Abstract
Rice (Oryza sativa L.), a major dietary source, is often cultivated in soils poor in available inorganic orthophosphate (Pi), which is a key nutrient for growth and development. Poor soils are amended by phosphorus (P) fertilizer, which is derived from the non-renewable rock phosphate reserves. Therefore, there is a need for developing rice varieties with high productivity under low P conditions. At the ICAR-IIRR, ethyl methanesulfonate (EMS) mutagenized rice genotype Nagina22 (N22) were screened for high grain yield in Pi-deprived soil, which led to the identification of ~ 10 gain-of-function mutants including NH787. Here, detailed comparative morphophysiological, biochemical, and molecular analyses of N22 and NH787 were carried out in hydroponics and potting soil under different Pi regimes. Under Pi-deprived condition, compared with N22, NH787 exhibited higher root and vegetative biomass, the number of tillers, and grain yield. The augmented agronomic traits of NH787 were corroborated with significantly higher photosynthetic rate, pollen fertility, stigma receptivity, and the activities of antioxidant enzymes superoxide dismutase (SOD) and catalase (CAT). Further, several genes involved in the maintenance of Pi homeostasis (GPH) were differentially regulated. The study thus revealed a wide-spectrum influence of the mutation in NH787 that contributed towards its higher Pi use efficiency (PUE).
Similar content being viewed by others
Introduction
Rice (Oryza sativa L.) is the staple food and a major source of dietary energy supply for more than half of the world’s 7.85 billion population (www.worldometers.info/world-population). Rice is consumed ∼ 90% in Asia (www.irri.org/rice-today). China is the world's biggest rice producer among the top 20 rice-producing countries in the world of which 70% are from Asia (Fig. 1A; Table 1). India is the second-largest producer and consumer of rice with ~ 44 million hectares under cultivation and West Bengal, Punjab, and Uttar Pradesh are the top three states in rice production1 (Fig. 1B; www.agriexchange.apeda.gov.in). The world population is projected to reach 9.7 billion by 2050 (www.populationpyramid.net/2050) and the percent increase in population in some of the rice-producing Asian countries ranges from 1.80% (Nepal), 15.81% (India) to 34.65% (Pakistan) (Table 1). Therefore, scaling up rice production to achieve sustainable food security for the burgeoning population is warranted.
Rice producing (metric tons) (A) top 20 countries (www.worldagriculturalproduction.com/crops/rice.aspx) and (B) top 10 states in India (www.mapsofindia.com/top-ten/india-crops/rice.html). (c) Available soil P content (high, medium, and low) in top 10 states in India producing rice (www.iiss.nic.in/districtmap.html). Photoshop 7.0 version was used to prepare the figure A–C.
Phosphorus (P), one of the essential macroelements, is a building block of various organic molecules such as ATP, nucleic acids, and phospholipids, and also plays a key role in energy transfer, signal transduction, metabolic pathways, and thus indispensable for the proper growth and development of plants2,3,4,5,6. In the rhizosphere, P is largely available in the form of inorganic orthophosphate (Pi) and its acquisition by the roots and subsequent translocation to various parts of the plants is mediated by a suite of Pi transporters7,8,9. However, rice is often cultivated in a rain-fed system on soils subjected to various abiotic stresses including poor availability and/or fixing of P, which adversely affects yield potential10,11. Rice in India is normally produced in soils poor in Pi availability and largely amended by application of P fertilizer12,13 (Fig. 1C). P fertilizer is produced from the non-renewable and finite rock phosphate (phosphorite) reserves likely to be exhausted in the next 50–100 years at the current rate of its usage across the globe14. Therefore, there is an urgent need to identify or develop rice varieties with higher PUE under low P conditions15,16.
Sequencing of the whole rice genome and an efficient transformation system has made it a favored model monocotyledonous plant17,18,19. The arduous task of the post-genomic era has been to systematically evaluate the function of an array of diverse GPH in rice. Loss-of-function mutagenesis (T-DNA and Tos17)-mediated reverse genetics has significantly contributed to the functional genomics of rice20,21. RNAi-mediated gene-silencing has also been an attractive approach for functional genomics22. A programmable CRISPR/Cas9 system emerged as a promising molecular tool for genome editing23and Jennifer Doudna and Emmanuelle Charpentier were awarded the 2020 Nobel prize in Chemistry for developing this versatile technology. CRISPR/Cas9 system is now a favored technology for generating transgene-free rice plants24,25,26. A gain-of-function mutagenesis is an alternative approach based on the ectopic overexpression of transgenes under the control of a strong constitutive CaMV35S or ubiquitin promoter27. Functional characterization of several GPH by reverse and/or forward genetic approach has thus led to the identification of several key positive and negative regulators of sensing and signaling cascades governing the maintenance of Pi homeostasis3,6,28 (Table 2). However, plants generated by these forward and reverse genetics approaches are often deemed as a potential transgene and are regulated by stringent country-specific ethical legislations, and often fail to comply with the biosafety regulations29,30,31. Although CRISPR-edited rice was considered to comply with the regulatory approval for commercial applications24, recently Court of Justice of the European Union has clubbed them with GM plants32,33,34. One of the classical controversial cases is the Golden rice, which was engineered to produce seeds enriched with β-carotene to mitigate vitamin A deficiency in the millions of poor people35 but has been embroiled in polarized debate over its ethicality36. On the contrary, mutation breeding by exposure to mutagens such as EMS or irradiation by X-rays is environmentally benign, has good safety records, and is not regulated worldwide29. EMS-induced mutagenesis is an attractive strategy for inducing genetic variations in the genome37,38 and has facilitated the development of a rich repository of rice mutants that exhibit tolerance to different biotic and/or abiotic stresses39. N22 is an upland and short aus genotype and is tolerant to heat and drought40,41. An initiative was launched by the Department of Biotechnology, Govt. of India, for generating EMS-mutagenized M2 populations (~ 85,000) in the background of N2241. At the ICAR-IIRR, efforts are underway for more than a decade to screen N22 EMS mutants that exhibited altered PUE under field condition, which led to the identification of several loss-of-function and gain-of-function mutants42,43,44,45,46,47. Among these mutants, detailed morphophysiological and molecular analyses were carried out for the loss-of-function mutant NH101, which revealed several traits that were affected contributing towards its lower PUE compared with N2247.
However, gain-of-function N22 mutant that shows significantly higher PUE has not been characterized as yet. Therefore, in this study, detailed comparative morphophysiological, biochemical, and molecular analyses of N22 and NH787 mutant were carried out in the hydroponics and potting soil under different Pi regimes. The analysis revealed several traits that contributed towards the higher PUE of the gain-of-function NH787 mutant.
Materials and methods
Plant materials and experimental conditions
Rice (Oryza sativa L. ssp indica) genotype N22 were mutagenized with EMS and several gain-of-function mutants i.e., NH363, NH514, NH686, NH719, NH776, and NH787 were identified, which exhibited high grain yield in Pi-deprived soil under field condition compared with N2245. From these gain-of-function mutants, NH787 was selected for detailed morphophysiological, biochemical, and molecular analyses. About 15 seeds each of N22 and NH787 were placed equidistant on a Pteri plate lined with germination paper soaked with deionized distilled water and wrapped in aluminum foil and kept for germination in a growth chamber (28–30 °C) for 4 days. N22 and NH787 seedlings showed significant variation in their radicle length. Therefore, the seedlings were scanned and their radicle was measured by employing the ImageJ program48 and only those in the range of 2.0–3.0 cm were selected and transferred to the hydroponic system as described49 containing P+ (0.3 mM NaH2PO4) and P− (0 mM NaH2PO4) media as described50 for 7 days. For the potting soil experiment, N22 and NH787 were grown initially under normal soil condition for 14 days. Subsequently, the seedlings were transplanted in earthen pots which were filled with 8 kg of normal soil (P+) and low P soil (P−) with the Olsen P values of 24 kg/ha and 1.8 kg/ha for P+ and P− soil, respectively. P+ and P− soils were fertilized as described47. All plant experiments were carried out in accordance with the guidelines and permission was obtained to collect the rice seeds.
