Abstract
Pseudomonas aeruginosa uses quorum sensing (QS) to modulate the expression of several virulence factors that enable it to establish severe infections. The QS system in P. aeruginosa is complex, intricate and is dominated by two main N-acyl-homoserine lactone circuits, LasRI and RhlRI. These two QS systems work in a hierarchical fashion with LasRI at the top, directly regulating RhlRI. Together these QS circuits regulate several virulence associated genes, metabolites, and enzymes in P. aeruginosa. Paradoxically, LasR mutants are frequently isolated from chronic P. aeruginosa infections, typically among cystic fibrosis (CF) patients. This suggests P. aeruginosa can undergo significant evolutionary pathoadaptation to persist in long term chronic infections. In contrast, mutations in the RhlRI system are less common. Here, we have isolated a clinical strain of P. aeruginosa from a CF patient that has deleted the transcriptional regulator RhlR entirely. Whole genome sequencing shows the rhlR locus is deleted in PA80 alongside a few non-synonymous mutations in virulence factors including protease lasA and rhamnolipid rhlA, rhlB, rhlC. Importantly we did not observe any mutations in the LasRI QS system. PA80 does not appear to have an accumulation of mutations typically associated with several hallmark pathoadaptive genes (i.e., mexT, mucA, algR, rpoN, exsS, ampR). Whole genome comparisons show that P. aeruginosa strain PA80 is closely related to the hypervirulent Liverpool epidemic strain (LES) LESB58. PA80 also contains several genomic islands (GI’s) encoding virulence and/or resistance determinants homologous to LESB58. To further understand the effect of these mutations in PA80 QS regulatory and virulence associated genes, we compared transcriptional expression of genes and phenotypic effects with isogenic mutants in the genetic reference strain PAO1. In PAO1, we show that deletion of rhlR has a much more significant impact on the expression of a wide range of virulence associated factors rather than deletion of lasR. In PA80, no QS regulatory genes were expressed, which we attribute to the inactivation of the RhlRI QS system by deletion of rhlR and mutation of rhlI. This study demonstrates that inactivation of the LasRI system does not impact RhlRI regulated virulence factors. PA80 has bypassed the common pathoadaptive mutations observed in LasR by targeting the RhlRI system. This suggests that RhlRI is a significant target for the long-term persistence of P. aeruginosa in chronic CF patients. This raises important questions in targeting QS systems for therapeutic interventions.
Similar content being viewed by others
Introduction
Quorum sensing (QS) is a cell density dependent signal transduction mechanism used by prokaryotes to regulate population level gene expression1. Pseudomonas aeruginosa has a sophisticated QS network that is orchestrated by three main signalling pathways namely las, rhl and pqs2,3. Each of these pathways are composed of a synthase protein that produces signal molecules called autoinducers (AI). AI molecules bind to the corresponding receptor proteins, this AI-ligand bound receptor complex regulates several genes in bacteria4. Two of the most well-studied QS systems in P. aeruginosa are the acyl homoserine lactone based las and rhl systems. The autoinducer synthase LasI catalyses the formation of the autoinducer N-(3-oxododecanoyl)-L-homoserine lactone (C12-HSL) which binds to the transcription regulator LasR to form a transcriptional activator complex (LasR:C12-HSL) that regulates the expression of QS regulatory genes rhlR and pqsR of the rhl and pqs pathways, respectively5,6,7. The transcription regulator RhlR forms a complex with the rhlI catalysed product N-butanoyl-L-homoserine lactone (C4-HSL). The RhlR:C4-HSL complex binds to conserved rhl sites in the promoter regions of the target genes including rhlI to trigger a second autoinduction-forward loop8,9. In studies with standard laboratory strains like PAO1 and PA14, mostly in nutrient rich medium, LasR has been shown to directly induce las, rhl and pqs systems thereby often being considered as the de facto QS regulator controlling QS mediated virulence in P. aeruginosa5,10,11.
QS regulated virulence is mediated by multiple factors including pyocyanin, rhamnolipid and exopeptidases12. Pyocyanin, one of the main phenazines found in CF patients has redox activity that can increase the production of reactive oxygen (ROS) species thereby producing hydrogen peroxide with detrimental effects to the host cell13. Exopeptidases like elastase have been shown to degrade opsonizing lung surfactant proteins14 and inactivate antimicrobial peptides LL-3715. Proteases have also been shown to cause damage to the lung epithelial lining and the degradation of complement proteins, fibrinogen and immunoglobulins16,17,18. Another widely studied QS metabolite, rhamnolipid, has been shown to cause necrosis of human polymorphonuclear leukocytes19, to support bacterial twitching and swarming motilities20,21 and to maintain biofilm architecture22,23. All these QS regulated metabolites are essential to early establishment of P. aeruginosa infection in lungs24,25,26 with mutations in QS phenotypes becoming common as the P. aeruginosa adapts to their host environment during chronic stages of infection27,28,29.
