16S-based studies21. We calculated co-enrichment for each of the three datasets. Overall, TCGA (RNA-seq and WGS) results were highly concordant (Table S5), and showed a consistent co-enrichment of certain taxa, with Fusobacterium and Prevotella co-enriched across all cohorts (Fig. 3A–C, Figure S5). These two taxa were also enriched with Leptotrichia and Veillonella, consistent with prior reports in colorectal cancer22,23. The NCI-MD cohort beta diversity suggested that Streptococcus-enriched samples were divergent from those with Campylobacter, Fusobacterium, and Prevotella (Figure S1E). We confirmed a negative co-enrichment between Streptococcus and Campylobacter and Fusobacterium in this cohort (Fig. 3A). Given this negative association, we used our TCGA WGS data to investigate species-level differences, and found a significant enrichment in S. oralis, F. nucleatum, P. denticola and P. intermedia in tumors as compared to non-tumors (Figure S6). Regardless of cohort, we found these four taxa were negatively associated with Acinetobacter, Brevundimonas, Klebsiella, Pseudomonas, and Xanthomonas (Fig. 3A–C). These data indicate that co-occurrence of Fusobacterium and Prevotella are a common feature of ESCA and may be important in ESCA pathology.
Fusobacterium and Prevotella are consistently co-associated across cohorts. (A) Taxa co-enrichment networks within the NCI-MD case control study. Taxa abundance was permuted through SparCC with 100 iterations, and correlation coefficients were filtered for X < − 0.2 and X > 0.2. Gold edges indicate positive coefficients demonstrating co-enrichment while blue edges indicate negative coefficients demonstrating exclusion. Edge thickness represents normalized coefficient values. (B) Taxa co-enrichment networks within TCGA RNA-seq. (C) Taxa co-enrichment networks within TCGA WGS. Networks for (B) and (C) were filtered for correlation coefficients X < − 0.3 and X > 0.3, otherwise networks were constructed as described for (A).
Since Fusobacterium spp. have demonstrated effects on gene expression changes within tumor epithelial and immune cells, we predicted immune cell infiltration from NCI-MD and TCGA RNA-seq data using the deconvolution algorithm xCell24, and then compared their abundances with our four co-enriched taxa. We identified a depletion of megakaryocyte-erythroid progenitor (MEP) cells in tumors with presence of any of the four bacteria (Fig. 4A,B), which was significant in the RNA-seq TCGA dataset (p < 0.001) (Figure S7-S8). Furthermore, we found a modest enrichment of platelets in tumors with Campylobactor or Fusobacterium (p < 0.06) (Fig. 4A,B, Figure S9). These data suggest that intratumoral presence of these bacterial genera results in loss of MEPs by promoting their terminal differentiation to platelets.
Megakaryocyte–erythroid progenitor cells are depleted in tumors with high carriage of ESCA-enriched taxa. (A) RNA-sequencing was performed on NCI-MD patients (n = 23; non-tumor = 13, tumor = 10) and samples were analyzed for predicted cell infiltration using xCell (citation). Cell infiltrates and taxa abundance were correlated using Spearman’s coefficient. (B) Correlation of xCell predicted cell infiltration in TCGA RNA-seq patients with taxa abundance. *Statistical significance analysis in Fig. S7-8.
