Abstract
Rho family GTPases regulate both linear and branched actin dynamics by activating downstream effectors to facilitate the assembly and function of complex cellular structures such as lamellipodia and contractile actomyosin rings. Wiskott-Aldrich Syndrome (WAS) family proteins are downstream effectors of Rho family GTPases that usually function in a one-to-one correspondence to regulate branched actin nucleation. In particular, the WAS protein Scar/WAVE has been shown to exhibit one-to-one correspondence with Rac GTPase. Here we show that Rac and SCAR are recruited to cell wounds in the Drosophila repair model and are required for the proper formation and maintenance of the dynamic actomyosin ring formed at the wound periphery. Interestingly, we find that SCAR is recruited to wounds earlier than Rac and is still recruited to the wound periphery in the presence of a potent Rac inhibitor. We also show that while Rac is important for actin recruitment to the actomyosin ring, SCAR serves to organize the actomyosin ring and facilitate its anchoring to the overlying plasma membrane. These differing spatiotemporal recruitment patterns and wound repair phenotypes highlight the Rac-independent functions of SCAR and provide an exciting new context in which to investigate these newly uncovered SCAR functions.
Similar content being viewed by others
Introduction
Cells encounter physical stresses daily leading to breaks in their cortex that must be rapidly repaired to maintain cell integrity and function1,2,3,4. One of the major features of cell wound repair is the assembly of a Rho family GTPase-dependent actomyosin ring at the wound periphery that attaches the cortical cytoskeleton to the overlying plasma membrane, followed by its dynamic translocation inward to pull the cell cortex breach closed5,6,7,8,9,10.
The three major Rho family GTPases—Rho, Rac, and Cdc42—are recruited to wounds in spatially and temporally distinct patterns to carry out specific functions during the repair process7,8,9,11. This dynamic recruitment of Rho family GTPases to the wound site varies somewhat among cell wound repair model systems. In the Xenopus model, all three GTPases are immediately recruited to the wound where they form two concentric rings with an interior ring consisting of Rho and an outer ring containing both Cdc42 and Rac8,9,12. In the Drosophila model, Rho GTPases are also rapidly recruited to wounds in concentric rings, but they exhibit both spatial and temporal differences7,13. Rho1 accumulates first at 30 s post-wounding, and similar to that observed in the Xenopus model, becomes enriched in a ring inside of and just overlapping with the inner edge of the actin ring. Cdc42 accumulates next (30–60 s post-wounding), whereas Rac 1 and Rac 2 accumulate last (60–90 s post-wounding). While both Cdc42 and Rac recruitment overlap with the actin ring, Rac also shows an additional broad region of slightly elevated accumulation overlapping the actin halo region.
While all three Rho family GTPases are needed to achieve proper cell wound repair, their functions also vary somewhat between cell wound repair models. In the Xenopus model, RhoA is needed for cortical flow, whereas Cdc42 regulates actin ring formation, and both are involved in actin ring translocation8,9,12. Rac function in this model has not been reported. In the Drosophila model, Rac proteins are necessary for actin recruitment to the wound edge, Cdc42 is needed to stabilize the actin ring at the wound periphery, and Rho1 is involved in actomyosin ring assembly/stabilization8,13.
Rho family GTPases are known to regulate actin dynamics through their regulation of linear and branched actin nucleation factors14,15,16,17,18,19. One such family of actin nucleation factors, the Wiskott-Aldrich Syndrome (WAS) family (WASp, Scar/WAVE, WASH), works with the Arp2/3 complex to promote branched actin nucleation15,19,20,21,22,23,24,25. In the context of cell wound repair, WAS family proteins have been shown to contribute to actin filament orientation and nucleate branched actin to serve as a scaffold to assemble and maintain the contractile actomyosin cable at the wound periphery10,19.
As Rho family GTPases usually exhibit a one-to-one correspondence with WAS family proteins (Cdc42 > WASp; Rac > Scar/WAVE; Rho1 > Wash)22,26,27,28,29,30,31,32,33,34,35,36, we expected Rac and SCAR to show similar recruitment to wounds and repair phenotypes. Surprisingly, we find that SCAR is recruited to wounds prior to Rac, does not require Rac for its recruitment to wounds, and exhibits Rac-independent repair phenotypes, suggesting that Rac and SCAR are functioning outside of their usual relationship in the cell wound repair context.
Results
SCAR is recruited to cell wounds before Rac GTPase
In the Drosophila cell wound repair model, actin is recruited to cell wounds in a dense ring around the wound periphery and in a less dense actin halo at 60 ± 6.3 s post-wounding (Fig. 1A-A’,H)37. Wounds were generated by laser ablation on the lateral side of nuclear cycle 4–6 Drosophila syncytial embryos expressing an actin reporter (sGMCA or sChMCA;38) (see “Methods”). Consistent with its requirement for cell wound repair, Rac is recruited to the wound edge. Wounding embryos expressing fluorescently tagged Rac1 (ChFP-Rac1) or Rac2 (GFP-Rac2) under the control of their endogenous promoters results in the strong accumulation of these proteins beginning at 60 ± 6.3 and 78 ± 4.9 s post-wounding, respectively, in a ring encircling the wound and in a less intense concentric ring corresponding to the actin ring and halo regions (Fig. 1B,C,E,F,H; Table 1)7,13.