The importance of QS has also been demonstrated in transmissible lineages of P. aeruginosa. The hypervirulent Liverpool epidemic strain (LESB58) was shown to upregulate and overproduce QS phenotypes, potentially contributing for enhanced morbidity and successful spread of the LES lineage throughout the CF population in the UK30,31. Therefore, with QS being strongly associated with virulence in clinical outcomes, ongoing research has been investigating alternative methods to attenuate bacterial virulence, especially now when antibiotics are a limited resource. A promising alternative to antibiotics has been in the use of anti-QS compounds such as trans-cinnamaldehyde (CA) and salicylic acid (SA)32 which reduce expression of QS associated virulence factors by targeting the master regulator, LasR, in P. aeruginosa33,34. Although these LasR anti-QS inhibitors have worked well with laboratory strains by decreasing the QS mediated virulence factor expression35,36, its effect on isolates collected from chronic patients has not been fully validated. There is a growing consensus of LasR mutations are frequent among chronic clinical isolates29,37,38. These LasR mutants were shown to induce exaggerated host inflammatory response, neutrophil degradation and immunopathology in animal models38. Paradoxically, it has been shown that in these LasR mutants, the QS hierarchy has shifted such that RhlR QS is independent of direct LasR regulation39. These findings therefore question how rigidly maintained is the established QS hierarchy in P. aeruginosa and whether LasR is the best target in anti-QS strategies.
In this study we have characterised a clinical isolate of P. aeruginosa that is deficient in RhlRI QS. RhlR mutants are very rarely isolated from CF patients and are mostly associated with hypermutability28. To understand the effect of this RhlR mutant on QS regulation and virulence gene expression, we have compared this isolate (PA80) with isogenic QS mutants in P. aeruginosa PAO1. We show that PA80 is completely deficient in expression of QS regulatory genes and QS regulated virulence genes. PA80 is closely related to the hypervirulent Liverpool epidemic strain LESB58. We attribute the complete loss of virulence expression and QS related activity to the inactivation of the RhlRI system as PA80 does not maintain any non-synonymous mutations in common pathoadaptive genes typically mutated in P. aeruginosa CF isolates. PA80 is an example of the diversity of long-term pathoadapted P. aeruginosa CF isolates.
Results
P. aeruginosa CF isolate PA80 is a RhlRI mutant
Whole genome sequencing (WGS) was carried out on P. aeruginosa isolate PA80, a clinical isolate collected from the sputa of a Cystic Fibrosis patient attending Regional Cystic Fibrosis Centre in Northern Ireland24. de novo assembly of PA80 generated 97 contigs, the genome size of PA80 is 6,500,365 bp with an average G.C content of 66.39%, similar to other published P. aeruginosa genomes. Whole genome alignment (WGA) showed that PA80 is 91.26% identical to the reference strain PAO1 and 90.74% similar to PA14. However, a high similarity was found between PA80 and P. aeruginosa Liverpool epidemic strain LESB58 with a 99.53% sequence match (Fig. 1B). PA80 lacks the large inversion (~ 4.5 Mb) observed in LESB58 (Fig. 1A). To understand the lineage of PA80 we performed a pan-genome analysis in comparison with 265 publicly available genomes from Pseudomonas.com. The core genome was determined using the core gene alignment output from roary. Core genome phylogeny reveals two distinct groups of P. aeruginosa, similar to previously published studies40,41,42,43. PA80 is found in a clade with LES isolates LESB58 and LES431 in both the core and accessory genome (Fig. 2A and Fig. S1).
Comparative genomics of PA80. (A) Whole genome alignment of PA80, PAO1 and LESB58. PA80 shares 99.53% sequence identity to LESB58. (B) Whole genome comparison of PA80 with LESB58 and PAO1. LESB58 genomic islands are highlighted in blue and prophage region in red. Coordinates were mapped to LESB58 genome from Jani et al., 201688. PA80 contains only one unique region relative to LESB58. PAO1 lack several of the GI’s and prophages identified in LESB58. (C) Genomic context of the main non-synonymous mutations in PA80. Shaded red indicates complete deletion of the rhlR gene.
Core genome phylogeny of P. aeruginosa. (A) Maximum likelihood phylogenetic tree of P. aeruginosa core genome. PA80 clusters with hypervirulent Liverpool Epidemic strains LESB58 and LES431. The widely referenced PAO1 strain is distinctly separate from more virulent strains that have been isolated from Cystic Fibrosis patients. Important P. aeruginosa strains are highlighted by red dot, strain PA80 is indicated by a cyan dot. The number of genes determined in the core across 95% of the P. aeruginosa isolates are shown inset in the pie chart. (B) A STRING interaction network to show common pathoadaptive genes (n = 47) that can be mutated in P. aeruginosa isolates from CF patients. Genes are colored based on Pseudocap annotations. Note—the following proteins are not found in the STRING database vqsR, rsaL, pprB, PA4851, PA152, and are not shown in the network.
