Abstract
This study aimed to identify shared specific genes associated with rheumatoid arthritis (RA) and inflammatory bowel disease (IBD) through bioinformatic analysis and to examine the role of the gut microbiome in RA. The data were extracted from the 3 RA and 1 IBD gene expression datasets and 1 RA gut microbiome metagenomic dataset. Weighted correlation network analysis (WGCNA) and machine learnings was performed to identify candidate genes associated with RA and IBD. Differential analysis and two different machine learning algorithms were used to investigate RA’s gut microbiome characteristics. Subsequently, the shared specific genes related to the gut microbiome in RA were identified, and an interaction network was constructed utilizing the gutMGene, STITCH, and STRING databases. We identified 15 candidates shared genes through a joint analysis of the WGCNA for RA and IBD. The candidate gene CXCL10 was identified as the shared hub gene by the interaction network analysis of the corresponding WGCNA module gene to each disease, and CXCL10 was further identified as the shared specific gene by two machine learning algorithms. Additionally, we identified 3 RA-associated characteristic intestinal flora (Prevotella, Ruminococcus, and Ruminococcus bromii) and built a network of interactions between the microbiomes, genes, and pathways. Finally, it was discovered that the gene CXCL10 shared between IBD and RA was associated with the three gut microbiomes mentioned above. This study demonstrates the relationship between RA and IBD and provides a reference for research into the role of the gut microbiome in RA.
Similar content being viewed by others
Introduction
Rheumatoid arthritis (RA) is an autoimmune inflammatory condition that primarily affects the joints. Individuals at risk for and those with RA experience gut dysbiosis1,2,3. A small-sample study on patients with RA reported that most patients had subclinical intestinal inflammation4. Intestinal inflammation can be caused by and sustained through gut dysbiosis5,6. Inflammatory bowel disease (IBD), which encompasses Crohn’s disease (CD) and ulcerative colitis (UC), is a chronic, recurrent inflammatory illness of the gut with immune system disturbance7. Despite having different target organs, IBD and autoimmune rheumatic illnesses share a genetic foundation8,9,10.
According to a population-based study from South Korea, RA is strongly associated with IBD11. Patients with RA have aberrant intestinal barrier permeability, which is consistent with the intestinal alterations observed in patients with IBD12. The IL-23/IL-17 inflammatory axis is activated during the development of both RA and IBD13. The expression of ENA78/CXCL5 is increased in tissues that are inflamed during RA and IBD14,15. Additionally, patients with RA and IBD have higher IL-6 levels16,17,18. These findings highlight the role of the gastrointestinal system in the development of RA and suggest a shared pathogenic mechanism between IBD and RA. However, the role of the gut microbiome in the pathogenesis of RA and the genetic interactions and molecular mechanisms underlying the relationship between RA and IBD remain unclear.
Machine learning technologies have been widely applied in the study of inflammatory diseases in recent years. Using machine learning and deep learning, Maria Giovanna Danieli et al. investigated how intravenous and subcutaneous immunoglobulin treatment affects patients with idiopathic inflammatory myopathies19. Isabelle Ayoub et al. used machine learning to assess the treatment response for lupus nephritis using standard clinical data with novel biomarkers20.
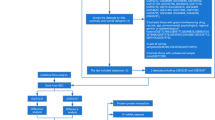
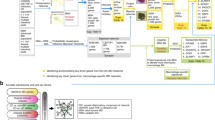
In this study, bioinformatic tools were used to identify the common specific genes and processes between RA and IBD and examine the relationship between the gut microbiome in RA and the shared specific genes. We comprehensively analyzed four gene expression datasets from the Gene Expression Omnibus (GEO) database (GSE55235, GSE55457, GSE179285, and GSE77298) and an RA-related metagenomic sequencing dataset, PRJEB6997, from the GMrepo database. Weighted correlation network analysis (WGCNA) was performed to identify candidate genes associated with RA and IBD. Gene set enrichment analysis (GSEA) was performed to assess changes at the pathway level. The shared specific genes between RA and IBD were screened using two types of machine learning algorithms and receiver operating characteristic (ROC) curves. The characteristics of the gut microbiome in RA were examined using differential analysis, two types of machine learning algorithms, and ROC curves. Subsequently, the shared specific genes related to the gut microbiome in RA were identified, and an interaction network of these genes and those related to shared GSEA pathways were constructed using the gutMGene, STITCH, and STRING databases. The Spearman correlation analysis was used to determine the connection between these genes and immune cells. To the best of our knowledge, this study is the first to report the common genes associated with both RA and IBD and their relationship with the gut microbiome in RA using a systematic bioinformatic approach. Figure 1 demonstrates the study design.
Materials and methods
Search strategy for datasets
For RA, 127 datasets were systematically retrieved from the GEO database (https://www.ncbi.nlm.nih.gov/geo/) using the keywords: ((((rheumatoid arthritis[MeSH Terms]) OR rheumatoid arthritis) AND human[Organism]) AND Expression profiling by array[Filter]) AND (“2012/01/01” [Publication Date]: “2022/01/01” [Publication Date]). Exclusion criteria: (1) Excluded those with a small sample size (sample size < 20), (2) Excluded datasets that were irrelevant (it is not rheumatoid arthritis), (3) Excluded blood samples and/or cell samples, (4) Excluded samples with significant drug, vaccine, age, environmental, psychological, regional genetic, or epidemiological factors, and (5) Excluded samples lacking normal samples (listed in Fig. 1).
For IBD, 112 data sets were systematically retrieved from the GEO database (https://www.ncbi.nlm.nih.gov/geo/) using the keywords: ((((inflammatory bowel disease[MeSH Terms]) OR inflammatory bowel disease) AND human[Organism]) AND Expression profiling by array[Filter]) AND (“2012/01/01” [Publication Date]: “2022/01/01” [Publication Date]). Exclusion criteria: (1) Excluded those with a small sample size (sample size < 50), (2) Excluded datasets that were irrelevant (it is not inflammatory bowel disease), (3) Excluded impure mRNA or non-mRNA transcriptome datasets, (4) Excluded samples with significant drug, age, environmental, psychological, regional genetic, or epidemiological factors, (5) Excluded blood samples and/or cell samples, (6) Excluded samples lacking normal samples, (7) Excluded single disease datasets, and (8) Excluded datasets with ambiguous disease sample site types. Inclusion criteria (Novelty assessment): The one with the most recent publication date and the fewest studies on PubMed was selected based on the exclusion criteria (listed in Fig. 1).
For the intestinal flora of RA, three datasets were systematically retrieved from the GMrepo database21 (https://gmrepo.humangut.info/home) using the keywords: [Arthritis, Rheumatoid]. Exclusion criteria: (1) Excluded datasets with 16 s rDNA, (2) Excluded datasets without publications in NCBI BioProjects (listed in Fig. 1).
