Abstract
Mucinous carcinomas can arise in any organ with epithelial cells that produce mucus. While mucinous tumors from different organs are histologically similar, it remains to be elucidated whether they share molecular alterations. Here we analyzed a total of 902 patients across six cancer types by comparing mucinous and non-mucinous samples, integrating text mining of pathology reports, gene expression, methylation, mutational and copy-number profiling. We found that, in addition to genes involved in mucin processing and secretion, MUC2 up-regulation is a multi-cancer biomarker of mucinous histology and is regulated by DNA methylation in colorectal, breast and stomach cancer. The majority of carcinomas with mucinous differentiation had fewer DNA copy-number alterations than non-mucinous tumors. The tumor mutational burden was lower in breast and lung with mucinous differentiation compared to their non-mucinous counterparts. We found several differences in the frequency of oncogenic gene and pathway alterations between mucinous and non-mucinous carcinomas, including a lower frequency of p53 pathway alterations in colorectal and lung cancer, and a lower frequency of PI-3-Kinase/Akt pathway alterations in breast and stomach cancer with mucinous differentiation. This study shows that carcinomas with mucinous differentiation originating from different organs share transcriptomic and genomic similarities. These results might pave the way for a more biologically relevant taxonomy for these rare cancers.
Similar content being viewed by others
Introduction
Mucinous carcinomas are rare histological types of cancer characterized by mucin production that can arise in any epithelial tissue that produces mucus. Mucinous tumors mostly occur in the digestive tract, including the appendix, and the breast, but also in the lung, cervix, ovaries and pancreas. Previous molecular studies were focused on individual cancer types and were often limited by sample size and the lack of multi-omics data integration1,2,3,4,5,6,7. At the transcriptomic level, MUC2, the most abundant secreted mucin, has been previously shown to be up-regulated in different mucinous carcinomas1,8,9. We and others have previously reported a lower tumor mutation burden (TMB), lower fraction of genome altered (FGA) and lower frequency of PIK3CA mutation in mucinous breast cancer as compared to non-mucinous breast cancer10,11. Other genomic studies have shown a lower FGA and a lower frequency of TP53 mutations in mucinous colorectal carcinoma12,13. Despite their histological similarities, it is not known whether mucinous carcinomas from different cancer types share molecular features. Therefore, re-analysis of publicly available datasets that provides multi-omics data, represents a great opportunity to investigate the complex biology of mucinous tumors across multiple layers. Here, we investigated the molecular landscape of carcinomas with mucinous differentiation from six different cancer types by integrating text mining of pathology reports, gene expression, methylation and mutational profiling from The Cancer Genome Atlas (TCGA).
Methods
Data acquisition, patients and study design
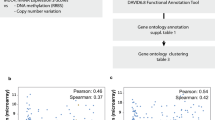
All data were obtained from the TCGA Pan-Cancer Atlas (PanCanAtlas)14,15 dataset available at (https://gdc.cancer.gov/about-data/publications/pancanatlas). All available pathology reports from TCGA were downloaded using GDC Data Transfer Tool (https://gdc.cancer.gov/access-data/gdc-data-transfer-tool). The curated genomic alteration matrices were obtained from Sanchez-Vega et al.15. We used a machine learning optical character recognition method using the tesseract R package to convert the scanned pathology reports into a text file and used ontology-based text mining to identify carcinomas with mucinous features. We manually inspected every pathology report that included the following keywords; “mucin”, “mucous” and “colloid”. Whenever possible, the Hematoxylin and eosin (H&E) slides corresponding to the identified mucinous cases (143/233) were manually reviewed by a trained pathologist (D.L.). Except for CEAD, which did not have sufficient controls, each mucinous case was matched to 3 non-mucinous counterpart controls (corresponding histotype) according to gender (except for BRCA and CESC, which were all female), age at diagnosis, date of diagnosis and pathological tumor-node-metastasis (pTNM) staging. We used additional matching rules such as breast cancer subtype (ER, PR and HER2 status) for BRCA, tobacco smoking history for LUAD, organ site (colon vs. rectum) and laterality (left vs. right) for CRC. The matching was performed using the nearest neighbor matching algorithm as implemented in the MatchIt R package16. A total of 902 samples were included, representing 285 colorectal adenocarcinoma (CRC), 95 colorectal mucinous adenocarcinoma (CRC-muc), 108 infiltrating ductal carcinoma breast cancer (BRCA), 36 mucinous carcinomas of the breast (BRCA-muc), 108 lung adenocarcinoma (LUAD), 36 mucinous (colloid) adenocarcinoma of the lung, 99 stomach adenocarcinoma (STAD), 33 mucinous adenocarcinoma of the stomach (STAD-muc), 24 endocervical adenocarcinoma (CEAD), 18 mucinous adenocarcinoma of endocervical type (CEAD-muc), 45 pancreas-adenocarcinoma ductal type (PAAD) and 15 pancreas-colloid (mucinous non-cystic) carcinoma (PAAD-muc).
