Abstract
The accuracy of sixteen commonly used internal reference genes was assessed in skeletal muscle-derived satellite cells of Qinchuan cattle at different stages of proliferation and induction of differentiation to determine the most suitable ones. Quantitative real-time PCR and three commonly used algorithmic programs, GeNorm, NormFinder and BestKeeper, were used to evaluate the stability of expression of the candidate internal reference genes (GAPDH, ACTB, PPIA, LRP10, HPRT1, YWHAZ, B2M, TBP, EIF3K , RPS9, UXT, 18S rRNA, RPLP0, MARVELD, EMD and RPS15A) in skeletal muscle-derived satellite cells at 0, 12, 24, 36 and 48 h of growth and after differentiation for 0, 2, 4, 6 and 8 days. The expression of two satellite cell marker genes, CCNA2 and MYF5, was used for validation analysis. The results of the software analyses showed that GAPDH and RPS15A were the most stable reference gene combinations during in vitro proliferation of bovine skeletal muscle-derived satellite cells, RPS15A and RPS9 were the most stable reference gene combinations during in vitro induction of differentiation of the cells, and PPIA was the least stable reference gene during proliferation and differentiation and was not recommended. This study lays the foundation for the selection of reference genes for qRT-PCR during the proliferation and induction of differentiation of bovine skeletal muscle-derived satellite cells.
Similar content being viewed by others
Introduction
The quantity and quality of livestock skeletal muscle are key factors in determining the quality of meat and are critical factors in the management of animal husbandry1. The process of growth and development of skeletal muscle is complex. It requires proliferation, differentiation and fusion of myoblasts into muscle fibers, and involves a large number of changes in gene expression2,3. Since it is difficult to study muscle cell proliferation and differentiation in vivo, primary cultured myoblasts are commonly used as a model because of their excellent species specificity. Studying the proliferation and differentiation of bovine skeletal muscle-derived satellite cells in vitro is of great importance in animal husbandry and the food industry. The quantitative real-time PCR (qRT-PCR) method is a widely used technique for measuring relative gene expression with high throughput, accuracy, sensitivity and reproducibility4. It is used in a quantitative way to evaluate relative gene expression, but the accuracy of qRT-PCR results depends on the stability of the reference gene5. Ideally, the genes selected for reference should show similar mRNA levels at different developmental stages of an organism, and different tissues or cells should not vary due to environmental factors or bioassay treatments6. However, different species, different tissues of the same species, different cells of the same tissue, different stages of development of the same cells and different experimental treatments can all affect the expression of reference genes7. There is no universal reference gene that can be used for all cell and tissue types8, and choosing an inappropriate reference gene may yield inaccurate results and erroneous conclusions. Therefore, the optimal reference gene should be carefully determined for each organism and experiment. Although many studies have investigated changes in gene expression in the proliferation and differentiation of bovine skeletal muscle-derived satellite cells (bovine SMSCs), few have assessed the stability, suitability and reliability of reference genes for qRT-PCR standardization. Evidence has shown that expression of the traditional internal reference genes sometimes changes significantly under different experimental conditions7. The reference genes commonly used to study the development of SMSCs are β-actin (ACTB), 3-phosphoglyceraldehyde dehydrogenase (GAPDH) and 18 s ribosomal RNA (18 s rRNA), but they may not be ideal and reliable9,10. The selection of a suitable reference gene is crucial in qRT-PCR studies for accuracy of the calculations of target gene expression.
In this study, three commonly used algorithms, GeNorm11, NormFinder12 and BestKeeper13, were used to identify and validate the expression stability of sixteen candidate reference genes commonly used for qRT-PCR, to provide a better understanding of the proliferation and differentiation of bovine primary skeletal muscle cell models in vitro.
Results
Proliferation and differentiation of bovine SMSCs in vitro
Accurately determining how well bovine SMSCs proliferated and differentiated into myotubes in vitro was one goal of this study. As shown in Fig. 1A, skeletal muscle cell cultures at a confluence of about 50% were defined as 0 h to measure proliferation. The morphology of SMSCs was normal at 0 h, with a distinctive shuttle shape and good ductility. As time progressed, the number of skeletal muscle cells increased significantly with no significant morphological changes, and, by 48 h, there was local contact inhibition and increased cell polarity. This indicated that the skeletal muscle cells were in a good proliferative state and could be used for subsequent experiments. As shown in Fig. 1B, day zero (d 0) for determining differentiation of skeletal muscle cells was defined as about 80% confluence. The SMSCs in differentiation medium exhibited normal morphology at d 0, but myotubes began to appear by d 2 under differentiation induction. By d 8, most of the skeletal muscle cells had fully differentiated and fused into mature myotubes, and the length of the myotubes increased significantly. As shown in Fig. 1B(4d) (IF), immunofluorescence was performed with myosin heavy chain (MYHC) antibody, and the observed myotubes displayed strong fluorescence. This indicated that they were differentiated from myocytes and that the skeletal muscle cells used in this study had good myogenic differentiation potential and could be used in subsequent experiments.