Discussion
Globally rising rates of ESCA suggest that potentially novel drivers are partially responsible, however identification of these factors remains a significant problem in diagnosis and treatment. In this study we asked if the microbiome, a known driver of various GI malignancies25, was altered in ESCA in comparison to non-tumor adjacent esophageal tissue. We found enrichment of the taxa Campylobacter, Fusobacterium, Prevotella, and Streptococcus in tumor tissue. These findings are consistent with other studies examining the ESCA microbiome8, although our study is the first to report the co-enrichment of all four taxa in the same cohorts. Additionally, as these taxa exist within a community, abundance changes in one taxon are associated with changes in others, and we found consistent co-enrichment networks with the above four taxa common across cohorts, including associations of Fusobacterium, Prevotella, Leptotrichia, and Veillonella. Interestingly, these taxa may also invoke terminal differentiation of MEPs into platelets within the tumor microenvironment. Streptococcus spp. induce platelet activation and secretion through FcγRIIA signaling26. Lipopolysaccharide, the major outer membrane component in Gram-negative microbes such as Campylobacter, Fusobacterium, and Prevotella, also activate platelets through TLR4 signaling and induces proinflammatory cytokine secretion26,27,28. This suggests that the intratumoral presence of these bacteria may result in MEP to platelet differentiation. Expanded platelet counts are known to play a role in esophageal cancer development and metastasis29. Platelet-derived growth factor A (PDGFA) increases proliferation and invasion of multiple cancer types and high expression is a poor prognostic factor in ESCA30. Platelets may also promote metastasis through increasing interactions between primary tumor and endothelial cells, as occurs in colorectal cancer30,31, suggesting these taxa may contribute to increased platelet counts and poorer ESCA prognosis.
A common finding among several cancers of the GI tract, from oral squamous cell carcinoma to colorectal cancer, is the enrichment of Fusobacterium spp., including Fusobacterium nucleatum32. This species in particular has been shown in many studies to be not only enriched in the tumor but also in adjacent biofilms33. Fusobacterium is known to be an important bridging species during dysbiosis development, by colonizing early and allowing other taxa, including Streptococcus, to form links to additional species to create and establish biofilms32. Biofilms are seen in multiple cancer types, including oral and colon, and create opportunities for the growth of more virulent or pathogenic strains of certain Gram-negative microbes11,33. This increased virulence allows for the attachment of Fusobacteria and other oral and GI enriched microbes to adhere to sugar molecules and proteins on the surface of epithelial and immune cells34,35. These attachments create opportunities for invasion of Fusobacterium and Porphyromonas spp. to invade epithelial cells, promote inflammatory cell signals, and enhance epithelial cell motility to promote extraversion and metastasis36. However, it is important to recognize that more than one pathogen, like F. nucleatum, is likely required to initiate or promote cancer development or metastasis37. Thus, the coordinate efforts between bridging species such as F. nucleatum, and others like those we found co-enriched in ESCA tumors, can create the opportunity for other pathogens to work together to promote cancer.
These findings suggest that therapeutic anti-cancer strategies targeting one genus or species are likely to fail as other associated taxa may offset the loss of those taxa. Instead, it is likely a multi-targeted strategy, based on presence of intratumoral taxa, for modulating microbial dysbiosis is required to improve treatment and patient outcome. Further research is needed, however, to better understand the mechanisms driving enrichment of these taxa and immune cells in ESCA and other cancers.
Methods
University of Maryland (UMD) esophageal samples: collection and DNA extraction
DNA was extracted from 212 esophageal tissue samples, one plain water control (Mo Bio Laboratories, Inc, Carlsbad, CA, USA), one water control (Mo Bio Laboratories, Inc, Carlsbad, CA, USA) that was carried through the DNA extraction process, and one mock community (BEI resources, Manassas, VA, USA). Esophageal tissue was collected at the University of Maryland Medical Center (Baltimore, MD, USA) under an IRB-approved collection protocol (OH98CN027/ FWA00005897, National Institutes of Health Institutional Review Board) where all surgical subjects gave informed, written consent prior to collection, and all procedures were performed in accordance with the Declaration of Helsinki31,32. The study was approved by National Institutes of Health Institutional Review Board. Samples were flash-frozen and stored at − 80 °C until DNA extraction. Tumor stage, histology, and Barrett’s Esophagus status were determined from the pathology report. All work areas were cleaned with 70% ethanol and 10% bleach prior to DNA extraction. DNA extraction was carried out by lysing the microbes in fresh frozen tissue samples using Yeast Cell Lysis Buffer (Epicentre, Madison, WI, USA) and bead beating. Proteinase K and RNAse A were added to the samples to remove proteins and RNA, and to enrich for DNA. Samples were processed through gDNA column (Invitrogen, Carlsbad, CA, USA) and eluted in certified DNA- and RNA-free water (Mo Bio Laboratories, Inc, Carlsbad, CA, USA)25.
