Abstract
Osteoporosis (OP) has plagued many women for years, and bone density loss is an indicator of OP. The purpose of this study was to evaluate the relationship between the polymorphism of the rs7586085, CCDC170 and GALNT3 gene polymorphisms and the risk of OP in the Chinese Han population. Using the Agena MassArray method, we identified six candidate SNPs on chromosomes 2 and 6 in 515 patients with OP and 511 healthy controls. Genetic model analysis was performed to evaluate the significant association between variation and OP risk, and meanwhile, the multiple tests were corrected by false discovery rate (FDR). Haploview 4.2 was used for haplotype analysis. In stratified analysis of BMI ˃ 24, rs7586085, rs6726821, rs6710518, rs1346004, and rs1038304 were associated with the risk of OP based on the results of genetic models among females even after the correction of FDR (qd < 0.05). In people at age ≤ 60 years, rs1038304 was associated with an increased risk of OP under genetic models after the correction of FDR (qd < 0.05). Our study reported that GALNT3 and CCDC170 gene polymorphisms and rs7586085 are the effective risk factors for OP in the Chinese Han population.
Similar content being viewed by others
Introduction
Osteoporosis (OP) is one of the most common and impactful metabolic diseases of elders1 with the clinical features of the reduced bone mineral density (BMD) and bone structure destruction leading to an increased risk of fracture2. Age and sex are the two most relevant hazard factors for OP3. Elders, especially in postmenopausal women, are at a high risk for it due to accelerated bone loss2,3. It has been 8.9 million fractures worldwide for the increasing and prevalence of OP4. Patients with brittle fractures are hospitalized more than 400,000 times a year and treated 2.5 million times a year, which are placed a huge financial burden on patients and their families5. It is genetic and environmental factors that contribute to OP6,7. Twin studies have shown a BMD heritability of 0.51 to 0.76 for different bones. Previous genome-wide association analyses (GWASs) have identified more than 60 loci related to bone density and OP, many of which are thought to play important roles in bone, such as RANKL, OPG, ESR1, and LRP52. GWASs have identified certain SNPs at risk for OP8.
GALNT3, located in 2q24-31, encodes UDP-N-acetyl-ɑ-d-galactosamine-polypeptide: polypeptide N-acetylgalactosaminyltransferase-3 (ppGalNacT3)9, and initiates the glycosylation of O-GalNAC. A GWAS by Duncan et al. covered an association between the GALNT3 gene and BMD and fracture risk in postmenopausal women10. The substances encoded by the GALNT3 gene in the human body are mainly involved in bone metabolism and related processes11. Studies have shown that GALNT3 gene mutation can cause hyperphosphatemic familial tumoral calcinosis that is an autosomal recessive genetic disease and that can lead to the symptoms of hyperphosphatemia12,13. Moreover, the abnormalities of ppGalNacT3 can cause a disorder of phosphorus regulation, thereby affecting bone mineralization and BMD, which is one of the most important indicative indexes of primary osteoporotic fractures14. GWASs have been confirmed that it has been very successful in identifying common genetic variations related to bone density. A GWAS study of BMD found that CCDC170 was strongly associated with BMD15. CCDC170 encodes the protein CCDC170, which is a predicted protein containing the coiled helical domain (CCDC), is associated with the golgi body, stabilizes peri-nuclear microtubules (MTs), and plays an vital role in the known process of mt-dependent golgi structure16.
In this study, we selected samples from Chinese Han ethnicity from Xi’an 630 Hospital and People's Hospital of Wanning to study the relationship between rs7586085, GALNT3 (rs6726821, rs6710518, and rs1346004) and CCDC170 (rs4869739 and rs1038304) gene polymorphisms and the OP phenotype in postmenopausal women in China. These findings are expected to elucidate important new pathways in bone metabolism and to contribute to the development of new therapies, which may have prognostic value.
Materials and methods
Study participants
The case–control study was collected from hospitals included 515 patients with OP and 511 healthy controls from April 2019 to April 2020. Subjects with OP were recruited from the Xi’an 630 Hospital, Yanliang, Xi’an, Shaanxi, China and People's Hospital of Wanning, Hainan Province, China. The control group was those who went to the two hospitals for general inspection, who had no history of cancer or any disease related to bone organs. BMD at the lumbar spine (l2-4) and femoral neck of all subjects were determined using a dual-energy X-ray absorptiometry (lunar specialist 1313). We diagnosed OP in strict accordance with the criteria of the World Health Organization17.