Proliferating and differentiating bovine skeletal muscle-derived satellite cells in vitro. (A) Cell proliferation at 0, 24 and 48 h (400 ×). (B-i~v) Cells at 0, 2, 4, 6 and 8 days of induced differentiation (40 ×). (B-vi) immunofluorescence, DAPI staining is shown in blue and MYHC is shown in red (100 ×).
Primer specificity and amplification efficiency
qRT-PCR amplification of candidate reference genes and gel electrophoresis showed primer-specific amplification of the target genes (Figure S1), and fusion curve analysis also demonstrated the specificity of the primers (Figure S2).The standard curves of the candidate internal reference genes obtained by fivefold dilution gradient were linear, with amplification efficiency ranging from 90.4% to 120.8% and coefficients of determination (R2) ranging from 0.969 to 1.000, indicating that the primers worked well under the qRT-PCR amplification conditions and yielded accurate and reliable results.
qRT-PCR Ct values of reference genes
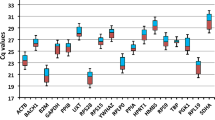
Ct values showed the expression levels of the candidate reference genes in proliferating and differentiating bovine SMSCs (Fig. 2). Higher Ct values indicate lower expression levels, and significant differences in expression between genes were found. The means of the Ct values of the candidate reference genes in SMSCs at different stages of proliferation and induced differentiation ranged from 13.58 to 30.16 and 14.23 to 30.85, respectively. Minimal differences in RPS15A expression in the proliferation stage and in TBP expression in the differentiation stage were observed. The highest expression of 18 s rRNA and the lowest expression of UXT were observed in the proliferation and differentiation stages, and PPIA showed the greatest variation. The statistical comparison of Ct values showed that PPIA was the least stable reference gene.
Stability analysis of internal reference genes
GeNorm analysis
The expression stability value (M, Fig. 3) of each candidate gene was calculated by GeNorm based on the pairwise variation of the test genes, and a smaller M value equates to greater stability. The results show that all the reference genes had high stability; M values ranged from 0.172 to 0.498 during proliferation and from 0.198 to 0.663 during differentiation of bovine SMSCs. The two most stable internal reference genes were GAPDH and RPS15A for proliferation (Fig. 3A), and the two most stable internal reference genes for differentiation were RPS15A and RPS9 (Fig. 3C). The software considered PPIA as the least stable reference gene after analysis during proliferation and differentiation (Fig. 3A,C). The pairwise difference values Vn/Vn + 1 of the internal reference genes (Fig. 3B,D) showed that V2/3 = 0.063 < 0.15 for proliferation and V2/3 = 0.084 < 0.15 for differentiation, indicating that the two reference genes were sufficient to accurately normalize the expression of the target genes.
Expression stability and optimal reference genes among the candidate reference genes were determined using the GeNorm program. (A) Stability of candidate reference genes during proliferation of bovine SMSCs. (B) Optimal reference genes for proliferation of SMSCs. (C) Stability of candidate reference genes during differentiation of bovine SMSCs. (D) Optimal reference genes for use during in vitro induction of differentiation of SMSCs.
NormFinder analysis
NormFinder software calculates arbitrary stability values and standard errors of the reference genes while taking into account the intra- and extra-group variation of each internal reference gene. According to the results given by NormFinder (Fig. 4), the stability value, M, of the reference genes ranged from 0.030 to 0.645 in the proliferation stage and from 0.129 to 0.814 in the differentiation stage, all of which reflected high stability. The two most stable internal reference genes screened by the software in the proliferation assay were UXT (0.030) and RPLP0 (0.094) (Fig. 4A). In the differentiation assay, the two most stable internal reference genes were GAPDH (0.129) and RPLP0 (0.160) (Fig. 4B). These results differed slightly from those of the GeNorm software analysis. In the proliferation and differentiation assays, the software identified the least stable gene as PPIA (0.645, 0.663), which is consistent with GeNorm software analysis.
Bestkeeper analysis
Bestkeeper software estimates the stability of a reference gene based on the variance and provides information about the coefficient of variation and correlation coefficients between samples, as the coefficient of variation (CV) ± standard deviation (SD). According to the stability rankings given by the software, the SD values of the internal reference genes during proliferation of SMSCs were between 0.54 and 0.91 (except for PPIA at 1.25). The two most stable genes screened by the software were EIF3K (1.51 ± 0.35) and RPS15A (2.01 ± 0.39) (Table 1). At the differentiation stage of SMSCs, the stability ranking results given by the software showed that twelve of the sixteen internal reference genes had SD values between 0.47 and 0.92, and four internal reference genes were > 1.00. The two most stable genes screened by the software were TBP (1.67 ± 0.47) and EMD (2.35 ± 0.61) (Table 2). The results were slightly different from those obtained in the GeNorm and NormFinder software analyses. The least stable internal reference gene screened by the software was PPIA (5.75 ± 1.25, 7.62 ± 1.88) (Table 1, 2) in proliferation and differentiation stages, which was consistent with the results obtained from GeNorm and NormFinder software.