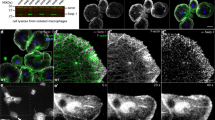
SCAR is recruited to cell wounds faster than Rac1 or Rac2. Confocal projection images of embryos expressing an actin reporter (sGMCA; A-A’) or co-expressing mCherry-Rac1 and sGMCA (B), GFP-Rac2 and sStMCA (C), and GFP-SCAR and sStMCA (D). Times post-wounding are indicated. UW = unwounded. Arrows indicate recruitment to the wound periphery. (E–G) Fluorescence intensity (arbitrary units) profiles across the wound area over time for the images shown in (B–D), respectively. (H) Dotplot of initial actin, Rac1, Rac2, and SCAR recruitment to wounds. Scale bars: 20 μm.
As Rho family GTPases usually exhibit a one-to-one correspondence with WAS family proteins (Rac1 > Scar/WAVE), we expected SCAR to show similar spatial and slightly delayed temporal recruitment to wounds as Rac. Wounding embryos expressing GFP-tagged SCAR (GFP-SCAR) under the control of its endogenous promoter resulted in the recruitment of SCAR to the wound periphery (Fig. 1D,G,H; Table 1). Surprisingly, SCAR is recruited to wounds earlier (33 ± 3 s post-wounding) than that observed for Rac1/Rac2 (60 s and 78 s post-wounding, respectively) (Fig. 1E,G,H; Table 1) and only overlapping the intense actin ring encircling the wound (i.e., absent from the actin halo region). Thus surprisingly, Rac1/Rac2 and SCAR recruitment to wounds is both temporally distinct and only partially overlapping spatially, suggesting that SCAR has Rac-independent functions in the context of cell wound repair.
SCAR recruitment to cell wounds does not require Rac activity
To further investigate the relationship between Rac and SCAR, we asked if recruitment of SCAR to cell wounds requires Rac. As Drosophila has three Rac genes (Rac1, Rac2, and Mtl), we modulated Rac activity by treating embryos with the potent Rac inhibitor NSC 237667,39. We examined GFP-SCAR recruitment to wounds in embryos injected with NSC 23766 (Fig. 2). Consistent with their different temporal recruitment patterns, GFP-SCAR was still recruited to wounds in these Rac inhibited embryos. The GFP-SCAR that is recruited to cell wounds does not form a well-defined actin ring, likely due to reduced actin recruitment to wounds upon Rac inhibition. Our results indicate that SCAR function is Rac-independent during cell wound repair.
SCAR is recruited to cell wounds in the absence of Rac activity. (A-A””) Confocal projection images of buffer-injected embryos co-expressing GFP-SCAR and an actin reporter (sStMCA) at the time points indicated. (B-B””) Fluorescence intensity (arbitrary units) profiles across the wound area in (A-A””), respectively. (C–C””) Confocal projection images of NSC 23766-injected embryos co-expressing GFP-SCAR and an actin reporter (sStMCA) at the time points indicated. (D-D””) Fluorescence intensity (arbitrary units) profiles across the wound area in (C–C””), respectively. (E) Dotplot of GFP-SCAR recruitment to wounds in control (buffer-injected) and NSC 23766-injected embryos at 240 s post-wounding. Scale bars: 20 μm.
Knockdown of Rac or SCAR results in distinct defects in wound healing dynamics
All three Rho family GTPases are required non-redundantly for cell wound repair. In particular, Rac is needed for the recruitment of actin to the wound edge, while Rho1 and Cdc42 are crucial for the formation and stability of the actin ring7,8,9,13. Interestingly, some actin organization persisted at the wound edge in embryos where Rho1 or Diaphanous (Dia; Drosophila formin protein and Rho1 downstream effector) were knocked down7, consistent with the differing roles for Rac and/or Cdc42 in the repair process.
We compared actin dynamics between embryos injected with NSC 23766 and SCAR knockdowns (Fig. 3; Table 2). In control (buffer injected) embryos, actin accumulates similarly to wildtype: in a highly enriched actomyosin ring bordering the wound edge and an elevated actin halo encircling the actin ring (Fig. 3A-C, J-L; Table 3; Video 1). As shown previously, Rac inhibition results in severely reduced recruitment of actin to the wound edge such that an actin ring is not formed, and significantly slower wound closure (control: 7.15 ± 0.25 μm2/s; NSC 23766: 5.62 ± 0.34 μm2/s, p < 0.05) (Fig. 3D–F,J–N; Table 3; Video 1)7. Rac inhibition also results in overexpansion of wounds (control: 1.74 ± 0.02 fold; NSC 23766: 2.59 ± 0.09 fold, p < 0.0001) (Fig. 3D-D’,K; Table 3). These phenotypes would be consistent with a role for branched actin nucleation factors as downstream effectors of Rac (and Cdc42) in this process.