Methylation analysis
We used data generated from Human Methylation 450 K arrays (Illumina, San Diego, CA). DNA methylation data was processed in R using the minfi package. Beta-values were normalized (preprocessQuantile) and the function dmpFinder (option; “shrinkVar = TRUE, type = categorical”), which use an F-test, was used to identify differentially methylated positions between carcinomas-muc and their non-mucinous counterparts within each cancer type. Differentially methylated CpGs were identified using a cut-off of FDR < 0.05. Methylation probes were mapped to genes using the illuminaHumanMethylation450kanno.ilmn12.hg19 Bioconductor package17.
Mucins gene expression signatures
As previously reported11, the metagene signature of gel-forming (“Gel-MUC”) and membrane-bound (“Membrane-MUC”) mucins was calculated by taking the mean RPKM expression level of (MUC2, MUC5B, MUC5AC, MUC6, MUC19) and (MUC1, MUC3A, MUC3B, MUC4, MUC12, MUC13, MUC14, MUC15, MUC16, MUC17, MUC20, MUC21, MUC22) respectively, scaled to a standard deviation of one and centered around zero.
Genomic analysis
For each cancer type, recurrent oncogenic alterations were defined as being oncogenic as per OncoKB annotation18 (version August 28, 2019) and present in at least 1%. Tumor mutation burden was calculated for each sample as the total number of non-synonymous mutations divided by the number of bases sequenced. Segmented copy number data were processed using CNtools package v1.4. Fraction of genome altered was calculated for each sample as the percentage of genome with log2 copy ratios > 0.2 or < − 0.2. Thresholds for copy number gain and loss were set at log2 copy ratios of > 0.2 and < − 0.2, respectively. The canonical oncogenic pathway level alterations were computed using the curated pathway templates provided in Sanchez-Vega et al.15.
Statistical analysis
Differences in clinicopathological characteristics between groups were analyzed using the χ2 test or the Fisher exact test when appropriate. All statistical tests comparing groups were done using the non-parametric Mann–Whitney U test and the χ2 test or the Fisher exact test when appropriate for continuous and categorical variables, respectively. All correlations were calculated using the non-parametric Spearman’s rho coefficient. Overall survival (OS) and progression-free interval (PFI) was obtained for each patient from the TCGA Pan-Cancer Clinical Data Resource19. Survival curves were analyzed using the Kaplan–Meier method and compared by the log-rank test. The prognostic impact of mucinous features on OS and PFI was evaluated using univariable and multivariable Cox proportional hazards regression models and expressed as hazard ratio (HR) with 95% confidence interval (95CI). Multivariable analyses were adjusted for standard clinical prognostic factors (age at diagnosis (dichotomized using the median), year of diagnosis (dichotomized using the median), pathological size, node involvement and metastasis status). All interaction and multivariable tests were performed using analysis of variance to compare the models with and without the extra term. Differential expression analysis was performed with DESeq2 v.1.14.1 R/Bioconductor package20 on raw count data. Significantly differentially expressed genes were selected with an FDR < 0.05. To estimate the probability of observing the 102 genes (N) that were consistently differentially expressed between carcinomas-muc and controls across cancer type, we used a Monte-Carlo simulation. We simulated 105 scenarios in which the differentially expressed genes for each cancer type were randomly permuted and intersected.
