Abstract
Severe acute respiratory coronavirus 2 (SARS-CoV-2) has spread globally since its emergence in 2019. Most SARS-CoV-2 infections generate immune responses leading to rising levels of immunoglobulins (Ig) M, A and G which can be detected using diagnostic tests including enzyme-linked immunosorbent assays (ELISA). Whilst implying previous SARS-CoV-2 infection, the detection of Ig by ELISA does not guarantee the presence of neutralising antibodies (NAb) that can prevent the virus infecting cells. Plaque reduction neutralisation tests (PRNT) detect NAb, but are not amenable to mass testing as they take several days and require use of SARS-CoV-2 in high biocontainment laboratories. We evaluated the ability of IgG and IgM ELISAs targeting SARS-CoV-2 spike subunit 1 receptor binding domain (S1-RBD), and spike subunit 2 (S2) and nucleocapsid protein (NP), at predicting the presence and magnitude of NAb determined by PRNT. IgG S2 + NP ELISA was 96.8% [95% CI 83.8–99.9] sensitive and 88.9% [95% CI 51.8–99.7] specific at predicting the presence of NAbs (PRNT80 > 1:40). IgG and IgM S1-RBD ELISAs correlated with PRNT titre, with higher ELISA results increasing the likelihood of a robust neutralising response. The IgM S1-RBD assay can be used as a rapid, high throughput test to approximate the magnitude of NAb titre.
Similar content being viewed by others
Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a novel, pandemic betacoronavirus that began spreading globally in early 2020. To date, there have been over 198 million reported infections and more than 4.2 million deaths1.
Most individuals infected with SARS-CoV-2 develop humoral immune responses, characterised by rising titres of immunoglobulins (Ig) M, A and G, within the first 2–3 weeks of infection2,3, which are detectable using enzyme-linked immunosorbent assays (ELISA). The presence of SARS-CoV-2 specific Ig therefore provides evidence of previous infection4, although their detection does not guarantee the presence of functional immunity against the virus5. For example, the viral nucleocapsid protein (NP), an abundant viral antigen, generates robust antibody responses, and is therefore a good antigen for diagnostic serological assays6, however these antibodies are not neutralising7,8.
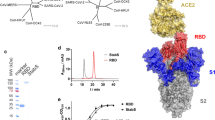
SARS-CoV-2 neutralising antibodies (NAb) primarily bind the receptor-binding domain of the spike (S) protein and disrupt virus entry by blocking interaction with the angiotensin converting enzyme 2 (ACE2) receptor of host cells7,9. The activity of these functional antibodies can be measured using the plaque reduction neutralisation test (PRNT). However, this method is not amenable to mass testing, as the process takes several days and requires working with SARS-CoV-2 in high biocontainment laboratories. Pseudotyped virus models may be used to detect neutralising antibodies under reduced biocontainment, but still lack the speed and reproducibility of ELISA assays10.
Previous studies have reported that NAb levels correlate with IgG and IgM titres11,12,13,14, but this relationship is variable, depending on the timing of sampling in the course of the infection and the antigen targets of the serological assays15. Here we evaluate the ability of SARS-CoV-2 IgG and IgM ELISAs to predict the presence and magnitude of SARS-CoV-2 NAbs in hospitalised COVID-19 patients.
Methods
Ethical statement
The study was conducted in accordance with relevant UK guidelines and regulations. Ethics approval was provided by the Institutional Review Board (South Central—Oxford C Research Ethics Committee, Research Development and Assessment of Rapid Testing for SARS-CoV-2 outbreak study; Integrated Research Application System project ID:282104; Research Ethics Committee Reference 20/SC/0171; registered at clinicaltrial.gov NCT04351646). The approved protocol permitted the analysis of antibody responses using anonymised excess diagnostic material (EDM) from the pathology laboratory of patients with and without PCR-confirmed SARS-CoV-2 infection. Informed consent was not required under the ethical approval status of the work and due to the nature of the samples.