Rac and SCAR knockdowns exhibit different cell wound repair phenotypes. (A–I) Confocal projection images of wounds generated in embryos expressing an actin marker (sGMCA) in control (buffer only; A–C), NSC 23766 injected (potent Rac inhibitor; D–F), or SCAR RNAi knockdowns (G–I). Actin ring and halo are indicated in (A). (A’,D’,G’) Kymographs across the wound area in (A,D,G), respectively. Wound expansion is noted with yellow lines; actomyosin ring formation with red arrows; and actomyosin ring disassembly with yellow arrows. (B,E,H) XY projection image at 0–150 s post wounding showing cortical flow of actin to the wound edge. (C,F,I) Vector maps from PIV (Particle Image Velocimetry) analysis depicting actin flow from 60 to 90 s for (A,B,D,E,G,H) respectively. (J) Quantification of the wound area over time for control (buffer injected), NSC 23766 injected, and SCAR RNAi knockdowns. (K–N) Quantification of fold wound expansion (K), wound contraction rate (L), actin ring width (M), and actin ring intensity (N). Black line and error bars represent mean ± SEM. Red dotted line and square represent mean ± 95% CI from control. Kruskal–Wallis test (K–L) and Welch’s t-test (M–N) were performed with * is p < 0.05, ** is p < 0.01, *** is p < 0.001, **** is p < 0.0001, and ns is not significant. Scale bars: 20 μm.
SCAR has been shown to contribute to actin filament orientation during actomyosin ring formation and translocation40, which would necessarily occur after actin recruitment to the wound periphery. Consistent with this and its Rac-independent recruitment to wounds, SCAR knockdowns exhibit different wound repair dynamics phenotypes from that of Rac inhibition. SCAR knockdowns also result in wound over-expansion (2.45 ± 0.11 fold, p < 0.001), but they exhibit a faster wound closure rate (9.12 ± 0.62 μm2/sec) than both control (p < 0.05) and Rac inhibited (p < 0.0001) embryos (Fig. 3G-G’, J-L; Table 3; Video 1). Importantly, unlike Rac inhibition, actin is recruited to the wound edge in SCAR knockdowns (Fig. 3H–I; Table 3). Despite this actin recruitment, the actomyosin ring that is formed is not as tightly organized as that in wildtype (Fig. 3G-H,M–N; Table 3), and the actomyosin ring undergoes premature disassembly (Fig. 3G’; Table 3).
Discussion
While Rac and SCAR are both needed for proper cell wound repair, they are doing so outside of their “Rac is the canonical activator of SCAR” relationship in this context. We find that Rac is necessary for actin recruitment to cell wounds, whereas SCAR is recruited to cell wounds in the absence of Rac activity and affects actin ring organization and contraction.
Scar/WAVE forms a heteropentameric complex with the Wave Regulatory Complex (WRC) composed of Sra1/PIR121, Nap1, Abi, and HSPC30025,28,41. Scar/WAVE, along with the Arp2/3 complex, can generate branched actin networks26,27,42,43. The Scar/WAVE–WRC complex is thought to be trans-inhibited44,45,46. Rac binds to the WRC to relieve this trans-inhibition, allowing Scar/WAVE to work with Arp2/3 to nucleate branched actin27,28,29,44. Rac does not regulate Scar/WAVE by binding directly to it, but rather interacts with it through binding to the Sra1 subunit of its WRC29. Actin is recruited to the wound periphery in SCAR knockdowns, but does not assemble into a robust actomyosin ring. Thus, in the context of cell wound repair, SCAR could bypass the need for Rac by a currently unknown mechanism such that it can use its branched actin nucleation activity. In this scenario, SCAR may need to be present at the wound edge quickly to put a branched actin scaffolding in place, such that additional actin can be organized into a robust ring upon its arrival at the wound edge. Alternatively, SCAR could be using an, as yet undescribed, alternate biochemical activity when at the wound edge. For example, in addition to its branched actin nucleation activity, the related Wiskott-Aldrich Syndrome family member WASH has been shown to bundle actin, bundle microtubules, and crosslink them35.
Surprisingly, despite the disruption of the actin ring, wounds in SCAR knockdowns close faster than in controls. A recent study showed that actomyosin contraction force is dependent on F-actin architectures47. While branched actin nucleation stabilizes F-actin networks by increasing their density and connectivity, this stabilization reduces actomyosin contractility by limiting movement of the myosin motor on F-actin. The three major branched actin nucleators (WASp, SCAR, and WASH) are required to form a robust actomyosin ring during cell wound repair in the Drosophila model19. While the actomyosin ring is disrupted in SCAR knockdowns, WASp and WASH could still generate branched F-actin to support the formation of an actomyosin ring at the wound edge. This disrupted actomyosin ring could result in increased actomyosin contractility due to its reduced F-actin network density and connectivity, thereby leading to the faster wound closure in SCAR knockdowns.