Quantitative analysis of the root traits
Seedlings grown in the hydroponic system were removed along with the mesh after 7 days treatment under P+ and P− conditions and placed in an inverted position in a Petri plate containing a pool of water. Under the stereomicroscope, roots were separated at the shoot: hypocotyl junction and transferred to a Petri plate containing 1% (w/v) agar. Adventitious, seminal, and lateral roots were spread gently with a camel hair brush to reveal the root system architecture (RSA). Spread-out roots were then scanned at 1000 dpi using a desktop scanner. Scanned images were then used for documenting the number and length of different root traits by using the ImageJ program48.
Quantitative analysis of agronomic traits
Plants grown in the potting soil (P+ and P−) up to maturity were quantitatively analyzed for the growth performance, biomass and length of root, vegetative biomass, number of tillers, filled spikelets/panicle, unfilled spikelets/panicle, 100-grain weight, and yield as described47.
Quantitative analysis of physiological traits
Plants were grown in the potting soil (P+ and P−) up to 50% flowering and flag leaf was assayed for photosynthetic rate (PN), stomatal conductance (gs), intercellular CO2 concentration (Ci), and transpiration rate (E) by using portable photosynthesis system LI-6400XT (LI-COR Biosciences, USA) set at 1200 μmol m–2 s–1 photosynthetically active radiation (PAR) and 387 ± 6 ppm CO2 concentration. Coefficient of photochemical quenching (qP), coefficient of non-photochemical quenching (qN), electron transport rate (ETR), and maximum efficiency of PSII photochemistry (Fv/Fm) were quantified by employing portable chlorophyll fluorometer PAM-2100 (Heinz Walz GmbH, Germany). Chlorophyll a, b, and carotenoids were extracted and their concentrations were quantified as described51, 52.
Quantification of soluble Pi
Harvested root and shoot were rinsed thoroughly 4–5 times with deionized distilled water, blotted-dry, frozen in liquid nitrogen, ground to a fine powder, and stored at – 80 °C till further use. Ground tissue (~ 25 mg) was homogenized in 200 μl of 1% (v/v) glacial acetic acid, vortexed, and centrifuged at 10,000 rpm for 10 min to remove the debris. The supernatant was collected for the quantification of soluble Pi by phosphomolybdate colorimetric assay as described53.
Quantification of APase enzyme activity
APase enzyme activity was quantified as described54 with minor modifications. Freshly harvested root and shoot tissues (~ 0.1 g) were ground in a chilled citrate buffer (0.1 M, pH 5.2), centrifuged at 12,000 rpm at 4 °C for 15 min, and the supernatant was assayed for APase enzyme activity. The reaction mixture comprised 0.1 ml supernatant, 0.4 ml chilled citrate buffer (0.1 M, pH 5.2), and 0.5 ml p-nitrophenol (pNP) (10 mM, pH 5.2). The reaction mixture was incubated at room temperature for 10 min and the reaction was then terminated by adding 2 ml of Na2CO3 (0.2 M). The standard curve was prepared with the known concentrations of pNP and APase enzyme activity was computed by estimating the accumulation of pNP at 405 nm.
Quantification of antioxidant enzyme activities
Freshly harvested root and shoot tissues (~ 0.1 g) were ground in phosphate buffer (0.1 M, pH 7.5) containing EDTA (0.5 mM) and centrifuged at 12,000 rpm at 4 °C for 15 min. The supernatant was collected for assaying the activities of different antioxidant enzymes. Superoxide dismutase (SOD) was assayed as described55. The reaction mixture (1.5 ml phosphate buffer [100 mM, pH 7.8], 0.2 ml methionine [200 mM], and 0.1 ml each of the plant extract, Na2CO3 [1.5 M], EDTA [3.0 mM], NBT [2.25 mM], and riboflavin [60 μM]) was incubated under a fluorescent lamp (15 W) for 15 min. SOD activity was determined by a 50% decrease in the absorbance at 560 nm due to rapid inhibition of O2− with NBT. Peroxidase (POD) activity was assayed as described56. The reaction mixture comprised 1.0 ml phosphate buffer (100 mM, pH 6.1), 0.5 ml each of guaiacol (96 mM), H2O2 (12 mM), and 0.1 ml of the enzyme extract. The absorbance was taken at 470 nm at different time intervals (0, 1, 2, and 3 min). Catalase (CAT) was assayed as described57. The reaction mixture comprised 1.5 ml phosphate buffer (100 mM, pH 7.0), 0.5 ml H2O2 (75 mM), and 0.05 ml of the enzyme extract. A temporal disappearance of H2O2 was recorded at an interval of 30 s for 2 min at 240 nm. Ascorbate peroxidase (APX) activity was assayed as described58. The root and shoot tissues were ground in a solution containing 1.5 ml phosphate buffer (100 mM, pH 7.0) containing ascorbic acid (1 mM), and EDTA (0.5 mM). The solution was centrifuged at 12,000 rpm at 4 °C for 20 min and the supernatant was collected for the assay. The reaction mixture comprised 1.5 ml phosphate buffer (100 mM, pH 7.0), 0.1 ml each of EDTA (3.0 mM), H2O2 (3.0 mM), 0.5 ml ascorbic acid (3 mM), and 0.05 ml of the enzyme extract. The APX activity was measured by monitoring the gradual decrease in the absorbance value at an interval of 30 s for 2 min at 290 nm.
Quantification of H2O2 content
H2O2 content was estimated as described59. Freshly harvested root and shoot tissues (~ 0.5 g) were ground in 10 ml of trichloroacetic acid, centrifuged at 12,000 rpm at 4 °C for 15 min, and the supernatant was collected for the assay. The reaction mixture comprised 0.5 ml of phosphate buffer (10 mM, pH 7.0), 2 ml of KI (1 M), and 0.5 ml of the supernatant. The reaction mixture was vortexed for 1 min, incubated in dark for 30 min, and H2O2 content was quantified at 390 nm.
Assay for pollen viability
The anthers from the spikelets were collected just before anthesis, crushed in Lugol's (I2-KI) solution, and observed under a light stereomicroscope as described60. Sterile and fertile pollens were unstained and stained, respectively and their images were captured using a compound microscope (10X).
Quality traits
Harvested grains were threshed, cleaned, and dried at 45 °C for 3 days to achieve identical moisture content. Grains (~ 25 g) were dehulled using a sheller (Satake Co. Ltd. Japan). The hulling rate was computed as described61. Brown rice was milled by employing Pearlest grain polisher (Kett, USA) and the milling rate was calculated as described61. The head rice recovery was calculated by weighing polished rice and separating head rice (≥ ¾ length of the brown rice) manually from the broken fractions. Gel consistency (GC) was computed as described62. Gelatinization temperature was calculated based on the alkali spread score of the milled rice as described61. Amylose content was estimated from the ground rice flour calorimetrically as described63. Length, width, and area of grains were measured by using the ImageJ program48.
qRT-PCR analysis
Total RNA (~ 2 μg) was isolated from the ground tissue using Trizol reagent and treated with RNase-free DNase. First-strand cDNA was synthesized by using oligo (dT)-18 primer and Superscript II Reverse Transcriptase (Invitrogen). OsActin (LOC_Os03g50885) was used as an internal control. The qRT-PCR analysis was performed in triplicate using SYBR Premix Ex TaqII (TaKaRa) in a StepOnePlus Real-time PCR system (Applied Biosystems). Relative expression levels of the genes were computed by the 2–ΔΔCT method of relative quantification64. Gene-specific primers are listed in Supplementary Table S1.
Statistical analysis
Two-way analysis of variance (ANOVA) was performed using open-source software R65 with agricolae package. Statistical significance of the parameter means was determined by performing Fisher's LSD test.
Ethical approval
The authors declare that the experiments comply with the current laws of the country in which they were performed and in compliance with ethical standards.