Finally, the empirical p-value was calculated by using the Monte-Carlo procedure21
where the r is the number of simulations that produced at least N genes and n is the number simulations.
Gene ontology enrichment analysis was restricted to biological process and performed using the topGO R package version 2.34.0 (Adrian A. and Jorg R.). Reported p-values were two-sided, and differences were considered significant when the p-value was less than 0.05. When applicable, multiple testing correction was performed using the false discovery rate method (FDR) (Benjamini and Hochberg, 1995), FDR below 0.05 was considered significant. All analyses were performed using R software version 3.5.2 (available at www.r-project.org) and Bioconductor version 3.8.
Results
Patients and survival
To identify carcinoma with mucinous differentiation, we used an ontology-based text mining approach on the available pathology reports (see “Methods”). A total of 902 patients (233 with mucinous features and 669 without mucinous features) representing six cancer types (95 colorectal adenocarcinomas; CRC, 36 invasive ductal breast carcinomas; BRCA, 36 lung adenocarcinomas; LUAD, 33 stomach adenocarcinomas; STAD, 18 cervical adenocarcinomas; CEAD and 15 pancreatic adenocarcinomas; PAAD) were included in this study (Supplementary Table 1). To reduce confounding factors, for each cancer type, each mucinous case was matched to three controls according to common cancer-type specific clinicopathological features (see “Methods”). Clinicopathological characteristics of carcinomas with mucinous differentiation (hereinafter referred to as “carcinomas-muc”) and non-mucinous carcinomas (hereinafter referred to as “controls”) are detailed in Supplement Table 1. We did not observe differences in overall survival (OS) or progression-free interval (PFI) within each cancer type, except for STAD-muc having a longer OS (median OS; 4.8 years vs 1.6 years, P = 0.03, adjusted hazard ratio, aHR = 0.4; 95% confidence interval, 95CI; 0.18–0.9, P = 0.015), and CEAD-muc having a median PFI of 3.1 vs. the median PFI was not reached in CEAD (P = 0.03) (aHR = 5.5, 95CI; 0.96–31.2, P = 0.04) (Supplementary Table 2 and Supplementary Fig. S1).
Transcriptomic similarities across carcinomas with mucinous differentiation from different cancer types
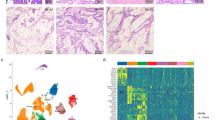
To better understand the intrinsic biology of carcinomas-muc, we performed a differential gene expression analysis between carcinomas-muc and their respective controls. For each cancer type, we identified a list of statistically significantly differentially expressed genes (7046, 4826, 4556, 2594, 2366, 478, respectively for CRC, BRCA, LUAD, STAD, CEAD and PAAD, Supplementary Table 3). The intersection of these lists revealed that three genes (MUC2, CRACR2A, SEC16A, Fig. 1a) were significantly up-regulated in carcinomas-muc of all tested cancer types compared to controls. We previously reported two different mucin gene expression signatures, composed of secreted gel-forming mucins and membrane-bound mucins, and observed that only the gel-forming signature was elevated in mucinous breast cancer11. Here, we observed that in every cancer type except PAAD, the secreted gel-forming mucins signature was statistically significantly upregulated in mucinous carcinoma compared to controls (Fig. 1a). Of note, the membrane-bound mucins signature was significantly up-regulated in CRC-muc, LUAD-muc and STAD-muc but downregulated in BRCA-muc. When restricting the analysis to the four largest cohorts (CRC, BRCA, LUAD and STAD), a total of 102 genes were consistently differentially expressed between carcinomas-muc and controls (Fig. 1b), a number that is statistically significantly higher than expected by chance (P < 0.001, based on Monte-Carlo simulations). MUC5B, another gel-forming mucin, was identified to be up-regulated in all four cancer types (Supplementary Fig. S2A). We previously demonstrated that expression of MUC2 in mucinous breast cancer was regulated by DNA methylation10,11. We interrogated if a similar mechanism was present in other carcinomas-muc and found that the expression of MUC2 was regulated by DNA methylation in CRC-muc, BRCA-muc and STAD-muc (Supplementary Fig. S2B,C and Supplementary Table 4). Gene ontology enrichment analysis using the 102 previously identified genes revealed enrichment of processes related to cellular divalent cation homeostasis and calcium (Ca++) transmembrane transport (Fig. 1c and Supplementary Fig. S2D). Taken together, these results support the notion that carcinomas with mucinous differentiation from different cancer types share transcriptomic similarities.