Serum samples
Anonymized EDM serum samples from hospital patients with SARS-CoV-2 infection confirmed by reverse transcription—quantitative polymerase chain reaction (RT-qPCR) were used for this study and were selected from 645 EDM serum samples that were collected from a pool of 177 patients treated at St George’s Hospital, London UK16. Where possible, samples were selected from patients at least 10 days post-RT-qPCR confirmation. Samples were grouped based on their normalised optical density (NOD) values derived from an anti-SARS-CoV-2 IgG S2 and NP ELISA carried out previously16. These were grouped into “negative NOD” values (< 0; indicating the patient had not seroconverted), “low NOD” (0 to 0.5), “medium NOD” (0.9 to 1.1); and “high NOD” (> 1.5). The final sample available from all patients was chosen for this study. The narrow “medium NOD” window was purposely selected to reduce sample numbers in this grouping, as the grouping 0.5 to 1.5 contained 5–6 times more samples than the other groupings. A single sample was then selected from any patient with at least 3 samples with NOD values remaining in one NOD grouping (i.e. indicating a stable antibody response). The serum sample selected for any given patient was that collected furthest from the swab taken for confirmation of SARS-CoV-2 infection (and at least 10 days post-swab). The approach resulted in 9, 9, 11 and 12 patient samples in each group (41 single patient serum samples in total).
All participants were confirmed as positive for SARS-CoV-2 using RT-PCR from nose/throat swabs (in Sigma Virocult®, Corsham, UK) and Roche RNA extraction kits (Magnapure, West Sussex, UK) followed by Altona Diagnostics RealStar® SARS-CoV-2 RT-PCR (S and E target genes, Hamburg, Germany) or Roche cobas® SARS-CoV-2 Test (E and ORF target genes).
ELISA to detect anti-SARS-CoV-2 IgM binding S2 and NP antigens
An anti-SARS-CoV-2 IgM ELISA (Mologic, Bedfordshire, UK), which targets the NP and S2 antigens, were used to measure antibodies, as per the manufacturer’s instructions. Briefly, sera were diluted (1:200) and incubated on a S2/NP pre-coated plate (30 min) at room temperature and then washed three times. Conjugated antibody (anti-human IgM) was then applied to each well and incubated (30 min) at room temperature. Following washing (× 4), TMB substrate was added and incubated for 10 min at room temperature before addition of stop solution. Optical densities (OD) were read at 450 nm within 10 min of addition of the stop solution.
ELISA to detect anti-SARS-CoV-2 IgG binding the receptor binding domain of S1 (S1-RBD)
An anti-SARS-CoV-2 IgG ELISA (Abcam, Cambridgeshire, UK), which targets S1-RBD antigen, was used to measure human antibodies, as per the manufacturer’s instructions. Briefly, samples (sera diluted 1 in 100), a calibrator (Recombinant Human anti-RBD IgG), positive control (Recombinant Human anti-RBD IgG) and negative control (diluent) were added to an S1-RBD pre-coated plate. The plate was sealed with film and incubated (30 min) at room temperature and then washed five times. HRP Conjugated antibody (Goat Anti-Human IgG-Fc) was then applied to each, the plate sealed and incubated (30 min) at room temperature. Following washing (× 5), TMB substrate was added and the plate incubated for 15 min in the dark at room temperature before addition of stop solution. Optical densities (OD) were read at 450 nm and 570 nm within 10 min of addition of the stop solution.
ELISA to detect anti-SARS-CoV-2 IgM binding the receptor binding domain of S1 (S1-RBD)
An anti-SARS-CoV-2 IgM ELISA (Abcam, Cambridgeshire, UK), which targets S1-RBD antigen, was also used to measure human antibodies, following the same procedure but with HRP conjugated Goat Anti-Human IgM-Fc antibody and Recombinant Human anti-RBD IgM controls.