Results and discussion
Selection of the uniformly grown seedlings for treatment under different Pi regimes in a hydroponic system
Easy-to-assemble, element-contamination-free, and the aseptic hydroponic system is suitable for documenting the developmental responses of different root traits of the rice seedlings grown under different Pi regimes45,47,49. The seed area of N22 and NH787 was documented by employing the ImageJ program48. There was no significant variation in the seed area of N22 and NH787 (Fig. S1a). Relatively, the seed area was marginally higher (~ 5%) in N22 EMS mutant NH10147. This suggested a variable effect of EMS mutagenesis on the seed area of N22 EMS mutants. Seeds (~ 20) of N22 and NH787 were placed equidistant on a Petri plate lined with a germination paper soaked with deionized distilled water, wrapped in aluminum foil, and maintained in a growth chamber (28–30 °C) for 4 days. Rice seed with radicle length > 0.5 cm was considered germinated47. The images of germinated seedlings (~ 200 each of N22 and NH787) spread over 10 Petri plates were captured by using a desktop scanner (Fig. S1b). A significant variation was apparent in the radicle length of the germinated seedlings of both N22 and NH787. Earlier studies had suggested selecting only those rice seedlings whose radicle length falls within a fairly comparable size range (~ 2 to 3 cm) for subsequent transfer to a hydroponic system under different Pi regime to circumvent any erroneous interpretations47,49. In the model plant Arabidopsis thaliana also, the selection of uniformly grown seedlings with primary root length in the range of ~ 1.5 to 2.5 cm was recommended to minimize the effect of intrinsic variability on the subsequent treatments under different Pi regimes66,67,68. Therefore, the radicle length of the germinated N22 and NH787 seedlings was measured by using the ImageJ program48 and categorized into different groups based on their radicle length (Fig. S1c). The size distribution pattern of N22 and NH787 radicle length is represented by the red (≤ 0.50 cm), black (0.51–2.0 cm), green (2.01–3.00 cm), and yellow (3.01–5.50) histograms, which exhibited a typical Gaussian curve and a noticeable variation between the genotypes. The number of seedlings with radicle length in the size range of 2.01–3.00 cm was significantly higher (49.38%) in NH787 compared with N22 (21.88%). These seedlings were eventually selected for transfer to the hydroponic system containing P+ and P− media and the rest of the seedlings (< 2.01 cm and > 3.00 cm) were discarded (Fig. S1d).
Responses of ontogenetically distinct root traits under different Pi regimes in a hydroponic system
The root system of rice comprises ontogenetically distinct embryonically developed primary and seminal roots that play a key role during the seedling stage and post-embryonically developed adventitious roots constitute the bulk of the functional root system in a mature plant69,70. N22 and NH787 seedlings (4-day-old) with radicle length in the size range of 2.01–3.00 cm were transferred to the hydroponic system containing P+ and P− media and grown for 7 d. After the treatment, roots of N22 and NH787 were separated at the shoot: hypocotyl junction and spread gently to reveal the architectural details of the embryonically and post-embryonically developed traits under P+ and P− conditions. Images of the spread-out roots were captured by using a desktop scanner and the ImageJ program48 was then used for quantitative documentation of the effects of P+ (Fig. S2B) and P− (Fig. 2b–g) treatments on different root traits. There was a significant reduction (31.23%) in the primary root length (PRL) of N22 under P− condition (data not shown) and the result was consistent with earlier studies on N2247,49, 71. On the contrary, PRL of NH787 was comparable under P+ and P− conditions (data not shown). Although PRL of N22 and NH787 was comparable under P+ condition, it was significantly higher (21.56%) in the latter compared with the former under P− condition (Fig. 2a,b). The number of lateral roots (NLR) was significantly reduced (47.31%) in NH787 compared with N22 under P+ condition (Fig. S2A,B) but was comparable under P− condition (data not shown). Pi deficiency triggered a significant reduction (72.01%) in the total length of the lateral roots (TLLR) on primary, seminal, and adventitious roots of N22 (data not shown) and agreed with earlier studies on N2247,49. Relatively, Pi deficiency-mediated reduction of TLLR in NH787 was 48.64%, which was significantly lower compared with N22 (data not shown). This suggested that the effect of Pi deficiency on TLLR was more aggravated on N22 than NH787. Although TLLR of N22 and NH787 was comparable under P+ condition (Fig. S2A), it was significantly higher (43.42%) in the latter compared with the former under P− condition (Fig. 2a,c). In rice, elongation of the seminal root plays a key role in the acquisition of nutrients such as Pi and nitrogen (N)72. Therefore, the effect of Pi deficiency was investigated on the number of seminal roots (NSR) and the total length of seminal roots (TLSR) of N22 and NH787. The effect of Pi deficiency was evident on the developmental response of the seminal roots of N22, which revealed significant reductions by 90.03% and 84.68% in their NSR and TLSR, respectively compared with P+ condition (data not shown) and was congruent with studies on N2247,49. The effects of Pi deficiency on both NSR and TLSR of NH787 were relatively less aggravated and resulted in reductions by 69.88% and 55.99%, respectively (data not shown). NSR and TLSR of N22 and NH787 were comparable under P+ condition (Fig. S2a). However, under P− condition the NSR and TLSR of NH787 were 2.6-fold and 2.9-fold higher, respectively compared with N22 (Fig. 2a,d,e). Pi deficiency has also been shown to exert an attenuating influence on the seminal root length of rice varieties O. rufipogon (IRGC 105491) and Curinga72. The total length of adventitious roots (TLAR) increased significantly (25.52%) in N22 during Pi deficiency (data not shown) and the result was in agreement with earlier studies on N2247,49. Relatively, the increase in TLAR was only 12.29% in Pi-deprived NH787 (data not shown). TLAR of N22 and NH787 was comparable under P+ condition (Fig. 2a) but was significantly higher (65.53%) in the latter compared with the former under P− condition (Fig. 2a,f). Finally, the total root length (TRL) was computed by summation of PRL, TLLR, TLSR, and TLAR. Pi deficiency exerted a significant (63.38%) attenuating effect on the TRL of N22 (data not shown). Earlier studies also reported the inhibitory effect of Pi deficiency on TRL of rice varieties N2247,49 and IR64 (transgenics [null] and NILs with [+] or without [−] Pup1)10. Comparatively, the effect of Pi deprivation was less aggravated on TRL of NH787 and exhibited a 36.74% reduction compared with P+ condition (data not shown). Although TRL of N22 and NH787 was comparable under P+ condition (Fig. S2A), it was significantly higher (65.68%) in the latter compared with the former under P− condition (Fig. 2a–g). Together, the detailed analyses of different root traits revealed that the effects of Pi deficiency were more aggravated on N22 than NH787. Further, the Pearson correlation was presented as a correlogram to determine the relationship across the developmental responses of ontogenetically distinct root traits of N22 and NH787 under different Pi regimes (Fig. S2C, Fig. 2h). Under P+ condition, TLLR was positively and significantly correlated with TRL and NSR in N22, whereas a significant positive correlation was observed between NLR, TLLR, and TRL and NAR and TLAR in NH787 (Fig. S2C). Under P− condition, NSR and TLSR in N22, and TLSR, NAR, TLAR, and TRL in NH787 exhibited a significant positive correlation (Fig. 2h). The analysis revealed that NSR, TLSR, and TLLR were positively and significantly correlated with TRL in both Pi-deprived N22 and NH787. Pearson correlation has also been used in earlier studies on various morpho-biochemical traits at various developmental stages of N22 and its EMS mutants under different Pi regimes43,44,47.
Effects of Pi deficiency on different RSA traits. N22 and NH787 seedlings (4-day-old) were grown hydroponically under P− condition for 7 day. (a) Roots were spread gently and scanned to reveal the architectural details. The primary root tip is indicated by a blue dot. (b–g) Data presented for (b) primary root length (PRL), (c) total length of the lateral roots (TLLR), (d) number of seminal roots (NSR), (e) total length of seminal roots (TLSR), (f) total length of adventitious roots (TLAR), and (g) total root length (TRL). Values (n = 12) are means ± SE and different letters on the histograms indicate that the means differ significantly (P < 0.05). (h) Correlogram of the RSA traits in Pi-deprived N22 and NH787. The scale represents Pearson correlation values with reddish and bluish shades indicate positive and negative correlation, respectively.