Transcriptomic similarities across carcinomas with mucinous differentiation from different cancer types. (a) Comparison of MUC2, SEC16A, CRACR2A, the gel-forming mucins signature (Gel-muc) and the membrane-bound mucins signature (Membrane-muc) between carcinomas-muc and controls for each cancer type (*; FDR < 0.05). Statistical significance was measured using the pairwise Mann–Whitney U tests adjusted for multiple comparisons. (b) Number of shared and unique differentially expressed genes between carcinoma-muc and control across each cancer type. (c) Top 20 enriched gene ontology terms associated with the 102 common differentially expressed genes. CRC colorectal adenocarcinoma, CRC-muc colorectal mucinous adenocarcinoma, BRCA infiltrating ductal carcinoma breast cancer, BRCA-muc mucinous carcinomas of the breast, LUAD lung adenocarcinoma, LUAD-muc mucinous (colloid) adenocarcinoma of the lung, STAD stomach adenocarcinoma, STAD-muc mucinous adenocarcinoma of the stomach, CEAD endocervical adenocarcinoma, CEAD-muc mucinous adenocarcinoma of endocervical type, PAAD pancreas-adenocarcinoma ductal type, PAAD-muc pancreas-colloid (mucinous non-cystic) carcinoma.
The unique genomic landscape of carcinomas with mucinous differentiation across different cancer types
To chart the somatic genomic landscape of carcinomas with mucinous differentiation, we systematically profiled broad genomic features, recurrent oncogenic alterations and pathway alterations in carcinomas-muc and controls. As previously shown1, we observed that CRC-muc is enriched for microsatellite unstable (MSI-H) tumors (26/73 (35.6%) vs. 27/215 (12.6%), P < 0.001). To reduce the heterogeneity of the cohort, we chose to exclude MSI-H cases (N = 75) in subsequent genomic analyses. When interrogating broad genomic features, we found that tumor mutational burden (TMB) was significantly lower in BRCA-muc and LUAD-muc and that fraction of genome altered (FGA), a measure for the degree of copy-number changes, was significantly lower in CRC-muc, BRCA-muc, LUAD-muc and STAD-muc than in controls (Fig. 2a). We also compared the number of driver alterations between carcinomas-muc and their non-mucinous counterparts within each cancer type. BRCA-muc and LUAD-muc had a significantly lower number of driver alterations, which is consistent with the lower TMB and FGA observations. Intriguingly, CRC-muc had a slightly higher number of driver alterations than CRC (Fig. 2a). Next, we compared the frequency of genome-wide copy number alterations (CNA) between carcinomas-muc and controls in CRC, BRCA, LUAD and STAD. Compared to their control counterparts, carcinomas-muc had a lower