Extraction and preprocessing of GEO data
The microarray datasets GSE5523522, GSE5545722, GSE17928523, and GSE7729824 were extracted from the GEO database (https://www.ncbi.nlm.nih.gov/geo/) using the R package GEOquery25. Additionally, Homo sapiens samples for the GSE55235 and GSE55457 datasets were generated using the GPL96 [HG-U133A] Affymetrix Human Genome U133A Array platform; those for the GSE179285 dataset were generated using the Agilent 014850 Whole Human Genome Microarray 4 × 44 K G4112F (Probe Name version) platform; and those for the GSE77298 dataset were generated using the GPL570 [HG-U133_Plus_2] Affymetrix Human Genome U133 Plus 2.0 Array platform. The GSE55235 dataset contained 10 synovial tissue samples from patients with RA and 10 samples from healthy individuals; the GSE55457 dataset contained 10 synovial tissue samples from healthy individuals and 13 samples from patients with RA; the GSE179285 dataset contained 14 inflamed colon tissue samples from patients with CD, 23 inflamed colon tissue samples from patients with UC, and 23 healthy colon tissue samples from healthy individuals; the GSE77298 dataset contained 16 synovial tissue samples from patients with RA and 7 samples from healthy individuals. The data from the GSE55235, GSE55457, and GSE77298 datasets were normalized using the RMA algorithm in the Affy package26 in R. The training set was created by merging the data from the GSE55235 and GSE55457 datasets, and the validation set was created using the data from the GSE77298 dataset. Additionally, the batch effects of the combined GSE55235 and GSE55457 datasets were removed using the Combat function of the sva package27. The principal component analysis (PCA) was used to evaluate the batch effect correction of the combined data28. Data from the GSE179285 dataset were downloaded, standardized, and analyzed using GEOquery25, Biobase29, and limma packages30. We used a distinct function of the “dplyr” package to remove the duplicate genes from each data set31. R software (version 4.3.0) was used for all data processing and analysis. Table 1 shows the comprehensive information for each dataset.
Weighted gene co-expression network analysis
WGCNA is a standard method for processing large amounts of data that permits the grouping and modularization of a collection of genes most closely related to disease onset. The “WGCNA” R package was used in the study to build the gene co-expression network32. The gene expression matrix is first entered into the R software to check for missing data and identify outliers. Next, we developed a scale-free network to select a soft threshold value for each disease, which is used as the parameter cut-off value for creating the adjacency and topology matrices. Next, gene co-expression modules for each disease were identified using the block modules function and module division analysis. Each module was associated with these diseases (RA, CD, and UC), and the Pearson correlation coefficients were used to filter the most relevant modules. The genes in these modules were classified as genes associated with diseases. Finally, the Venn diagram package33 was used to overlap module genes associated with RA, CD, and UC to screen for shared candidate genes.The analysis images of WGCNA are generated by R software. R software is open source software, and the various packages are free and open source.
Building a protein–protein interaction (PPI) network for module genes and identifying hub genes
The PPI network of module genes for each disease was investigated using the interaction relation in the database STRING (https://string-db.org/)34. The string result table was then entered into Cytoscape35. We performed Degree analysis to predict important nodes (or hub genes) using Cytoscape’s cytoHubba plugin36.
Functional analysis and gene set enrichment analysis of candidate genes
The clusterProfiler package37 was used for GO and KEGG analyses of the candidate genes. Significant differences were defined as an adjusted P-value of ≤ 0.05. The STRING (https://string-db.org/)34 database was used to construct a PPI network of the candidate genes. Additionally, GSEA was performed using the clusterProfiler package to analyze genes associated with RA, CD, and UC (previously ranked based on their log2FC values between the analyzed groups). The “c2.cp.kegg.v7.5.1.symbols.gmt” gene set was used to identify significantly enriched genes with a nominal false discovery rate (FDR) of < 0.25 and P < 0.05.
Screening and validation of the shared specific genes
LASSO38 was performed using the glmnet package39 to identify genes in the RA training set. In addition, the SVM-RFE algorithm40 in the e1071 package41 was used to select genes. A Venn diagram was created using the machine learning mentioned above to further identify key genes by overlapping genes in the two modules. Immediately, a boxplot demonstrating the expression of these genes in the RA training set, CD dataset, and RA validation set was created using the ggplot242 and ggpubr43 packages. Furthermore, the function of these significant genes in the RA training set, CD dataset, UC dataset, and RA validation set was independently assessed by creating ROC curves in RStudio using the pROC package44. The shared specific genes in RA and IBD were selected as those with significant differences (P < 0.05) and AUC values of > 0.7 in all datasets.
Identification of the gut microbiome in RA
The RA-related metagenomic sequencing dataset PRJEB699745 was retrieved from the GMrepo database (https://gmrepo.humangut.info/home)21. A total of 147 samples, including 92 samples from patients with RA and 55 samples from healthy individuals, were selected from the PRJEB6997 database. The limma package in R was used for analyzing the differential abundance of gut microbes. Bacteria with significantly differential abundance were selected based on |log2 FC| values of > 3 and P < 0.05. The ggplot2 and ggrepel46 packages in R were used to create a volcano map to demonstrate bacteria with differential abundance. LASSO was performed using the glmnet package to identify potential specific bacteria. Additionally, the SVM-RFE algorithm in the e1071 package was used to select potential specific bacteria. Subsequently, a Venn diagram was developed to identify the markers of gut microbiota by overlapping bacteria in the two modules of LASSO and SVM-RFE. To examine the relationship between the gut microbial markers and RA, the pROC package was used to analyze ROC curves in RStudio, and the ggplot2 and ggpubr packages were used to construct a boxplot in RStudio to compare gene expression among groups.
Identification of shared specific genes related to the gut microbiome in RA
The GutMGene (http://bio-annotation.cn/gutmgene/)47 database was used to identify metabolites from the markers of gut microbiota. The STITCH database (http://stitch.embl.de/)48 was used to identify shared specific genes directly associated with these metabolites.
Construction of an interaction network among shared specific genes related to the gut microbiome in RA, genes related to shared GSEA pathways, and gut microbial markers
Venn diagrams were constructed to overlap pathways identified via GSEA in the RA training set, CD dataset, UC dataset, and RA validation set and to overlap genes associated with these pathways in the RA training set, CD dataset, UC dataset, and RA validation set. The relationship between the metabolites and markers of gut microbiota was analyzed using the gutMGene (http://bio-annotation.cn/gutmgene/) database. The STITCH database (http://stitch.embl.de/) was used to investigate the link between metabolites and genes(including genes associated with pathways identified via GSEA and the shared specific genes related to the gut microbiome in RA), and the STRING (https://string-db.org/)34 database was used for analyzing the interaction between genes associated with pathways identified via GSEA and the shared specific genes related to the gut microbiome in RA. Finally, the Cytoscape35 program was being used to combine and display the previously indicated network links, resulting in an interaction network of shared specific genes related to the gut microbiome in RA, genes associated with pathways identified via GSEA, and the markers of the gut microbiome.