Figure 2B highlights a list of common pathoadaptive genes that are known to mutate in P. aeruginosa CF isolates during adaptation and evolution to the lung environment. In PA80 we identified several SNPs, most of which are synonymous SNPs found in these common functional pathoadaptive mutations among P. aeruginosa (Table 1). For a more extensive list please see Table S1. However, the main feature of PA80 is the deletion of the rhlR gene. The entire rhlR gene has been deleted along with the C-terminal of rhlB and the N-terminal of rhlI (Fig. 1C). The core genome is generally conserved with sequence diversity only ranging between 0.5 and 0.7%44. Pseudomonas aeruginosa diversification arises from the accessory genome which is reflected in the varying genome sizes ranging from 5.5 to 7 Mbp45. PA80 accessory genome shares > 99% sequence identity with LESB58 and contains several LES genomic island and prophage regions (Fig. 1B). PA80 is larger in genome size compared to PAO1 indicative of a larger accessory genome (Fig. S1A). Phylogeny of the accessory genomes shows that PA80 is most closely related to LESB58 (Fig. 2A). Functional annotation using RAST46 revealed that the region contained 15 genes involved in phage replication, packaging, capsid proteins and lysis; 5 genes involved with copper transport system and 11 genes responsible for resistance to antimicrobial and toxic compounds like cobalt, zinc and mercury (Fig. S1B). This suggests that PA80 is a derivative of the P. aeruginosa LES lineage and to the best of our knowledge this is the first report of a LES rhlR mutant isolated from a CF patient.
A functional autoinducer is essential for QS gene expression
To understand the role of RhlRI in P. aeruginosa QS, we investigated individual isogenic mutants in both the LasRI and RhlRI QS systems in the reference strain PAO1. Inactivation of either las or rhl QS does not significantly affect the growth of PAO1 under phosphate limiting conditions (Fig. 3A) except for a slight lag in ΔrhlI. It has been reported that the rhl system can work independently of las regulation through the observation that LasRI null mutants are frequently isolated from CF patients29,39,47.To assess if these systems can work independently, we examined the gene expression of QS regulatory and virulence associated genes using the PAO1 QS mutants. We show inactivation of RhlRI abolishes expression of both las and rhl regulatory genes (Fig. 3B–E). However, inactivation of lasR did not affect expression of RhlR (Fig. 3C) demonstrating that RhlR expression can be independent of LasRI. Deletion of either LasRI and RhlRI systems can be complemented by restoring a functional protein or in the case of the synthases protein (lasI and rhlI) by exogenous autoinducer molecules48,49. Recent studies have elegantly demonstrated evolutionary trajectories for LasR null mutants where a functional RhlRI system independent of LasRI quickly emerges48,49,50. This was an interesting finding considering the QS hierarchy puts lasR at the top of the QS system with the LasRI activated complex inducing expression of the rhlR regulator. However, we do note that phosphate limitation is known to induce expression of key QS regulatory genes. Our results support recent work by Meng et al., 2020 which report increased expression of rhlR in a LasR mutant strain of PAO1 under phosphate-depleted conditions51. Similar to our results this increased expression was not observed for rhlI (Fig. 3E). Meng et al., 2020 show this increased expression is due to activation by the PhoR/PhoB two- component regulatory system51. The unaltered rhlR expression in the absence of a functional lasR therefore challenges the established QS hierarchy and further supports recent claims of other regulatory pathways capable of inducing rhl QS, by bypassing the las QS system in a nutrient deprived environment50. However, in ΔlasR, expression of rhlI was completely switched off to levels observed for ΔrhlI and all other QS mutants (Fig. 3E).
Inactivation of QS regulatory genes reduces both las and rhl QS systems. (A) Growth phenotypes of PAO1 and isogenic QS regulatory mutants in a phosphate limited media (PPGAS). Relative expression levels of QS regulatory genes (B) lasR, (C) rhlR, (D) lasI, and (E) rhlI in both PAO1 and QS mutants. Relative expression levels were quantified in the stationary phase (indicated by red arrow) of growth by qRT-PCR. Error bars represent S.D. ± (n = 3 biological replicates). All mutant data was analysed relative to PAO1 expression values. Significance was determined by a one-way ANOVA followed by Dunnett’s multiple comparison test (*p < 0.05, **p < 0.01, ***p < 0.001).
To understand the effect of knocking out the different QS systems, we measured the expression of virulence genes under the control of either las or rhl. Rhamnolipids are low molecular weight glycolipids that play important role in P. aeruginosa pathogenesis52. RLs are synthesised de novo in P. aeruginosa by the biosynthetic genes rhlABC that are directly regulated by the RhlRI QS system53. rhlAB is not expressed in ΔrhlR whereas inactivation of ΔlasR does not affect their transcriptional expression (Fig. 4A,B). Expression of the protease LasA and the elastase LasB is positively regulated by LasR54. We show expression of lasA is completely abolished in all QS regulatory mutants (Fig. 4D). Expression of lasB was downregulated in ΔlasR but not completely inhibited as observed in the other mutants (Fig. 4E). It is well established that RhlR mediated activity can be uncoupled from LasR regulation by the isolation of LasR mutants that are RhlRI active28,39. However, we note that signal negative mutants (ΔlasI ΔrhlI) have a significant effect on expression of QS regulated genes (Fig. 4). Expression of lasA, lasB and rhlABC is completely inhibited in signal negative ΔlasI and significantly reduced in ΔrhlI (Fig. 4). This suggests that production of the signal molecules may be the most critical part for a functional QS system.