frequency of CNA, evenly distributed across the genome (Fig. 2b). Then, for each cohort, we identified recurrent oncogenic alterations and compared their frequencies in controls and carcinomas-muc. Compared to their control counterparts, we observed a lower frequency of TP53 mutations in CRC-muc and LUAD-muc (12/48 (25%) vs. 146/189 (77.2%), FDR < 0.001 and 5/35 (14.3%) vs. 49/101 (48.5%), FDR = 0.02, respectively), a higher frequency of SMAD4 and SMAD2 mutations in CRC-muc (17/48 (35.4%) vs. 14/189 (7.4%), FDR = 0.0001 and 8/48 (16.7%) vs. 2/189 (1.1%), FDR = 0.001, respectively) and a lower frequency of PIK3CA mutations in BRCA-muc (3/32 (9.4%) vs. 45/98 (45.9%), FDR = 0.006). Of note, we detected a lower frequency of TP53 mutation in BRCA-muc and PAAD-muc, a higher frequency of KRAS mutation in CRC-muc and LUAD-muc and a higher frequency of methylation silencing of TCF7 in CRC-muc and BRCA-muc and TCF7L2 and CDKN1B in BRCA-muc, although these observations did not reach statistical significance after correcting for multiple testing (Fig. 2c). Next, we compared the frequency of ten canonical oncogenic pathways commonly altered in cancer (RTK-RAS, p53, cell cycle, β-catenin/Wnt, PI-3-Kinase/Akt, Notch, TGFβ signaling, Myc, Hippo and Nrf2, as described in15) in controls and carcinomas-muc. In CRC-muc, we observed a lower frequency of p53 pathway alteration and a higher frequency of TGFβ signaling and RTK-RAS pathways alterations (19/48 (39.6%) vs. 153/189 (81%) FDR < 0.001, 28/48 (58.3%) vs. 35/189 (18.5%) FDR < 0.001, 43/48 (89.6%) vs. 140/189, FDR = 0.07). In BRCA-muc, we observed a lower frequency of PI-3-Kinase/Akt pathway alterations and a higher frequency of β-catenin/Wnt pathway alterations (8/32 (25%) vs. 58/98 (59.1%), P = 0.001, FDR = 0.01 and 11/32 (34.4%) vs. 13/98 (13.3%), P = 0.02, FDR = 0.08). In LUAD-muc, we observed a lower frequency of p53 and TGFβ signaling pathways alterations (8/35 (22.9%) vs. 63/101 (62.4%), P < 0.001, FDR < 0.001 and 0/35 (0%) vs. 12/101 (11.8%), P = 0.03, FDR = 0.18). Finally, in STAD-muc, we observed a lower frequency of PI-3-Kinase/Akt pathway alterations (2/22 (9%) vs. 28/69 (40.6%), P = 0.008, FDR = 0.07) (Fig. 2d). These data indicate that the genomic landscape of carcinomas-muc is different from that of their control counterparts. Carcinomas with mucinous differentiation, however, share genomic similarities across different cancer types, such as quieter copy-number alteration profiles in CRC-muc, BRCA-muc, LUAD-muc and STAD-muc, a lower frequency of p53 pathway alterations in CRC-muc and LUAD-muc and a lower frequency of PIK3CA pathway alterations in BRCA-muc and STAD-muc.