Clinical data and demographic information
We used standardized epidemiological questionnaires, including area of residence, age, sex, BMI, ethnicity, and family history, to collect personal data in face-to-face interviews. The 5 mL venous blood was taken from each subject for DNA extraction. All the volunteers signed an informed consent that stated the purpose of this study and the experiment. The protocol (approved number: hnwnrmyy-2020-yxk-05) was approved by the ethics committee of People's Hospital of Wanning, and was in accordance with the Declaration of Helsinki.
SNPs selection and genotyping
We selected carefully rs7586085, GALNT3 and CCDC170 SNPs from 1000 Genomes Project (http://www.internationalgenome.org/) and the SNPs were in conformity with the minor allele frequency (MAF) ˃ 5%. The distribution of SNPs genotypes in the control group was in accordance with Hardy–Weinberg equilibrium (HWE) (p > 0.05). We genotyped SNPs using Agena MassARRAY RS1000. Moreover, the call rate of our results was greater than 95%. Ten percent of the samples were genotyped repeatedly and the concordance rate was 100%. The investigators who genotyped the samples were unknown the status of the sample. Then, using the Haploview 4.2, the pairwise linkage disequilibrium (LD) of rs7586085, GALNT3 and CCDC170 gene polymorphisms was estimated. After finished the steps mentioned above, we selected six SNPs rs7586085, rs6726821, rs6710518, rs1346004, rs4869739 and rs1038304 as the gene variation to study. Genomic DNA was extracted from peripheral blood with the Gold Mag-Mini genomic DNA purification kit (Gold Mag Co. Ltd., Xi’an, China) and quantified with the Nano Drop spectrophotometer 2000C (Thermo Scientific, Waltham, Massachusetts, USA). SNPs genotyping of the Agena MassARRAY RS1000 instrument (Shanghai, China) system was performed in accordance with the standard scheme recommended by the manufacturer. The experimental data were managed and analyzed using Agena Typer 4.0 software. Primers of each SNP are presented in Supplementary Table S1.
Statistical analyses
First, the HWE of each SNP in the control group was inspected by the goodness-of-fit chi-square test. In this study, all p values were bilateral, and p value less than 0.05 was regarded as the cut-off value, which was considered statistically significant. The chi-square test was used to compare the allele frequency and genotype frequency of each SNP in the patients and controls. Odds ratio (OR) and 95% confidence interval (95% CI) were obtained by unconditional logistic regression analysis adjusted for BMI and age. To account for multiple comparisons at each genetic model, we further considered FDR adjusted p value (qd) < 0.05 as significance. The relationship between genotypes and OP risk was tested in different genetic models (co-dominant, dominant, recessive, and additive) using PLINK 1.9. The demographic characteristics were experimented using SPSS statistical software package, version 19.0 (SPSS Inc., Chicago, Illinois, USA). Haploview 4.2 was used to perform the LD and haplotype analysis of these six polymorphisms to OP risk.
Ethical approval
All procedures completed in this study were in keeping with the ethical standards of the ethics committee of People's Hospital of Wanning and with the 1964 Helsinki declaration and its later amendments.
Results
Population characteristics
A total of 515 female patients with OP and 511 female controls were enrolled in our study. The mean age [± standard deviation (SD)] of the case group was 63.72 ± 5.58 years at diagnosis and that of the control group was 62.87 ± 4.68 years at recruitment.
SNPs and OP risk
The essential information and allele frequencies of GALNT3 and CCDC170 gene polymorphisms and rs7586085 are displayed in Table 1. The six SNPs were all conformed to the HWE without deviation in the control group. The minor allele of each SNP was considered as a risk factor. Results of the four genetic models analyses are shown in Table 2. We used logistic regression to analyze SNPs of four genetic models. The results showed that there were no significant loci.