Plaque reduction neutralisation tests
Vero E6 cells were seeded into 24-well cell culture plates at a density of 250,000 cells/ml and incubated (24 h, 37 °C, 5% CO2). The following day serum samples were heated to inactivate complement (56 °C for 1 h). Heat-inactivated serum samples were twofold serially diluted in infection media (DMEM with 2% v/v FBS and 1:1000 50 mg/ml gentamicin). Under biosafety level 3 conditions, SARS-CoV-2 isolate REMRQ0001/Human/2020/Liverpool17 was added to an equal volume of diluted patient serum, at a titre of 800 pfu/ml, to achieve 12 final serum dilutions from 1:20 to 1:40,960 for each patient sample. Following incubation (1 h, 37 °C), the virus-serum mixture (100 µl) was inoculated onto Vero E6 cells and incubated (1 h, 37 °C, 5% CO2) before applying an overlay of infection media containing agarose (0.4% w/v). Infected cells were then incubated (48 h, 37 °C, 5% CO2). The assays were fixed with formaldehyde (37% w/v), stained with crystal violet solution (0.25% w/v) and allowed to air dry. The PRNT80 was determined as the lowest dilution of serum that produced a ≥ 80% reduction in the number of plaques compared to controls that contained no patient serum. The investigators were blinded to the ELISA status of the samples when performing the PRNTs.
Western blots
Western blots were conducted to investigate the antigen binding profiles associated with the neutralising responses. Recombinant S1, S2 and NP (Native Antigen Company, Kidlington, UK) were prepared in NuPAGE LDS sample buffer (Invitrogen, Carlsbad, USA) with 50 mM dithiothreitol (Sigma, St Louis, USA), and heated for 10 min at 70 °C. Protein (1 µg) was loaded on a Mini-PROTEAN TGX 12% PAGE gel (BioRad, Hercules, USA) and run under reducing conditions in Tris/Glycine/SDS Buffer (BioRad, Hercules, USA). Proteins were transferred to 0.2 µm nitrocellulose membrane using BioRad TransBlot Turbo system mixed MW programme, and blocked in 5% (v/v) goat serum (Sigma, St Louis, USA) in PBS (Gibco, Waltham, USA) with 0.1% (v/v) Tween 20 (Sigma, St Louis, USA) overnight at 4 °C with gentle shaking. Blots were incubated with sera diluted 1 in 200 in blocking solution for 1 h at room temperature with gentle shaking and washed three times for 5 mins with PBS-0.1% (v/v) Tween 20. Blots were then incubated with goat anti-human Kappa-AP and anti-human Lambda-AP (Southern Biotech, Birmingham, USA), both diluted 1 in 1000 in blocking solution, for 1 h at room temperature with gentle shaking, followed by three washes as above. Blots were developed with BCIP/NBT (Sigma, St Louis, USA) for 1 min and stopped with H2O. The assays were scored visually as positive or negative by two independent observers who were blinded to the ELISA status of the samples. In the event of a disagreement between observers, a third independent observer determined the result. One IgG ELISA negative sample was excluded because there was not enough serum remaining for western blot.
Statistical analysis
We model the PRNT80 outcome for the i-th patient, denoted by \({Y}_{i}\), as an ordinal variable taking the values from k = 1, corresponding to a titre less than 1:20, to k = 10, for a titre of 1:2560 or more. We assume that the observed titre values correspond to the discretization of a continuous antibody distribution, \({Y}_{i}^{*}\), which we assume to follow a log-Gaussian distribution with mean \({\vartheta }_{i}\) and variance \({\sigma }^{2}\).
Hence, we can write:
where \({a}_{1}=0\), \({a}_{2}=20\), \({a}_{3}=40\) and so forth up \({a}_{10}=2560\) and with the convention \({a}_{11}=\infty \). The probability in (1) is computed using the cumulative density function of a log-Gaussian distribution.
We model the mean concentration \({\vartheta }_{i}\) of PRNT80 as a log-linear regression on: \({a}_{i}\), a covariate which defined as IgG S1-RBD, IgM S1-RBD, IgG S2 + NP and IgM S1 + NP, in four different fitted models; and the days post-symptom, \({b}_{i}\). Hence, the linear predictor for \({\vartheta }_{i}\) is:
where the \({\alpha }\) is the intercept; \({\beta }\) express the strength of the selected antibody concentrations on the log-mean levels of PRNT80; similarly, \(\gamma \) indicates the effect of the sampling time days post-symptom onset. All four models are fitted via maximum likelihood using the R software environment (Supplementary Data 1). Since the variable days post-symptom was not found to be significant at the conventional 95% confidence level, Tables 1 and 2 show the estimates from the best fitting models according to the likelihood values, after excluding that variable.