Effects of different Pi regimes on various morpho-agronomic traits of N22 and NH787 grown to maturity in potting soil
Growth performance and morpho-agronomic traits of N22 and NH787 plants grown to maturity (50% flowering) in Pi-replete (P+) and low Pi (P−) potting soil were determined (Fig. S3, Fig. 3). Pi deficiency exerted inhibitory effects on various morpho-agronomic traits of both N22 and NH787, which resulted in stunted phenotype, and significant reductions in the root biomass (N22 [88.37%], NH787 [24.59%]), vegetative biomass (N22 [73.48%], NH787 [12.03%]), filled spikelets/panicle (N22 [50.18%], NH787 [17.53%]), 100-grain weight (N22 [25.00%], NH787 [8.69%]), and yield (N22 [83.08%], NH787 [23.59%]) (data not shown). Whereas, during Pi deficiency, the unfilled spikelets/panicle was significantly higher in N22 (52.74%) but was comparable in NH787 with P+ condition (data not shown). It was evident from the analysis that the effects of Pi deprivation were relatively more aggravated in N22 than NH787. Earlier studies also reported the inhibitory effects of Pi deficiency on various morpho-agronomic traits of N22 and its EMS mutants42,43,44,46,47,71. Under P+ condition, there was no apparent difference in the phenotype of N22 and NH787 (Fig. S3A). However, the phenotype of the root, panicles, and grain was more robust in NH787 compared with N22 (Fig. S3B-D). This was reflected in significantly higher root biomass (41.86%), vegetative biomass (19.70%), filled spikelets/panicle (89.79%), 100-grain weight (15.00%), and yield (36.96%) of NH787 compared with N22 (Fig. S3E–G,I,J). On the contrary, unfilled spikelets/panicle was significantly higher (61.64%) in N22 compared with NH787 (Fig. S3H). However, under P+ condition the root length and number of tillers in N22 and NH787 were comparable (data not shown). Further, Pearson analysis revealed a positive and significant correlation of yield with root and vegetative biomass, number of tillers, and filled spikelets/panicle in both N22 and NH787 under P+ condition (Fig. S3K). Under P− condition, the phenotype of the plant, root, panicles, and grain were more robust in NH787 than N22 (Fig. 3a–d). The phenotypic observation was substantiated with significantly higher root biomass (9.2 folds), root length (12.21%), vegetative biomass (3.9 folds), number of tillers (2.37 folds), filled spikelets/panicle (3.15 folds), 100-grain weight (40.00%), and yield (6.42 folds) of NH787 compared with N22 (Fig. 3e–i,k,i). However, unfilled spikelets/panicle was significantly higher (73.99%) in N22 than NH787 (Fig. 3j). Similar to P+ condition, under P− condition also Pearson analysis showed a positive and significant correlation of yield with root and vegetative biomass, number of tillers, and filled spikelets/panicle in both N22 and NH787 (Fig. 3h).
Effects of Pi deficiency on the growth performance and agronomic traits. N22 and NH787 seedlings (15-day-old) were grown in a low Pi (P−) potting soil up to maturity. (a–d) Growth performance (a) and phenotype of the root (b), panicle (c), and seed (d) of Pi-deprived N22 and NH787. The photographs (a–d) are representatives of 12 independent biological replicates. (e–l) Data presented for (e) root biomass, (f) root length, (g) vegetative biomass, (h) number of tillers, (i) filled spikelets/panicle, (j) unfilled spikelets/panicle, (k) 100-grain weight, and (l) yield. Values (n = 12) are means ± SE and different letters on the histograms indicate that the means differ significantly (P < 0.05). Correlogram of agronomic traits i.e., filled spikelets/panicle (FSP), number of tillers (NT), root biomass (RB), root length (RL), vegetative biomass (VB), unfilled spikelets/panicle (USP), and 100-Grain weight (100 GW) in Pi-deprived N22 and NH787. The scale represents Pearson correlation values with reddish and bluish shades indicate positive and negative correlation, respectively.
Photosynthetic and chlorophyll fluorescence traits of N22 and NH787 grown to maturity in potting soil
Pi deficiency adversely affects photosynthetic and chlorophyll fluorescence traits in rice73,74. Therefore, photosynthetic and fluorescence traits were assayed in N22 and NH787 grown to maturity under different Pi regimes (Fig. S4, Fig. 4). Pi deficiency triggered significant reductions in the photosynthetic rate (PN) (N22 [21.84%], NH787 [26.39%]), stomatal conductance (gs) (N22 [88.24%], NH787 [42.23%]), transpiration rate (E) (N22 [15.31%], NH787 [27.90%]), maximum efficiency of PSII photochemistry (Fv/Fm) (N22 [15.79%], NH787 [8.73%]), electron transport rate (ETR) (N22 [26.69%], NH787 [18.32%]), coefficient of photochemical quenching (qP) (N22 [21.26%], NH787 [11.26%]), and coefficient of non-photochemical quenching (qN) (N22 [28.66%], NH787 [15.60%]) (data not shown). Earlier studies also reported the inhibitory effects of Pi deficiency on various photosynthetic and chlorophyll fluorescence traits in the rice genotypes73,74, N22 and its EMS mutants43. On the contrary, Pi deficiency excreted significant increase in the contents of intercellular CO2 (Ci) (N22 [26.02%], NH787 [26.32%]), chlorophyll a (N22 [35.18%], NH787 [16.53%]), chlorophyll b (N22 [32.47%], NH787 [28.70%]), and carotenoid (N22 [18.39%], NH787 [7.22%]) (data not shown). The result was consistent with an earlier study showing Pi deficiency-mediated elevated content of Ci74. Relatively, the augmenting effects of Pi deficiency on chlorophyll a, b, and carotenoid were significantly lower in NH787 compared with N22 (data not shown). Under P+ condition, PN (27.32%), gs (74.66%), Ci (3.99%), E (29.02%), Fv/Fm (3.79%), ETR (15.22%), qN (14.03%), and contents of chlorophyll a (32.74%), chlorophyll b (16.75%), and carotenoid (21.35%) were significantly higher in NH787 than N22 (Fig. S4A–J). A similar trend was also observed during Pi deficiency where these values (PN [19.90%], gs [31.97%], Ci [4.24%], E [9.83%], Fv/Fm [12.48%], ETR [28.39%], qP [15.67%], qN [2.46%], and contents of chlorophyll a [14.42%], chlorophyll b [13.43%], and carotenoid [9.91%]) were significantly higher in NH787 compared with N22 (Fig. 4a–k). The analyses revealed that NH787 maintained higher photosynthetic and chlorophyll fluorescence traits than N22 under different Pi regimes (Fig. S4, Fig. 4).
Effects of Pi deficiency on the photosynthetic and fluorescence traits in N22 and NH787. (a–k) Data are presented for (a) Photosynthetic rate (PN), (b) Stomatal conductance (gs), (c) Intercellular CO2 concentration (Ci), (d) Transpiration rate (E), (e) Maximum efficiency of PSII photochemistry (Fv/Fm), (f) Electron transport rate (ETR), (g) Coefficient of photochemical quenching (qP), (h) Coefficient of non-photochemical quenching (qN), and contents of (i) Chlorophyll a, (j) Chlorophyll b, and (k) carotenoid. Values (n = 12) are means ± SE and different letters on the histograms indicate that the means differ significantly (P < 0.05).
