Abstract
To systematically explore and analyze the microbial composition and function of microbial consortium M44 with straw degradation in the process of subculture at low temperature. In this study, straw degradation characteristics of samples in different culture stages were determined. MiSeq high-throughput sequencing technology was used to analyze the evolution of community structure and its relationship with degradation characteristics of microbial consortium in different culture periods, and the PICRUSt function prediction analysis was performed. The results showed that straw degradation rate, endoglucanase activity, and filter paper enzyme activity of M44 generally decreased with increasing culture algebra. The activities of xylanase, laccase, and lignin peroxidase, as well as VFA content, showing a single-peak curve change with first an increase and then decrease. In the process of subculture, Proteobacteria, Bacteroidetes, and Firmicutes were dominant in different culture stages. Pseudomonas, Flavobacterium, Devosia, Brevundimonas, Trichococcus, Acinetobacter, Dysgonomonas, and Rhizobium were functional bacteria in different culture stages. It was found by PICRUSt function prediction that the functions were concentrated in amino acid transport and metabolism, carbohydrate transship and metabolism related genes, which may contain a large number of fibers and lignin degrading enzyme genes. In this study, the microbial community succession and the gene function in different culture periods were clarified and provide a theoretical basis for screening and rational utilization of microbial consortia.
Similar content being viewed by others
Introduction
Lignocellulose is one of the most abundant renewable carbon sources in the biosphere, and its resource utilization efficiency has potential significance for sustainable development and environmental protection1,2. However, inefficient lignocellulose deconstruction is a primary bottleneck for its economic conversion and further utilization (i.e., of hemicellulose and cellulose, which are enclosed by lignin)3,4, especially under low-temperature conditions. Biodegradation, accomplished through coordination of various microorganisms, is currently considered a highly efficient method for lignocellulosic degradation5,6. Previous studies have shown that more efficient and suitable strains can be screened from similar ecological environments according to application purposes7. And complete degradation of lignocellulose requires the synergistic action of various microorganisms in the natural environment8,9,10. Zheng et al.11 obtained the lignocellulose-degrading bacterium LTF-27 from cold perennial forest soil, which mainly composed of Parabacteroides, Alcaligenes, Lysinibacillus, Sphingobacterium, and Clostridium. Alessi et al.12 showed that Asticcacaulis, Leadbetterella, and Truepera played a key role in wheat straw degradation. Wang et al.'s13 study showed that a lignin-degrading composite bacteria LDC was obtained from the root soil of rotten reed straw by restricted enrichment culture method, Pseudomonas, Pannonibacter, Thauera, Ruminofilibacter and Anaerocolumna were main bacteria that have played an important role in the process of corn straw degradation14.
To promote the in situ return of corn straw to the field in the low-temperature growing area in northern of China, the dried dung from the low temperature (−7 to 8 ℃) ecological environment was used as the original material for screening corn straw degradation microbial consortium at low temperature by our research team. Finally, a microbial consortium M44 was obtained by low temperature restriction subculture, and the straw degradation rate in laboratory was more than 30%15. However, the correlation of species composition and the succession rule of microbial community of the M44 in different culture periods are not clear. Therefore, in this study, original samples and series of bacteria in different subculture periods of the screened microbial consortium M44 were used as materials to measure its degradation characteristics and the composition and relative abundance of microbial classification units in the process of subculture at low temperature were analyzed by using 16S rRNA gene amplification method. And the relationship between the microbial community composition and the degradation characteristics of the M44 in different culture stages was revealed, and the gene function was explored preliminary.
Materials and methods
Experiment materials
Air-dried sheep dung (S) was taken from Chenbarhu Banner, Hulunbuir City, Inner Mongolia, China (125°07′ E, 46°28′ N). The tested straw degradation microbial consortium M44 was screened by our laboratory15.
Corn straw was taken from the experimental field of the Corn Center of Inner Mongolia Agricultural University (110°28′ E, 40°32′ N), and the cellulose, hemicellulose and lignin contents were determined to be 47.23%, 34.34%, and 16.77%, respectively. Corn straw of moderate thickness and with no pests or disease was selected, washed and dried (60 °C), and cut into small pieces of 2–3 cm for use.