We collected the height and weight of the individuals. Then, stratified analysis was performed whether BMI was greater than 24 (Table 3). Individuals with BMI > 24 were considered overweight. Stratified analysis by BMI indicated that the rs7586085 polymorphism was significantly related to an increased risk of OP in BMI ˃ 24 (heterozygote: OR 2.13, 95% CI 1.28–3.57, p = 0.004, qd = 0.036; additive: OR 1.55, 95% CI 1.10–2.18, p = 0.012, qd = 0.048; alleles: OR 1.49, 95% CI 1.07–2.07, p = 0.018, qd = 0.05). The polymorphism of rs6726821 was significantly associated with an increased risk of OP in BMI ˃ 24 (heterozygote: OR 2.13, 95% CI 1.28–3.57, p = 0.004, qd = 0.029; alleles: OR 1.55, 95% CI 1.07–2.07, p = 0.018, qd = 0.046). The polymorphism of rs6710518 was significantly associated with an increased risk of OP in BMI ˃ 24 (heterozygote: OR 2.11, 95% CI 1.27–3.51, p = 0.004, qd = 0.024; dominant: OR 2.02, 95% CI 1.25–3.28, p = 0.004, qd = 0.021). The polymorphism of rs1346004 was significantly associated with an increased risk of OP in BMI ˃ 24 (heterozygote: OR 2.13, 95% CI 1.28–3.57, p = 0.004, qd = 0.018; dominant: OR 2.10, 95% CI 1.29–3.41, p = 0.003, qd = 0.036; additive: OR 1.55, 95% CI 1.10–2.18, p = 0.012, qd = 0.043; alleles: OR 1.49, 95% CI 1.07–2.07, p = 0.018, qd = 0.043). The polymorphism of rs1038304 was significantly associated with an increased risk of OP in BMI ˃ 24 (homozygote: OR 2.41, 95% CI 1.18–4.91, p = 0.016, qd = 0.048; recessive: OR 2.24, 95% CI 1.19–4.19, p = 0.012, qd = 0.039). After FDR correction, significant association remained among rs7586085, rs6726821, rs6710518, rs1346004, rs1038304 and increased risk of OP. Rs4869739 polymorphism was not observed significance with OP in BMI ˃ 24 after FDR correction.
We also investigated the relationship of six SNPs with OP risk under age subgroup. As summarized in Table 4, the polymorphism of rs1038304 was found to significantly increase the risk of OP at age ≤ 60 years even after FDR correction (homozygote: OR 2.99, 95% CI 1.50–6.00, p = 0.002, qd = 0.024; dominant: OR 2.49, 95% CI 1.43–4.34, p = 0.001, qd = 0.036; additive: OR 1.73, 95% CI 1.23–2.44, p = 0.002, qd = 0.018; alleles: OR 1.64, 95% CI 1.21–2.23, p = 0.001, qd = 0.018). There was no significant association observed in other SNPs.
Association of haplotype with OP
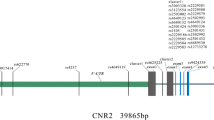
We further explored the LD and haplotype analyses of those SNPs. A haplotype block with strong LD is presented in Fig. 1 with four SNPs including rs7586085, rs6726821, rs6710518, and rs1346004. The distribution of frequencies for haplotypes in the cases and controls are observed in Table 5. The haplotype results show a remarkable associations of ‘GGCA’ haplotypes with an increased risk of OP (OR 2.74, 95% CI 1.20–6.22, p = 0.016) (Table 5).
Discussion
The main characteristic of OP is that it can decrease the risk in bone density. BMD is defined as the amount of minerals in bone and is associated with estrogen18. The majority of BMD-related SNPs identified by GWASs are located in non-coding regions of the genome2. Our study provides an extensive evidence that SNPs (rs7586085, rs6726821, rs6710518, rs1346004, rs4869739, and rs1038304) located on chromosomes 2 and 6 can serve as multiple loci which were associated with an increased risk of OP. We demonstrated that risk SNPs loci were significantly associated with an increased OP risk in various genetic models, and haplotype ‘GGCA’ consisting of four SNPs was also associated with increasing the risk of OP. Additionally, it turns out that all the five SNPs, which were associated with increasing the risk of OP in people with BMI ˃ 24, were obviously some risk loci in overweight people. Rs1038304 was associated with an increased risk of OP in people at age ≤ 60 years. The six SNPs what we studied were in the non-coding region of the gene. Rs7586085 was close to the GALNT3 gene, but located on an unknown gene.