While Rac has known effectors other than Scar/WAVE that could work downstream to mediate its role in actin recruitment to wounds (cf.17,48,49), less is known about Scar/WAVE regulation outside of its activation through Rac GTPases. Strikingly, our data suggest that SCAR localization and its activity are not always dependent on Rac GTPase, which raises new mechanistic questions for SCAR/WAVE function. What protein(s) regulate SCAR recruitment to the cell cortex without Rac GTPase? While active Rac GTPase regulates SCAR activity through binding to Sra1 (WRC subunit), what other proteins can change the conformation of SCAR to release its VCA domain to promote branched actin nucleation? The linear actin nucleation factor Diaphanous (Dia) has been shown to function upstream of SCAR to regulate WRC localization and activity during Drosophila myoblast fusion50,51. Consistent with this possible regulation, Dia is rapidly recruited to the wound edge at 30 s post wounding, similar to that of SCAR40. SCAR may also depend on another Rho family GTPase. Cip4, a Cdc42 downstream effector, has been shown to function upstream of SCAR via associating with the WRC to control Dynamin-dependent cell polarization in the Drosophila wing52. Interestingly, recent studies have also implicated other means of activating the WRC in specific contexts such as lamellipodia formation through interaction with factors including Arf-family GTPases (Arf1, Arf6), other Rho family GTPases (Cdc42, RhoG), various kinases, phospholipids, or membrane receptors25,31,53,54,55,56,57. The identification of these instances of Rac-independent regulation for Scar/WAVE provides exciting new entry points for investigating the upstream control of this essential branched actin nucleation promoting factor.
Materials and methods
Fly stocks and genetics
Flies were cultured and crossed at 25 °C on yeast-cornmeal-molasses-malt extract medium. Flies used in this study are: ChFP-Rac1 (BDSC #76266)13, GFP-Rac2 (BDSC #52286)7; SCAR-GFP40, Vermillion RNAi (BDSC #50641; TRiP.HMC03041), and SCAR RNAi (BDSC #51803; TRiP.HMC0336). RNAi lines were driven using the maternally expressed GAL4-UAS driver, Pmatalpha-GAL-VP16V37 (BDSC #7063).
An actin reporter, sGMCA (spaghetti squash driven, moesin-alpha-helical-coiled and actin binding site fused to GFP) reporter38, the mCherry fluorescent equivalent, sChMCA (BDSC #35520), or the mScarlet-i fluorescent equivalent, sStMCA (BDSC #90928)58, was used to follow wound repair dynamics of the cortical cytoskeleton.
In this study, we used an actin reporter + maternal GAL4 driver + vermilion RNAi (unrelated fly RNAi) + injection buffer as the control.
Localization patterns and mutant analyses were performed at least twice with independent genetic crosses and ≥ 10 embryos were examined. Images representing the average phenotype were selected for figures.
Western Blotting
To generate embryo whole cell lysates, 10 nuclear cycles 4–6 embryos were collected, dechorionated, and then homogenized in 2X sample buffer (125 mM Tris–Cl pH 6.8, 4% SDS, 0.1% Bromophenol blue, 20% glycerol). Western blotting was performed according to standard procedures using anti-SCAR (P1C1, 1:10)59 antibodies, with anti-ATP5A (15H4C4; 1:50,000; Abcam) for the loading control.
Embryo handling and preparation
Nuclear cycle 4–6 embryos were collected for 30 min at 25 °C and harvested at room temperature (22 °C). Collected embryos were dechorionated by hand, mounted onto No. 1.5 coverslips coated with glue, and covered with Series 700 halocarbon oil (Halocarbon Products Corp.) as previously described37.
Drug injections
The pan Rac GTPase inhibitor NSC 23766 (50 mM; Tocris Bioscience) was injected into NC4-6 staged Drosophila embryos, incubated at room temperature (22 °C) for 5 min, and then subjected to laser wounding. NSC23766 was prepared in injection buffer (5 mM KCl, 0.1 mM NaP pH6.8). Injection buffer alone was used as the control.
Laser wounding
All wounds were generated using a pulsed nitrogen N2 micropoint laster (Andor Technology Ltd.) set to 435 nm and focused on the lateral surface of the embryo. An 18 × 18 μm circular region was set as the target site along the lateral midsection of the embryo, and ablation was controlled by MetaMorph software (Molecular Devices). Average ablation time was less than 3 s and time-lapse image acquisition was initiated immediately after ablation.