Medium and culture conditions
Mandels medium (M medium) was composed of K2HPO4 (3.0 g), NaNO3 (3.0 g), CaCl2 (0.5 g), MgSO4·7H2O (0.5 g), Fe2SO4·7H2O (7.5 mg), MnSO4·H2O (2.5 mg), ZnSO4 (2.0 mg), CoCl2 (3.0 mg), and distilled water (1 L). Then, 40 mL M medium and 1.0 g corn straw were added to a 100-mL triangular flask for subsequent subculture, and the corn straw degradation ratio and enzyme activity were measured. Following this, the mixtures were sterilized at 121 °C for 20 min and set aside 15.
Cultivation of microbial consortium
2 g dried dung was put into a triangular bottle filled with 40 mL sterile distilled water and glass beads and placed on a shaker at 15 °C for 2 h. Then, 5% (V/V) supernatant was absorbed in 40 mL M medium, corn straw was used as the substrate carbon source, and cultured at 15 °C for 21 days. After culturing for 21 days, the fermentation liquid at an inoculation rate of 5% (V/V) was transferred to new M medium and cultured successively until the 11th generation (F11). Among them, the F1, F5, F8, and F11 generations of M44 were stored at –80 °C for later use.
Determination of straw degradation characteristics
The fermentation broth of microbial consortium M44 at F1, F5, F8, and F11 generations was inoculated into 40 mL M medium at 5% (V/V) and cultured at 15 °C for 21 days. Then, the corn straw degradation ratio was determined using the weight loss method. 5 mL of fermentation material was centrifuged at 12,000 rpm at 4 °C for 10 min and the supernatants were used as extracellular crude enzyme samples to analyze the enzyme activities and volatile fatty acid (VFA) content in F1, F5, F8, and F11 generations. Filter paper enzyme activity and endonuclease 1,4-β-glucanase activity were assessed using the DNS method15, and xylanase activity was established by DNS method16. The activity of laccase was assessed using the ABTS method, and that of lignin peroxidase was examined according to the resveratrol method17. One milliliter supernatant was integrated into a 1.5-mL centrifuge tube, and an acid adsorbent was added to an Agilent GC-6890 N meteorological chromatographic column was an Agilent l9091F-112 (length 30 m, inner diameter 0.32 mm, film thickness 0.5 µm). The column temperature was raised gradually. The gasification chamber temperature was 220 °C, the detector temperature was 240 °C, the carrier gas was hydrogen, the flow rate was 30.0 mL min–1, and the tail was blowing at 30 mL min−1 injection volumes 1 µL.
Microbial composition analysis of S, F1, F5, F8 and F11 generations
Under aseptic conditions, genomic DNA from the original sample(S) and different culture periods (F1, F5, F8, F11) of M44 was extracted using a bacterial genomic DNA extraction kit (China, Tiengen Biochemical Technology Co., Ltd.), and a 1% agarose gel was used for electrophoresis. A NanoDrop 2000 UV-V spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA) was used to determine the concentration and purity of DNA. The hypervariable region V3–V5 of the bacterial 16S rRNA gene was amplified using primer pair 338F (5'-ACTCCTACGGGAGCAG-3') and 806R (5'-GGACTACHVGGGTWTCTAAT-3') with an ABI Gene Amp® 9700 PCR thermonuclear system (Applied Biosystems, Thermo Fischer Scientific)18. PCR amplification system: TransStart Fastpfu DNA Polymerase, 20 μL reaction system, including 4 μL 5× Fast Pfu Buffer, 2 μL 2.5 mmol dNTPs, 0.8 μL Forward Primer(5 μmol/L), 0.8 μL Reverse Primer (5 μmol/L), 0.4 μL FastPfu Polymerase, 0.2 μL BSA, 10 ng Template DNA, supplement ddH2O to 20 μL. PCR reaction parameters: (a) 1× (3 min at 95 ℃); (b) recurring number × (30 s at 95 ℃); 30 s at 55 ℃; 45 s at 72 ℃); (c)10 min at 72 ℃, 10 ℃ until halted by user. PCR amplification products were sent to Shanghai Meiji Biomedical Technology Co., Ltd. for sequencing.