Results
Patient demographics and symptoms
Twenty-five (61.0%) of the 41 patients were male, and the median age was 63 (IQR 55–71) years. Seventeen patients (41.5%) were classified as white, 13 (31.7%) non-white, and 11 (26.8%) were of unknown or ‘other’ ethnicity. Twenty-three patients (56.1%) had one or more comorbidities. Ten patients (24.4%) were obese, with a body mass index (BMI) > 30. These patients come from a subset already described16.
Of the thirty-nine patients for whom data were available, 35 (89.7%) were symptomatic at the time of their initial swab. Thirty-three of these patients (94.3%) had one or more of the classic triad of symptoms: cough, fever, and shortness of breath, 12 (34.3%) had gastrointestinal symptoms, including nine patients with diarrhoea. Three patients (7.3%) had an incidental positive swab, taken prior to admission to a rehabilitation facility, and three patients were swabbed following contact with a patient who had tested positive. The median interval between the onset of symptoms and date of the first positive swab was four days (IQR 3–7 days). Seven (17%) patients died within 28 days of their first positive swab. The median timing of serum sampling post symptom onset was 29 days (range 13–60 days). The sample timing post-symptom onset was not available for six patients.
Diagnostic sensitivity of ELISAs at identifying a PRNT80 ≥ 1:40
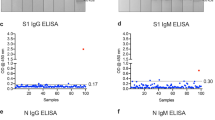
The correlation of IgG and IgM ELISAs and NAb responses are shown in Fig. 1. The IgG S1-RBD and IgG S2 + NP ELISAs identified a PRNT80 ≥ 1:40 with 100% [95% CI 89.1–100.0]) and 96.8% [95% CI 83.8–99.9] sensitivity, respectively (Table 3). This was higher than either of the IgM ELISAs (IgM S1-RBD sensitivity 62.5% [95% CI 43.7–78.9]; IgM S2 + NP sensitivity 62.5% [95% CI 43.7–78.9]). However, the IgG S1-RBD ELISA demonstrated poor specificity (44.4% [95% CI 13.7–78.8]) compared to the IgM S1-RBD (100.0% [95% CI 66.4–100.0]), IgG S2 + NP 88.9% [95% CI 51.8–99.7]), and IgM S2 + NP ELISAs (88.9% [95% CI 51.8–99.7]).
Four samples that were only ELISA positive by the IgG S1-RBD assay, did not exhibit a neutralising response (PRNT ≥ 1:40). In addition, one sample was positive by both IgG S1-RBD, and IgG S2 + NP ELISA but did not show a neutralising response (PRNT80 ≥ 1:40). Both IgM ELISAs were negative for this sample. One non-neutralising sample (PRNT80 < 1:40) was only positive by IgM S2 + NP ELISA. One sample that had a PRNT80 of 1:40 was negative by all ELISAs except IgG S1-RBD. Several samples with neutralising responses (PRNT80 ≥ 1:40) were negative by both IgM ELISAs, but positive by both IgG assays.
Correlation of ELISA and PRNT
Separate ordinal outcomes models were fitted to explore the relationship of each of the IgG and IgM ELISAs, and the timing of serum sampling post-onset of symptoms (denoted \(\gamma \)) on the observed neutralising titre (PRNT80). The maximum likelihood estimates from these models are shown in Tables 4, 5, 6 and 7. The IgG S1-RBD, IgM S1-RBD and IgG S2 + NP ELISAs were significant predictors of PRNT80 (0.191 [95% CI 0.117–0.266], 0.507 [95% CI 0.337–0.676], and 0.690 [95% CI 0.393–0.986], respectively). The IgM S2 + NP ELISA result was not a significant predictor of PRNT80 (− 0.053 [95% CI − 0.873–0.768]). The value of the likelihood functions for each of these models show that the IgG S1-RBD (-78.338) and IgM S1-RBD (− 77.414) assays best predicted PRNT80 (Table 8). The IgG and IgM S1-RBD maximum likelihood estimates, after excluding the non-significant \(\gamma \) variable, indicated that each unit increase in antibody titre by ELISA was associated with a 0.187 [95% CI 0.112–0.262], and 0.454 [95% CI 0.302–0.607)-fold increased likelihood of being in a higher PRNT80 category, respectively.