University of Maryland (UMD) esophageal samples: PCR amplification and MiSeq sequencing
PCR amplification of the V3-4 region of the 16S rRNA gene in each sample was completed at three different dilutions of genomic DNA (1×, 10× and 100×), and the PCR reaction with the highest yield was carried forth to sequencing as previously described27. This process is designed to overcome the inhibitory effect of a large amount of human DNA in esophageal tissue samples. Paired-end DNA sequencing of the amplicons from all samples and both variable regions were completed in the same run on a MiSeq machine (M04141, Illumina, San Diego, CA, USA) using the 2 × 300 base pair chemistry (Reagent barcode: MS3917443-600V3) and 50 unique sample barcodes. All dilutions, PCR amplification and sequencing were completed at the University of Minnesota Genomics Center (Minneapolis, MN, USA).
16S rRNA Sequence analysis
Raw read pairs from the MiSeq platform were trimmed for quality using Trimmomatic38 with a target final error rate of 0.5%, and merged into consensus fragments with FLASH39. High-quality unmerged forward reads (≥ 200bp after trimming) were also included for downstream analysis to increase sample coverage. PhiX spike-in fragments were detected using BLASTN40 and removed. Sequences associated with PCR chimeras were identified using UCLUST41 and filtered. Human genome contaminant identification was performed by aligning sequences against hg19 using Bowtie242, and mitochondria and chloroplast removal utilized assignments by the RDP classifier43. Of the 212 original samples extracted and sequenced, 154 remained after performing the above filtering and contamination steps. Barrett’s esophagus samples were removed from downstream analysis due to low quality reads, leaving only 5 samples remaining, which was too few for analysis. Passing 16S rRNA gene sequences were assigned a high-resolution taxonomic lineage using Resphera Insight, a custom 16S rRNA bioinformatics pipeline that utilizes both the SILVA and Greengenes databases for alignment33,44. This analysis utilized a data processing, checking and exploration 9-step process as described in https://greathouselab.github.io/esoph-micro-cancer-workflow/data_processing_nci_umd.html.
To filter out contaminant organisms associated with DNA extraction kit reagents and other sources, we first reviewed negative controls / blank samples prepared with original tissue samples, and developed a set of dominant indicator contaminant species including Bradyrhizobium spp., Propionibacterium_acnes, Agrobacterium_tumefaciens, Delftia spp. and Ralstonia spp. We then performed a correlation analysis between all species/OTUs and these indicator species. Any species/OTU with a nonparametric Spearman correlation ≥ 0.25 was then considered to be a contaminant and was removed; however 10 species/OTUs with known human body site associations were retained including: Faecalibacterium prausnitzii, Prevotella copri, Collinsella aerofaciens, Lactobacillus rhamnosus, Prevotella nigrescens, Prevotella disiens, and Finegoldia magna. To filter very low frequency contaminants, we further removed all members associated with a set of genera known to be contaminants from prior literature45: Bradyrhizobium, Ralstonia, Delftia, Agrobacterium, Janthinobacterium, Halomonas, Methylobacterium, Aquamicrobium, Diaphorobacter, Herbaspirillum, and Variovorax. After contaminant removal, samples were normalized through rarefaction to 500 sequences per sample. Alpha and beta-diversity analysis performed with QIIME46.