PICRUSt function predictive analysis
In order to explore the function of dominant microbial community in straw degradation, PICRUSt was used to predict the metagenomic function composition of 16S RNA amplicon database19. In this method, the 16S RNA ampland database obtained by high-throughput sequencing was compared with the Greengenes database to obtain the functional information of OUT corresponding species. The functional composition of colonies was predicted based on the latest Kyoto Encyclopedia of Genes and Genomes (KEGG) database information.
Data processing
Data on straw degradation characteristics were analyzed in IBM SPSS Statistics 25.0 (IBM Inc., Armonk, NY, USA, https://www.ibm.com/cn-zh/analytics/spss-statistics-software), R (V3.6.1) (https://www.r-project.org/) and Origin 2018 (https://www.Originlab.com) were used to create figures.
Results and analysis
Changes of straw degradation characteristics at different culture stages
Corn straw degradation ratio
Corn straw weight loss in M44 at F1 reached 35.90% at 15 ℃ for 21 days, which was greater than that at F5, F8, and F11 by 2.33%, 3.01%, and 3.35%, respectively. There were no significant differences between F8 and F11(Fig. 1).
Enzyme activities
The highest endoglucanase and filter paper enzyme activities were 2.01 and 2.16 U mL−1 in F1, respectively, which was significantly higher than that of F8 and F11(Fig. 2a,b). Xylanase activities was the highest in F5, with enzyme activities of 21.50 U mL−1, and the enzyme activities of F1 and F5 were significantly higher than those of F8 and F11(Fig. 2c). Laccase and lignin peroxidase activity reached 101.02 and 80.37 U L−1 at F5, which was greatly different than that of other algebras (Fig. 2d,e).
VFA content
Acetic acid, prophetic acid, and butyric acid contents were all the highest at F5. Acetic acid and propionic acid contents were 143.91 mmol L−1 and 6.70 mmol L−1 at F5, respectively, which were significantly different from those at F1 and F8 (Fig. 3a,b). Butyric acid was 3.80 mmol L−1 in F5, which was significantly different from that in F1 but not from that in F8 and F11(Fig. 3c).
Alpha diversity of microorganisms at different culture stages
Alpha diversity was used as a measure of microbial community diversity within the sample. The Ace and Sobs indexes for the original samples(S) were 1358.22 and 1101.33, respectively, which were significantly higher than those of F1, F5, F8, F11. Shannon and Simpson indexes showed the opposite trend, with values of 2.59 and 0.161 in the F5 generation, respectively, indicating that the microbial community was more abundant and diverse in this culture stage (Fig. 4).
Beta diversity of microorganisms at different culture stages
A principal component analysis (PCA) was conducted on the bacterial community in each group of samples, and the results was shown in Fig. 5. The contribution rates of PC1 and PC2 were 42.64% and 23.3% of the total, respectively. Samples of S and F1, F8, and F11 clustered together, indicating that the composition of microbial communities in these two groups was similar. On the other hand, samples from F5 clustered far from each other and into a single cluster, indicating that the microbial community compositions of the F5 samples were significantly different from those of other periods. To further define the differences, ANOSIM and PERMANOVA were performed at the OTU level based on the Bray–Curtis distance algorithm. The results showed that there were significant differences between different stages (p < 0.05; N = 999 permutations).
Taxonomic composition analyses at different culture stages
Phylum level
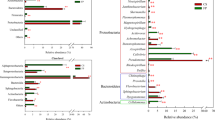
The relative abundance of bacterial groups according to classification level was shown in Fig. 6. At the phylum level, microbial consortium M44 was mainly composed of Proteobacteria, Bacteroidetes, Firmicutes, Actinobacteria, and Verrucomicrobia. Among these, Proteobacteria was dominant in M44, with abundances of 56.84%, 87.09%, 61.64%, and 53.94% at F1, F5, F8, and F11, respectively. Bacteroidota accounted for 32.11% of the total bacterial content in the F1, which was significantly higher than that of S, F5, F8, F11. The relative abundance of Firmicutes increased steadily, accounting for 26.80% of the total in F11, which was considerably higher than that at F1 (5.83%), F5 (7.68%), and F8 (17.37%). The relative abundance of Actinobacteriota in the original sample(S) was 41.19%, but decreased with subculture of the microbial consortium. The relative abundance of Verrucomicrobiota fluctuated with increasing culture stage, increasing to 3.03% in F11.