In a meta-analysis, gender- and age-adjusted variants of the CCDC170/ESR1 gene were found to be associated with BMD19. Other studies found that rs1038304 polymorphism on CCDC170 gene was associated with fracture and vertebral fracture risk in postmenopausal women in China20. CCDC170 gene polymorphism may not only play an important role in bone metabolism. Previous studies have found a significant association between vertebral fracture risk and rs1038304 and a protective effect20, and other study has found that rs1038304 is related to BMD21. Therefore, we studied the relationship between CCDC170 gene polymorphism and OP risk, and found that the SNPs were associated with increasing the risk of OP. Rs1038304 was in the intron region of CCDC170 gene and was associated with increasing the risk of OP.
Previous studies have suggested a relationship between GALNT3 gene polymorphism and the OP phenotype in postmenopausal women in China9. GALNT3 is an enzyme involved in the glycosylation of serine and threonine residues, whose process is critical to the integrity and viability of fibroblast growth factor-23 (FGF23). A functional copy of GALNT3 may be sufficient to secrete complete FGF23 and appropriately regulate serum phosphate22. FGF23 is a phosphorus-promoting hormone produced by bones which enhances the reabsorption of calcium and sodium into the kidney23. The polymorphisms of GALNT3 and FGF23 can cause familial neoplastic calcinosis in hyperphosphatemia24. Furthermore, Runx2 is an important transcription factor for chondrocyte maturation25. GALNT3 is one of the downstream genes of Runx2 in chondrocytes, however many GALNT family genes are expressed in cartilage tissue. Galnt3 mice showed short stature and shortened limbs. GALNT3 has non-redundant function during chondrocyte maturation25. Ichikawa et al. found increased bone density Galnt3-deficient mice22. Generally speaking, polymorphism in the GALNT3 gene plays an important role in BMD loss. A significant relationship between the polymorphism of rs6710518 and BMD has been discovered9. Therefore, polymorphisms of GALNT3 gene were detected to be the risk factor to OP, leading to new findings on the pathological mechanism of OP.
Although we successfully identified individual trait correlations and pleiotropic SNPs of OP, our study still had some potential limitations. First, we only found some polymorphisms in some of the non-coding genes on chromosome 2 and chromosome 6, and it may have other polymorphisms around. Second, the sample size of the case and control groups was small, which was only limited to the population of Northwest China. Therefore, we need to continue to expand the sample size and further study the mechanisms at the cellular level and in vivo.
Conclusion
Taken together, our study uncovered a new association between genetic polymorphisms on chromosomes 2 and 6 and the risk of OP in the Chinese Han population. These outcomes are helpful to further study the mechanism of polymorphism affecting the pathogenesis of OP. The larger sample sizes were, the more cellular and in vivo studies to further explore and confirm the function of these polymorphisms in increasing the risk of femoral OP were needed, which will provide new insights on prevention and treatment of OP.
Data availability
The data that support the findings of this study are available on request from the corresponding author.
References
Almeida, M. et al. Estrogens and androgens in skeletal physiology and pathophysiology. Physiol. Rev. 97(1), 135–187 (2017).
Chen, X. F. et al. An osteoporosis risk SNP at 1p36.12 acts as an allele-specific enhancer to modulate LINC00339 expression via long-range loop formation. Am. J. Hum. Genet. 102(5), 776–793 (2018).
Zhang, D. et al. Genetic association study identified a 20 kb regulatory element in WLS associated with osteoporosis and bone mineral density in Han Chinese. Sci. Rep. 7(1), 13668 (2017).
Pisani, P. et al. Major osteoporotic fragility fractures: Risk factor updates and societal impact. World J. Orthop. 7(3), 171–181 (2016).
Lane, J. M., Russell, L. & Khan, S. N. Osteoporosis. Clin. Orthop. Relat. Res. 372, 139–150 (2000).
Karasik, D. & Cohen-Zinder, M. Osteoporosis genetics: Year 2011 in review. Bonekey Rep. 1, 114 (2012).
Özbaş, H., Tutgun Onrat, S. & Özdamar, K. Genetic and environmental factors in human osteoporosis. Mol. Biol. Rep. 39(12), 11289–11296 (2012).
Farber, C. R. & Lusis, A. J. Future of osteoporosis genetics: Enhancing genome-wide association studies. J. Bone Miner. Res. 24(12), 1937–1942 (2009).
Li, N. et al. Association of GALNT3 gene polymorphisms with bone mineral density in Chinese postmenopausal women: The Peking Vertebral Fracture study. Menopause 21(5), 515–521 (2014).