Comprehensive analysis of candidate internal reference genes
To combine the results of analysis by the three procedures, the stability ranks of the candidate reference genes were calculated and ranked by geometric mean for both proliferation and induced differentiation stages of bovine SMSCs (Fig. 3). In the proliferation stage, the comprehensive stability rankings showed that GAPDH (2.47) and RPS15A (2.52) were the two most stable internal reference genes (Table 3). After induction differentiation, the comprehensive stability ranking results showed that RPS15A (3.11) and RPS9 (3.43) were the two most stable internal reference genes (Table 4). The least stable reference gene in the comprehensive analysis of both proliferation and differentiation was PPIA (Tables 3 and 4). Therefore, GAPDH and RPS15A were the best reference gene combination to normalize the expression of target genes during proliferation, and RPS15A and RPS9 were the best reference gene combination to normalize the expression of target genes during the induction of differentiation.
Expression validation of candidate internal reference genes
The effects of the most and least stable internal reference genes on gene expression during proliferation and induced differentiation of bovine SMSCs were investigated. The mRNA expression of the marker genes was similar when the two most stable reference genes were used alone or in combination, whereas the mRNA expression of the marker genes was different when the least stable reference gene was used (Fig. 5). Cyclin A2 (CCNA2) is a ubiquitously expressed member of the cyclin family14,15. It plays an indispensable role in regulating the G1/S transition and in the process of mitosis through the activation of kinases15. CCNA2 is frequently related to cell proliferation and usually considered as a marker of cell proliferation15. During the proliferation of bovine SMSCs, the mRNA expression of CCNA2 first increased (0 h to 24 h) and then decreased (24 h to 48 h) when the geometric mean of GAPDH and RPS15A was used together or separately (Fig. 5). When PPIA was used as the correction factor, the mRNA expression of CCNA2 decreased from 0 to 12 h, then increased from 12 to 36 h, and finally decreased from 36 to 48 h. Myogenic regulatory factor 5 (MYF5) is a key transcription factor that plays a central role in transcriptional regulation during muscle formation16. MYF5 is implicated in the initial steps of myoblast differentiation17 and, therefore, is often used as a marker gene for differentiation of SMSCs. During the induced differentiation of bovine SMSCs, the mRNA expression of MYF5 tended to decrease (0 d-2 d), then increase (2 d-4 d), then decrease (4 d-6 d), and finally increase (6 d-8 d), when the geometric means of RPS15A and RPS9 were used as correction factors together or separately. When PPIA was used as a correction factor, the mRNA expression of MYF5 was different from the former, showing an increase (0 d to 2 d), followed by a decrease (2 d to 4 d), then an increase (4 d to 6 d), and finally a decrease (6 d to 8 d). The trend was opposite to that when the two most stable reference genes were used alone or in combination and had a larger magnitude of change, peaking at 2 d with a 4.19-fold upregulation. It is clear that use of inappropriate reference genes may cause misinterpretation of target gene expression. Therefore, it is important to use the appropriate stable reference gene for relative gene expression.
Effect of reference gene normalization on CCNA2 and MYF5 gene expression in bovine skeletal muscle-derived satellite cells. (A) Expression of CCNA2 during proliferation was normalized to the geometric means of GAPDH, RPS15A, PPIA and GAPDH + RPS15A. (B) Expression of MYF5 during the induced differentiation phase was normalized to the geometric means of RPS15A, RPS9, PPIA, and RPS15A + RPS9.
Discussion
Although qRT-PCR techniques have been widely used to elucidate gene expression in the proliferation and differentiation of bovine skeletal muscle-derived satellite cells in vitro, there is no unequivocal data on the most stable reference genes to use for gene expression measurement. In this study, we investigated the proliferation and differentiation of bovine skeletal muscle-derived satellite cells in vitro and compared their Ct values by qRT-PCR using sixteen commonly used reference genes, and then assessed the stability of Ct values using three commonly used algorithms (GeNorm, Normfinder and Bestkeeper). Our study found that GAPDH and RPS15A were the most stable reference genes in the proliferation of bovine SMSCs, while RPS15A and RPS9 were the two most stable reference genes in the induction differentiation of bovine SMSCs. The least stable internal reference gene in proliferation and differentiation was PPIA. These reference genes validated under each experimental condition are a prerequisite for reliable assessment of gene expression by qRT-PCR, as the selection of inappropriate reference genes may yield inaccurate values or even erroneous or contradictory results18,19,20. To the best of our knowledge, this study is the first to validate the stability of reference genes in the proliferation and differentiation of bovine SMSCs in vitro.