Genus level
At the genus level (Fig. 7), the relative abundance of Pseudomonas was 0.46% in the original sample(S), with its abundance was shown increased first and decreased then with culture time, reaching 8.75% in F11. The relative abundance of Brevundimonas was highest in the F1, was 10.79%, and which was significantly different from that in F5, F8 and F11. The relative abundance of Flavobacteria in the original sample(S) was 0.53%, which increased first and decreased then across culture stages. The relative abundance of Devosia was 3.09%, with its highest abundance at 1.71% in the F1 generation, after decreased with time. The relative abundances of Achromobacter and Ochrobactrum in F1 were 5.60% and 6.56%, respectively, but bacteria in these genera were rarely found in the original samples(S). Their relative abundances decreased with increasing culture time. In addition, Trichococcus, Acinetobacter and Azospirillum were found in F5, and the relative abundances of F11 were 19.65%, 13.01%, and 2.96%, respectively.
Correlation analysis of microbial community at different culture stages
The analysis of the dominant microbial community in the microbial consortium at different culture periods showed (Fig. 8) that Proteobacteria contained the largest number of genera with relatively distant evolution, and was mainly composed of 12 genera, including Pseudomonas, Azospirillum, Brevundimonas and Ochrobactrum. Among them, the abundance of Pseudomonas was dominant in the F1, F5, F8 and F11 generation samples, which played a key role in the degradation process of straw. Bacteroidetes was composed of Flavobacterium, Dysgonomonas and Taibaiella with similar evolution. It could be concluded that Proteobacteria was the main functional bacteria involved in straw degradation in samples of different culture periods.
The correlation network diagram was used to study the interrelationship between straw degrading microorganisms of M44 in different subculture periods (Fig. 9). Rhizobium, Acinetobacter, Trichococcus, Dysgonomonas, Azospirillum, Enterobacter were strongly correlated with each other and positively correlated with other bacteria. The results showed that there were significant interactions among different genera in the samples of different culture periods, and a variety of microorganisms synergistically degraded corn straw.
Correlation analyses of physicochemical characteristics and dominant genera
Correlation analysis between the TOP20 genera in M44 and straw degradation characteristics (Fig. 10) showed that endoglucanase activity was positively correlated with Brevundimonas, Achromobacter, Hydrogenophaga, Chryseobacterium, Sphingobacterium, and some bacteria that degrade lignocellulosic or intermediate products in the colony. Dysgonomonas had a significant negative correlation with filter paper enzyme activity, Acinetobacter had a significant negative correlation with xylanase activity, and Pseudomonas and Enterobacter had a significant positive correlation with laccase and lignin peroxidase activity. Rhizobium and Proteiniphilum were positively correlated with acetic acid, prophetic acid, and butyric acid contents.
Correlation analysis of dominant genera and straw degradation characteristics at different culture stages. Z1: Pseudomonas, Z2: Trichococcus, Z3: Flavobacterium, Z4: Acinetobacter, Z5: Azospirillum, Z6: Brevundimonas, Z7: Achromobacter, Z8: Enterobacter, Z9: Ochrobactrum, Z10: Acidovorax, Z11: Dysgonomonas, Z12: Rhizobium, Z13: Enterococcus, Z14: Hydrogenophaga, Z15: Chryseobacterium, Z16: Proteiniphilum, Z17: Devosia, Z18: Paracoccus, Z19: Sphingobacterium, Z20: Bacillus, Z21: Endoglucanase, Z22: FPase, Z23: Xlyanase, Z24: Laccase, Z25: Lignin peroxidase, Z26:Acetic acid, Z27:Propanoic acid, Z28:Butyric acid. * and ** indicate significant correlations at the 0.05 and 0.01 levels.
Functional prediction analysis
The COG database comparison
Based on COG database comparison results (Fig. 11), it was found that the function of the M44 was mainly concentrated Amino acid transport and metabolism, General functional prediction only, Transcription, Carbohydrate transport and metabolism and Cell wall/membrane/envelope biogenesis and so on in different culture stages. It could be predicted that M44 may contain abundant genes related to protein decomposition, transport and metabolism enzymes, as well as a large number of genes related to cellulose and lignin degradation enzymes during subculture at low temperature.