Duncan, E. L. et al. Genome-wide association study using extreme truncate selection identifies novel genes affecting bone mineral density and fracture risk. PLoS Genet. 7(4), e1001372 (2011).
Wang, X. Y. et al. Research on correlation between GALNT3 gene and osteoporosis. Eur. Rev. Med. Pharmacol. Sci. 22(1 Suppl), 69–75 (2018).
Sun, L. et al. Identification of two novel mutations in the GALNT3 gene in a Chinese family with hyperphosphatemic familial tumoral calcinosis. Bone Res. 4, 16038 (2016).
Ramnitz, M. S. et al. Phenotypic and genotypic characterization and treatment of a cohort with familial tumoral calcinosis/hyperostosis–hyperphosphatemia syndrome. J. Bone Miner. Res. 31(10), 1845–1854 (2016).
Wang, G. et al. Association of genetic polymorphisms of GALNT3 and VDR with osteoporosis in postmenopausal women. Exp. Ther. Med. 12(4), 2629–2633 (2016).
Villalobos-Comparán, M. & Jiménez-Ortega, R. F. A pilot genome-wide association study in postmenopausal Mexican-Mestizo women implicates the RMND1/CCDC170 locus is associated with bone mineral density. Int. J. Genom. 2017, 5831020 (2017).
Jiang, P. et al. The protein encoded by the CCDC170 breast cancer gene functions to organize the golgi-microtubule network. EBioMedicine 22, 28–43 (2017).
Kanis, J. A. Assessment of fracture risk and its application to screening for postmenopausal osteoporosis: Synopsis of a WHO report. WHO Study Group. Osteoporos Int. 4(6), 368–381 (1994).
Scalco, R. C. et al. ESR1 polymorphism (rs2234693) influences femoral bone mass in patients with Turner syndrome. Endocr. Connect. 8(11), 1513–1519 (2019).
Mullin, B. H. et al. Genome-wide association study using family-based cohorts identifies the WLS and CCDC170/ESR1 loci as associated with bone mineral density. BMC Genom. 17, 136 (2016).
Luo, L. et al. Association of ESR1 and C6orf97 gene polymorphism with osteoporosis in postmenopausal women. Mol. Biol. Rep. 41(5), 3235–3243 (2014).
Peng, C. et al. Enhanced identification of potential pleiotropic genetic variants for bone mineral density and breast cancer. Calcif. Tissue Int. 101(5), 489–500 (2017).
Ichikawa, S. et al. Ablation of the Galnt3 gene leads to low-circulating intact fibroblast growth factor 23 (Fgf23) concentrations and hyperphosphatemia despite increased Fgf23 expression. Endocrinology 150(6), 2543–2550 (2009).
Fukumoto, S. FGF23 and bone and mineral metabolism. Handb. Exp. Pharmacol. 262, 281– 308 (2019).
Emecen Sanli, M. et al. Familial hyperphosphatemic tumoral calcinosis in an unusual and usual sites and dramatic improvement with the treatment of acetazolamide, sevelamer and topical sodium thiosulfate. J. Pediatr. Endocrinol. Metab. 34(6), 813–816 (2021).
Yoshida, C. A. et al. Overexpression of Galnt3 in chondrocytes resulted in dwarfism due to the increase of mucin-type O-glycans and reduction of glycosaminoglycans. J. Biol. Chem. 289(38), 26584–26596 (2014).
Acknowledgements
We thank all patients and individuals for their participation. Thank you to the Xi’an 630 hospital, Yanliang, Xi’an, Shaanxi, China and People's Hospital of Wanning, Hainan Province, China for samples.
Author information
Authors and Affiliations
Contributions
J.Z. and Q.C. conceived and designed the project. J.Z. and W.C. collected study samples. W.C. and M.H. selected the SNPs and designed primers. M.H. and Q.C. performed the experiments. Q.C. and R.G. analyzed the data. J.Z. and T.J. wrote and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, J., Cai, Q., Chen, W. et al. Relationship between rs7586085, GALNT3 and CCDC170 gene polymorphisms and the risk of osteoporosis among the Chinese Han population. Sci Rep 12, 6089 (2022). https://doi.org/10.1038/s41598-022-09755-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-09755-z
- Springer Nature Limited