Pi, Apase and the enzymes involved in ROS scavenging of N22 and NH787 grown to maturity in potting soil
Pi deficiency exerts an attenuating effect on the concentration of Pi, while its effect is augmenting on the activities of Apase and ROS scavenging pathway (APX, CAT, H2O2, POD, and SOD) in the root and shoot of rice47. Therefore, the concentration of Pi and the activities of Apase and ROS scavenging enzymes were assayed in N22 and NH787 grown to maturity under different Pi regimes (Fig. S5, Fig. 5). Pi deficiency triggered significant reductions in the concentration of Pi in the root (N22 [61.30%], NH787 [55.77%]) and in shoot (N22 [66.86%], NH787 [65.31%]) (data not shown). The result was consistent with earlier studies reporting Pi deficiency-mediated reduction in the concentration of Pi in the root and shoot of N22 and its EMS mutants45,47. It was apparent from this analysis that the effect of Pi deficiency on the concentration of Pi in root was relatively more aggravated in N22 than NH787 but was comparable in the shoot. Concentration of Pi in the root (P+ [14.90%], P− [31.32%]) and shoot (P+ [15.22%], P− [20.60%]) were significantly higher in NH787 than N22 (Fig. S5A, Fig. 5a). On the contrary, the activity of Apase increased significantly during Pi deficiency in the root (N22 [2.17 fold], NH787 [2.86 fold]) and in the shoot (N22 [4.31 fold], NH787 [4.46 fold]) (data not shown) and was coherent with earlier studies on N22 and its EMS mutants46,47. Although the augmenting effect of Pi deficiency on Apase activity was significantly higher in the root of NH787 compared with N22, it was comparable in the shoot of these two genotypes. The activity of Apase in the root (P+ [36.51%], P− [16.28%]) and shoot (P+ [22.17%], P− [19.50%]) were significantly higher in N22 than NH787 (Fig. S5b, Fig. 5b). Significant augmenting effects of Pi deficiency were also evident in the root and shoot of N22 and NH787 on different components of ROS pathway comprising SOD (root [41.86% in N22 and 33.59% in NH787] and shoot [40.36% in N22 and 50.19% in NH787]), H2O2 (root [2.87 fold in N22 and 2.97 fold in NH787] and shoot [69.11% in N22 and 54.03% in NH787]), POD (root [40.03% in N22 and 44.97% in NH787] and shoot [82.65% in N22 and 63.84% in NH787]), APX (root [77.63% in N22 and 44.16% in NH787] and shoot [35.90% in N22 and 22.10% in NH787]), and CAT (root [74.25% in N22 and 87.84% in NH787] and shoot [2.00 fold in N22 and 2.18 fold in NH787]) (data not shown). The analysis revealed that the values in N22 were significantly higher (root [SOD], shoot [H2O2 and POD], and root and shoot [APX]) or lower (root [H2O2 and POD], shoot [SOD], and root and shoot [CAT]) compared with NH787 (data not shown). Under P+ condition, the values were significantly lower (root and shoot [H2O2 and POD] and shoot [SOD]), higher (root and shoot [APX and CAT]), and non-significant (root [SOD]) in NH787 compared with N22 (Fig. S5C–G). Almost a similar trend was observed under P− condition with values significantly lower (root and shoot [SOD, H2O2, and POD] and root [CAT]) and higher (root and shoot [APX] and shoot [CAT]) in NH787 compared with N22 (Fig. 5c–g). The results highlighted differential effects on ROS-mediated redox signaling and oxidative stress in NH787 compared with N22 under different Pi regimes. Earlier studies also showed the Pi-dependent differential effects on ROS homeostasis in the EMS mutants of N2243,47.
Effects of Pi deficiency on the concentrations of Pi, Apase, and the enzymes involved in ROS scavenging. N22 and NH787 were grown in a potting soil up to 50% flowering under P+ and P− conditions. (a–g) Data are presented for the concentrations of (a) Pi, (b) Apase, (c) SOD, (d) H2O2, (e) POD, (f) APX, and (g) CAT. Values (n = 12) are means ± SE and different letters on the histograms indicate that the means differ significantly (P < 0.05). Ps potting soil.
Reproductive traits of N22 and NH787 grown to maturity in potting soil
In earlier studies, significant inhibitory effects of Pi deficiency were observed on the yield potential of N22 and its EMS mutants42,43,44,45,46,47. Therefore, the effects of Pi deprivation were investigated on the male reproductive traits of N22 and NH787 grown under different Pi regimes in a potting soil up to maturity (Fig. 6). I2-KI staining was used for determining the viability of pollen collected after anthesis from N22 and NH787 grown under P+ and P− conditions (Fig. 6a–d). Pollen viability was significantly higher (P+ [14.73%], P− [23.76%]) in NH787 compared with N22, which suggested that Pi deficiency-mediated effect on pollen viability was more aggravated in the latter than the former (Fig. 6a–E). Further, there were significant reductions in the activities of SOD (P+ [29.48%], P− [29.64%]), POD (P+ [12.58%], P− [29.02%]), and APX (P+ [5.48%], P− [13.63%]) in the anthers of NH787 compared with N22 irrespective of Pi regimes (Fig. 6f–h). On the contrary, the activity of CAT in NH787 was significantly lower and higher under P+ (19.05%) and P− (47.05%) conditions, respectively compared with N22 (Fig. 6I). The analysis revealed differential effects on antioxidant enzyme activities of N22 and NH787 under different Pi regimes. Under Pi-deprived condition, NH787 exhibited significantly higher CAT activity in the anthers compared with N22, which triggered the efficient conversion of H2O2 into water and oxygen possibly favoring higher pollen fertility and yield (Fig. S6). Further, the effects of Pi deficiency were determined on various grain parameters of N22 and NH787 grown in P+ and P− potting soil up to maturity (Fig. 7). Single grain weight of N22 and NH787 collected from P+ and P− plants were categorized into a week (6–10 mg and 10–15 mg) and robust (16–20 mg, 21–25 mg, and 26–30 mg) categories and the frequency of weight distribution pattern in these categories was computed, which revealed a typical Gaussian curve (Fig. 7a). Under P+ condition, the frequency of single grain weight under different categories was comparable between N22 and NH787 (Fig. 7a). However, significant variation in the frequency of single grain weight was observed under P− condition for N22 compared with NH787 ranging from higher (6–10 mg and 10–15 mg), lower (16–20 mg and 21–25 mg), and comparable (26–30 mg) values (Fig. 7a). The analysis revealed that NH787 seeds were comparatively more robust than N22 when grown under Pi-deprived condition. Under both P+ and P− conditions, several grain quality parameters of NH787 were significantly higher than N22 comprising hulling (P+ [8.07%], P− [24.18%]) (Fig. 7b), milling (P+ [9.48%], P− [21.21%]) (Fig. 7c), percent head rice recovery (P+ [11.79%], P− [26.54%]) (Fig. 7d), percent amylose content (P+ [18.23%], P− [2.21 fold]) (Fig. 7e), grain length (P+ [7.07%], P− [8.47%]) (Fig. 7f), grain width (P+ [20.30%], P− [27.71%]) (Fig. 7g), and grain area (P+ [28.72%], P− [38.42%]) (Fig. 7h). Different parameters such as alkali spread value, gelatinization temperature, and gel consistency are conventionally used for determining the gain quality traits of rice75,76,77. Therefore, alkali spread value, gelatinization temperature, and gel consistency were assayed for P+ and P− seeds of N22 and NH787, which revealed that these traits were superior in the latter compared with the former (Table S2). Although α-amylase activity in the spikelets of N22 and NH787 was comparable under P+ condition, it was significantly lower in the latter compared with the former under P− condition (Table S3). Low α-amylase activity in the rice spikelets has been correlated with the grain weight and yield78,79. Pearson correlation analysis was carried out to determine the relationship across agronomical and quality traits (yield, TN, GA, GW, GL, AC, HRR, milling, and hulling), pollen fertility (PF), and the activities of antioxidant enzymes (CAT, APX, POD, and SOD) and α-amylase (AA) in the anther and spikelets of N22 and NH787 under different Pi regimes (Fig. 8). Under P+ condition, a positive and significant correlation in N22 was observed with yield, TN, AA, and AC and that of NH787 with yield, TN, PF, AC, and HRR. Whereas, under P− condition, a positive and significant correlation in N22 was detected with yield, TN, APX, AA, AC, and hulling and that of NH787 with yield, TN, PF, POD, APX, GA, GW, AC, and milling. The analysis revealed that TN, AC, AA, POD, APX, and PF were correlated positively and significantly with the yield of N22 and NH787 under different Pi regimes.
Effects of Pi deficiency on pollen viability and antioxidant enzyme activities in the anther. N22 and NH787 were grown in a potting soil up to 50% flowering under P+ and P− conditions. (a–d) Pollen viability was assayed by staining with I2-KI and the images were captured with a stereomicroscope. The data are presented for (e) Percent pollen viability and (f–i) the ROS scavenging enzyme activities in the anthers of (f) SOD, (g) POD, (h) APX, and (i) CAT. Values (n = 6) are means ± SE and different letters on the histograms indicate that the means differ significantly (P < 0.05).