Processing of The Cancer Genome Atlas (TCGA) samples
RNA-seq and WGS bam files reflecting cancer and non-cancer samples from esophageal carcinoma patients available from TCGA were identified using the Genomic Data Commons (GDC) portal and downloaded using the GDC data transfer client (http://portal.gdc.cancer.gov/; Link: https://nam02.safelinks.protection.outlook.com/?url=https%3A%2F%2Fwww.ncbi.nlm.nih.gov%2Fgeo%2Fquery%2Facc.cgi%3Facc%3DGSE234304&data=05%7C01%7CLeigh_Greathouse%40baylor.edu%7C9b71dac5157d46f4d99008dbae42459d%7C22d2fb35256a459bbcf4dc23d42dc0a4%7C0%7C0%7C638295371882327356%7CUnknown%7CTWFpbGZsb3d8eyJWIjoiMC4wLjAwMDAiLCJQIjoiV2luMzIiLCJBTiI6Ik1haWwiLCJXVCI6Mn0%3D%7C3000%7C%7C%7C&sdata=GBWGhA7NtppWJL7UcfC5M0x3JM%2B3Z9Tei7nL1fzT6eM%3D&reserved=0 Token (password)—ctchamkodpurjqf). Barrett’s esophagus status, tumor stage, gender, race and survival information were also retrieved when available from the GDC.
Quality control and identification of microbial DNA
Unmapped sequences from the raw RNA-seq and WGS bam files were converted to FASTQ format using Samtools47 and trimmed for quality with Trimmomatic38 to remove error-prone reads. Additionally, in order to remove unmapped spliced transcripts and other poorly aligning sequences, we performed a local alignment to the human reference (hg19) using Bowtie242. Clean sequences passing all filters were assigned to a taxonomic lineage using Pathoscope (v1.0)48,49. To filter out contaminant organisms associated with DNA extraction kit reagents and other laboratory sources, we developed a set of 10 dominant indicator contaminant species including members of Bradyrhizobium, Propionibacterium, Pseudomonas, and Arthrobacter. We then performed analysis between all species/OTUs and these indicator species across WGS tumor, WGS normal and RNA-seq samples. Any species detected in at least 58 of RNA-seq samples, or 55 of WGS tumor or 55 of WGS normal samples was often found to show a strong Spearman correlation with one or more of the indicator contaminant species, and were thus assigned putative contaminant status. We further removed all species associated with a set of higher taxa known to be contaminants from published literature45 or that were also highly recurrent across most samples including members of Pseudomonadales, Comamonadaceae, Rhizobiales, Burkholderiales, Paenibacillaceae, Propionibacterium acnes, Escherichia, and Bacillaceae.
Integration of 16S rRNA and TCGA microbial profiles
In order to provide a direct comparison between the 16S rRNA and TCGA WGS/RNA-seq microbial profiles, we first performed a concordance study at the species level across all technologies. Manual examination of the WGS/RNA-seq and 16S rRNA data revealed that some species in WGS previously determined to be contaminant were more likely to reflect true oral and upper respiratory tract species (such as Rothia mucilaginosa and Streptococcus mitis). Therefore, we revisited the contaminant removal process for our data integration of 16S rRNA WGS / RNA-seq data, and rescued species that were present in the 16S rRNA contaminant-free dataset, or those reported in a second esophageal tissue 16S rRNA study by Gall et al.50. This effort confirmed consistent taxonomic profiles for joint interpretation across genomic data types.