This study showed that GAPDH was ranked 4th overall among sixteen candidate reference genes in the in vitro differentiation of bovine SMSCs and had good stability. It was also one of the most stable reference genes in the in vitro proliferation of bovine SMSCs. One study found that GAPDH was a stable reference gene for C2C12 myoblast differentiation, which is consistent with the present study21,22. Previous studies found that GAPDH was not suitable for the normalization of skeletal muscle development in cattle, and similar experimental results were obtained in pigs, goats and mice10,23,24,25. However, those experiments were conducted on the development of muscle tissue, whereas the present study on the proliferation and induction of differentiation of bovine primary skeletal muscle in vitro may have inconsistent results with GAPDH as an internal reference gene. In other tissue studies in cattle, GAPDH as a reference gene was stably expressed in buffalo oocytes, bovine endometrium and bovine blastocysts collected in winter and summer26,27,28, while it was expressed with low stability in bovine ovaries, corpus luteum and myometrium29,30,31, indicating that GAPDH had different stability as an internal reference gene in different tissues of cattle. The results of this study showed that GAPDH has good stability in both proliferation and induced differentiation of bovine SMSCs in vitro. This differs from studies of different tissues in cattle, suggesting that there may be differences in the selection of internal reference genes between in vitro cultures of cells and studies of tissues.
Our study also revealed that RPS15A was one of the most stable internal references in the proliferation of bovine SMSCs in vitro; RPS15A and RPS9 were the two most stable RGs in the induction of differentiation of bovine SMSCs in vitro. This indicates that RPS15A has good stability in the proliferation or induced differentiation in vitro. RPS15A and RPS9 belong to the ribosomal protein family. Several studies have been conducted with other bovine specimens to verify the stability of ribosomal protein family genes as internal reference genes under different experimental conditions. It has been shown that RPS15A and RPS9 were the most stable internal reference genes in the Indian buffalo and bovine peripheral blood mononuclear cells7. RPL15 was also the most stable internal reference in bovine oocytes collected in winter and summer28, while RPL4 was the most stable internal reference gene in bovine bone marrow mesenchymal stem cell differentiation32. RPS9 and RPL19 in bovine maternal reproductive tissues and fetal tissues were found to be stably expressed33. In other species studies, RPL13A and RPL4 were stable internal reference genes in long-term in vitro cultures of human bone marrow MSCs20,34. RPL13 was the most stably expressed internal reference gene in visceral tissues of laying hens, broilers, and turkeys35. Similar to the results of the present study, these findings suggest the potential of ribosomal protein family genes to serve as broadly stable reference genes. However, it has also been shown that although ribosomal protein family genes were the most stably expressed, they may also not be truly stable reference genes36. The ribosomal protein family genes need to be tested to determine if they are stably expressed based on experimental data.
PPIA was the least stable internal reference gene in this study for both in vitro proliferation and induction of differentiation. It should be noted that the stability of PPIA in other tissues of cattle is different. PPIA was previously shown to be least stable in bovine adipose tissue, muscle, mammary gland and liver37, whereas it was stably expressed in oocytes of Bos indicus cattle, in bovine oocytes collected in winter and summer and bovine maternal reproductive and fetal tissues28,33,38. Similar results have been found in reference gene studies in other species, where PPIA was shown to be the least stable reference gene in the quadriceps muscle of energy-restricted mice39. However, PPIA was the most stable reference gene for gene expression in the longissimus dorsi (LD) muscle of postnatal Yorkshire pigs10. The differences in stability of reference genes were considered to be due to differences in experimental conditions, and verifying reference gene stability under each experimental condition was considered a necessary step before analyzing bovine gene expression by qRT-PCR18,19,40.
In this study, we performed an extensive literature search and selected sixteen candidate internal reference genes commonly used in cattle. Although we selected a large number of genes, these candidates may be limited, and there may be more stable internal reference genes that we did not select. The results of transcriptome sequencing to select the candidate reference gene may be more satisfactory, and this may be the direction of stable reference gene selection in the future. In our present study, we selected skeletal muscle-derived satellite cells from the longest dorsal muscle of Qinchuan cattle, however, different sites of skeletal muscle-derived satellite cells may show different results7,41.
In summary, this is the first time that the expression stability of a set of candidate reference genes was validated in vitro for studies of proliferation and induced differentiation of bovine skeletal muscle-derived satellite cells. Three different statistical calculations showed slight differences in the final ranking of reference genes; however, by combining the data, we determined the best combination of stably expressed and least stably expressed internal reference genes for proliferation and induced differentiation of bovine SMSCs under the present experimental conditions. Our results provide an important reference for the selection of stable internal reference genes for bovine gene expression by qRT-PCR analysis in future studies of in vitro proliferation and induced differentiation of bovine SMSCs.