Correlation between shared specific genes associated with RA-specific gut microbiome and immune infiltration
We performed two immunoinvasive correlation analyses for this study. First, we combined the tagged genomes of several immune cell subpopulations in the CIBERSORT49, a deconvolution method to determine the proportion of 22 types of immune cells (reported in previous studies). In the meanwhile, we quantified the infiltration abundances of the 24 immune cells (reported in earlier studies) in these samples using ssGSEA50 based on the R package “GSVA”51. Finally, the association between immune cell infiltration in the RA training set, CD dataset, UC dataset, and RA validation set and shared specific genes connected to the gut microbiota in RA was analyzed using Spearman analysis.
Results
Screening of candidate genes associated with RA and IBD
Before analysis, batch effect reduction was applied to the GSE55235 and GSE55457 data sets (the RA training set), and PCA was used to evaluate and compare the features. Before the batch effect was removed, the data were dispersed as data sets, and it was visible (Fig. 2A). The overall expression of the data was distributed in the form of sample treatment (normal and RA) and published more evenly than before after the batch effect was removed (Fig. 2B).
Principal component analysis (PCA) of combined data sets before and after batch effect removal. (A) PCA analysis was performed before batch effect elimination. (B) PCA analysis was performed after batch effect elimination. The GSE55235 dataset is red, and the GSE55457 dataset is blue. The triangular dots and the circular dots indicate samples from the RA and normal groups, respectively.
To construct a RA scale-free network, the soft threshold (R2 = 0.85) was set to 9 (Fig. 3A). To construct a CD scale-free network, the soft threshold (R2 = 0.85) was set to 5 (Fig. 3C). To construct a UC scale-free network, the soft threshold (R2 = 0.85) was set to 6 (Fig. 3E). Finally, the WGCNA revealed 28 gene modules associated with the occurrence of RA in the RA training set (Fig. 3B). Each module was identified using a different color. Genes in the “blue” module had a significant positive association with RA (blue module: r = 0.82, P = 1e − 11; Fig. 3G; Supplementary Tables S1, S2, Supplementary Spreadsheet S3). The “royal blue” module was among the 52 modules identified in the CD dataset (Fig. 3D) that had a significant positive association with CD (royal blue module: r = 0.74, P = 4e − 07; Fig. 3H; Supplementary Tables S4, S5, Supplementary Spreadsheet S6). The “yellow-green” module was among the 57 modules identified in the UC dataset (Fig. 3F) that had a significant positive association with UC (yellow-green module: r = 0.78, P = 3e − 10; Fig. 3I; Supplementary Tables S7, S8, Supplementary Spreadsheet S9). A total of 15 candidate genes associated with both RA and IBD were identified after the intersection of genes in the abovementioned target modules: CXCL9, CCL18, CXCL10, S100A9, MMP9, RARRES3, S100A8, FCN1, ISG20, LILRB2, IDO1, CD19, CIITA, SIRPG, and DUOX2 (Fig. 3J).
Potential genes implicated in both rheumatoid arthritis (RA) and inflammatory bowel disease (IBD) were discovered using WGCNA52. (A) Analysis of the network topology for RA utilizing various soft-threshold powers. (B) Determination of the gene modules that RA co-expresses. The 28 modules comprising the dendrogram’s branches are each assigned a different color. (C) A study of network topology for Crohn’s disease (CD) utilizing various soft-threshold powers. (D) Determination of the gene modules that CD co-expresses. The 52 modules comprising the dendrogram’s branches are each assigned a different color. (E) Analysis of the network topology for ulcerative colitis (UC) utilizing various soft-threshold powers. (F) Identification of the gene modules co-expressed by UC. The 57 modules comprising the dendrogram’s branches are each assigned a different color. (G) Heatmap depicting the association between the prevalence of RA and module genes. (H) Heatmap depicting the association between the prevalence of CD and module genes. (I) Heatmap depicting the association between the prevalence of UC and module genes. Red and blue show a positive and negative association, respectively, with the hue’s depth indicating each’s strength. (J) Venn diagram demonstrating the overlap between candidate genes of two IBD (CD and UC) modules and those of one RA module.
Construction of the PPI network of module genes and identification of hub genes
We entered 1266 module genes from the RA’s corresponding WGCNA module (Fig. 4A), 239 module genes from the CD’s corresponding WGCNA module (Fig. 4B), and 111 module genes from the UC’s corresponding WGCNA module (Fig. 4C) into the STRING database to visualize the PPI network to investigate further whether the 15 candidate shared genes are the hub genes of the corresponding WGCNA module of each disease. Subsequently, the string result table was entered into Cytoscape. The cytoHubba tool was used to search hub nodes in the network, and the top 5% of genes were selected as hub gene nodes using Degree. The score increases as the node color darkens, and the number of edge interactions increases as the line color increases. The hub genes of the WGCNA module corresponding to RA include CD19 and CXCL10 (Fig. 4D; Supplementary Spreadsheet S10). The hub genes of the WGCNA module corresponding to CD include CXCL10, LILRB2, and MMP9 (Fig. 4E; Supplementary Spreadsheet S11). The hub genes of the WGCNA module corresponding to UC include CXCL10 and MMP9 (Fig. 4F; Supplementary Spreadsheet S12).
Protein–protein interaction network analysis of the corresponding WGCNA module genes for rheumatoid disease (RA) and inflammatory bowel disease (IBD) (Crohn’s disease [CD] and ulcerative colitis [UC]). (A) Protein interaction network of RA’s corresponding WGCNA module genes. (B) Protein interaction network of CD’s corresponding WGCNA module genes. (C) Protein interaction network of UC’s corresponding WGCNA module genes. (D) Network diagram of the hub nodes from RA. (E) Network diagram of the hub nodes from CD. (F) Network diagram of the hub nodes from UC.
Functional annotation of candidate genes and identification of pathways associated with RA and IBD
The abovementioned 15 candidate genes were subjected to GO (Table 2) and KEGG (Table 3) functional enrichment analyses. The results of GO analysis revealed that the candidate genes were primarily associated with neutrophil chemotaxis, collagen-containing extracellular matrix, and chemokine activity (Fig. 5A). The results of KEGG analysis revealed the candidate genes were primarily associated with the IL-17 signaling pathway, chemokine signaling pathway, and cytokine–cytokine receptor interaction (Fig. 5B). Subsequently, the STRING database was used to construct a PPI network to visualize the interaction among the 15 candidate genes (Fig. 5C). The pathways associated with these genes in the RA training (Table 4), CD (Table 5) and UC (Table 6) cohorts were identified via GSEA. In the RA training cohort, pathways related to the intestinal immune network for IgA production, allograft rejection, and antigen processing and presentation were activated, whereas those related to retinol metabolism, regulation of lipolysis in adipocytes, and tyrosine metabolism were inhibited (Fig. 5D,G). In the CD cohort, pathways related to IBD, the intestinal immune network for IgA production, and asthma were activated, whereas those related to the metabolism of xenobiotics by cytochrome P450, metabolism of drugs by cytochrome P450, and butanoate metabolism were inhibited (Fig. 5E,H). In the UC cohort, pathways related to IBD, asthma, and the intestinal immune network for IgA production were activated, whereas those related to the metabolism of xenobiotics by cytochrome P450 and metabolism of drugs by cytochrome P450 were inhibited (Fig. 5F,I).