Inactivation of LasRI and RhlRI QS systems reduces virulence factor expression. Relative expression levels of virulence factor genes (A) rhlA, (B) rhlB, (C) rhlC, (D) lasA and (E) lasB. Relative expression levels were quantified in the stationary phase (indicated by red arrow) of growth by qRT-PCR. Error bars represent S.D. ± (n = 3 biological replicates). All mutant data was analysed relative to PAO1 expression values. Significance was determined by a one-way ANOVA followed by Dunnett’s multiple comparison test (*p < 0.05, **p < 0.01, ***p < 0.001).
Deletion of RhlR inhibits QS activity in clinical isolate PA80
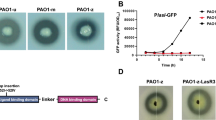
We also determined the effect of the rhlR deletion in PA80 by examining the expression of QS regulatory genes and virulence gene expression. Similar to the signal negative ΔrhlI, the clinical isolate PA80 too showed a significant delay in log phase growth compared to PAO1 (Fig. 5A). This could suggest that a functional rhl system is important for the growth of P. aeruginosa. PAO1 shows a typical AHL-dependent expression of QS genes, where expression correlates with an increase in cell density (Fig. 5). As expected, we observe expression of lasR in PA80 during the stationary phase similar to PAO1 (Fig. 5B). The expression of both rhlR and rhlI was completely abolished in PA80 (Fig. 5C,E). Interestingly we did not detect expression of lasI in PA80 (Fig. 5D), this was surprising as no mutations were detected in lasI or upstream in the promoter region (Table 1). In correlation with the lack of QS gene expression in PA80, the virulence factor expression was completely switched off (Fig. 6) while PAO1 expressed the virulence associated genes in a typical growth phase dependent manner (Fig. 6)32.
CF isolate PA80 does not express any QS regulatory genes. (A) Growth of PAO1 in comparison with rhlR mutant isolate PA80. Relative expression of PAO1 and PA80 during log and stationary growth phases. Relative expression levels of QS regulatory genes (B) lasR, (C) rhlR (D) lasI and (E) rhlI in both PAO1 and PA80. Expression levels are shown as the mean relative expression ratios to log phase levels (i.e. 6 h). Error bars represent the S.D (biological triplicates). Data was analysed using a one-way ANOVA followed by a Dunnett’s multiple comparison test comparing each time point to log phase levels (i.e. 6 h) (*p < 0.05, **p < 0.01, ***p < 0.001).
Repression of QS genes attenuates virulence factor production in PA80. Relative expression levels of QS regulated virulence genes (A) rhlA, (B) rhlB, (C) rhlC and (D) lasA and (E) lasB in both PAO1 and PA80. Expression levels are shown as the mean relative expression ratios to log phase levels (i.e. 6 h). Error bars represent the S.D (biological triplicates). Data was analysed using anone-way ANOVA followed by a Dunnett’s multiple comparison test comparing each time point to log phase levels (i.e. 6 h) (*p < 0.05, **p < 0.01, ***p < 0.001).
Inactivation of QS attenuates virulence factor production
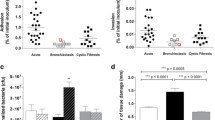
We measured the production of extracellular virulence factors in both PAO1 and PA80 (Figs. 7, 8). In correlation with the lack of rhamnolipid gene expression, PA80 does not produce any rhamnolipids (Fig. 7). Production of extracellular rhamnolipids was quantified by HPLC–MS from crude extracts for both PAO1 and PA80. We also quantified rhamnolipid production in both QS and rhlABC mutants (ΔlasR, ΔlasI, ΔrhlR, ΔrhlI, ΔrhlA, ΔrhlB and ΔrhlC). PAO1 produced both mono- and di-rhamnolipids with the di-rhamnolipid congeners Rha-Rha-C10-C10 (m/z 649) and Rha-Rha-C10-C12 (m/z 677) being most abundant (Fig. 7A). Production of rhamnolipids is under the control of the rhl QS system, inactivation of lasRI does not affect extracellular production of rhamnolipids (Fig. 7A). However, inactivation of rhlR abrogates rhamnolipid production in both PA80 and ΔrhlR. While di-RLs are predominant, its production is dependent on the conversion of mono-RLs to di -RLs. Therefore, when we delete rhlC, which is responsible for di-RL production, we only detect mono-RL congeners (Fig. 7B). These data clearly support that production of RLs is stringently regulated in P. aeruginosa55 and rhlR is essential for its production.