Conclusions
In summary, we evaluated the stability of sixteen reference genes in skeletal muscle-derived satellite cells during in vitro proliferation and induced differentiation using three assays (geNorm, NormFinder, and BestKeeper) to identify the most stable reference genes under different conditions. Our findings suggested that two reference genes were sufficient for accurate normalization in most conditions, although in some cases more than two reference genes may be required to accurately assess the changes in gene expression levels. Importantly, our data indicated that GAPDH and RPS15A were the most suitable reference genes during proliferation in skeletal muscle-derived satellite cells; and that RPS15A and RPS9 were the most suitable reference genes during differentiation. The gene, PPIA was the most variable and least suitable for normalization of SMSCs during in vitro proliferation and induction of differentiation. These findings provide important data for selecting suitable reference genes in future studies. Our data provide guidelines for the selection of appropriate reference genes for studies on skeletal muscle development in mammals.
Materials and Methods
Ethical affirmation
In this study, three-day-old healthy Qinchuan beef cattle were used for myogenic cell isolation in accordance with the guidelines established by the Regulations on the Management of Laboratory Animal Affairs (Ministry of Science and Technology, China, 2004) and approved by the Institutional Animal Care and Use Committee (School of Animal Science and Technology, Northwest Agriculture and Forestry University). Cattles were raised under free-range conditions and samples were collected after euthanasia at the National Beef Cattle Improvement Center (Yangling, China). This study was carried out in compliance with the ARRIVE guidelines.
Isolation and culture of bovine skeletal muscle-derived satellite cells (SMSCs)
SMSCs were obtained from three-day-old healthy Qinchuan beef cattle and isolated as previously described by Wang Yaning, et al.42. The cells were cultured in DMEM/F-12 (Gibco, Shanghai, China), supplemented with 20% FBS (Gibco, Shanghai, China) and 1% penicillin/streptomycin (Hyclone, Thermo-Fisher Scientific, Shanghai. China). When the confluence reached about 80%, the cells were seeded into six-well plates and in vitro proliferation of SMSCs was measured starting from 0 h at a confluence of about 50%. The cells were collected at 0, 12, 24, 36 and 48 h, with three biological replicates for each time point. To study induction of differentiation, SMSCs were grown in culture medium to 80% confluence in six-well plates, and on day-zero (0 d) the medium was changed to differentiation medium containing DMEM/F-12, 2% horse serum (Gibco, Shanghai, China) and 1% penicillin/streptomycin. The cells were collected at 0, 2 , 4 , 6 , and 8 d, with three biological replicates at each time point.
Immunofluorescence assay for myocyte-specific protein
SMSCs were cultured in 12-well plates, fixed with 4% paraformaldehyde for 15 min at room temperature, washed with PBS, permeabilized with 0.2% Triton X-100 for 15 min and then incubated in 10% (vol/vol) normal donkey serum/1% BSA (Sigma) /0.3 M glycine (Sigma) for 1 h to block non-specific protein–protein interactions at room temperature. For immunofluorescence, the cells were incubated with the primary antibody (diluted in 10% normal donkey serum/1% BSA/0.3 M glycine) overnight at 4˚C. The cells were then washed with PBS and incubated with secondary antibody at 37˚C for 1 h (protected from light). The nuclei were stained with DAPI (Sigma) at room temperature for 15 min (protected from light). The antibodies used were as follows: anti-MYHC (1:300, GeneTex), and donkey anti-mouse IgG H&L (AlexaFluor1 555) (1:1000, Abcam). DAPI was used at a concentration of 1 μg/ml. Immunofluorescence images were obtained using an Evos-fl-auto2 microscopy imaging system (Thermo Scientific, USA).
RNA extraction and reverse transcription to cDNA
Total RNA was extracted from the third cell passage of bovine SMSCs using a kit containing TRIzol (Takara) according to instructions. The quality and concentration of the RNA were determined by electrophoresis of aliquots on a 1% agarose gel and measuring A260/A280 with a microplate reader. Using the PrimeScript RT kit with gDNA Eraser (Perfect Real Time; Takara), 1 µg of total RNA was reverse transcribed to cDNA and stored at -80 °C for backup.
Primer design
The primer sequences of relevant reference genes in bovine skeletal muscle cells from previous studies were checked by the Primer-Blast program (NCBI tools). The primers with good specificity were used directly, while those with poor specificity were redesigned using Primer Premier 5, according to primer length, annealing temperature, base composition and 3'-end stability; specificity of the primers was checked using the Primer-Blast program (NCBI tools) to ensure primer specificity (Table 5).