Functional annotation of candidate genes and identification of pathways associated with RA and IBD. (A) Results of GO enrichment analysis of 15 candidate genes identified via WGCNA. (B) Results of KEGG enrichment analysis53 of 15 candidate genes identified via WGCNA. (C) PPI network of 15 candidate genes. (D) Upregulated enriched pathways identified via GSEA in the RA training cohort. (E) Upregulated enriched pathways identified via GSEA in the CD cohort. (F) Upregulated enriched pathways identified via GSEA in the UC cohort. (G) Downregulated enriched pathways identified via GSEA in the RA training cohort. (H) Downregulated enriched pathways identified via GSEA in the CD cohort. (I) Downregulated enriched pathways identified via GSEA in the UC cohort.
Machine learning algorithm-based screening and validation of shared specific genes
The RA training set was selected to screen for key genes between RA and IBD using two different machine learning algorithms. Of the 15 candidate genes, 4 were identified using the SVM-RFE algorithm (Fig. 6A,B), and 7 were identified using the LASSO regression algorithm (Fig. 6C,D). Eventually, three key genes (CXCL10, DUOX2, and CCL18) that were commonly identified using these two algorithms were selected (Fig. 6E). We performed differential expression and ROC curve discriminative efficacy demonstration to determine whether these three key genes are shared specific genes of RA and IBD. In the RA training set, CXCL10 and CCL18 had a high expression difference, while DUOX2 had a low expression difference (Fig. 6F); CXCL10, DUOX2, and CCL18 expressed fair discriminative efficiency (AUC > 0.7) (Fig. 6J). In the CD dataset, CXCL10, DUOX2, and CCL18 had high expression difference (Fig. 6G); CXCL10, DUOX2, and CCL18 expressed fair discriminative efficiency (AUC > 0.7) (Fig. 6K). In the UC dataset, CXCL10, DUOX2, and CCL18 had high expression difference (Fig. 6H); CXCL10, DUOX2, and CCL18 expressed fair discriminative efficiency (AUC > 0.7) (Fig. 6L). In the RA validation set, CXCL10 and CCL18 showed high expression difference, whereas DUOX2 showed no significant difference (Fig. 6I); CXCL10 and CCL18 expressed fair diagnostic efficiency (AUC > 0.7), whereas DUOX2 expressed poor discriminative efficiency (AUC < 0.7) (Fig. 6M). The shared specific genes in RA and IBD were selected as those with significant differences (P < 0.05) and AUC values of > 0.7 in all datasets for further analysis (CXCL10 and CCL18).
Machine learning-based identification and validation of potential shared specific genes. (A,B) Four genes were identified using the SVM-RFE algorithm in the RA training set. (C) LASSO coefficient profiles of 15 candidate genes in the RA training set. (D) LASSO coefficient profiles of 7 genes were selected as optimal (lambda) in the RA training set. (E) Venn diagram depicting the three key genes related to IBD in the RA training set. (F) Expression of CXCL10, DUOX2, and CCL18 in the RA training set. (G) Expression of CXCL10, DUOX2, and CCL18 in the CD dataset. (H) Expression of CXCL10, DUOX2, and CCL18 in the UC dataset. (I) Expression of CXCL10, DUOX2, and CCL18 in the RA validation set. (J) ROC curve for the verification of discriminative efficiency in the RA training set. (K) ROC curve for the verification of discriminative efficiency in the CD dataset. (L) ROC curve for the verification of discriminative efficiency in the UC dataset. (M) ROC curve for the verification of discriminative efficiency in the RA validation set (****P < 0.0001; ***P < 0.001; **P < 0.01; *P < 0.05).
Identification of the gut microbiome in RA based on machine learning
Differential analysis revealed two intestinal microbes at the genus (Prevotella and Ruminococcus) and two intestinal microbes at the species (Prevotella copri and Ruminococcus bromii) levels in the PRJEB6997 dataset (Fig. 7A; Supplementary Spreadsheet S13). Gut microbes associated with RA were screened using two machine learning algorithms in the PRJEB6997 dataset. The LASSO regression algorithm revealed four microbial groups associated with RA (Fig. 7B,C), whereas the SVM-RFE algorithm revealed three microbial groups (Fig. 7D,E). The three overlapping microbial groups (Prevotella, Ruminococcus, and Ruminococcus bromii) identified using the two methods were selected (Fig. 7F), and their diagnostic efficacy and abundance were examined. Prevotella, Ruminococcus, and Ruminococcus bromii exhibited lower diagnostic values (0.5 < AUC < 0.7) (Fig. 7G). The abundance of these three bacterial groups was different between healthy and RA. The abundance of Prevotella was high and that of Ruminococcus and Ruminococcus bromii was low among patients with RA (Fig. 7H).
Identification of gut microbes associated with RA. (A) Volcano map demonstrating the differential abundance of intestinal microbes based on the criteria of |log2 FC| values of > 3 and P < 0.05. (B) LASSO coefficients of four intestinal microbes in the PRJEB6997 dataset. (C) LASSO coefficients of four microbes selected as optimal (lambda) in the PRJEB6997 dataset. (D,E) The PRJEB6997 dataset was screened using the SVM-RFE algorithm to identify three diagnostic indicators. (F) Venn diagram demonstrating the three ideal diagnostic biomarkers in the PRJEB6997 dataset. (G) ROC curve for the verification of diagnostic efficiency in the PRJEB6997 dataset. (H) Relative abundance of three bacterial groups (Prevotella, Ruminococcus, and Ruminococcus bromii) in the PRJEB6997 dataset (*P < 0.05).
Construction of an interaction network among shared specific gene associated with the gut microbiome in RA, genes related to shared pathways identified via GSEA, and RA-specific gut microbiome
Based on the previous results and data extracted from the gutMGene database, gut microbes associated with RA were identified at the genus (Ruminococcus) and species (Prevotella copri, Ruminococcus bromii, Ruminococcus flavefaciens, Ruminococcus gnavus, and Ruminococcus champanellensis 18P13[T]) levels. The gutMGene database was used to identify metabolites associated with the abovementioned microbes (butyrate, alanine, leucine, isoleucine, glycine, proline, tartaric acid, glycocholic acid, fructose, propionate, glycerol, ursodeoxycholic acid, acetate, and succinate) (Supplementary Spreadsheet S14). Subsequently, the STITCH database was used to identify a single gene (CXCL10) directly associated with the metabolites as the shared specific genes related to the gut microbiome in RA (Fig. 8A).