The KEGG database comparison
According to KEGG level 1 (Table 1), genes in samples of different culture stages were mainly enriched in Metabolism, Environmental Information Processing, Genetic Information Processing, Cellular Processes, etc. Among them, Metabolism accounted for the highest proportion, and the proportion of original samples was 78.91%, which showed no significant difference with F1, F5, F8 and F11 generation samples. There were 46 metabolic pathways in KEGG Level 2. Table 2 showed the results of the top 15 pathway abundance values in different culture stages. The main metabolic pathways included Global and overview maps, Carbohydrate metabolism, Amino acid metabolism, Energy metabolism, Metabolism of cofactors and vitamins, Membrane transport and Signal transduction, etc. The top 30 enzymes in abundance were further analyzed (Table 3), the results showed that the relative abundance of DNA-directed DNA polymerase, DNA helicase, Peptidylprolyl isomerase, NADH:ubiquinone reductase (H( +)-translocating), 3-oxoacyl-[acyl-carrier-protein] reductase were high. In addition, the Peroxiredoxin, Acetyl-CoA carboxylase were present in different culture periods, and the abundance is obviously different.
Discussion
Many studies had shown that by simulating the decomposition process of lignocellulose under natural conditions (i.e., taking the original environmental samples as the inoculum and adopting restrictive culture techniques), composite flora that could efficiently degrade filter paper, rice straw, and pulp waste could be identified. In this study, the original material was taken from the dried dung sample of Hulunbuir city, Inner Mongolia, and mainly composed of Actinobacteria, Proteobacteria, Bacteroidetes, Firmicutes and Chloroflexi, which was rich in degraded cellulose, hemicellulose, lignin. The M44 was screened out from the original material by long-term restricted subculture, and the corn straw degradation rate was 35.90% at 15 °C for 21 days. The microbial community structure of the original samples was significantly different from that of the microbial consortia obtained after a long period of restricted subculture. The subculture process was not only a process of eliminating bacteria unrelated to straw degradation or not adapted to the medium conditions, but also a process of enriching lignocellulose-degrading bacteria, with randomness.
Microbial diversity of microbial consortium with straw degradation
The M44 was a complex microbial that mix composed of aerobic bacteria, anaerobic bacteria, and strict anaerobic bacteria, and its microbiome structure changed considerably during low-temperature subculture. The Proteobacteria and Firmicutes were vital bacterial in different culture generation. As a reported, these types of bacteria were common in rice straw compost20, decaying wood21, and rumen22, which could produce laccase and degrade Kraft lignin23, and degrading lignin monoaryls, biaryls, and phenolic intermediates using extracellular laccases and peroxidases24,25. It is reported that Clostridium are anaerobic bacteria with a superior ability to decompose lignocellulosic materials and digest cellulosic waste in a methanogenic bioreactor26,27,28. However, it was not detected in this study, which may be due to the conditions of subculture and the unsuitable medium for its mass reproduction and growth.
Acinetobacter, Azospirillum, Pseudomonas, Brevundimonas, Devosia, Achromobacter, and Chryseobacterium played an important role in the straw degradation. Among them, Acinetobacter was found in cellulose-containing agricultural waste as the only carbon source, and efficiently secreted extracellular cellulase and hemicellulose enzymes29,30; Azospirillum had been shown to produce hydrogen peroxide enzymes, oxidase, methyl cellulase, and produced acetic acid, butyric acid, and lactic acid, and to participate in straw degradation metabolism31,32; Dye-decolonizing peroxidases (DYPs) secreted by Pseudomonas had the ability to degrade lignin and lignin model compounds33,34 and had high laccase and lignin peroxidase activities35. They were believed to be important functional bacteria for degradation of straw lignin. Studies had shown that Brevundimonas secreted oxidase and catalase to promote the decomposition of cellulose36; Devosia decomposed catalase and utilizes xylose, glyceraldehyde, cellulose, etc37; Achromobacter could oxidize xylose, secreted oxidase and xylanase, which effectively degraded cellulose and hemicellulose38; and Chryseobacterium decomposed cellulase and protease, degrading cell walls, and could cooperate with Pseudomonas to degrade cellulose and hemicellulose39,40, which correlated with xylanase activity. They were speculated to be functional bacterium for straw cellulose and hemicellulose degradation.