The unique genomic landscape of carcinomas with mucinous differentiation from different cancer types. (a) Comparison of TMB (left), FGA (middle) and the number of driver alterations (right) between carcinoma-muc and control for each cancer type (*; FDR < 0.05). Statistical significance was measured using the pairwise Mann–Whitney U tests adjusted for multiple comparisons. (b) Comparison of the CNAs frequencies between carcinoma-muc (colored) and control (grey) for each cancer type. (c) Comparison of the frequency of recurrent oncogenic alteration present in each cancer type. Statistical significance was measured using the Fisher’s exact test adjusted for multiple comparisons. (d) Percentages of patients with alterations in canonical oncogenic signaling pathways between carcinoma-muc (colored) and control (grey) for each cancer type (*P < 0.05, **FDR < 0.05). Note that the NRF2 pathway was not altered in PAAD. Statistical significance was measured using the Fisher’s exact test adjusted for multiple comparisons. TMB tumor mutation burden, FGA fraction of genome altered, CRC colorectal adenocarcinoma, CRC-muc colorectal mucinous adenocarcinoma, BRCA infiltrating ductal carcinoma breast cancer, BRCA-muc mucinous carcinomas of the breast, LUAD lung adenocarcinoma, LUAD-muc mucinous (colloid) adenocarcinoma of the lung, STAD stomach adenocarcinoma, STAD-muc mucinous adenocarcinoma of the stomach, CEAD endocervical adenocarcinoma, CEAD-muc mucinous adenocarcinoma of endocervical type, PAAD pancreas-adenocarcinoma ductal type, PAAD-muc pancreas-colloid (mucinous non-cystic) carcinoma.
Discussion
To our knowledge, this is the first systematic attempt to characterize carcinomas with mucinous features across different cancer types. In this study, we have demonstrated that the up-regulation of MUC2, SEC16A and CRACR2A is the common denominator of all carcinomas with mucinous differentiation studied here. The up-regulation of MUC2, the most abundant secreted mucin, has been previously shown in different cancer types1,8,9. Several studies have reported a possible role of DNA methylation in regulating the expression of some gel-forming mucins22 and the epigenetic regulation of MUC2 has been shown in mucinous colorectal cancer and mucinous breast cancer10,11,23. We demonstrated that MUC2 expression is also regulated by DNA methylation in stomach cancer. The fact that we did not find an association between MUC2 expression and DNA methylation in LUAD, CEAD and PAAD might be due to low statistical power or other mechanisms of mRNA expression regulation such as histone methylation24. Nevertheless, our results suggest that up-regulation of MUC2 and the gel-forming mucins signature might be considered as pan-cancer biomarkers of mucinous histology. SEC16A encodes a protein required for protein trafficking to the cell membrane25. We hypothesize that carcinomas-muc up-regulates SEC16A to sustain mucin secretion. Differential gene expression analysis revealed up-regulation of genes related to divalent cation homeostasis and Ca++ transmembrane transport such as CRACR2A, encoding a rab GTPase Ca++ binding protein and acting as a cytoplasmic calcium-sensor. Interestingly, it has been shown that packing and releasing of gel-forming MUC2 is a pH-dependent mechanism that relies on Ca++ release regulation26. We show that, independent of cancer type, carcinomas with mucinous differentiation not only display elevated expression of MUC2 but also complementary genes involved in MUC2 packing, folding and transport.
At the genomic level, we found that a flat CNA landscape was a common trait across CRC-muc, BRCA-muc, LUAD-muc and STAD-muc. Thus, we hypothesize that carcinomas-muc are less likely driven by copy number alterations. Higher frequencies of SMAD4, BRAF and KRAS mutations have been previously reported in CRC-muc1,2,27. Here, we confirmed these observations and found that CRC-muc was also associated with a higher frequency of SMAD2 and lower frequency of TP53 mutations. The slightly higher median number of driver alterations observed in CRC-muc might be explained by the higher frequency of mutation in genes belonging to TGF-Beta and RTK-RAS pathways in CRC-muc. We and others have previously reported a lower TMB, FGA and lower frequency of PIK3CA mutation in mucinous breast cancer10,11. Here we found that BRCA-muc were associated with a higher frequency of β-catenin/Wnt pathway alterations, mostly caused by epigenetic silencing of TCF7L2 and TCF7. We have also observed a higher frequency of KRAS mutations and lower frequency of TP53 mutations in LUAD-muc, consistent with previous findings3,28.