Live image acquisition
All live imaging was performed at room temperature with the following microscope:
-
(1)
Revolution WD systems (Andor Technology Ltd.) mounted on a Leica DMi8 (Leica Microsystems Inc.) with a 63x/1.4 NA objective lens under the control of MetaMorph software (Molecular devices). Images were captured using 488 nm and/or 561 nm lasers with a Yokogawa CSU-W1 confocal spinning disk head attached to an Andor iXon Ultra 897 EMCCD camera (Andor Technology Ltd.). Time-lapse images were acquired with 17–20 µm stacks/0.25 µm steps. Images were acquired every 30 s for 15 min and then every 60 s for 15 min.
-
(2)
UltraVIEW VoX Confocal Imaging System (Perkin Elmer, Waltham, MA, USA) mounted on a Nikon Eclipse Ti (Nikon Instruments, Melville NY,USA) with a 60x/1.4 NA objective lens under the control of Volocity software(v.5.3.0, Perkin-Elmer). Images were captured using 488 nm and/or 561 nm lasers with a Yokogawa CSU-X1 confocal spinning disk head attached to a Hamamatsu C9100-13 EMCCD camera (Perkin-Elmer, Waltham,MA,USA). Time-lapse images were acquired with 17–20 µm stacks/0.25 µm steps. Images were acquired every 30 s for 15 min and then every 60 s for 15 min.
Image processing, analysis, and quantification
Image processing was performed using FIJI software60. In all images, the top side is anterior and the bottom side is posterior of embryos. Kymographs were generated using the crop feature to select ROIs of 5.3 × 94.9 µm. Wound area was manually measured using Fiji and the values were imported into Prism 8.2.1 (GraphPad Software Inc.) to construct corresponding graphs.
For fluorescent lineplots, the mean fluorescence profile intensities were calculated from 51 equally spaced radial profiles anchored at the center of the wound, swept from 0° to 180°40. Radial profiles of 301-pixel diameter were used. Fluorescence intensity profiles were calculated and averaged using an in house code using MATLAB R2020b (MathWorks) (available at: https://github.com/FredHutch/wound_radial_lineplot), then plotted using MATLAB R2020b. For dynamic lineplots, we generated fluorescent profile plots from each timepoint and then concatenated them. The lines represent the averaged fluorescent intensity and gray area is the 95% confidence interval.
Quantification of the width and average intensity of actin ring, wound expansion, and closure rate was performed as follows: the width of actin ring was calculated with two measurements, the feret diameters of the outer and inner edge of actin ring at 90 s post-wounding. Using these measurements, the width of actin ring was calculated with (outer feret diameter − inner feret dimeter)/2. The average intensity of actin ring was calculated with two measurements. Instead of measuring feret diameters, we measured area and integrated intensity in same regions as described in ring width. Using these measurements, the average intensity in the actin ring was calculated with (outer integrated intensity − inner integrated intensity)/(outer area − inner area). To calculate relative intensity for unwounded (UW) time point, average intensity at UW was measured with 50 × 50 pixels at the center of embryos and then averaged intensity of actin ring at each timepoint was divided by average intensity of UW. Wound expansion was calculated with max wound area/initial wound size. Closure rate was calculated with two time points, one is tmax that is the time of reaching maximum wound area, the other is t < half that is the time of reaching 50–35% size of max wound since the slope of wound area curve changes after t < half. Using these time points, average speed was calculated with (wound area at tmax − wound area at t < half)/tmax-t < half.
Figures were assembled in Canvas Draw 6 for Mac (Canvas GFX, Inc.).
Statistical analysis
All statistical analysis was done using Prism 8.2.1 (GraphPad, San Diego, CA). Statistical significance was calculated using the Kruskal–Wallis or Welch’s t tests with * is p < 0.05, ** is p < 0.01, *** is p < 0.001, **** is p < 0.0001, and ns is not significant.
Data availability
All data generated and/or analyzed during this study are included in this published article and its supplementary video.
References
Cooper, S. T. & McNeil, P. L. Membrane repair: Mechanisms and pathophysiology. Physiol. Rev. 95, 1205–1240. https://doi.org/10.1152/physrev.00037.2014 (2015).
Howard, A. C., McNeil, A. K., Xiong, F., Xiong, W. C. & McNeil, P. L. A novel cellular defect in diabetes: Membrane repair failure. Diabetes 60, 3034–3043. https://doi.org/10.2337/db11-0851 (2011).
Galan, J. E. & Bliska, J. B. Cross-talk between bacterial pathogens and their host cells. Annu. Rev. Cell Dev. Biol. 12, 221–255. https://doi.org/10.1146/annurev.cellbio.12.1.221 (1996).
Velnar, T., Bailey, T. & Smrkolj, V. The wound healing process: An overview of the cellular and molecular mechanisms. J. Int. Med. Res. 37, 1528–1542. https://doi.org/10.1177/147323000903700531 (2009).
Sonnemann, K. J. & Bement, W. M. Wound repair: Toward understanding and integration of single-cell and multicellular wound responses. Annu. Rev. Cell Dev. Biol. 27, 237–263. https://doi.org/10.1146/annurev-cellbio-092910-154251 (2011).