Predicted PRNT80 based on IgM S1-RBD and IgG S1-RBD ELISA
It was assumed that the relationship between IgM and IgG S1-RBD titres, and PRNT80 was unlikely to be linear across all the ranges of observed ELISA results. Therefore, the model predictions for PRNT80 based on IgG and IgM S1-RBD were considered for the first quartile, median, and third quartile of the antibody titres as measured by ELISA (see Fig. 2). For the first quartile, there was a low probability (< 0.15) of observing neutralising titres ≥ 1:20 for both IgM and IgG S1-RBD (Fig. 2A). For median titres, the highest probability (0.15) was for PRNT80 values in the range of 1:80–1:160 (Fig. 2B). Model predictions for IgG and IgM S1-RBD ELISA results in the third quartile indicated that PRNT80 values ≥ 1:320 were most likely to be observed for both IgG and IgM S1-RBD ELISAs (Fig. 2C).
Across the range of IgG S1-RBD titres detected by ELISA, we predicted that there was a low probability of observing an increasing PRNT80 value with increasing IgG S1-RBD, with predicted PRNT80 values remaining low despite increasing antibody binding detected by ELISA (Fig. 2D). IgM S1-RBD ELISA result demonstrated a stronger predicted quantifiable relationship with PRNT80, albeit with increasing uncertainty at higher IgM S1-RBD ELISA results (Fig. 2D).
Antibody binding in western blot assays to all three of the S1, S2 and NP antigens, or to both the S2 and NP antigens, were seen in all samples that had high neutralising titres (PRNT80 ≥ 1:80) (Fig. 3). In samples with low PRNT80 (< 1:40) there was greater variability in antigen binding, with a larger proportion demonstrating antibody binding to single antigens, or combinations of two antigens involving S1. Three of the six samples that did not achieve PRNT80 ≥ 1:20 demonstrated a lack of binding to S1, S2 or NP. The full suite of ELISA, PRNT and Western blot results for each serum sample are available in Supplementary Data 2. Images of all Western blots are included in Supplementary Data 3.
Discussion
Our data show that the IgG S2 + NP ELISA provides a sensitive (96.8%) and specific (88.9%) test to predict the presence of SARS-CoV-2 neutralising antibodies (PRNT80 ≥ 1:40). This is an interesting finding given that antibodies directed against S2 and NP are not considered to be highly neutralising8. However, NP-only IgG ELISAs have performed similarly to those targeting only the S1 antigen in previous evaluations18,19. This suggests that anti-NP antibodies are raised as part of a broad immune response, and whilst not directly neutralising, are indicative of the presence of other neutralising immunoglobulins. This is supported by our western blot analysis which revealed that the majority of serum samples with neutralising activity had antibodies directed against the NP, S1 and S2.
The IgG S1-RBD ELISA was also highly sensitive (100%) but was much less specific (44.4%). Our findings show there was a substantial chance of receiving a positive IgG S1-RBD ELISA result when there was no corresponding neutralising activity. Both IgG ELISAs were more sensitive than the IgM S1-RBD (62.5%) and IgM S2-RBD (62.5%) assays. However, the IgM S1-RBD ELISA demonstrated high specificity (100.0%). Previously, IgM ELISAs have been reported to be more predictive of neutralising titres than IgG, but given the short duration of IgM expression, the timing of serum sampling post-infection is an important determinant of these relationships14,20,21.