PA80 does not produce rhamnolipids. (A) HPLC–MS chromatogram of rhamnolipids (RLs) detected in PAO1, ΔlasR, ΔrhlR and PA80. Rhamnolipids are produced independent of lasR but it is essential to have a functional rhlR present. PA80 lacking rhlR does not produces RLs. (B) Comparison of the RLs congeners produced in wild-type PAO1 and mutant strains. ΔlasR produces RLs congeners in the same composition as PAO1.
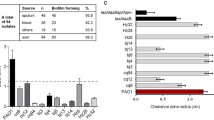
PA80 has reduced virulence factor production. (A) Swarming motility of PAO1 and PA80. (B) PA80 does not reduce surface tension due to lack of RLs produced. PAO1 typically reduces surface tension of water to ~ 38 (N/m). Production of (C) elastase (D) protease (E) pyocyanin and (F) biofilm in PAO1 and PA80.
Similar to other pathoadapted P. aeruginosa isolates, PA80 has reduced production of several virulence factors56. RLs are essential for motility in P. aeruginosa, as expected PA80 exhibits no motility (Fig. 8A). Consistent with the lack of RL production in PA80, is the inability to decrease the surface tension of water (Fig. 8B). We also show that PA80 does not produce virulence associated secretions for elastase, protease and pyocyanin (Fig. 8C–E). PA80 is also a weak biofilm producer in comparison to PAO1 (Fig. 8F). Loss of these virulence phenotypes has been reported for both LasR−ve and RhlR−ve mutants, Chen et al., 2019 show that deletion of lasR in PAO1 and mutation of RhlR in strain E80 abolishes pyocyanin and protease production49. Similar phenotypes have been reported Kostylev et al., 2020 with LasR mutants50. Here we show loss of function for both las and rhl controlled virulence factors even though PA80 maintains a functional LasRI system, thereby suggesting that RhlRI could be the linchpin in the P. aeruginosa QS hierarchy.
Discussion
A hallmark of P. aeruginosa evolution in the CF lung is loss of function in key virulence associated phenotypes such as quorum sensing (QS). It is now well established that during chronic CF infections, P. aeruginosa can rewire its QS hierarchy such that LasR, the prime regulator that directly or indirectly controls the expression of other key regulators in the QS pathway is no longer functional49. This enables RhlR to act independently of LasR regulation and makes its vital for long term survival within the host49. LasR null mutants are commonly isolated from chronic CF infections27,29,39,57,58, these isolates typically have attenuated virulence but maintain an active RhlRI QS system59,60. As a result, LasR was considered a viable therapeutic target for anti-virulence drugs61. Emergence of LasR mutants in P. aeruginosa CF isolates is regarded as evidence for adaptive evolution to the CF lung environment56 however the benefit of these mutants still remains unclear. It has been suggested that these loss of function in QS mutants may have an advantage in nutrient acquisition in the CF lung environment, act as social cheats or maybe a functional QS system is only required for establishing infection62. Regardless, isolation of LasR mutants in P. aeruginosa are frequent among chronic CF infections, whereas mutations in RhlRI are not common with only a few reported cases from very late stages of infection28.
Here we report a P. aeruginosa isolate PA80 that is a RhlR null LasR+ve mutant. Based on phylogeny we propose that PA80 belongs to the divergent lineage of the Liverpool epidemic Strain (LES) (Fig. 1), the most common strain to infect the CF population in the UK63. PA80 shares 99.53% core genome identity with LESB58. Functional annotation of the accessory genome in PA80 revealed the presence of several prophage genes which have been linked to enhanced competitiveness and fitness of LESB5863. These prophage elements are known to be important for early stage infection63,64 while also being significant reservoirs for horizontal gene transfer (HGT) of several antimicrobial resistant determinants which is characteristic among P. aeruginosa strains65. The unique feature of PA80 is that this strain is a highly pathoadapted LES variant that has completely inactivated its QS system due to the deletion of the rhlR locus.
WGS identified several mutations in PA80, most of which were synonymous mutations among several key global regulators (Table 1). It is interesting to note, PA80 contains a predicted functional MexT by the clean deletion of the duplicated 8 bp sequence found in PAO1 (Table 1). In several PAO1 lineages the presence of this duplicated 8 bp produces a truncated MexT polypeptide (89aa) which lack six terminal residues of the HTH DNA binding domain, thus in P. aeruginosa strains containing this 8 bp duplication, the MexEF-OprN operon is not expressed66. PA80 also contains the common MexS-D249N mutation observed in other clinical P. aeruginosa isolates67. Most of the mutations observed in PA80 are synonymous with no major deleterious or loss of function effects with the exception of the targeted deletion of the rhlR locus. To date, the paradigm for P. aeruginosa evolution in the CF lung has more or less adhered to the following pathway: (1) initial colonisation of wild type P. aeruginosa, (2) emergence of LasR−VE cheats with RhlRI functionality via mutation of mexT, (3) PQS null mutants in LasR−VE MexT−VE isolates. PA80 is a late-stage chronic CF isolate, it represents a distinct pathoadapted variant of LES in that it does not contain any of the expected genotypes as mentioned above, rather the rhlR locus seems to be targeted for complete deletion. Not only are RhlR mutants rare, they are typically restricted to late stage chronic CF infections and strongly correlate with precursor lasR mutations28. RhlRI is critical for the regulation of several virulence associated phenotypes that are required for colonisation and acute infection in CF patients48.