Nakamura, M. et al. Into the breach: How cells cope with wounds. Open Biol. 8, 180135. https://doi.org/10.1098/rsob.180135 (2018).
Abreu-Blanco, M. T., Verboon, J. M. & Parkhurst, S. M. Coordination of Rho family GTPase activities to orchestrate cytoskeleton responses during cell wound repair. Curr. Biol. 24, 144–155. https://doi.org/10.1016/j.cub.2013.11.048 (2014).
Benink, H. A. & Bement, W. M. Concentric zones of active RhoA and Cdc42 around single cell wounds. J. Cell Biol. 168, 429–439. https://doi.org/10.1083/jcb.200411109 (2005).
Moe, A. et al. Cross-talk-dependent cortical patterning of Rho GTPases during cell repair. Mol. Biol. Cell 32, 1417–1432. https://doi.org/10.1091/mbc.E20-07-0481 (2021).
Hui, J., Stjepic, V., Nakamura, M. & Parkhurst, S. M. Wrangling actin assemblies: Actin ring dynamics during cell wound repair. Cells https://doi.org/10.3390/cells11182777 (2022).
Simon, C. M., Vaughan, E. M., Bement, W. M. & Edelstein-Keshet, L. Pattern formation of Rho GTPases in single cell wound healing. Mol. Biol. Cell 24, 421–432. https://doi.org/10.1091/mbc.E12-08-0634 (2013).
Golding, A. E., Visco, I., Bieling, P. & Bement, W. M. Extraction of active RhoGTPases by RhoGDI regulates spatiotemporal patterning of RhoGTPases. Elife 8, 50471. https://doi.org/10.7554/eLife.50471 (2019).
Nakamura, M., Verboon, J. M. & Parkhurst, S. M. Prepatterning by RhoGEFs governs Rho GTPase spatiotemporal dynamics during wound repair. J. Cell Biol. 216, 3959–3969. https://doi.org/10.1083/jcb.201704145 (2017).
Goley, E. D. & Welch, M. D. The ARP2/3 complex: An actin nucleator comes of age. Nat. Rev. Mol. Cell Biol. 7, 713–726. https://doi.org/10.1038/nrm2026 (2006).
Campellone, K. G. & Welch, M. D. A nucleator arms race: Cellular control of actin assembly. Nat. Rev. Mol. Cell Biol. 11, 237–251. https://doi.org/10.1038/nrm2867 (2010).
Olson, M. F. Rho GTPases, their post-translational modifications, disease-associated mutations and pharmacological inhibitors. Small GTPases 9, 203–215. https://doi.org/10.1080/21541248.2016.1218407 (2018).
Mosaddeghzadeh, N. & Ahmadian, M. R. The RHO family GTPases: Mechanisms of regulation and signaling. Cells 10, 1831. https://doi.org/10.3390/cells10071831 (2021).
Narumiya, S. & Thumkeo, D. Rho signaling research: History, current status and future directions. FEBS Lett. 592, 1763–1776. https://doi.org/10.1002/1873-3468.13087 (2018).
Hui, J., Nakamura, M., Dubrulle, J. & Parkhurst, S. M. Coordinated efforts of different actin filament populations are needed for optimal cell wound repair. Mol. Biol. Cell 34, ar15. https://doi.org/10.1091/mbc.E22-05-0155 (2023).
Millard, T. H., Sharp, S. J. & Machesky, L. M. Signalling to actin assembly via the WASP (Wiskott-Aldrich syndrome protein)-family proteins and the Arp2/3 complex. Biochem. J. 380, 1–17. https://doi.org/10.1042/BJ20040176 (2004).
Takenawa, T. & Suetsugu, S. The WASP-WAVE protein network: Connecting the membrane to the cytoskeleton. Nat. Rev. Mol. Cell Biol. 8, 37–48. https://doi.org/10.1038/nrm2069 (2007).
Burianek, L. E. & Soderling, S. H. Under lock and key: Spatiotemporal regulation of WASP family proteins coordinates separate dynamic cellular processes. Semin. Cell Dev. Biol. 24, 258–266. https://doi.org/10.1016/j.semcdb.2012.12.005 (2013).
Massaad, M. J., Ramesh, N. & Geha, R. S. Wiskott-Aldrich syndrome: A comprehensive review. Ann. N. Y. Acad. Sci. 1285, 26–43. https://doi.org/10.1111/nyas.12049 (2013).
Gautreau, A. M., Fregoso, F. E., Simanov, G. & Dominguez, R. Nucleation, stabilization, and disassembly of branched actin networks. Trends Cell Biol. https://doi.org/10.1016/j.tcb.2021.10.006 (2021).
Kramer, D. A., Piper, H. K. & Chen, B. WASP family proteins: Molecular mechanisms and implications in human disease. Eur. J. Cell Biol. 101, 151244. https://doi.org/10.1016/j.ejcb.2022.151244 (2022).