Standard curve construction of qRT-PCR primers
A total of sixteen genes, including GAPDH, ACTB, PPIA, LRP10, HPRT1, YWHAZ, B2M, TBP, EIF3K, RPS9, UXT, 18S rRNA, RPLP0, MARVELD, EMD and RPS15A, were selected as candidate internal reference genes in this study. The primers were synthesized by Chengdu Prime Biotechnology Co., Ltd, and the cDNA template was diluted by 1, 5–1, 5–2, 5–threefold for qRT-PCR. The standard curve was plotted as the logarithm of the sample concentration on the x-axis and the ct value on the y-axia. The amplification efficiency was calculated according to the formula: E = 10(-1/slope) − 1.
Quantitative real-time PCR (qRT-PCR)
The cDNA stored at − 80 °C was diluted tenfold and qRT-PCR was performed using SYBR green real-time PCR master mix (Takara) in a CFX96 BIO-RAD thermocycler (USA). The final reaction volume of 25 µl contained 12.5 µl of SYBR Premix Ex Taq II (Tli RNaseH Plus) (2 ×), 1 µl each of forward and reverse primers (final primer concentration 0.4 µM), 2 µL of cDNA and 8.5 µL dd H2O. Thermocycling conditions were: denaturation at 95 °C for 5 min followed by forty amplification cycles with denaturing at 95 °C for 30 s, 60 °C for 30 s and 72 °C 30 s.
Stability analysis of reference genes
The expression stability of selected reference genes was evaluated using three programs: GeNorm11, Normfinder12, and Bestkeeper13, following the developer's instructions. GeNorm was also used to calculate pairwise variations (V/V values) and to determine the minimum number of reference genes required for accurate normalization. The geometric mean of the ranking values of the internal reference genes in different software was used for integrated stability analysis to obtain a reliable agreement.
Validation of reference gene expression
The stability of the tested reference genes was verified by measuring the expression of CCNA2 at different times during proliferation and the expression of MYF5 at different times during the differentiation of bovine SMSCs in vitro. The expression of CCNA2, MYF5 was normalized using the most stable candidate reference gene group and the least stable candidate reference gene.
References
Li, W. et al. A multiallelic indel in the promoter region of the Cyclin-dependent kinase inhibitor 3 gene is significantly associated with body weight and carcass traits in chickens. Poult. Sci. 98, 556–565. https://doi.org/10.3382/ps/pey404 (2019).
Molkentin, J. D. & Olson, E. N. Defining the regulatory networks for muscle development. Curr. Opin. Genet. Dev. 6, 445–453. https://doi.org/10.1016/s0959-437x(96)80066-9 (1996).
Yin, H. D. et al. Housing system influences abundance of Pax3 and Pax7 in postnatal chicken skeletal muscles. Poult. Sci. 93, 1337–1343. https://doi.org/10.3382/ps.2013-03555 (2014).
Bustin, S. A. Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): Trends and problems. J. Mol. Endocrinol. 29, 23–39. https://doi.org/10.1677/jme.0.0290023 (2002).
Yuan, J. S., Reed, A., Chen, F. & Stewart, C. N. Jr. Statistical analysis of real-time PCR data. BMC Bioinform. 7, 85. https://doi.org/10.1186/1471-2105-7-85 (2006).
Zhang, J. et al. Identification of suitable reference genes for quantitative RT-PCR during 3T3-L1 adipocyte differentiation. Int. J. Mol. Med. 33, 1209–1218 (2014).
Kishore, A. et al. Selection of stable reference genes in heat stressed peripheral blood mononuclear cells of tropically adapted Indian cattle and buffaloes. Mol. Cell Probes 27, 140–144. https://doi.org/10.1016/j.mcp.2013.02.003 (2013).
Bustin, S. A. et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622. https://doi.org/10.1373/clinchem.2008.112797 (2009).
Li, Y., Chen, W., Wang, Q., Wang, N. & Wu, Y. F. Assessment of reference genes for quantitative real-time PCR gene expression normalization in periwinkle during Wheat Blue Dwarf phytoplasma infection. Australas. Plant Pathol. 43, 477–485 (2014).
Niu, G. et al. Identifying suitable reference genes for gene expression analysis in developing skeletal muscle in pigs. PeerJ 4, e2428. https://doi.org/10.7717/peerj.2428 (2016).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, Research0034. https://doi.org/10.1186/gb-2002-3-7-research0034 (2002).
Andersen, C. L., Jensen, J. L. & Orntoft, T. F. Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Can. Res. 64, 5245–5250. https://doi.org/10.1158/0008-5472.can-04-0496 (2004).
Pfaffl, M. W., Tichopad, A., Prgomet, C. & Neuvians, T. P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper–Excel-based tool using pair-wise correlations. Biotech. Lett. 26, 509–515. https://doi.org/10.1023/b:bile.0000019559.84305.47 (2004).
Bendris, N. et al. Cyclin A2: A genuine cell cycle regulator?. Biomol. Concepts 3, 535–543 (2012).