Our ordinal outcomes model demonstrated that both IgM S1-RBD and IgG S1-RBD ELISAs had the strongest correlation with PRNT80. Whilst our findings suggest that there is an increased probability of being in a higher PRNT80 category with increasing ELISA IgM or IgG S1-RBD titre, there was considerable variation in ELISA titre within each PRNT80 category. Consequently, it is not possible to make accurate predictions on the magnitude of NAb response based on the IgG S1-RBD ELISA. The model predictions indicate that IgM S1-RBD ELISA is the best predictor of the scale of the neutralising response, but there is significant uncertainty, especially at higher ELISA titres. Previous studies have reported a significant correlation between ELISAs targeting anti-spike and anti-RBD antigens of both IgG and IgM subclasses with the neutralisation titres established by microneutralisation tests, PRNTs and pseudotyped virus neutralisation assays6,15,20,22,23,24. However, all have noted a similar broad range of ELISA antibody titres within each neutralisation category.
The main limitation of our study is the small sample size. As the relationship between serological assays and neutralising titres appears to be variable, a larger sample size would have sufficient power to reveal overall trends and minimise the effects of outliers. IgA ELISA results have been reported to correlate well with neutralisation titres24. As we did not measure IgA, it is possible that neutralising activity due to IgA had a confounding effect on the trends observed. Moreover, our cohort only includes hospitalised patients with severe COVID-19 and therefore the findings of this study may not be applicable to individuals with mild and asymptomatic infections, or those who have been vaccinated against SARS-CoV-2, in whom antibody responses may be different.
In conclusion, ELISAs can be used as a proxy for PRNT to both predict the presence, and estimate the magnitude, of a neutralising antibody response. IgG ELISA targeting S2 + NP provides a sensitive and specific test to determine the presence of NAb. Whilst accurate predictions of precise PRNT80 values based on IgM S1-RBD ELISA are not possible, broad inferences on the magnitude of the neutralising response can be made using this assay. Assessment of NAb titre by PRNT is the gold standard and should be performed where feasible. However, given the biosafety requirements and low throughput nature of this technique, there is likely to be benefit in using more rapid and high throughput ELISAs to obtain approximations of NAb titre.
References
WHO. WHO Coronavirus Disease (COVID-19) Dashboard. https://covid19.who.int/. Accessed 22 Feb 2021.
To, K.K.-W. et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: An observational cohort study. Lancet. Infect. Dis 20(5), 565–574 (2020).
Long, Q.-X. et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat. Med. 26(6), 845–848 (2020).
Adams, E. et al. Evaluation of antibody testing for SARS-CoV-2 using ELISA and lateral flow immunoassays. MedRxiv 2020, 20066407 (2020).
VanBlargan, L. A., Goo, L. & Pierson, T. C. Deconstructing the antiviral neutralizing-antibody response: Implications for vaccine development and immunity. Microbiol. Mol. Biol. Rev. 80(4), 989–1010 (2016).
Wajnberg, A. et al. SARS-CoV-2 infection induces robust, neutralizing antibody responses that are stable for at least three months. MedRxiv 2020, 20151126 (2020).
Ju, B. et al. Human neutralizing antibodies elicited by SARS-CoV-2 infection. Nature 584(7819), 115–119 (2020).
Ni, L. et al. Detection of SARS-CoV-2-Specific humoral and cellular immunity in COVID-19 convalescent individuals. Immunity 52(6), 971-977.e3 (2020).
Shi, R. et al. A human neutralizing antibody targets the receptor-binding site of SARS-CoV-2. Nature 584(7819), 120–124 (2020).
Hyseni, I. et al. Characterisation of SARS-CoV-2 lentiviral pseudotypes and correlation between pseudotype-based neutralisation assays and live virus-based micro neutralisation assays. Viruses 12(9), 1011 (2020).
Wang, K. et al. Longitudinal dynamics of the neutralizing antibody response to severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection. Clin. Infect. Dis. 73, e531–e539 (2020).
Wang, Y. et al. Kinetics of viral load and antibody response in relation to COVID-19 severity. J. Clin. Investig. 130(10), 5235–5244 (2020).
Deeks, J. J. et al. Antibody tests for identification of current and past infection with SARS-CoV-2. Cochrane Database Syst. Rev. 6(6), 013652 (2020).