Effects of Pi deficiency on different grain parameters. N22 and NH787 were grown in a potting soil up to 50% flowering under P+ and P− conditions and after harvesting, grains were threshed, cleaned, and dried under natural conditions. Data are presented for (a) frequency of single grain weight, (b) percent hulling, (c) percent milling (d) percent head rice recovery, (e) percent amylose content, (f) grain length, (g) grain width, and (h) grain area. Values (n = 3, 6, and 20 for (a,b–e), and (f–h), respectively) are means ± SE and different letters on the histograms indicate that the means differ significantly (P < 0.05).
Correlogram of the agronomical and quality traits, pollen fertility, and the activities of antioxidant enzymes and α-amylase in the anther and spikelets, respectively in N22 and NH787 grown in a potting soil up to 50% flowering under P+ and P− conditions. The scale represents Pearson correlation values with brownish and bluish shades indicate positive and negative correlation, respectively. TN Tiller number, PF pollen fertility, GA grain area, GW grain width, GL grain length, AA α-amylase, AC amylose content, HRR head rice recovery.
Relative expression levels of GPH in N22 and NH787 grown to maturity in potting soil
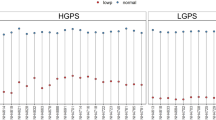
The qRT-PCR assay was employed to decipher Pi deficiency-mediated effects on the relative expression levels of functionally diverse GPH in the roots of N22 and NH787 grown to maturity in potting soil under P+ and P− conditions (Fig. 9). For this experiment, only those GPH were selected, which had been functionally characterized either by overexpression under the constitutive promoter or by mutation (T-DNA, Tos17, RNAi, or CRISPR-cas9) and implicated in their tissue-specific key roles in the sensing and signaling cascades governing the maintenance of Pi homeostasis under Pi regimes (Table 2). In Pi-deprived roots of NH787 compared with N22, the relative expression levels of several GPH were significantly higher that are implicated in the transcriptional regulation of signaling pathway (OsPHR280), regulation by systemic and local Pi signaling and hormones (OsIPS81), regulation of Fe transport by integrating Pi and Zn deficiency signaling (OsPHO1;182), inhibition of Pi starvation responses by interacting with OsPHR2 in a Pi-dependent manner (OsSPX283), and uptake and/or mobilization of Pi by low-and high-affinity Pi transporters (OsPht1;1, OsPht1;2, OsPht1;4, OsPht1;6, OsPht1;8, and OsPht1;950,84,85,86,87,88,89 (Fig. 9a). Higher relative expression levels of these GPHs in NH787 could be assumed to play a pivotal role in contributing towards its higher yield potential (Fig. 3). On the contrary, the relative expression levels of several GPH were significantly reduced in Pi-deprived roots of NH787 compared with N22 that are involved in the uptake of Pi by high-affinity Pi transporter (OsPht1;1087), utilization of extracellular organic P (OsPAP10a90), regulation of the growth during Pi deficiency via a negative feedback loop and by interacting with OsPHR2 in a Pi-dependent manner (OsSPX183,91), regulation of Pi starvation signal transduction (OsmiRNA399a92), growth, development, and maintenance of Pi homeostasis (OsLPR593), regulation of Pi starvation responses (OsPHO294), and post-translational SUMOylation of proteins (OsSIZ195) (Fig. 9b). In Arabidopsis thaliana, electrophoretic mobility shift assay revealed the binding of the transcription factor PHR1 as a dimer to an imperfect palindromic 8-bp sequence (5′-GNATATNC-3′) named as PHR1 binding sequence (P1BS) found in the promoter (2 kb upstream of ATG start codon) of several genes involved in Pi deficiency-mediated responses96,97. Therefore, the P1BS (GNATATNC) motif was analyzed in the promoter (3 kb upstream of ATG initiation site) of the 17 GPH revealing its presence in 14 of them, which suggested their potential regulation by OsPHR2 (Table 2). In this context, significantly higher relative expression of OsPHR2 in Pi-deprived roots of NH787 compared with N22 (Fig. 9a) suggested its potential regulatory influence on the expression of several GPH that play a key role in the maintenance of Pi homeostasis under different Pi regimes.
Relative expression levels of GPH in Pi-deprived roots of N22 and NH787. N22 and NH787 were grown in potting soil under P+ and P− conditions up to 50% flowering stage and their roots were harvested. Quantitative real-time RT-PCR (qRT-PCR) was employed for determining the relative expression levels of GPH genes. (a) GPH genes induced in NH787 (green bars) compared with N22 (black bars). (b) GPH genes were suppressed in NH787 (red bars) compared with N22 (black bars). OsActin (LOC_Os03g50885) was used as an internal control. Values are means ± SE (n = 6) and different letters on the histograms indicate that the values differ significantly (P < 0.05).
Conclusions
The results provided empirical evidence towards the differential effects of the EMS mutagenesis on various morphophysiological, biochemical, and molecular traits of NH787 that conferred higher PUE under low Pi soil condition compared with N22. A schematic diagram is presented highlighting the efficacy of the EMS mutants for screening the rice variety NH787 with higher PUE in an environment-friendly manner for sustainable production (Fig. 10). NH787 is now used as a donor in breeding programs for developing low P tolerant varieties with superior grain quality and is also being evaluated in larger plots at multiple locations with variable agroclimatic conditions. Efforts are also underway to identify the candidate genes in NH787 responsible for higher PUE by employing quantitative trait loci (QTL) mapping and MutMap approach in the F2 populations revealing a discernible phenotype98. MutMap approach has been used in an earlier study for identifying the candidate genes conferring salt tolerance in the F2 populations of EMS mutant hitomebore salt tolerant 1 (hst1) of rice variety Hitomebore99.
Data availability
All data generated or analyzed during this study are included in this published article (and its Supplementary Information files). The sequence data is available on request.
References
Singh, S. K., Bhati, P. K., Sharma, A. & Sahu, V. Super hybrid rice in China and India: current status and future prospects. Int. J. Agric. Biol. 17, 221–232 (2015).
Chien, P.-S., Chiang, C.-P., Leong, S. J. & Chiou, T.-J. Sensing and signaling of phosphate starvation: from local to long distance. Plant Cell Physiol. 59, 1714–1722 (2018).
Chiou, T.-J. & Lin, S.-I. Signaling network in sensing phosphate availability in plants. Annu. Rev. Plant Biol. 62, 185–206 (2011).
Crombez, H., Motte, H. & Beeckman, T. Tackling plant phosphate starvation by the roots. Dev. Cell 48, 599–615 (2019).
Gutiérrez-Alanís, D., Ojeda-Rivera, J. O., Yong-Villalobos, L., Cárdenas-Torres, L. & Herrera-Estrella, L. Adaptation to phosphate scarcity: tips from Arabidopsis roots. Trends Plant Sci. 23, 721–730 (2018).
López-Arredondo, D. L., Leyva-González, M. A., González-Morales, S. I., López-Bucio, J. & Herrera-Estrella, L. Phosphate nutrition: improving low-phosphate tolerance in crops. Annu. Rev. Plant Biol. 65, 95–123 (2014).
Gu, M., Chen, A., Sun, S. & Xu, G. Complex regulation of plant phosphate transporters and the gap between molecular mechanisms and practical application: what is missing?. Mol. Plant 9, 396–416 (2016).
Nussaume, L. et al. Phosphate import in plants: focus on the PHT1 transporters. Front. Plant Sci. 2, 83 (2011).
Raghothama, K. G. Phosphate acquisition. Annu. Rev. Plant Physiol. Plant Mol. Biol. 50, 665–693 (1999).
Gamuyao, R. et al. The protein kinase Pstol1 from traditional rice confers tolerance of phosphorus deficiency. Nature 488, 535–539 (2012).
Haefele, S. M., Nelson, A. & Hijmans, R. J. Soil quality and constraints in global rice production. Geoderma 235–236, 250–259 (2014).
Krishnamurthy, P., Sreedevi, B., Ram, T. & Latha, P. C. Evaluation of rice genotypes for phosphorus use efficiency under soil mineral stress conditions. J. Rice Res. 7, 53–61 (2014).
Prasad, R. Fertilizers and manures. Curr. Sci. 102, 894–898 (2012).
Cordell, D., Drangert, J.-O. & White, S. The story of phosphorus: global food security and food for thought. Global Environ. Change 19, 292–305 (2009).