One of the most striking features of PA80 is that there are no mutations associated in the lasR-lasI genomic region, whereas the rhlR locus has been targeted for deletion alongside the N-terminal of rhlI and C-terminal of rhlB (Fig. 1). To understand the loss of function in RhlRI QS in PA80, we initially analysed PAO1 isogenic QS mutants for expression and production of QS regulated phenotypes. Consistent with previous reports of RhlR QS independent of LasR regulation we observed expression of rhl regulated rhamnolipid and elastase genes in ΔlasR. We show that expression of the virulence factors rhamnolipids and elastase are not affected by inactivation of LasR. However, we do note that inactivation of the AHL signal gene lasI had a significant impact on both QS regulatory and virulence gene expression in PAO1. Much less attention has been focused on inhibition of QS signal molecules as a therapeutic target in comparison to LasR. However, as we learn more about the intricate and complex network of QS in P. aeruginosa it is obvious that LasR is not a viable therapeutic target therefore research has now shifted to RhlR as a potential target68,69.
We observe complete downregulation of LasRI in PA80 even though there are no mutations in this genomic region or with any associated global regulators that drive expression of LasR (Table 1). This correlated with the downregulation of lasB which is stringently regulated by the las system70,71. This suggests that inactivating the RhlRI system has significant impact on LasRI expression. We also show this in the isogenic QS mutants in wild-type PAO1, inactivation of either rhlR or rhlI represses expression of both lasRI (Fig. 3). We do not know the mechanism by which inactivation of RhlRI in PA80 has abolished LasRI activity. There could be unknown regulatory elements that maybe upregulated or repressing expression of lasRI—however this does suggest that PA80 is a highly pathoadapted strain that has loss of function in both las and rhl QS systems. In several LasR mutants isolated from the CF lung environment, the QS hierarchy seems to be readily reprogrammed such that RhlRI independent from LasR can be achieved by simple genetic changes in the global regulator mexT49,72. While it seems typical to rewire the QS circuitry such that RhlRI is independent of a functional las system, the inverse however does not seem to be true.
Recently Chen et al. 201949 demonstrated in vitro by experimental evolution that RhlR mutants do not readily emerge in LasR−VE mutants, rather mutations emerge in the non-AHL Pseudomonas quinolone signal (PQS) and the related 2-alkylquinolone (HHQ) molecules. Chen et al., 2019 also showed that constructed RhlRI null mutants have a significant fitness cost that is outcompeted by the wild-type and are completely sensitive to cyanide which is synthesised by P. aeruginosa. Production of cyanide in P. aeruginosa is considered a metabolic policing mechanism by which it monitors cheaters (mutant in public goods) which are typically more susceptible to cyanide toxicity in comparison to wild-type73. We also observe a significant fitness cost in PA80 when grown in phosphate limiting peptide rich media (Fig. 5). Interestingly we also show that ΔrhlI-PAO1 has a similar growth defect in comparison to wild-type PAO1 (Fig. 3). This suggests that RhlRI is essential in P. aeruginosa even in heterogenous populations that undergo rapid evolutionary changes during CF infection and evolution. A functional RhlRI system is necessary to allow for the emergence of LasR cheats, where such strains become dependent on C4-HSL secreted by cells with QS intact, as a result of this dependence mutations are less likely to arise in RhlRI. However, this selection pressure is most likely observed during the early stages of infection where P. aeruginosa undergoes rapid microevolution. Late-stage chronic infections are characterised by high genotypic and phenotypic diversity, that is reflective of highly adapted lineages that persist long term. These variants have attenuated virulence, that helps evade immune recognition and enables long term P. aeruginosa persistence and survival74.
Our study is significantly limited to the examination of a single isolate and standard laboratory growth conditions compared to the dynamic selection pressures and polymicrobial conditions of the CF lung environment. However, we can glean some significant insights to P. aeruginosa evolution from PA80. PA80 was isolated from late-stage chronic infection, we however do not know anything about the evolutionary dynamics that selected for the deletion of the rhlR locus. It is unlikely that PA80 would emerge in isolation, most likely there would have been a divergent clonal population with several mutant populations/cheats (i.e. LasR−VE, LasR−VEMexT−VE) to enable the emergence of cheats while in the presence of other QS active wild-types and intermediates.