Padoan, A. et al. Evaluation of an ELISA for SARS-CoV-2 antibody testing: Clinical performances and correlation with plaque reduction neutralization titer. Clin. Chem. Lab. Med. 58(11), e247–e249 (2020).
Luchsinger, L. L. et al. Serological assays estimate highly variable SARS-CoV-2 neutralizing antibody activity in recovered COVID-19 patients. J. Clin. Microbiol. 58(12), e02005-e2020 (2020).
Staines, H. M. et al. IgG seroconversion and pathophysiology in severe acute respiratory syndrome coronavirus 2 infection. Emerg. Infect. Dis. 27(1), 85–91 (2021).
Patterson, E. I. et al. Methods of inactivation of SARS-CoV-2 for downstream biological assays. J. Infect. Dis. 222(9), 1462–1467 (2020).
Kohmer, N. et al. Brief clinical evaluation of six high-throughput SARS-CoV-2 IgG antibody assays. J. Clin. Virol. 129, 104480–104480 (2020).
Tang, M. S. et al. Association between SARS-CoV-2 neutralizing antibodies and commercial serological assays. Clin. Chem. 66(12), 1538–1547 (2020).
Perera, R. A. et al. Serological assays for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), March 2020. Eurosurveillance 25(16), 2000421 (2020).
Padoan, A. et al. Clinical performances of an ELISA for SARS-CoV-2 antibody assay and correlation with neutralization activity. Clin. Chim. Acta 510, 654–655 (2020).
Jungbauer, C. et al. Characterization of 100 sequential SARS-CoV-2 convalescent plasma donations. Transfusion.
Lau, E. H. Y. et al. Neutralizing antibody titres in SARS-CoV-2 infections. Nat. Commun. 12(1), 63 (2021).
Mazzini, L. et al. Comparative analyses of SARS-CoV-2 binding (IgG, IgM, IgA) and neutralizing antibodies from human serum samples. J. Immunol. Methods 489, 112937 (2021).
Funding
This work is supported by grants from DFID/Wellcome Trust Epidemic Preparedness coronavirus Grant (220764/Z/20/Z) to ERA and LEC and the Rosetrees Trust and the John Black Charitable Foundation (grant number M959) to HMS, TP, SK, DJC, JRAF. HMS is supported by the Wellcome Trust Institutional Strategic Support Fund (204809/Z/16/Z) awarded to St. George’s University of London. This work is also supported by the NIHR Health Protection Research Unit (HPRU) in Emerging and Zoonotic Infections at University of Liverpool in partnership with Public Health England (PHE), the Liverpool School of Tropical Medicine and the University of Oxford (Award 200907).
Author information
Authors and Affiliations
Contributions
Conceptualization: S.K., H.M.S., D.J.C., T.P., E.R.A., L.E.C. Methodology: S.K., H.M.S., D.J.C., T.P., E.R.A., L.E.C., G.L.H., E.I.P. Investigation: G.A.K., S.I.O., C.T.W., S.M., D.J.C., B.M.O.D., N.M.E., D.E.K., H.M.S. Formal Analysis: G.A.K., E.G. Writing—Original Draft: G.A.K., S.I.O., C.T.W., S.M. Writing—Review & Editing: All.
Corresponding authors
Ethics declarations
Competing interests
SK and HMS are shareholders and SK is advisor to QuantuMDx, a molecular nucleic acid test-based diagnostic company. SK is also member of the Scientific Advisory Committee for the Foundation for Innovative New Diagnostics (FIND), a not-for-profit organisation that produces global guidance on affordable diagnostics. GAK, SIO, EG, CTW, SM, DJC, LC, BMOD, NME, GLH, DEK, EIP, TP and ERA report no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kay, G.A., Owen, S.I., Giorgi, E. et al. SARS-CoV-2 enzyme-linked immunosorbent assays as proxies for plaque reduction neutralisation tests. Sci Rep 12, 3351 (2022). https://doi.org/10.1038/s41598-022-07263-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-07263-8
- Springer Nature Limited