Machesky, L. M. & Insall, R. H. Scar1 and the related Wiskott-Aldrich syndrome protein, WASP, regulate the actin cytoskeleton through the Arp2/3 complex. Curr. Biol. 8, 1347–1356. https://doi.org/10.1016/s0960-9822(98)00015-3 (1998).
Miki, H., Suetsugu, S. & Takenawa, T. WAVE, a novel WASP-family protein involved in actin reorganization induced by Rac. EMBO J. 17, 6932–6941. https://doi.org/10.1093/emboj/17.23.6932 (1998).
Eden, S., Rohatgi, R., Podtelejnikov, A. V., Mann, M. & Kirschner, M. W. Mechanism of regulation of WAVE1-induced actin nucleation by Rac1 and Nck. Nature 418, 790–793. https://doi.org/10.1038/nature00859 (2002).
Chen, B. et al. Rac1 GTPase activates the WAVE regulatory complex through two distinct binding sites. Elife 6, 29795. https://doi.org/10.7554/eLife.29795 (2017).
Schaks, M. et al. Distinct interaction sites of Rac GTPase with WAVE regulatory complex have non-redundant functions in vivo. Curr. Biol. 28, 3674–3684. https://doi.org/10.1016/j.cub.2018.10.002 (2018).
Rottner, K., Stradal, T. E. B. & Chen, B. WAVE regulatory complex. Curr. Biol. 31, R512–R517. https://doi.org/10.1016/j.cub.2021.01.086 (2021).
Ding, B. et al. Structures reveal a key mechanism of WAVE regulatory complex activation by Rac1 GTPase. Nat. Commun. 13, 5444. https://doi.org/10.1038/s41467-022-33174-3 (2022).
Steffen, A. et al. Rac function is crucial for cell migration but is not required for spreading and focal adhesion formation. J. Cell Sci. 126, 4572–4588. https://doi.org/10.1242/jcs.118232 (2013).
Buracco, S., Claydon, S. & Insall, R. Control of actin dynamics during cell motility. F1000Res 8, 18669. https://doi.org/10.12688/f1000research.18669.1 (2019).
Liu, R. et al. Wash functions downstream of Rho and links linear and branched actin nucleation factors. Development 136, 2849–2860. https://doi.org/10.1242/dev.035246 (2009).
Verboon, J. M., Rahe, T. K., Rodriguez-Mesa, E. & Parkhurst, S. M. Wash functions downstream of Rho1 GTPase in a subset of Drosophila immune cell developmental migrations. Mol. Biol. Cell 26, 1665–1674. https://doi.org/10.1091/mbc.E14-08-1266 (2015).
Abreu-Blanco, M. T., Verboon, J. M. & Parkhurst, S. M. Cell wound repair in Drosophila occurs through three distinct phases of membrane and cytoskeletal remodeling. J. Cell Biol. 193, 455–464. https://doi.org/10.1083/jcb.201011018 (2011).
Kiehart, D. P., Galbraith, C. G., Edwards, K. A., Rickoll, W. L. & Montague, R. A. Multiple forces contribute to cell sheet morphogenesis for dorsal closure in Drosophila. J. Cell Biol. 149, 471–490. https://doi.org/10.1083/jcb.149.2.471 (2000).
Gao, Y., Dickerson, J. B., Guo, F., Zheng, J. & Zheng, Y. Rational design and characterization of a Rac GTPase-specific small molecule inhibitor. Proc. Natl. Acad. Sci. U.S.A. 101, 7618–7623. https://doi.org/10.1073/pnas.0307512101 (2004).
Hui, J., Nakamura, M., Dubrulle, J. & Parkhurst, S. M. Cell wound repair requires the coordinated action of linear and branched actin nucleation factors. bioRxiv https://doi.org/10.1101/2022.04.14.488397 (2022).
Stovold, C. F., Millard, T. H. & Machesky, L. M. Inclusion of Scar/WAVE3 in a similar complex to Scar/WAVE1 and 2. BMC Cell Biol 6, 11. https://doi.org/10.1186/1471-2121-6-11 (2005).
Pollitt, A. Y. & Insall, R. H. WASP and SCAR/WAVE proteins: The drivers of actin assembly. J. Cell Sci. 122, 2575–2578. https://doi.org/10.1242/jcs.023879 (2009).
Bear, J. E., Rawls, J. F. & Saxe, C. L. 3rd. SCAR, a WASP-related protein, isolated as a suppressor of receptor defects in late Dictyostelium development. J. Cell Biol. 142, 1325–1335. https://doi.org/10.1083/jcb.142.5.1325 (1998).
Rotty, J. D., Wu, C. & Bear, J. E. New insights into the regulation and cellular functions of the ARP2/3 complex. Nat. Rev. Mol. Cell Biol. 14, 7–12. https://doi.org/10.1038/nrm3492 (2013).