Functional prediction of microbial consortium with straw degradation
The degradation of lignocellulose requires the joint action of a variety of enzymes produced by different microorganisms to attack the complex structure of its biomass41 and produce a fully complementary enzymatic system. Therefore, it is very important to explore some functional genes closely related to lignocellulose degradation. It was reported that the microorganism with high levels of carbon hydration gene had high degradation activity. Zhang et al.'s42 study showed that through the COG database compared, the gene for Carbohydrate transport and metabolism was the most; Singh et al43 found that Buffalo's rumen microbial group had a large number of functional genes for polysaccharide degradation. Based on COG and the KEGG database analysis, this study mainly was concentrated in Amino acid transport and metabolism, General functional prediction only, Carbohydrate transport and metabolism and Cell wall/membrane/envelope biogenesis, etc., which were similar to those of the above studies. In addition, samples with different culture generations in M44 contained Peroxiredoxin and Acetyl-CoA carboxylase, etc. Studies have shown that the Lig K enzyme and peroxidase could catalyze the degradation of lignin intermediates to pyruvate and oxaloacetate, which were finally degraded by tricarboxylic acid cycle44,45. In this study, it was speculated that corn straw lignocellulose could produce small molecular substances through different metabolic pathways under the action of transaminases, lyases and dehydrogenases, and finally completely degrade into organic acids or carbon dioxide.
Conclusion
In nature, degradation of lignocellulose was coordinated by various active enzymes secreted by various microorganisms. In this study, Pseudomonas, Azospirillum, Brevundimonas, Ochrobactrum from Proteobacteria and Flavobacterium, Dysgonomonas, Taibaiella from Bacteroidetes, and Devosia, Trichococcus, Acinetobacter, Rihizobium, Achromobacter, Chryseobacterium were found to be the key bacteria for subculture progress. In different culture periods, the main metabolic pathways include Carbohydrate metabolism, Amino acid metabolism and Energy metabolism, etc. Furthermore, the M44 may contain a large amount of lignin biodegradable enzyme genes that could degrade the material, such as cellulose, hemicellulose and lignin. This study provided theoretical guidance for the selection of functional microorganisms of microbial consortium.
Data availability
The data used to analyze microbial diversity has been uploaded to the SRA database in NCBI with the entry number PRJNA777010.
References
Ma, H., Liu, W. W., Chen, X., Wu, Y. J. & Yu, Z. L. Enhanced enzymatic saccharification of rice straw by microwave pretreatment. Biores. Technol. 100, 1279–1284 (2009).
Fang, X. et al. Progress on cellulase and enzymatic hydrolysis of lignocellulosic biomass. Chin. J. Biotechnol. 26, 864–869 (2010).
Chen, H. et al. A review on the pretreatment of lignocellulose for high-value chemicals. Fuel Process. Technol. 160, 196–206 (2017).
Wang, Z. et al. Evaluation of methane production and energy conversion from corn stalk using furfural wastewater pretreatment for whole slurry anaerobic codigestion. Biores. Technol. 293, 121–962 (2019).
Zhang, Y. et al. Research progress on degradation of lignocellulosic biomass by screening microorganisms. China Biotechnol. 40, 100–105 (2020).
Pérez, J., Munoz-Dorado, J., De la Rubia, T. D. L. R. & Martinez, J. Biodegradation and biological treatments of cellulose, hemicellulose and lignin: An overview. Int. Microbiol. 5, 53–63 (2002).
Ulrich, A., Klimke, G. & Wirth, S. Diversity and activity of cellulose-decomposing bacteria, isolated from a sandy and a loamy soil after long-term manure application. Microb. Ecol. 55, 512–522 (2008).
Wang, W. D. et al. Characterization of a microbial consortium capable of degrading lignocellulose. Biores. Technol. 102, 9321–9324 (2011).
Wang, C. F. et al. Characterization and microbial community shifts of rice straw degrading microbial consortia. Acta Microbiol. Sin. 56, 1856–1868 (2016).
Hua, B. et al. Dynamic changes in the composite microbial system MC1 during and following its rapid degradation of lignocellulose. Appl. Biochem. Biotechnol. 172, 951–962 (2014).