Construction of an interaction network among shared specific gene associated with the gut microbiome in RA, genes related to shared pathways identified via GSEA, and the RA-specific gut microbiome. (A) Interaction network of shared specific gene and metabolites associated with the gut microbiome in RA; CXCL10 was directly associated with the metabolites. (B) Venn diagram demonstrating 7 shared high-expression pathways associated with RA and IBD identified via GSEA. (C) Venn diagram demonstrating 18 common genes associated with the 7 pathways. (D) An interaction network between the, shared specific gene associated with the gut microbiome in RA, shared pathways via GSEA, genes related to shared pathways identified via GSEA, and the RA-specific gut microbiome.
Based on the findings of GSEA and the high expression of CXCL10 in the RA and IBD samples, Venn diagrams were drawn to demonstrate 7 shared high-expression pathways identified via GSEA and 18 shared genes among these pathways (Fig. 8B,C). Furthermore, an interaction network was established based on the two gut microbial groups identified at the genus level (Ruminococcus and Prevotella), 5 gut microbial groups identified at the species level (Prevotella copri, Ruminococcus bromii, Ruminococcus flavefaciens, Ruminococcus gnavus, and Ruminococcus champanellensis 18P13[T]), 14 metabolites (butyrate, alanine, leucine, isoleucine, glycine, proline, tartaric acid, glycocholic acid, fructose, propionate, glycerol, ursodeoxycholic acid, acetate, and succinate) associated with the gut microbiome in RA, and the one shared specific gene related to the gut microbiome (CXCL10). This network contained 47 nodes and 231 edges (Fig. 8D). The results suggest that gut microbes associated with RA control the expression of CXCL10 by altering metabolite content in vivo, thereby regulating the intestinal immune network for IgA synthesis and other pathways.
Correlation between immune infiltration and shared specific gene related to the gut microbiome in RA
In the CIBERSORT algorithm, CXCL10 had a significant positive correlation with M1 macrophages, plasma cells, follicular helper T cells, naive B cells, and gamma-delta T cells, and a significant negative correlation with resting NK cells and activated Mast cells in the RA training set (Fig. 9A). In the ssGSEA algorithm, CXCL10 had a significant correlation with activated CD8 T cell, activated B cell, MDSC, activated CD4 T cell, and immature B cell and a significant inverse correlation with CD56dim natural killer cell, central memory CD4 T cell, plasmacytoid dendritic cell, immature dendritic cell, neutrophil, mast cell, monocytein, etc. in the RA training set (Fig. 9E). In the CIBERSORT algorithm, CXCL10 had a significant positive correlation with activated dendritic cells, eosinophils, M2 macrophages, activated NK cells, and resting mast cells and a significant negative correlation with naïve CD4 T cells (Fig. 9B) in the CD dataset. In the ssGSEA algorithm, CXCL10 had a significant positive correlation with activated dendritic cells, MDSC, effector memory CD8 T cell, gamma delta T cell, and monocyte and a significant negative correlation with neutrophil in the CD dataset (Fig. 9F). In the CIBERSORT algorithm, CXCL10 had a significant positive correlation with M0 macrophages, activated dendritic cells, M1 macrophages, and eosinophils in the UC dataset (Fig. 9C). In the ssGSEA algorithm, CXCL10 had a significant positive correlation with gamma delta T cell, immature B cell, activated dendritic cell, monocyte, and MDSC and a significant negative correlation with immature dendritic cell and neutrophil in the UC dataset (Fig. 9G). In the CIBERSORT algorithm, CXCL10 had a significant positive correlation with M1 macrophages, gamma-delta T cells, activated memory CD4 T cells, plasma cells, and T follicular helper cells and a significant negative correlation with resting NK cells, resting dendritic cells, and regulatory T cells (Tregs) in the RA validation set (Fig. 9D). In the ssGSEA algorithm, CXCL10 had a significant positive correlation with activated CD8 T cell, activated B cell, MDSC, activated CD4 T cell, immature B cell, and type 1 T helper cell and a significant negative correlation with immature dendritic cell, type 17 T helper cell, plasmacytoid dendritic cell, central memory CD4 T cell, and memory B cell in the RA validation set (Fig. 9H).
Correlation between immune infiltration and shared specific gene related to the gut microbiome in RA. (A) Through the use of the CIBERSORT algorithm, the CXCL10 expression and immune cells that are entering the body were correlated in the RA training set. (B) Through the use of the CIBERSORT algorithm, the CXCL10 expression and immune cells that are entering the body were correlated in the CD dataset. (C) Through the use of the CIBERSORT algorithm, the CXCL10 expression and immune cells that are entering the body were correlated in the UC dataset. (D) Through the use of the CIBERSORT algorithm, the CXCL10 expression and immune cells that are entering the body were correlated in the RA validation set. (E) Through the use of the ssGSEA algorithm, the CXCL10 expression and immune cells that are entering the body were correlated in the RA training set. (F) Through the use of the ssGSEA algorithm, the CXCL10 expression and immune cells that are entering the body were correlated in the CD dataset. (G) Through the use of the ssGSEA algorithm, the CXCL10 expression and immune cells that are entering the body were correlated in the UC dataset. (H) Through the use of the ssGSEA algorithm, the CXCL10 expression and immune cells that are entering the body were correlated in the RA validation set. The size of the dots indicates the strength of the association between gene expression and immune cell infiltration; the bigger the dots, the greater the correlation. The P-value is represented by the color of the dots; the greener the color, the lower the P-value. Statistical significance was defined as P < 0.05.
Discussion
In this study, shared specific genes between IBD and RA were identified via bioinformatic analysis. These genes are associated with the gut microbiome in RA. CXCL10 was the most relevant gene associated with IBD and RA, which also had a direct relationship with metabolites produced by gut microbes in RA. Additionally, metabolites associated with the gut microbiome in RA and pathways associated with RA and IBD were identified.
CXCL10 was the most significant shared specific gene between RA and IBD. CXCL10, also known as interferon-inducible protein-10 (IP-10), is an ELR-CXC chemokine54. It is mostly induced in humans when cell-mediated immune responses are elicited in pathological conditions such as infection, allograft rejection, and autoimmunity55. The expression of CXCL10 is very low in the colonic epithelium but substantially increases in colitis under the induction of IFN-γ56,57. Inhibiting CXCL10 reduces the frequency and severity of colitis and intestinal inflammation58,59. Additionally, patients with RA have higher CXCL10 expression in the synovial membrane60. High expression of CXCL10 mRNA and tissue infiltration of functional proteins in synovial tissue were also reported in CIA model rats with RA in a study on bone marrow mesenchymal stem cell therapy61. Increased CXCL10 mRNA has been observed in liver tissues of patients with HIV and HBV infection in various studies of inflammatory diseases, and its elevation is associated with disease activity62. Eldelumab (BMS-936557, formerly known as MDX-1100), a human monoclonal antibody against CXCL10, has reportedly been developed and has demonstrated effectiveness in the treatment of RA and IBD63,64.