Ismail, A. M., Padrick, S. B., Chen, B., Umetani, J. & Rosen, M. K. The WAVE regulatory complex is inhibited. Nat. Struct. Mol. Biol. 16, 561–563. https://doi.org/10.1038/nsmb.1587 (2009).
Derivery, E., Lombard, B., Loew, D. & Gautreau, A. The Wave complex is intrinsically inactive. Cell Motil. Cytoskelet. 66, 777–790. https://doi.org/10.1002/cm.20342 (2009).
Muresan, C. G. et al. F-actin architecture determines constraints on myosin thick filament motion. Nat. Commun. 13, 7008. https://doi.org/10.1038/s41467-022-34715-6 (2022).
Takai, Y., Sasaki, T. & Matozaki, T. Small GTP-binding proteins. Physiol. Rev. 81, 153–208. https://doi.org/10.1152/physrev.2001.81.1.153 (2001).
Bustelo, X. R., Sauzeau, V. & Berenjeno, I. M. GTP-binding proteins of the Rho/Rac family: Regulation, effectors and functions in vivo. BioEssays 29, 356–370. https://doi.org/10.1002/bies.20558 (2007).
Deng, S., Bothe, I. & Baylies, M. Diaphanous regulates SCAR complex localization during Drosophila myoblast fusion. Fly 10, 178–186. https://doi.org/10.1080/19336934.2016.1195938 (2016).
Deng, S., Bothe, I. & Baylies, M. K. The formin diaphanous regulates myoblast fusion through actin polymerization and Arp2/3 regulation. PLoS Genet. 11, e1005381. https://doi.org/10.1371/journal.pgen.1005381 (2015).
Fricke, R. et al. Drosophila Cip4/Toca-1 integrates membrane trafficking and actin dynamics through WASP and SCAR/WAVE. Curr. Biol. 19, 1429–1437. https://doi.org/10.1016/j.cub.2009.07.058 (2009).
Schaks, M. et al. RhoG and Cdc42 can contribute to Rac-dependent lamellipodia formation through WAVE regulatory complex-binding. Small GTPases 12, 122–132. https://doi.org/10.1080/21541248.2019.1657755 (2021).
Koronakis, V. et al. WAVE regulatory complex activation by cooperating GTPases Arf and Rac1. Proc. Natl. Acad. Sci. U.S.A. 108, 14449–14454. https://doi.org/10.1073/pnas.1107666108 (2011).
Singh, S. P. et al. Cell-substrate adhesion drives Scar/WAVE activation and phosphorylation by a Ste20-family kinase, which controls pseudopod lifetime. PLoS Biol. 18, e3000774. https://doi.org/10.1371/journal.pbio.3000774 (2020).
Singh, V., Davidson, A. C., Hume, P. J. & Koronakis, V. Arf6 Can Trigger Wave Regulatory Complex-Dependent Actin Assembly Independent of Arno. Int. J. Mol. Sci. 21, 2457. https://doi.org/10.3390/ijms21072457 (2020).
Buracco, S. et al. The Scar/WAVE complex drives normal actin protrusions without the Arp2/3 complex, but proline-rich domains are required. bioRxiv https://doi.org/10.1101/2022.05.14.491902 (2022).
Nakamura, M., Verboon, J. M., Prentiss, C. L. & Parkhurst, S. M. The kinesin-like protein Pavarotti functions noncanonically to regulate actin dynamics. J. Cell Biol. 219, 2117. https://doi.org/10.1083/jcb.201912117 (2020).
Rodriguez-Mesa, E., Abreu-Blanco, M. T., Rosales-Nieves, A. E. & Parkhurst, S. M. Developmental expression of Drosophila Wiskott-Aldrich Syndrome family proteins. Dev. Dyn. 241, 608–626. https://doi.org/10.1002/dvdy.23742 (2012).
Schindelin, J. et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 9, 676–682. https://doi.org/10.1038/nmeth.2019 (2012).
Acknowledgements
We thank the Fred Hutch/Leica Center of Excellence (RRID:SCR_022720), Murdoch Charitable Trust, FlyBase, the Bloomington Stock Center, the Harvard Transgenic RNAi Project, and the Developmental Studies Hybridoma Bank for information, microscopes, DNAs, flies, and other reagents used in this study.
Funding
This research was supported by NIH HD095798 and the Mark Groudine Chair for Outstanding Achievements in Science and Service (to SMP) and NIH/NCI Cancer Center Support Grant P30 CA015704 (for Cellular Imaging Shared Resource).
Author information
Authors and Affiliations
Contributions
All authors contributed to the design of the experiments, performed experiments, analyzed data, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Video 1.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nakamura, M., Hui, J., Stjepić, V. et al. Scar/WAVE has Rac GTPase-independent functions during cell wound repair. Sci Rep 13, 4763 (2023). https://doi.org/10.1038/s41598-023-31973-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-31973-2
- Springer Nature Limited