Abdelhalim, et al. Cyclin A2: At the crossroads of cell cycle and cell invasion. World J. Biol. Chem. 6, 346–350 (2015).
Bentzinger, C., Yu, X. W. & Rudnicki, M. A. Building muscle: Molecular regulation of myogenesis. Cold Spring Harb. Perspect. Biol. 4, 441–441 (2012).
Ustanina, S., Carvajal, J., Rigby, P. & Braun, T. The myogenic factor Myf5 supports efficient skeletal muscle regeneration by enabling transient myoblast amplification. Stem Cells 25, 2006–2016 (2010).
Jeon, R. H. et al. PPIA, HPRT1, and YWHAZ genes are suitable for normalization of mRNA expression in long-term expanded human mesenchymal stem cells. Biomed. Res. Int. 2019, 3093545–3093545 (2019).
Palombella, S. et al. Identification of reference genes for qPCR analysis during hASC long culture maintenance. PLoS ONE 12, e0170918. https://doi.org/10.1371/journal.pone.0170918 (2017).
Dheda, K. Validation of housekeeping genes for normalizing RNA expression in real-time PCR. Biotechniques 37, 118–119 (2004).
Nishimura, M., Nikawa, T., Kawano, Y., Nakayama, M. & Ikeda, M. Effects of dimethyl sulfoxide and dexamethasone on mRNA expression of housekeeping genes in cultures of C2C12 myotubes. Biochem. Biophys. Res. Commun. 367, 603–608. https://doi.org/10.1016/j.bbrc.2008.01.006 (2008).
Masilamani, T. J., Loiselle, J. J. & Sutherland, L. C. Assessment of reference genes for real-time quantitative PCR gene expression normalization during C2C12 and H9c2 skeletal muscle differentiation. Mol. Biotechnol. 56, 329–339. https://doi.org/10.1007/s12033-013-9712-2 (2014).
Saremi, B., Sauerwein, H., Danicke, S. & Mielenz, M. Technical note: Identification of reference genes for gene expression studies in different bovine tissues focusing on different fat depots. J. Dairy Sci. 95, 3131–3138. https://doi.org/10.3168/jds.2011-4803 (2012).
Najafpanah, M. J., Sadeghi, M. & Bakhtiarizadeh, M. R. Reference genes selection for quantitative real-time PCR using RankAggreg method in different tissues of Capra hircus. PLoS ONE 8, e83041. https://doi.org/10.1371/journal.pone.0083041 (2013).
Thomas, K. C. et al. Evidence based selection of commonly used RT-qPCR reference genes for the analysis of mouse skeletal muscle. PLoS ONE 9, e88653. https://doi.org/10.1371/journal.pone.0088653 (2014).
Luchsinger, C. et al. Stability of reference genes for normalization of reverse transcription quantitative real-time PCR (RT-qPCR) data in bovine blastocysts produced by IVF, ICSI and SCNT. Zygote (Cambridge, England) 22, 505–512. https://doi.org/10.1017/s0967199413000099 (2014).
Lesage-Padilla, A. et al. Maternal metabolism affects endometrial expression of oxidative stress and FOXL2 genes in cattle. PLoS ONE 12, e0189942. https://doi.org/10.1371/journal.pone.0189942 (2017).
Macabelli, C. H. et al. Reference gene selection for gene expression analysis of oocytes collected from dairy cattle and buffaloes during winter and summer. PLoS ONE 9, e93287. https://doi.org/10.1371/journal.pone.0093287 (2014).
Schoen, K., Plendl, J., Gabler, C. & Kaessmeyer, S. Identification of stably expressed reference genes for RT-qPCR data normalization in defined localizations of cyclic bovine ovaries. Anat. Histol. Embryol. 44, 200–211. https://doi.org/10.1111/ahe.12128 (2015).
Rekawiecki, R., Rutkowska, J. & Kotwica, J. Identification of optimal housekeeping genes for examination of gene expression in bovine corpus luteum. Reprod. Biol. 12, 362–367. https://doi.org/10.1016/j.repbio.2012.10.010 (2012).
Rekawiecki, R., Kowalik, M. K. & Kotwica, J. Validation of housekeeping genes for studying differential gene expression in the bovine myometrium. Acta Vet. Hung. 61, 505–516. https://doi.org/10.1556/AVet.2013.037 (2013).
Jang, S. J. et al. TATA box binding protein and ribosomal protein 4 are suitable reference genes for normalization during quantitative polymerase chain reaction study in bovine mesenchymal stem cells. Asian-Australas J. Anim. Sci. 33, 2021–2030. https://doi.org/10.5713/ajas.20.0238 (2020).
Cheng, L. et al. Identification of reliable reference genes for expression studies in maternal reproductive tissues and foetal tissues of pregnant cows. Reprod. Domest. Anim. 55, 1554–1564. https://doi.org/10.1111/rda.13808 (2020).