Our study has several limitations. First, the definition of mucinous carcinomas varies by cancer types and includes pure as well as mixed mucinous carcinomas, but due to the pan-cancer nature of this study and for practical reasons, we choose to include any carcinoma with mucinous differentiation. Of note, not all HE slides were available to be centrally reviewed. It should also be noted that the sample size was relatively low for CEAD and PAAD hence negative conclusions related to these cohorts should be interpreted with caution, given the limited statistical power. Finally, only samples that were part of the TCGA were analyzed and future studies that include ovarian and appendiceal cancer are needed to investigate if their biology are similar to other mucinous carcinomas.
In conclusion, here we provide important novel insights into the biology of carcinomas with mucinous differentiation suggesting transcriptional and genomic similarities across different cancer types and alternative oncogenic pathway alterations underlying their pathogenesis, paving the way for a more biologically relevant taxonomy for these cancers.
References
Hugen, N. et al. The molecular background of mucinous carcinoma beyond MUC2. J. Pathol. Clin. Res. 1, 3–17 (2015).
Fleming, N. I. et al. SMAD2, SMAD3 and SMAD4 mutations in colorectal cancer. Cancer Res. 73, 725–735 (2013).
Kadota, K. et al. Associations between mutations and histologic patterns of mucin in lung adenocarcinoma: Invasive mucinous pattern and extracellular mucin are associated with KRAS mutation. Am. J. Surg. Pathol. 38, 1118–1127 (2014).
Kehr, E. L. et al. Mucinous breast carcinomas lack PIK3CA and AKT1 mutations. Hum. Pathol. 43, 2207–2212 (2012).
Lacroix-Triki, M. et al. Mucinous carcinoma of the breast is genomically distinct from invasive ductal carcinomas of no special type. J. Pathol. 222, 282–298 (2010).
He, M. et al. KRAS mutations in mucinous lesions of the uterus. Am. J. Clin. Pathol. 143, 778–784 (2015).
Dal Molin, M. et al. Clinicopathological correlates of activating GNAS mutations in intraductal papillary mucinous neoplasm (IPMN) of the pancreas. Ann. Surg. Oncol. 20, 3802–3808 (2013).
İlhan, Ö., Han, Ü., Önal, B. & Çelık, S. Y. Prognostic significance of MUC1, MUC2 and MUC5AC expressions in gastric carcinoma. Turk. J. Gastroenterol. 21, 345–352 (2010).
Hanski, C. et al. Overexpression or ectopic expression of MUC2 is the common property of mucinous carcinomas of the colon, pancreas, breast, and ovary. J. Pathol. 182, 385–391 (1997).
Pareja, F. et al. The genomic landscape of mucinous breast cancer. JNCI J. Natl. Cancer Inst. 111, 1–5 (2019).
Nguyen, B. et al. Genomic, transcriptomic, epigenetic, and immune profiling of mucinous breast cancer. JNCI J. Natl. Cancer Inst. https://doi.org/10.1093/jnci/djz023 (2019).
Hugen, N. et al. Reduced rate of copy number aberrations in mucinous colorectal carcinoma. Oncotarget 6, 25715–25725 (2015).
Hu, Y. et al. Dynamics of genome alterations in Crohn’s disease-associated colorectal carcinogenesis. Clin. Cancer Res. 24, 4997–5011 (2018).
Hoadley, K. A. et al. Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell 173, 291-304.e6 (2018).
Sanchez-Vega, F. et al. Oncogenic signaling pathways in the cancer genome atlas. Cell 173, 321–337 (2018).
Imai, K., King, G., Stuart, E. A. & Ho, D. E. MatchIt: Nonparametric preprocessing for parametric causal inference. J. Stat. Softw. 42, 1–43 (2011).
Hansen, K. D. IlluminaHumanMethylation450kanno.ilmn12.hg19: Annotation for Illumina’s 450k methylation arrays. J. Stat. Softw. https://doi.org/10.18129/B9.bioc.IlluminaHumanMethylation450kanno.ilmn12.hg19 (2016).
Chakravarty, D., Gao, J., Phillips, S. & Kundra, R. OncoKB: A precision oncology knowledge base. Precis. Oncol. https://doi.org/10.1200/PO.17.00011 (2017).