Inferred microbial metabolism
The input files were a FASTA file of representative sequences and a BIOM table of the abundance of each ASV across each sample from the NCI-MD cohort. The steps of the pipeline used were (1) sequence placement, (2) hidden-state prediction of genomes, (3) metagenome prediction, and (4) pathway-level predictions. The following pipeline was followed to perform this analysis: https://github.com/picrust/picrust2/wiki/Full-pipeline-script
Statistical methods
Statistical comparisons were performed in R (cran.r-project.org). To establish associations of specific microbial members with tumor status, we utilized Generalized linear fixed effects models (GLMs) and Generalized linear mixed effects models (GLMMs) in which patient membership was considered a fixed effect, or random effect, respectively. The Mann–Whitney test for differential abundance was applied per each genomic data type independent as a supplement to GLM analyses. Fisher’s exact test was applied to evaluate differential frequencies of positive vs negative status for each microbial member in the integrated analysis. Generalized linear models – taxon % abundance modeled by Tumor / Normal status (fixed effect) and Patient ID (fixed effect) (stratified by genomic data type). Generalized linear mixed effects models – taxon % abundance modeled by Tumor / Normal status (fixed effect) and Patient ID (random effect) (stratified by genomic data type). Fisher’s exact test for positive status (stratified by genomic data type). Comparisons to adjust for the blood-derived normal samples in TCGA were also applied. Creation of heatmaps – cases were subset to only those without Barrett’s Esophagus with EAC. OTUs below a minimum threshold of average relative abundance were removed (e.g., an average of 1% relative abundance). The heatmaps plot the individual tissues along the X-axis and the genus abundance along the Y-axis after filtering for the minimum threshold of relative abundance, with cell shading based on the individual genus relative abundances. The scale of shading was adjusted for each data source due to differences in average relative abundance. Hierarchical clustering of tissues and genera was performed using the hclust(.) function in R with default method complete-linkage. Code to replicate the heatmaps is available in our code repository under the file “Fig. 1_heatmaps.R”. Heatmaps were generated using the pheatmap package (v1.0.12). All analyses for the main figures are located at https://github.com/GreathouseLab/esoph-micro-cancer-workflow
RNA-sequencing and immune infiltration
Total RNA was extracted from fresh-frozen esophageal tissues using TRIzol. RNA quality was validated by Agilent TapeStation and samples with RIN value ≥ 7.0 were selected for sequencing. Samples were sequenced on the DNBseq platform with 2 × 100 bp paired-end sequencing. Reads were aligned to the human genome (hg38) using HISAT and bowtie224,42,51. xCell was used to predict immune cell infiltration in each sample24 and predicted infiltrates were correlated with microbial abundance by the Spearman method.
Microbial co-abundance networks
For each cohort, taxa co-occurrence was calculated using SparCC with 100 iterations and default correlation method (not Pearson or Spearman)18. Correlation coefficients were filtered for X < − 0.2 and X > 0.2 (NCI-MD) or X < − 0.3 and X > 0.3 (TCGA). Networks were generated using Cytoscape v3.9.1.
Data availability
All de-identified data and code used to conduct analyses and generate figures for this manuscript are available from TCGA or at https://github.com/GreathouseLab/esoph-micro-cancer-workflow. All sequences generated during this study are deposited under the GEO accession #GSE234304. Any protocols will be made available at the request of the researcher by contacting Dr. Leigh Greathouse.
References
Arnold, M. et al. Global incidence of oesophageal cancer by histological subtype in 2012. Gut 64(3), 381–387 (2015).
Stabellini, N. et al. Sex differences in esophageal cancer overall and by histological subtype. Sci. Rep. 12(1), 5248 (2022).
Abnet, C. C., Arnold, M. & Wei, W.-Q. Epidemiology of esophageal squamous cell carcinoma. Gastroenterology 154(2), 360–373 (2018).
Dulak, A. M. et al. Exome and whole-genome sequencing of esophageal adenocarcinoma identifies recurrent driver events and mutational complexity. Nat. Genet. 45(5), 478–486 (2013).
Feakins, R. M. Obesity and metabolic syndrome: pathological effects on the gastrointestinal tract. Histopathology 68(5), 630–640 (2016).
Kim, J. et al. Integrated genomic characterization of oesophageal carcinoma. Nature 541(7636), 169–175 (2017).
Agrawal, K., Markert, R. J. & Agrawal, S. Risk factors for adenocarcinoma and squamous cell carcinoma of the esophagus and lung. AME Med. J. 3(3), 1 (2018).