Nonetheless, PA80 provides a unique evolutionary trajectory which, to our knowledge has not been reported to date. This is an important discovery as the focus shifts from developing inhibitors that target LasR to RhlR. Our data show the loss of function rhlR does render PA80 avirulent in both las and rhl regulated virulence. This taken together with the fact that a functional RhlRI is essential during early infection and cannot be easily rewired as seen with LasR, RhlR may be a better therapeutic target. However, mutants can arise in RhlR, therefore targeted inhibition should be aimed at early-stage acute infections rather than in long term chronic infections. PA80 provides another genome available for comparison of long term pathoadapted P. aeruginosa isolates from the CF lung. In the CF lung P. aeruginosa undergoes an evolutionary pathway that can take several directions, however as we build a better genomic map of these adaptations, it is clear that a multi-target approach is needed to treat the highly divergent P. aeruginosa lineages.
Materials and methods
Bacterial strains and growth media
The clinical isolate, PA80 was obtained from the culture bank maintained at the Ulster University in Coleraine campus. The isolate was initially collected from a cystic fibrosis patient (aged 20) attending the CF clinic in the Belfast City Hospital24. All the P. aeruginosa PAO1 mutants (ΔlasR, ΔlasI, ΔrhlR, ΔrhlI, ΔrhlA, ΔrhlB, ΔrhlC) were purchased from the P. aeruginosa mutant library maintained at Manoil Laboratory in the University of Washington75. The well-studied and fully sequenced P. aeruginosa PAO1 was used as the control strain in the experiments. Pseudomonas aeruginosa PAO1 is also QS proficient32. All the overnight cultures were prepared from the − 80 °C frozen culture stocks either in a LB or nutrient broth and cultivated under at 37 °C with shaking at 180 rpm. The overnight culture was used to inoculate the phosphate limited proteose-peptone-glucose-ammonium-salts (PPGAS) medium76 for growth and gene expression experiments. All experimental reagents were purchased from the Sigma-Aldrich, UK unless mentioned otherwise. Experiments performed in biological triplicates.
Nucleic acid extraction and quality-check
Genomic DNA: gDNA was isolated from a freshly prepared overnight culture using the Wizard Genomic DNA Purification Kits (Promega) following the manufacturer’s protocol. The Nanodrop 1000 spectrophotometer (Thermo Fisher Scientific) was used for DNA quantification and purity assessment. High quality DNA (A260/280 and A260/230 ~ 1.8 and ~ 2.0 respectively) were only considered for further experiments and were stored in small aliquots in nuclease free water at − 20 °C.
Total RNA: The cell pellets from the different bacterial cultures were collected at the experimental time points for RNA isolation using the JetGene RNA Purification Kit (Thermo Fisher Scientific). In brief, the cells were lysed in a buffer solution containing 1X TE buffer, 20 mg/ml proteinase K (Promega) and 15 mg/ml lysozyme. The lysed samples were then transferred to a 2 ml Lysing Matrix A tube (MP Biomedicals) containing RLT buffer from the kit and β-Mercaptoethanol (10 µl/ml). The contents in the Matrix A tube were homogenised using the FastPrep FP 200 cell disrupter at speed 5.5 for 30 s. Following centrifugation, the supernatant was transferred to RNeasy spin columns (Qiagen) for DNase treatment. Another round of lysing buffer treatment was performed before doing a second DNase treatment. The RNA extracted was quantified and assessed for purity similarly to DNA. The integrity of the RNAs isolated were ascertained through visualization of two sharp bands corresponding to 16S and 23S rRNA under UV light following electrophoretic separation on agarose gel. Additionally, the RNA samples were also checked for integrity with Agilent 2100 Bioanalyser.
Reverse transcription quantitative polymerase chain reaction
500 ng of target mRNA was added to a reaction mix consisting of 20–250 ng of random primers (Promega) and 10 mM dNTPs (Invitrogen). The reaction was incubated for 5 min at 65 °C for cDNA synthesis. After the incubation, the mix was centrifuged briefly and 5X stand buffer, 0.1 M DTT and RNase out (Invitrogen) were added in volumes corresponding to final concentrations of 1X, 10 µM and 40 units respectively. A second incubation at 25 °C for 2 min was performed before addition of Superscript II Reverse Transcriptase (200U final concentration). This was followed by a series of incubation steps: 25 °C for 10 min, 42 °C for 50 min and 70 °C for 15 min to give the first strand cDNA. The cDNA synthesis was performed for all biological replicates. A negative control without reverse transcriptase was added in every run. All newly synthesized cDNAs were stored at − 20 °C prior to use as template for real time PCR amplification.
Real time qPCR was performed with the ROCHE LightCycler LC480 system using SYBR-Green. Before the mRNA transcripts were quantified, the qPCR primers for the target genes were validated for specificity by generating a PCR calibration curve using PAO1 gDNA. For the mRNA quantification study, only those primers that gave a slope value of − 3.1 to − 3.6 and amplification efficiencies of 90–110% were selected. The primers binding specificity was further confirmed by the presence of a single sharp peak in the melt curve.