Veneklaas, E. J. et al. Opportunities for improving phosphorus-use efficiency in crop plants. New Phytol. 195, 306–320 (2012).
van de Wiel, C. C. M., van der Linden, C. G. & Scholten, O. E. Improving phosphorus use efficiency in agriculture: opportunities for breeding. Euphytica 207, 1–22 (2016).
International Rice Genome Sequencing Project. The map-based sequence of the rice genome. Nature 436, 793–800 (2005).
Jackson, S. A. Rice: the first crop genome. Rice 9, 14 (2016).
Jung, K.-H., An, G. & Ronald, P. C. Towards a better bowl of rice: assigning function to tens of thousands of rice genes. Nat. Rev. Genet. 9, 91–101 (2008).
Hirochika, H. Insertional mutagenesis with Tos17 for functional analysis of rice genes. Breed. Sci. 60, 486–492 (2010).
Hirochika, H. et al. Rice mutant resources for gene discovery. Plant Mol. Biol. 54, 325–334 (2004).
Wilson, R. & Doudna, J. A. Molecular mechanisms of RNA interference. Annu. Rev. Biophys. 42, 217–239 (2013).
Jinek, M. et al. A programmable dual RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
He, Y. et al. Programmed self-elimination of the CRISPR/Cas9 construct greatly accelerates the isolation of edited and transgene-free rice plants. Mol. Plant 11, 1210–1213 (2018).
Hu, X., Meng, X., Liu, Q., Li, J. & Wang, K. Increasing the efficiency of CRISPR-Cas9-VQR precise genome editing in rice. Plant Biotechnol. J. 16, 292–297 (2018).
Miao, C. et al. Mutations in a subfamily of abscisic acid receptor genes promote rice growth and productivity. Proc. Natl Acad. Sci. USA 115, 6058–6063 (2018).
Kondou, Y., Higuchi, M. & Matsui, M. High-throughput characterization of plant gene functions by using gain-of-function technology. Annu. Rev. Plant Biol. 61, 373–393 (2010).
Wu, P., Shou, H., Xu, G. & Lian, X. Improvement of phosphorus efficiency in rice on the basis of understanding phosphate signaling and homeostasis. Curr. Opin. Plant Biol. 16, 205–212 (2013).
Chen, K. & Gao, C. Genome-edited crops: how to move them from laboratory to market. Front. Agr. Sci. Eng. 7, 181–187 (2020).
Cunningham, F. J., Goh, N. S., Demirer, G. S., Matos, J. L. & Landry, M. P. Nanoparticle-mediated delivery towards advancing plant genetic engineering. Trends Biotechnol. 36, 882–897 (2018).
Schmidt, S. M., Belisle, M. & Frommer, W. B. The evolving landscape around genome editing in agriculture. EMBO Rep. 21, e50680 (2020).
Editorial. Gene-edited plants cross European event horizon, Nat. Biotechnol. 36, 776 (2018).
Ishii, T. Crop gene-editing: should we bypass or apply existing GMO policy?. Trends Plant Sci. 23, 947–950 (2018).
Purnhagen, K. P. et al. EU court casts new plant breeding techniques into regulatory limbo. Nat. Biotechnol. 36, 799–800 (2018).
Potrykus, I. Golden rice and beyond. Plant Physiol. 125, 1157–1161 (2001).
Kettenburg, A. J., Hanspach, J., Abson, D. J. & Fischer, J. From disagreements to dialogue: unpacking the golden rice debate. Sustain. Sci. 13, 1469–1482 (2018).
Till, B. J. et al. Discovery of chemically induced mutations in rice by TILLING. BMC Plant Biol. 7, 19 (2007).
Tsai, H. et al. Discovery of rare mutations in populations: TILLING by sequencing. Plant Physiol. 156, 1257–1268 (2011).
Jiang, S.-Y. & Ramachandran, S. Natural and artificial mutants as valuable resources for functional genomics and molecular breeding. Int. J. Biol. Sci. 6, 228–251 (2010).
Jagadish, S. V. K., Craufurd, P. Q. & Wheeler, T. R. Phenotyping parents of mapping populations of rice for heat tolerance during anthesis. Crop Sci. 48, 1140–1146 (2008).
Mithra, S. V. A. et al. DBT propelled national effort in creating mutant resource for functional genomics in rice. Curr. Sci. 110, 543–548 (2016).
Poli, Y. et al. Genotype × environment interactions of Nagina22 rice mutants for yield traits under low phosphorus, water limited and normal irrigated conditions. Sci. Rep. 8, 15530 (2018).
Poli, Y. et al. Increased catalase activity and maintenance of photosystem II distinguishes high-yield mutants from low-yield mutants of rice var Nagina22 under low-phosphorus stress. Front. Plant Sci. 9, 1543 (2018).
Yugandhar, P. et al. Identifying markers associated with yield traits in Nagina22 rice mutants grown in low phosphorus field or in alternate wet/dry conditions. Aust. J. Crop. Sci. 11, 548–556 (2017).
Yugandhar, P. et al. Comparing hydroponics, sand, and soil medium to evaluate contrasting rice Nagina22 mutants for tolerance to phosphorus deficiency. Crop Sci. 57, 2089–2097 (2017).
Yugandhar, P. et al. Nagina22 mutants tolerant or sensitive to low P in field show contrasting response to double P in hydroponics and pots. Arch. Agron. Soil Sci. 64, 1975–1987 (2018).
Yugandhar, P. et al. Characterization of the loss-of-function mutant NH101 for yield under phosphate deficiency from EMS-induced mutants of rice variety Nagina22. Plant Physiol. Biochem. 130, 1–13 (2018).
Collins, T. J. ImageJ for microscopy. Biotechniques 43, S25–S30 (2007).
Negi, M., Sanagala, R., Rai, V. & Jain, A. Deciphering phosphate deficiency-mediated temporal effects on different root traits in rice grown in a modified hydroponic system. Front. Plant Sci. 7, 550 (2016).
Jia, H. et al. The phosphate transporter gene OsPht1;8 is involved in phosphate homeostasis in rice. Plant Physiol. 156, 1164–1175 (2011).
Lichtenthaler, H. K. & Wellburn, A. R. Determinations of total carotenoids and chlorophylls a and b in leaf extracts in different solvents. Biochem. Soc. Trans. 11, 591–592 (1983).
Porra, P. J., Thompson, W. A. & Kriedemann, P. E. Determination of accurate extinction coefficients and simultaneous equations for assaying chlorophylls a and b extracted with four different solvents: verification of the concentration of chlorophyll standards by atomic absorption spectroscopy. Biochim. Biophys. Acta 975, 384–394 (1989).
Ames, B. N. Assay of inorganic phosphate, total phosphate and phosphatases. Methods Enzymol. 8, 115–118 (1966).
Johnson, C. B., Holloway, B. R., Smith, H. & Grierson, D. Isoenzymes of acid phosphatase in germinating peas. Planta 115, 1–10 (1973).
Dhindsa, R. S., Plumb-Dhindsa, P. & Thorpe, T. A. Leaf senescence: correlated with increased levels of membrane permeability and lipid peroxidation, and decreased levels of superoxide dismutase and catalase. J. Exp. Bot. 32, 93–101 (1981).
Castillo, F. J., Penel, C. & Greppin, H. Peroxidase release induced by ozone in Sedum album leaves Involvement of Ca2+. Plant Physiol. 74, 846–851 (1984).
Aebi, H. Catalase in vitro. Methods Enzymol. 105, 121–126 (1984).
Nakano, Y. & Asada, K. Hydrogen peroxide is scavenged by ascorbate-specific peroxidase in spinach chloroplasts. Plant Cell Physiol. 22, 867–880 (1981).
Alexieva, V., Sergiev, I., Mapelli, S. & Karanov, E. The effect of drought and ultraviolet radiation on growth and stress markers in pea and wheat. Plant Cell Environ. 24, 1337–1344 (2001).
Poli, Y. et al. Characterization of a Nagina22 rice mutant for heat tolerance and mapping of yield traits. Rice 6, 36 (2013).