Li, X., Yang, Q., Bai, J., Yang, Y. & Wang, Y. Identification of optimal reference genes for quantitative PCR studies on human mesenchymal stem cells. Mol. Med. Rep. 11, 1304–1311 (2015).
Mitra, T., Bilic, I., Hess, M. & Liebhart, D. The 60S ribosomal protein L13 is the most preferable reference gene to investigate gene expression in selected organs from turkeys and chickens, in context of different infection models. Vet. Res. 47, 105 (2016).
Thorrez, L. et al. Using ribosomal protein genes as reference: A tale of caution. PLoS ONE 3, e1854. https://doi.org/10.1371/journal.pone.0001854 (2008).
Bonnet, M., Bernard, L., Bes, S. & Leroux, C. Selection of reference genes for quantitative real-time PCR normalisation in adipose tissue, muscle, liver and mammary gland from ruminants. Anim. Int. J. Anim. Biosci. 7, 1344–1353 (2013).
Caetano, L. C., Miranda-Furtado, C. L., Batista, L. A., Pitangui-Molina, C. P. & Rosa-E-Silva, A. Validation of reference genes for gene expression studies in bovine oocytes and cumulus cells derived from in vitro maturation. Anim. Reprod. 16, 290–296 (2019).
Gong, H. et al. Evaluation of candidate reference genes for RT-qPCR studies in three metabolism related tissues of mice after caloric restriction. Sci. Rep. 6, 38513. https://doi.org/10.1038/srep38513 (2016).
Ragni, E., Viganò, M., Rebulla, P., Giordano, R. & Lazzari, L. What is beyond a qRT-PCR study on mesenchymal stem cell differentiation properties: How to choose the most reliable housekeeping genes. J. Cell Mol. Med. 17, 168–180 (2013).
Kozera, B. & Rapacz, M. Reference genes in real-time PCR. J. Appl. Genet. 54, 391–406 (2013).
Wang, Y. N. et al. Myocyte enhancer factor 2A promotes proliferation and its inhibition attenuates myogenic differentiation via myozenin 2 in bovine skeletal muscle myoblast. PLoS ONE 13, e0196255. https://doi.org/10.1371/journal.pone.0196255 (2018).
Divari, S., Berio, E., Biolatti, B. & Cannizzo, F. T. Reference gene selection and prednisolone target gene expression in adipose tissues of friesian cattle. J. Agric. Food Chem. 65, 11140–11145 (2017).
Cao, K. X. et al. Cold exposure induces the acquisition of brown adipocyte gene expression profiles in cattle inguinal fat normalized with a new set of reference genes for qRT-PCR. Res. Vet. Sci. 114, 1–5. https://doi.org/10.1016/j.rvsc.2017.02.021 (2017).
Roberts, S. L., Lancaster, P. A., DeSilva, U., Horn, G. W. & Krehbiel, C. R. Coordinated gene expression between skeletal muscle and intramuscular adipose tissue in growing beef cattle. J. Anim. Sci. 93, 4302–4311. https://doi.org/10.2527/jas.2015-8886 (2015).
Schering, L., Albrecht, E., Komolka, K., Kuhn, C. & Maak, S. Increased expression of thyroid hormone responsive protein (THRSP) is the result but not the cause of higher intramuscular fat content in cattle. Int. J. Biol. Sci. 13, 532–544. https://doi.org/10.7150/ijbs.18775 (2017).
Baddela, V. S., Baufeld, A., Yenuganti, V. R., Vanselow, J. & Singh, D. Suitable housekeeping genes for normalization of transcript abundance analysis by real-time RT-PCR in cultured bovine granulosa cells during hypoxia and differential cell plating density. Reprod. Biol. Endocrinol. 12, 1–7 (2014).
Acknowledgements
This research was funded by the National Key Research and Development Program of China (No. 2018YFD0501700), the National Modern Agricultural Industry Special Program (No. CARS-37), the National Natural Science Foundation of China (31972994), the Agricultural Science and Technology Innovation and Transformation Project of Shaanxi Province (NYKJ-2018-LY09) and the Key Research and Development Program of Ningxia (Grant No. 2019BEF02004).
Author information
Authors and Affiliations
Contributions
L.-S.Z., G.-H.W. and C.-c.L. led the experiments and designed the analytical strategy. L.-S.Z., G.C., and B.-z.L. performed animal work and prepared biological samples. Guo-Hua Wang wrote the manuscript. L.-S.Z., W.-Z.Z., C.-C.L., G.C., X.-Z.D., revised the manuscript. All the authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, GH., Liang, CC., Li, BZ. et al. Screening and validation of reference genes for qRT-PCR of bovine skeletal muscle-derived satellite cells. Sci Rep 12, 5653 (2022). https://doi.org/10.1038/s41598-022-09476-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-09476-3
- Springer Nature Limited