Epidemiological and translational studies have suggested an interaction among bacteria in dysbiotic microbiota, and mucosal locations may play a causative role in the onset of RA2,65,66. Considering that the human intestinal microflora is affected by many factors such as region, population, host genetic factors, environment, and diet and different diseases may have different microbial flora, this study aimed to examine the relationship between the intestinal microflora in only one RA dataset and identify shared specific genes between IBD and RA. In the original study of the PRJEB6997 dataset, the abundance of Lactobacillus salivarius, Enterococcus, and Bacteroides was found to be high and that of Haemophilus, Klebsiella, and Bifidobacteria was found to be low in patients with RA45. In this study, the abundance of Prevotella was high and that of Ruminococcus and Ruminococcus bromii was low in patients with RA. It’s reported that the abundance of Prevotella copri was higher in stool samples of individuals with untreated new-onset RA (between 6 weeks and 6 months after diagnosis), and its presence was associated with a decline in the abundance of Bacteroides species and a loss of purportedly beneficial microbes2. Ruminococcus species can suppress TNF-α, and its abundance is lower in patients with Crohn’s disease than in healthy individuals67,68. On the contrary, Ruminococcus and Ruminococcus bromii are less reported in RA. In this study, data extracted from the gutMGene and STITCH databases revealed some metabolites associated with Ruminococcus, and succinate found in Ruminococcus champanellensis 18P13(T) had a direct relationship with CXCL10. A crucial metabolite in both host and microbial activities is succinate. Although succinate is typically considered an intermediate, it gets accumulated in some pathological conditions, especially during inflammation and metabolic stress69. Succinate was considered a pro-inflammatory metabolite; however, a study showed that it exerts anti-inflammatory effects on inflammatory signaling in macrophages70.
CXCL10 was significantly linked with the invasion of M1 macrophages in both RA and UC. Macrophages play an important role in RA. They are commonly found at the cartilage–pannus junction and in inflammatory synovial membranes. The extensive proinflammatory, destructive, and remodeling abilities of macrophages play an important role in both acute and chronic phases of RA71. TNF-α and interleukin-1 are two pro-inflammatory cytokines secreted by M1 macrophages, and several experimental and clinical studies have demonstrated their importance in the pathophysiology of RA72,73. Additionally, macrophages have been associated with IBD because they play a crucial role in several IBD-related risk genes74,75. Macrophages are regionally concentrated and polarize to the M1 subtype in UC, leading to persistent and recurrent inflammation76,77,78.
Although this study had a relatively large sample size (GEO and GMrepo datasets), certain limitations should be noted. Clinical samples should be used to validate these findings. Owing to clinical research and ethical constraints, the present study was not completely rigorous. Moreover, this study focused on the relationship between intestinal microbiota and genes. However, it is challenging to collect or monitor the intestinal microbiota of a single patient at the same time, as the intestinal microbiota may change owing to various factors such as environmental change or growth and development. The use of microarray technology to evaluate gene expression presents another drawback. Since fluorescence-mediated gene expression assessment is a biased method in contrast to hypothesis-free sequencing technology, RNA sequencing is more frequently used for assessing broad gene expression than microarray. Furthermore, even if we attempted to eliminate the batch impact of the combined data using the combat function of the sva package, it is undeniable that any solution can only mitigate this effect. These important issues should be considered in future studies.
In conclusion, because CXCL10 is involved in the onset of RA and IBD, it can be used to diagnose these two conditions. In addition, the gut microbiome in RA and several pathways related to IBD and RA were also found to be regulated by CXCL10. The findings of this study revealed the mechanism underlying the association between RA and IBD and served as a reference for further investigation of the intestinal flora in RA.
Data availability
The datasets GSE55235, GSE55457, GSE179285, and GSE77298 for this study can be found in the Gene Expression Omnibus database [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE55235/GSE55457/GSE55584/GSE179285/GSE77298/GSE82107]. The RA-related metagenomic sequencing dataset, PRJEB6997, for this study can be found in the GMrepo database [https://gmrepo.humangut.info/data/project/PRJEB6997].
References
Aletaha, D. & Smolen, J. S. Diagnosis and management of rheumatoid arthritis: A review. JAMA 320(13), 1360–1372. https://doi.org/10.1001/jama.2018.13103 (2018).
Scher, J. U. et al. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. Elife 2, e01202. https://doi.org/10.7554/eLife.01202 (2013).
Alpizar-Rodriguez, D. et al. Prevotella copri in individuals at risk for rheumatoid arthritis. Ann. Rheum. Dis. 78(5), 590–593. https://doi.org/10.1136/annrheumdis-2018-214514 (2019).
Tajik, N. et al. Targeting zonulin and intestinal epithelial barrier function to prevent onset of arthritis. Nat. Commun. 11(1), 1995. https://doi.org/10.1038/s41467-020-15831-7 (2020).
Lee, Y. K. & Mazmanian, S. K. Has the microbiota played a critical role in the evolution of the adaptive immune system?. Science 330(6012), 1768–1773. https://doi.org/10.1126/science.1195568 (2010).
Sommer, F. & Bäckhed, F. The gut microbiota–masters of host development and physiology. Nat. Rev. Microbiol. 11(4), 227–238. https://doi.org/10.1038/nrmicro2974 (2013).
Lee, J. W. et al. Inflammatory bowel disease cohort studies in Korea: Present and future. Intest. Res. 13(3), 213–218. https://doi.org/10.5217/ir.2015.13.3.213 (2015).
Lettre, G. & Rioux, J. D. Autoimmune diseases: Insights from genome-wide association studies. Hum. Mol. Genet. 17(R2), R116–R121. https://doi.org/10.1093/hmg/ddn246 (2008).
Baranzini, S. E. The genetics of autoimmune diseases: a networked perspective. Curr. Opin. Immunol. 21(6), 596–605. https://doi.org/10.1016/j.coi.2009.09.014 (2009).
Lees, C. W. et al. New IBD genetics: Common pathways with other diseases. Gut 60(12), 1739–1753. https://doi.org/10.1136/gut.2009.199679 (2011).
Bae, J. M. et al. Association of inflammatory bowel disease with ankylosing spondylitis and rheumatoid arthritis: A nationwide population-based study. Mod. Rheumatol. 27(3), 435–440. https://doi.org/10.1080/14397595.2016.1211229 (2017).
Smith, M. D., Gibson, R. A. & Brooks, P. M. Abnormal bowel permeability in ankylosing spondylitis and rheumatoid arthritis. J. Rheumatol. 12(2), 299–3054032403 (1985).
Schinocca, C. et al. Role of the IL-23/IL-17 pathway in rheumatic diseases: An overview. Front. Immunol. 12, 637829. https://doi.org/10.3389/fimmu.2021.637829 (2021).