Baba, Y. et al. Review of the gut microbiome and esophageal cancer: Pathogenesis and potential clinical implications. Ann. Gastroenterol. Surg. 1(2), 99–104 (2017).
Lopetuso, L. R. et al. Esophageal microbiome signature in patients with Barrett’s esophagus and esophageal adenocarcinoma. PLOS ONE 15(5), e0231789 (2020).
Snider, E. J. et al. Alterations to the esophageal microbiome associated with progression from Barrett’s esophagus to esophageal adenocarcinoma. Cancer Epidemiol. Biomark. Prevent. 28(10), 1687–1693 (2019).
Blackett, K. L. et al. Oesophageal bacterial biofilm changes in gastro-oesophageal reflux disease, Barrett’s and oesophageal carcinoma: association or causality?. Aliment. Pharmacol. Therapeut. 37(11), 1084–1092 (2013).
Kaakoush, N. O. et al. Cross-talk among metabolic parameters, esophageal microbiota, and host gene expression following chronic exposure to an obesogenic diet. Sci. Rep. 7(1), 45753 (2017).
Greathouse, K. L. et al. Interaction between the microbiome and TP53 in human lung cancer. Genome Biol. 19(1), 123 (2018).
Enzinger, P. C. & Mayer, R. J. Esophageal cancer. N. Engl. J. Med. 349(23), 2241–2252 (2003).
Kolenbrander, P. E. et al. Oral multispecies biofilm development and the key role of cell–cell distance. Nat. Rev. Microbiol. 8(7), 471–480 (2010).
Shao, D. et al. Microbial characterization of esophageal squamous cell carcinoma and gastric cardia adenocarcinoma from a high-risk region of China. Cancer 125(22), 3993–4002 (2019).
Bonett, D. G. & Wright, T. A. Sample size requirements for estimating Pearson, Kendall and Spearman correlations. Psychometrika 65(1), 23–28 (2000).
Friedman, J. & Alm, E. J. Inferring correlation networks from genomic survey data. PLoS Comput. Biol. 8(9), e1002687 (2012).
Kurtz, Z. D. et al. Sparse and compositionally robust inference of microbial ecological networks. PLoS Comput. Biol. 11(5), e1004226 (2015).
Lovell, D. et al. Proportionality: A valid alternative to correlation for relative data. PLoS Comput. Biol. 11(3), e1004075 (2015).
Friedman, J. & Alm, E. J. Inferring correlation networks from genomic survey data. PLOS Computat. Biol. 8(9), e1002687 (2012).
Dohlman, A. B. et al. The cancer microbiome atlas: a pan-cancer comparative analysis to distinguish tissue-resident microbiota from contaminants. Cell Host Microbe 29(2), 281-298.e5 (2021).
Warren, R. L. et al. Co-occurrence of anaerobic bacteria in colorectal carcinomas. Microbiome 1(1), 16 (2013).
Aran, D., Hu, Z. & Butte, A. J. xCell: Digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 18(1), 220 (2017).
Elinav, E. et al. The cancer microbiome. Nat. Rev. Cancer 19(7), 371–376 (2019).
Arman, M. et al. Amplification of bacteria-induced platelet activation is triggered by FcγRIIA, integrin αIIbβ3, and platelet factor 4. Blood 123(20), 3166–3174 (2014).
Zhang, G. et al. Lipopolysaccharide stimulates platelet pecretion and potentiates platelet aggregation via TLR4/MyD88 and the cGMP-dependent protein kinase pathway. J. Immunol. 182(12), 7997–8004 (2009).
Galgano, L. et al. The controversial role of LPS in platelet activation in vitro. Int. J. Mol. Sci. 23(18), 10900 (2022).
Shimada, H. et al. Thrombocytosis associated with poor prognosis in patients with esophageal carcinoma. J. Am. Coll. Surg. 198(5), 737–741 (2004).