Cruz, N. D. & Khush, G. S. Rice grain quality evaluation procedures. In Aromatic Rices (eds Singh, R. K. et al.) 15–28 (Oxford and IBH Publishing Co Pvt Ltd, 2000).
Cagampang, G. B., Perez, C. M. & Juliano, B. O. A gel consistency test for eating quality of rice. J. Sci. Fd. Agric. 24, 1589–1594 (1973).
Juliano, B. O. A simplified assay for milled rice amylase. Cereal Sci. Today 16, 334–340 (1971).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2–ΔΔCT method. Methods 25, 402–408 (2001).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2012).
Jain, A. et al. Differential effects of sucrose and auxin on localized phosphate deficiency-induced modulation of different traits of root system architecture in Arabidopsis. Plant Physiol. 144, 232–247 (2007).
Jain, A. et al. Variations in the composition of gelling agents affect morphophysiological and molecular responses to deficiencies of phosphate and other nutrients. Plant Physiol. 150, 1033–1049 (2009).
Nagarajan, V. K. et al. Arabidopsis Pht1;5 mobilizes phosphate between source and sink organs and influences the interaction between phosphate homeostasis and ethylene signaling. Plant Physiol. 156, 1149–1163 (2011).
Hochholdinger, F., Park, W. J., Sauer, M. & Woll, K. From weeds to crops: genetic analysis of root development in cereals. Trends Plant Sci. 9, 42–48 (2004).
Hochholdinger, F. & Zimmermann, R. Conserved and diverse mechanisms in root development. Curr. Opin. Plant Biol. 11, 70–74 (2008).
Panigrahy, M. et al. Hydroponic experiment for identification of tolerance traits developed by rice Nagina 22 mutants to low-phosphorus in field condition. Arch. Agron. Soil Sci. 60, 565–576 (2014).
Ogawa, S. et al. N- and P-mediated seminal root elongation response in rice seedlings. Plant Soil 375, 303–315 (2014).
Veronica, N. et al. Influence of low phosphorus concentration on leaf photosynthetic characteristics and antioxidant response of rice genotypes. Photosynthetica 55, 285–293 (2017).
Xu, H. X., Weng, X. Y. & Yang, Y. Effect of phosphorus deficiency on the photosynthetic characteristics of rice plants. Russ. J. Plant Physiol. 54, 741–748 (2007).
Graham-Acquaah, S. et al. Variations in agronomic and grain quality traits of rice grown under irrigated lowland conditions in West Africa. Food Sci. Nutr. 6, 970–982 (2018).
Lapitan, V. C., Redoña, E. D., Abe, T. & Brar, D. S. Mapping of quantitative trait loci using a doubled-haploid population from the cross of Indica and Japonica cultivars of rice. Crop Sci. 49, 1620–1628 (2009).
Misra, G. et al. Dissecting the genome-wide genetic variants of milling and appearance quality traits in rice. J. Exp. Bot. 70, 5115–5130 (2019).
Hakata, M. et al. Suppression of α-amylase genes improves quality of rice grain ripened under high temperature. Plant Biotechnol. J. 10, 1110–1117 (2012).
Nakata, M. et al. High temperature-induced expression of rice α-amylases in developing endosperm produces chalky grains. Front. Plant Sci. 8, 2089 (2017).
Zhou, J. et al. OsPHR2 is involved in phosphate-starvation signaling and excessive phosphate accumulation in shoots of plants. Plant Physiol. 146, 1673–1686 (2008).
Hou, X. L. et al. Regulation of the expression of OsIPS1 and OsIPS2 in rice via systemic and local Pi signaling and hormones. Plant Cell Environ. 28, 353–364 (2005).
Saenchai, C. et al. The involvement of OsPHO1;1 in the regulation of iron transport through integration of phosphate and zinc deficiency signaling. Front. Plant Sci. 7, 396 (2016).
Wang, Z. et al. Rice SPX1 and SPX2 inhibit phosphate starvation responses through interacting with PHR2 in a phosphate-dependent manner. Proc. Natl Acad. Sci. USA 111, 14953–14958 (2014).
Ai, P. et al. Two rice phosphate transporters, OsPht1;2 and OsPht1;6, have different functions and kinetic properties in uptake and translocation. Plant J. 57, 798–809 (2009).
Jia, H. et al. OsPht1;8, a phosphate transporter, is involved in auxin and phosphate starvation response in rice. J. Exp. Bot. 68, 5057–5068 (2017).
Sun, S. et al. A constitutive expressed phosphate transporter, OsPht1;1, modulates phosphate uptake and translocation in phosphate-replete rice. Plant Physiol. 159, 1571–1581 (2012).
Wang, X. et al. Phosphate transporters OsPHT1;9 and OsPHT1;10 are involved in phosphate uptake in rice. Plant Cell Environ. 37, 1159–1170 (2014).
Zhang, F. et al. Overexpression of rice phosphate transporter gene OsPT6 enhances phosphate uptake and accumulation in transgenic rice plants. Plant Soil 384, 259–270 (2014).
Zhang, F. et al. Involvement of OsPht1;4 in phosphate acquisition and mobilization facilitates embryo development in rice. Plant J. 82, 556–569 (2015).
Tian, J. et al. Overexpression of OsPAP10a, a root-associated acid phosphatase, increased extracellular organic phosphorus utilization in rice. J. Integr. Plant Biol. 54, 631–639 (2012).
Wang, Z. et al. Regulation of OsSPX1 and OsSPX3 on expression of OsSPX domain genes and Pi-starvation signaling in rice. J. Integr. Plant Biol. 51, 663–674 (2009).
Hu, B. et al. MicroRNA399 is involved in multiple nutrient starvation responses in rice. Front. Plant Sci. 6, 188 (2015).
Ai, H. et al. The ferroxidase LPR5 functions in the maintenance of phosphate homeostasis and is required for normal growth and development of rice. J. Exp. Bot. 71, 4828–4842 (2020).
Hu, B. et al. LEAF TIP NECROSIS1 plays a key role in the regulation of multiple phosphate starvation responses in rice. Plant Physiol. 156, 1101–1115 (2011).
Wang, H. et al. OsSIZ1, a SUMO E3 ligase gene, is involved in the regulation of the responses to phosphate and nitrogen in rice. Plant Cell Physiol. 56, 2381–2395 (2015).
Misson, J. et al. A genome-wide transcriptional analysis using Arabidopsis thaliana Affymetrix gene chips determined plant responses to phosphate deprivation. Proc. Natl Acad. Sci. USA 102, 11934–11939 (2005).
Rubio, V. et al. A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev. 15, 2122–2133 (2001).
Abe, A. et al. Genome sequencing reveals agronomically important loci in rice using MutMap. Nat. Biotechnol. 30, 174–178 (2012).
Takagi, H. et al. MutMap accelerates breeding of a salt-tolerant rice cultivar. Nat. Biotechnol. 33, 445–449 (2015).
Acknowledgements
We thank Dr SR Voleti and Dr N Sarla (ICAR-Indian Institute of Rice Research, Hyderabad, India) for their valuable and critical suggestions throughout this study. The work was supported by the research Grants (BT/PR-9264/AGR/02/ 406(04)/2007 and BT/PR10787/AGIII/103/883/2014) by the Department of Biotechnology, Govt. of India to SKM. We also acknowledge the Fund for Improvement of S&T Infrastructure in Universities and Higher Educational Institutions (FIST) for 2017 by the Department of Science & Technology, Government of India) to Amity Institute of Biotechnology, Amity University Rajasthan (AUR) (SR/FST/LS-1/2017/56).
Author information
Authors and Affiliations
Contributions
A.J. developed the ideas, designed, and supervised all the experiments, and prepared the final manuscript. Y.P. performed most of the experiments, analyzed the data, and prepared the final manuscript. S.D. and S.K.M. supervised all the physiological and biochemical experiments. N.V. performed most of the experiments and analyzed the data. A.H. analyzed the molecular data. X.W. prepared the figures. M.D.G. carried out the literature search.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Poli, Y., Nallamothu, V., Hao, A. et al. NH787 EMS mutant of rice variety Nagina22 exhibits higher phosphate use efficiency. Sci Rep 11, 9156 (2021). https://doi.org/10.1038/s41598-021-88419-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-88419-w
- Springer Nature Limited