Liu, J. et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173, 400-416.e11 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
North, B. V., Curtis, D. & Sham, P. C. A note on the calculation of empirical P values from Monte Carlo procedures. Am. J. Hum. Genet. 71, 439–441 (2002).
Yamada, N. et al. Epigenetic regulation of mucin genes in human cancers. Clin. Epigenet. 2, 85–96 (2011).
Kim, K. et al. MUC2 gene promoter methylation in mucinous and non-mucinous colorectal cancer tissues. Int. J. Oncol. 36, 765–775 (2010).
Vincent, A. et al. Epigenetic regulation (DNA methylation, histone modifications) of the 11p15 mucin genes (MUC2, MUC5AC, MUC5B, MUC6) in epithelial cancer cells. Oncogene 26, 6566–6576 (2007).
Watson, P., Townley, A. K., Koka, P., Palmer, K. J. & Stephens, D. J. Sec16 defines endoplasmic reticulum exit sites and is required for secretory cargo export in mammalian cells. Traffic 7, 1678–1687 (2006).
Ambort, D. et al. Calcium and pH-dependent packing and release of the gel-forming MUC2 mucin. Proc. Natl. Acad. Sci. 109, 5645–5650 (2012).
Hugen, N., Brown, G., Glynne-Jones, R., De Wilt, J. H. W. & Nagtegaal, I. D. Advances in the care of patients with mucinous colorectal cancer. Nat. Rev. Clin. Oncol. 13, 361–369 (2016).
Shim, H. S. et al. Unique genetic and survival characteristics of invasive mucinous adenocarcinoma of the lung. J. Thorac. Oncol. 10, 1156–1162 (2015).
Funding
This work was supported by the Marie-Josée and Henry R. Kravis Center for Molecular Oncology, a National Cancer Institute Cancer Center Core Grant (P30-CA008748), the Thompson Family Foundation, the Robertson Foundation (N.S.), the Breast Cancer Research Foundation (J.S.R.-F.) and the Congressionally Directed Medical Research Program W81XWH-15-1-0547 (J.S.R.-F.).
Author information
Authors and Affiliations
Contributions
B.N., N.S., C.D., J.R.F., B.W., C.S. designed the study. B.N. performed the analysis and wrote the manuscript. B.N., F.S.V. performed computational and statistical analyses. F.S.V., C.F., W.K.C., A.M.B., F.P. performed the data preprocessing. D.L., reviewed the pathology reports and HE slides. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
B.N., F.S.V., W.K.C., A.M.B., F.P., C.D., B.W., N.S., C.F., D.L.: no disclosures. J.S.R.-F. reports receiving personal/consultancy fees from VolitionRx, Paige.AI, Goldman Sachs, REPARE Therapeutics, GRAIL, Ventana Medical Systems, Roche, Genentech and InVicro outside of the scope of the submitted work. C.S. has patents on gene expression and methylation signatures. He has served on advisory boards, and/or speaker at meetings, and/or recipient of travel support for participation in medical meetings for/from (in alphabetical order) from Amgen, Astellas, AstraZeneca, Bayer, Celgene, Nanostring Technologies, Novartis, Pfizer, Roche.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nguyen, B., Sanchez-Vega, F., Fong, C.J. et al. The genomic landscape of carcinomas with mucinous differentiation. Sci Rep 11, 9478 (2021). https://doi.org/10.1038/s41598-021-89099-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-89099-2
- Springer Nature Limited
This article is cited by
-
Integrated clinical and genomic analysis identifies driver events and molecular evolution of colitis-associated cancers
Nature Communications (2023)
-
Invasive lobular carcinoma with extracellular mucin (ILCEM): clinicopathologic and molecular characterization of a rare entity
Modern Pathology (2022)
-
Clinicopathologic and genomic characteristics of mucinous gastric adenocarcinoma
Gastric Cancer (2022)