Han, N. et al. High expression of PDGFA predicts poor prognosis of esophageal squamous cell carcinoma. Medicine 100(20), e25932 (2021).
Plantureux, L. et al. The interaction of platelets with colorectal cancer cells inhibits tumor growth but promotes metastasis. Cancer Res. 80(2), 291–303 (2020).
Brennan, C. A. & Garrett, W. S. Fusobacterium nucleatum—symbiont, opportunist and oncobacterium. Nat. Rev. Microbiol. 17(3), 156–166 (2019).
Drewes, J. L. et al. High-resolution bacterial 16S rRNA gene profile meta-analysis and biofilm status reveal common colorectal cancer consortia. NPJ Biofilms Microbiomes 3, 34 (2017).
Cavallucci, V., et al., Proinflammatory and Cancer-Promoting Pathobiont Fusobacterium nucleatum Directly Targets Colorectal Cancer Stem Cells. Biomolecules, 2022. 12(9).
Abed, J. et al. Fap2 mediates Fusobacterium nucleatum colorectal adenocarcinoma enrichment by binding to tumor-expressed Gal-GalNAc. Cell Host Microbe 20(2), 215–225 (2016).
Bullman, S. et al. Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science 358(6369), 1443–1448 (2017).
Queen, J. et al. Comparative analysis of colon cancer-derived fusobacterium nucleatum subspecies: inflammation and colon tumorigenesis in murine models. mBio 13(1), e0299121 (2021).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30(15), 2114–2120 (2014).
Magoc, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27(21), 2957–2963 (2011).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res 25(17), 3389–3402 (1997).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19), 2460–2461 (2010).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9(4), 357–359 (2012).
Wang, Q. et al. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73(16), 5261–5267 (2007).
Daquigan, N. et al. High-resolution profiling of the gut microbiome reveals the extent of Clostridium difficile burden. NPJ Biofilms Microbiomes 3, 35 (2017).
Salter, S. J. et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 12, 87 (2014).
Kuczynski, J., et al. Using QIIME to analyze 16S rRNA gene sequences from microbial communities. Curr. Protoc. Microbiol. Chapter 1: p. Unit 1E 5 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25(16), 2078–2079 (2009).
Francis, O. E. et al. Pathoscope: species identification and strain attribution with unassembled sequencing data. Genome Res. 23(10), 1721–1729 (2013).
Oh, J. et al. Biogeography and individuality shape function in the human skin metagenome. Nature 514(7520), 59–64 (2014).
Gall, A. et al. Bacterial composition of the human upper gastrointestinal tract microbiome is dynamic and associated with genomic instability in a Barrett’s Esophagus cohort. PLoS ONE 10(6), e0129055 (2015).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 12(4), 357–360 (2015).
Acknowledgements
We would like to thank all of the participants of the NCI-MD cohort study who kindly agreed to provide their data and samples for this research study.
Funding
This research was supported by the Intramural Research Program of the NIH, NCI/CCR. A. Choudhury is funded by a Postdoctoral Fellowship Award from Baylor University.
Author information
Authors and Affiliations
Contributions
C.H. and A.V. conceptualized study ideas; K.L.G., C.H., J.S. and A.V. and J.W. contributed to methodology, study design, and data curation, A.C., K.L.G., A.J., N.P., J.W. and J.S. contributed to formal analysis, bioinformatics, sequencing, visualization and validation of datasets; C.H., K.L.G., J.S., and A.V. contributed to original draft preparation and review/editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Greathouse, K.L., Stone, J.K., Vargas, A.J. et al. Co-enrichment of cancer-associated bacterial taxa is correlated with immune cell infiltrates in esophageal tumor tissue. Sci Rep 14, 2574 (2024). https://doi.org/10.1038/s41598-023-48862-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-48862-3
- Springer Nature Limited