Zheng, G. et al. Degradation of rice straw at low temperature using a novel microbial consortium LTF-27 with efficient ability. Biores. Technol. 304, 123064 (2020).
Alessi, A. M. et al. Defining functional diversity for lignocellulose degradation in a microbial community using multiomics studies. Biotechnol. Biofuels 11, 1–16 (2018).
Wang, Y. X. et al. A novel lignin degradation bacterial consortium for efficient pulping. Biores. Technol. 139, 113–119 (2013).
Su, X. et al. Microbial community succession associated with corn straw degradation in a bacterium consortium. Acta Microbiol. Sin. 60, 2675–2689 (2020).
Zhang, X. et al. Screening and composition of the microbial consortium with corn straw decomposition under low temperature. J. Agro-Environ. Sci. 40, 1565–1574 (2021).
Yu, I. S. et al. Substrate specificity of Stenotrophomonas nitritireducens in the hydroxylation of unsaturated fatty acid. Appl. Microbiol. Biotechnol. 78, 157–163 (2008).
Tian, L. S. Study on the standard method for the determination of enzymes related to lignin degradation. Anim. Husb. Feed Sci. 30, 13–15 (2009).
Robert, W. et al. Characterization of the rumen microbiota of preruminant calves using metagenomic tools. Environ. Microbiol. 14, 129–139 (2012).
Langille, M. G. I. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814–821 (2013).
Matsuyama, T. et al. Bacterial community in plant residues in a Japanese paddy field estimated by RFLP and DGGE analyses. Soil Biol. Biochem. 39, 463–472 (2007).
Zhang, H. B., Yang, M. X. & Tu, R. Unexpectedly high bacterial diversity in decaying wood of a conifer as revealed by a molecular method. Int. Biodeterior. Biodegrad. 62, 471–474 (2008).
Wang, A. et al. Enrichment strategy to select functional consortium from mixed cultures: Consortium from rumen liquor for simultaneous cellulose degradation and hydrogen production. Int. J. Hydrogen Energy 35, 13413–13418 (2010).
Huang, X. F. et al. Isolation and characterization of lignin-degrading bacteria from rainforest soils. Biotechnol. Bioeng. 110, 1616–1626 (2013).
Masai, E., Katayama, Y. & Fukuda, M. Genetic and biochemical investigations on bacterial catabolic pathways for lignin-derived aromatic compounds. Biosci. Biotechnol. Biochem. 23, 0612070214–0612070214 (2007).
Ahmad, M. et al. Development of novel assays for lignin degradation: comparative analysis of bacterial and fungal lignin degraders. Mol. BioSyst. 6, 815–821 (2010).
Chamkha, M., Garcia, J. L. & Labat, M. Metabolism of cinnamic acids by some Clostridiales and emendation of the descriptions of Clostridium aerotolerans, Clostridium celerecrescens and Clostridium xylanolyticum. Int. J. Syst. Evol. Microbiol. 51, 2105–2111 (2001).
Shiratori, H. et al. Isolation and characterization of a new Clostridium sp. that performs effective cellulosic waste digestion in a thermophilic methanogenic bioreactor. Appl. Environ. Microbiol. 72, 3702–3709 (2006).
Ren, Z., Ward, T. E., Logan, B. E. & Regan, J. M. Characterization of the cellulolytic and hydrogen-producing activities of six mesophilic Clostridium species. J. Appl. Microbiol. 103, 2258–2266 (2007).
Lo, Y. C., Lu, W. C., Chen, C. Y., Chen, W. M. & Chang, J. S. Characterization and high-level production of xylanase from an indigenous cellulolytic bacterium Acinetobacter junii F6–02 from southern Taiwan soil. Biochem. Eng. J. 53, 77–84 (2010).
Poomai, N., Siripornadulsil, W. & Siripornadulsil, S. Cellulase enzyme production from agricultural waste by Acinetobacter sp. KKU44. Adv. Mater. Res. 931, 1106–1110 (2014).
Anandham, R. et al. Azospirillum ramasamyi sp. nov., a novel diazotrophic bacterium isolated from fermented bovine products. Int. J. Syst. Evolut. Microbiol. 69, 1369–1375 (2019).
Eckert, B. et al. Azospirillum doebereinerae sp. nov., a nitrogen-fixing bacterium associated with the C4-grass Miscanthus. Int. J. Syst. Evolut. Microbiol. 51, 17–26 (2001).