Walz, A. et al. Regulation and function of the CXC chemokine ENA-78 in monocytes and its role in disease. J. Leukoc. Biol. 62(5), 604–611. https://doi.org/10.1002/jlb.62.5.604 (1997).
Z’Graggen, K. et al. The C-X-C chemokine ENA-78 is preferentially expressed in intestinal epithelium in inflammatory bowel disease. Gastroenterology 113(3), 808–816. https://doi.org/10.1016/s0016-5085(97)70175-6 (1997).
De Benedetti, F. et al. Serum soluble interleukin 6 (IL-6) receptor and IL-6/soluble IL-6 receptor complex in systemic juvenile rheumatoid arthritis. J. Clin. Investig. 93(5), 2114–2119. https://doi.org/10.1172/jci117206 (1994).
Bertani, L. et al. Assessment of serum cytokines predicts clinical and endoscopic outcomes to vedolizumab in ulcerative colitis patients. Br. J. Clin. Pharmacol. 86(7), 1296–1305. https://doi.org/10.1111/bcp.14235 (2020).
Gross, V. et al. Evidence for continuous stimulation of interleukin-6 production in Crohn’s disease. Gastroenterology 102(2), 514–519. https://doi.org/10.1016/0016-5085(92)90098-j (1992).
Danieli, M. G. et al. A machine learning analysis to predict the response to intravenous and subcutaneous immunoglobulin in inflammatory myopathies. A proposal for a future multi-omics approach in autoimmune diseases. Autoimmun. Rev. 21(6), 103105. https://doi.org/10.1016/j.autrev.2022.103105 (2022).
Ayoub, I. et al. Prediction models of treatment response in lupus nephritis. Kidney Int. 101(2), 379–389. https://doi.org/10.1016/j.kint.2021.11.014 (2022).
Dai, D. et al. GMrepo v2: A curated human gut microbiome database with special focus on disease markers and cross-dataset comparison. Nucleic Acids Res. 50(D1), D777-d784. https://doi.org/10.1093/nar/gkab1019 (2022).
Woetzel, D. et al. Identification of rheumatoid arthritis and osteoarthritis patients by transcriptome-based rule set generation. Arthritis Res. Ther. 16(2), R84. https://doi.org/10.1186/ar4526 (2014).
Keir, M. E. et al. Regulation and role of αE integrin and gut homing integrins in migration and retention of intestinal lymphocytes during inflammatory bowel disease. J. Immunol. 207(9), 2245–2254. https://doi.org/10.4049/jimmunol.2100220 (2021).
Broeren, M. G. et al. Disease-regulated gene therapy with anti-inflammatory interleukin-10 under the control of the CXCL10 promoter for the treatment of rheumatoid arthritis. Hum. Gene Ther. 27(3), 244–254. https://doi.org/10.1089/hum.2015.127 (2016).
Davis, S. & Meltzer, P. S. GEOquery: A bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23(14), 1846–1847. https://doi.org/10.1093/bioinformatics/btm254 (2007).
Gautier, L. et al. affy–analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20(3), 307–315. https://doi.org/10.1093/bioinformatics/btg405 (2004).
Leek, J. T. et al. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28(6), 882–883. https://doi.org/10.1093/bioinformatics/bts034 (2012).
Ringnér, M. What is principal component analysis?. Nat. Biotechnol. 26(3), 303–304. https://doi.org/10.1038/nbt0308-303 (2008).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods. 12(2), 115–121. https://doi.org/10.1038/nmeth.3252 (2015).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43(7), e47. https://doi.org/10.1093/nar/gkv007 (2015).
Wickham, F. R. H., Henry, L., Müller, K., Vaughan, D. dplyr: A Grammar of Data Manipulation_. R package version 1.1.2. (2023). https://CRAN.R-project.org/package=dplyr.
Langfelder, P. & Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 9, 559. https://doi.org/10.1186/1471-2105-9-559 (2008).
C. H. VennDiagram: Generate High-Resolution Venn and Euler Plots_.R package version 1.7.3.2022. https://CRAN.R-project.org/package=VennDiagram.
Szklarczyk, D. et al. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47(D1), D607-d613. https://doi.org/10.1093/nar/gky1131 (2019).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13(11), 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Bader, G. D. & Hogue, C. W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 4, 2. https://doi.org/10.1186/1471-2105-4-2 (2003).
Yu, G. et al. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16(5), 284–287. https://doi.org/10.1089/omi.2011.0118 (2012).
Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. Ser. B Methodol. 58(1), 267–288 (1996).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33(1), 1–2220808728 (2010).
Suykens, J. A. & Vandewalle, J. Least squares support vector machine classifiers. Neural Process. Lett. 9(3), 293–300 (1999).
Huang, M. L. et al. SVM-RFE based feature selection and Taguchi parameters optimization for multiclass SVM classifier. Sci. World J. 2014, 795624. https://doi.org/10.1155/2014/795624 (2014).
Villanueva, R. A. M. & Chen, Z. J. ggplot2: Elegant Graphics for Data Analysis (Springer, 2019).
K. A. ggpubr: 'ggplot2' Based Publication Ready Plots_.R package version 0.4.0.2020. https://CRAN.R-project.org/package=ggpubr.
Robin, X. et al. pROC: An open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform. 12, 77. https://doi.org/10.1186/1471-2105-12-77 (2011).
Zhang, X. et al. The oral and gut microbiomes are perturbed in rheumatoid arthritis and partly normalized after treatment. Nat. Med. 21(8), 895–905. https://doi.org/10.1038/nm.3914 (2015).
S. K.ggrepel: Automatically Position Non-Overlapping Text Labels with 'ggplot2'_.R package version 0.9.2.2022. https://CRAN.R-project.org/package=ggrepel.
Cheng, L. et al. gutMGene: A comprehensive database for target genes of gut microbes and microbial metabolites. Nucleic Acids Res. 50(D1), D795-d800. https://doi.org/10.1093/nar/gkab786 (2022).
Kuhn, M. et al. STITCH 4: Integration of protein-chemical interactions with user data. Nucleic Acids Res. 42(Database issue), D401–D407. https://doi.org/10.1093/nar/gkt1207 (2014).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods. 12(5), 453–457. https://doi.org/10.1038/nmeth.3337 (2015).
Bindea, G. et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39(4), 782–795. https://doi.org/10.1016/j.immuni.2013.10.003 (2013).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 14, 7. https://doi.org/10.1186/1471-2105-14-7 (2013).
R. C. Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria (2023). https://www.R-project.org/.
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Gao, J. et al. Role of chemokine (C-X-C Motif) ligand 10 (CXCL10) in renal diseases. Mediat. Inflamm. 2020, 6194864. https://doi.org/10.1155/2020/6194864 (2020).
Antonelli, A. et al. CXCR3, CXCL10 and type 1 diabetes. Cytokine Growth Factor Rev. 25(1), 57–65. https://doi.org/10.1016/j.cytogfr.2014.01.006 (2014).