Kumar, P., Maharjan, A., Jun, H. B. & Kim, B. S. Bioconversion of lignin and its derivatives into polyhydroxy alkanoates: Challenges and opportunities. Biotechnol. Appl. Biochem. 66, 153–162 (2019).
Rahmanpour, R. & Bugg, T. D. Characterization of Dyp-type peroxidases from Pseudomonas fluorescens Pf-5: Oxidation of Mn (II) and polymeric lignin by Dyp1B. Arch. Biochem. Biophys. 574, 93–98 (2015).
Yang, C. X., Wang, T., Gao, L. N., Yin, H. J. & Lü, X. Isolation, identification and characterization of lignin-degrading bacteria from Qinling, China. J. Appl. Microbiol. 123, 1447–1460 (2017).
Egorova, D. O., Nazarova, E. A. & Demakov, V. A. New lindane-degrading strains Achromobacter sp. NE1 and Brevundimonas sp. 242. Microbiology 90, 392–396 (2021).
Hassan, Y. I., Lepp, D. & Zhou, T. Genome assemblies of three soil-associated Devosia species: D. insulae, D. limi, and D. soli. Genome Announc. 3, 0051415 (2015).
Jiménez, D. J., Dini-Andreote, F. & Van Elsas, J. D. Metataxonomic profiling and prediction of functional behavior of wheat straw degrading microbial consortia. Biotechnol. Biofuels 7, 1–18 (2014).
Bhise, K. K., Bhagwat, P. K. & Dandge, P. B. Synergistic effect of Chryseobacterium gleum sp. SUK with ACC deaminase activity in alleviation of salt stress and plant growth promotion in Triticum aestivum L. 3 Biotech 7, 1–13 (2017).
Maki, M., Iskhakova, S., Zhang, T. & Qin, W. Bacterial consortia constructed for the decomposition of Agave biomass. Bioengineered 5, 165–172 (2014).
Van Dyk, J. S. & Pletschke, B. I. A review of lignocellulose bioconversion using enzymatic hydrolysis and synergistic cooperation between enzymes-factors affecting enzymes, conversion and synergy. Biotechnol. Adv. 30, 1458–1480 (2012).
Zhang, H. M. et al. Metagenomic analysis of microorganisms in rumen of Haizi buffalo. Chin. J. Anim. Nutr. 29, 4151–4161 (2017).
Singh, K.M. et al. High potential source for biomass degradation enzyme discovery and environmental aspects revealed through metagenomics of Indian buffalo rumen. BioMed Res. Int. 267189 (2014).
Masai, E. et al. Roles of the enantioselective glutathione S-transferases in cleavage of β-aryl ether. J. Bacteriol. 185, 1768–1775 (2003).
Hogancamp, T. N., Ogancamp, T. N., Mabanglo, M. F. & Raushel, F. M. Structure and reaction mechanism of the Lig J hydratase: An enzyme critical for the bacterial degradation of lignin in the protocatechuate 4,5-cleavage pathway. Biochemistry 57, 5841–5850 (2018).
Acknowledgements
This study was supported by the Inner Mongolia Natural Foundation (grant nos. 2020MS03086 and 2018ZD02), the National Natural Science Foundation of China (grant no. 32060434 and 31760353), the Science and Technology Xingmeng Project (KJXM2020001-06), the China Agriculture Research System of MOF and MARA (grant no. CARS-02-63), and the Crop Science Observation and Experiment Station in Loess Plateau of North China of Ministry of Agriculture (grant no. 25204120).
Author information
Authors and Affiliations
Contributions
Performed the experiments: X.Z., J.-L.G., S.-P.H., S.-C.H., and B.-Z.Z. Analyzed the data: X.Z. and Qinggeer Borjigin. Critically revised the manuscript for important intellectual content: J.-L.G. and X.-F.Y. Wrote the paper: X.Z. and Qinggeer Borjigin.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, X., Borjigin, Q., Gao, JL. et al. Community succession and functional prediction of microbial consortium with straw degradation during subculture at low temperature. Sci Rep 12, 20163 (2022). https://doi.org/10.1038/s41598-022-23507-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-23507-z
- Springer Nature Limited