Shibahara, T. et al. Characterization of epithelial chemoattractants for human intestinal intraepithelial lymphocytes. Gastroenterology 120(1), 60–70. https://doi.org/10.1053/gast.2001.20904 (2001).
Dwinell, M. B. et al. Regulated production of interferon-inducible T-cell chemoattractants by human intestinal epithelial cells. Gastroenterology 120(1), 49–59. https://doi.org/10.1053/gast.2001.20914 (2001).
Singh, U. P. et al. Inhibition of IFN-gamma-inducible protein-10 abrogates colitis in IL-10-/- mice. J. Immunol. 171(3), 1401–1406. https://doi.org/10.4049/jimmunol.171.3.1401 (2003).
Hyun, J. G. et al. Anti-interferon-inducible chemokine, CXCL10, reduces colitis by impairing T helper-1 induction and recruitment in mice. Inflamm. Bowel Dis. 11(9), 799–805. https://doi.org/10.1097/01.mib.0000178263.34099.89 (2005).
Patel, D. D., Zachariah, J. P. & Whichard, L. P. CXCR3 and CCR5 ligands in rheumatoid arthritis synovium. Clin. Immunol. 98(1), 39–45. https://doi.org/10.1006/clim.2000.4957 (2001).
Gao, J. et al. Bone marrow mesenchymal stem cells improve bone erosion in collagen-induced arthritis by inhibiting osteoclasia-related factors and differentiating into chondrocytes. Stem Cell Res. Ther. 11(1), 171. https://doi.org/10.1186/s13287-020-01684-w (2020).
Singh, K. P. et al. Intrahepatic CXCL10 is strongly associated with liver fibrosis in HIV-Hepatitis B co-infection. PLoS Pathog. 16(9), e1008744. https://doi.org/10.1371/journal.ppat.1008744 (2020).
Trivedi, P. J. & Adams, D. H. Chemokines and chemokine receptors as therapeutic targets in inflammatory bowel disease; Pitfalls and Promise. J. Crohns Colitis. 12(suppl_2), S641-s652. https://doi.org/10.1093/ecco-jcc/jjx145 (2018).
Yellin, M. et al. A phase II, randomized, double-blind, placebo-controlled study evaluating the efficacy and safety of MDX-1100, a fully human anti-CXCL10 monoclonal antibody, in combination with methotrexate in patients with rheumatoid arthritis. Arthritis Rheum. 64(6), 1730–1739. https://doi.org/10.1002/art.34330 (2012).
Catrina, A. I., Deane, K. D. & Scher, J. U. Gene, environment, microbiome and mucosal immune tolerance in rheumatoid arthritis. Rheumatology (Oxford) 55(3), 391–402. https://doi.org/10.1093/rheumatology/keu469 (2016).
Scher, J. U. et al. The lung microbiota in early rheumatoid arthritis and autoimmunity. Microbiome. 4(1), 60. https://doi.org/10.1186/s40168-016-0206-x (2016).
Ménard, S. et al. Lactic acid bacteria secrete metabolites retaining anti-inflammatory properties after intestinal transport. Gut 53(6), 821–828. https://doi.org/10.1136/gut.2003.026252 (2004).
Nagao-Kitamoto, H. & Kamada, N. Host-microbial cross-talk in inflammatory bowel disease. Immune Netw. 17(1), 1–12. https://doi.org/10.4110/in.2017.17.1.1 (2017).
Akram, M. Citric acid cycle and role of its intermediates in metabolism. Cell Biochem. Biophys. 68(3), 475–478. https://doi.org/10.1007/s12013-013-9750-1 (2014).
Harber, K. J. et al. Succinate is an inflammation-induced immunoregulatory metabolite in macrophages. Metabolites https://doi.org/10.3390/metabo10090372 (2020).
Basri, F. et al. CD1d deficiency limits tolerogenic properties of peritoneal macrophages. BMB Rep. 54(4), 209–214. https://doi.org/10.5483/BMBRep.2021.54.4.183 (2021).
Roberts, C. A., Dickinson, A. K. & Taams, L. S. The interplay between monocytes/macrophages and CD4(+) T cell subsets in rheumatoid arthritis. Front. Immunol. 6, 571. https://doi.org/10.3389/fimmu.2015.00571 (2015).
Rana, A. K. et al. Monocytes in rheumatoid arthritis: Circulating precursors of macrophages and osteoclasts and their heterogeneity and plasticity role in RA pathogenesis. Int. Immunopharmacol. 65, 348–359. https://doi.org/10.1016/j.intimp.2018.10.016 (2018).
Hedl, M. & Abraham, C. Distinct roles for Nod2 protein and autocrine interleukin-1beta in muramyl dipeptide-induced mitogen-activated protein kinase activation and cytokine secretion in human macrophages. J. Biol. Chem. 286(30), 26440–26449. https://doi.org/10.1074/jbc.M111.237495 (2011).
Lassen, K. G. et al. Genetic coding variant in GPR65 alters lysosomal pH and links lysosomal dysfunction with colitis risk. Immunity 44(6), 1392–1405. https://doi.org/10.1016/j.immuni.2016.05.007 (2016).
Baumgart, D. C. & Carding, S. R. Inflammatory bowel disease: Cause and immunobiology. Lancet 369(9573), 1627–1640. https://doi.org/10.1016/s0140-6736(07)60750-8 (2007).
Domínguez Conde, C. & Teichmann, S. A. Deciphering immunity at high plexity and resolution. Nat. Rev. Immunol. 20(2), 77–78. https://doi.org/10.1038/s41577-019-0254-0 (2020).
Zhu, W. et al. Disequilibrium of M1 and M2 macrophages correlates with the development of experimental inflammatory bowel diseases. Immunol. Investig. 43(7), 638–652. https://doi.org/10.3109/08820139.2014.909456 (2014).
Tisza, M. J. & Buck, C. B. A catalog of tens of thousands of viruses from human metagenomes reveals hidden associations with chronic diseases. Proc. Natl. Acad. Sci. U. S. A. https://doi.org/10.1073/pnas.2023202118 (2021).
Funding
This work was supported by the National Natural Science Foundation of China (Grant numbers 81973769) and Postgraduate Research & Practice Innovation Program of Jiangsu Province (Grant numbers KYCX22_1941).
Author information
Authors and Affiliations
Contributions
Y.G.: Methodology, investigation, writing-original draft, writing – review & editing, software, visualization. Y.Z.: Methodology, investigation. Y.Z.: Methodology, investigation. Y.W.: Conceptualization, supervision, writing—review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guan, Y., Zhang, Y., Zhu, Y. et al. CXCL10 as a shared specific marker in rheumatoid arthritis and inflammatory bowel disease and a clue involved in the mechanism of intestinal flora in rheumatoid arthritis. Sci Rep 13, 9754 (2023). https://doi.org/10.1038/s41598-023-36833-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-36833-7
- Springer Nature Limited