Each of qPCR reaction mix contained 2X SYBR Green master mix, 1 µM of forward and reverse primers, cDNA template and nuclease free water to make up the 10 µl volume. Negative controls in form of no reverse transcriptase (NRT) and no template DNA (NTD) and positive control in form of gDNA were included for accuracy. Cut-off values for residual gDNA and negative controls were set at greater than 35 and 40 cycles respectively. The qPCR amplification conditions used were: initial denaturation for 5 min at 95 °C, 40–50 cycles of denaturation for 10 s at 95 °C, annealing for 10 s at 59 °C, extension for 10 s at 72 °C.
Relative gene expression data analysis
The reference gene validation and selection were done using six candidate genes (gyrB, proC, cysG, rpoD, rpoB and 16S). Three different and independent software packages were used to select for the most stable genes as previously reported from our lab32. Based on the algorithms of these programs, the candidate genes rpoD and proC were selected as the most stable genes for use as reference genes in P. aeruginosa PAO1.
System (LC480 software, version 2) generated analysis was performed on the real time qPCR data following the steps outline by Ahmed et al. (2019)32. In brief, relative quantities (RQ) values were calculated using the threshold values (Cq) of the technical replicates. The RQ values of the target genes were divided by the geomean of the reference genes to generate the normalised relative quantity values (NRQ). The relative expression value at the early log (6 h) analysis was used as experimental calibration value to calculate the relative expression of the target genes at the different experimental time points for plotting.
QS virulence factors quantification
Overnight cultures of PAO1 and PA80 were grown in PPGAS medium for 24 h. Cell-free supernatants were collected through centrifugations, and filter sterilised for use in the following assays:
LasA protease Amount of protease production in the culture was assessed by incubating the reaction mixture containing 0.1 ml of the supernatant and 0.8% azocasein (in 500 µl of 50 mM K2HPO4) at 25 °C for 3 h. The reaction was stopped by first adding 0.5 ml of 1.5 M HCl and then cooling it on ice for 30 min. The tube was centrifuged, and the supernatant transferred in a fresh tube. 1 N NaOH was added to the collected supernatant in equal volumes and the concentration of the acid soluble azopeptides was measured at 440 nm using a UV–vis spectrophotometer.
LasB elastase In this assay, 2 ml reaction buffer containing 100 mM Tris-HCl, 1 mM CaCl2 and the enzyme substrate elastin-congo red was incubated with 1 ml of the overnight culture supernatant for 3 h at 37 °C for 3 h at 180 rpm. The reaction was stopped by first adding 2 ml of 0.7 M sodium phosphate buffer (pH 6) and then cooling it on ice for 15 min. The mixture was centrifuged, and the supernatant collected for spectrophotometric measurement of the congo-red dye released due to elastase activity in the supernatant at 495 nm.
Pyocyanin In a 50 ml tube, 7.5 ml of the culture supernatant was mixed vigorously with 4.5 ml of chloroform till the colour changed to greenish-blue. The mixture was spun and 3 ml of the resulting blue/pink colour solution from the bottom layer was transferred to a fresh tube containing 1.5 ml of 0.2 M HCl. The tube was vortexed vigorously and the resulting pink colour solution was collected for spectrometric measurement at 520 nm. The concentration (µg/ml) of pyocyanin was calculated as OD X 17.07277.
HPLC–MS analysis of rhamnolipid production
Estimation through surface tension reduction ability: 15 ml of the cell free culture supernatants were collected from the different experiment time points of growth. The surface tension was measured using the Du Nouy ring method with a digital tensiometer (Kruss, K10ST, Hambury, Germany)78. The ability of the supernatant to reduce the surface tension of the medium is indicative of the presence of the surface-active reducing agent rhamnolipid.
High-Performance Liquid Chromatography Mass Spectrometry/Mass Spectrometry (HPLC MS/MS): At first, crude rhamnolipid was extracted following the protocol outline by Smyth et al. (2010)79. Briefly, the cell-free supernatant from the different cultures were collected and acidified with 32% HCl to pH ~ 2. The acidification made the rhamnolipid less soluble in the aqueous state by causing protonation. The acidified supernatant was shaken vigorously in a separating funnel with equal amount of ethyl acetate until two distinct layers became visible; the aqueous layer containing unwanted compounds and the ethyl acetate organic layer containing the rhamnolipid. The rhamnolipid containing organic phase was dried with anhydrous MgSO4 and then filtered to collect the filtrate in a round bottomed flask. The organic solvent was evaporated in a rotary evaporator (Buchi, Flawil, Switzerland) to leave a yellowish oily residue containing the crude rhamnolipid.
The crude rhamnolipid was purified using solid phase extraction by passing the samples through a conditioned Strata SI-1 Silica (55 µm, 70A) Giga tubes (Phenomenex). Solvent mixture of chloroform and methanol in 5:0.3 ratio was passed through the column to elute the mono-rhamnolipids from the samples. The same mix again but in 5:0.5 ratio was now passed to elute the di-rhamnolipids from the samples, leaving any remaining impurities trapped in the column.
The pure rhamnolipid extract was analysed for congener composition using an LCQ quadrupole ion trap with a negative ESI interface linked to a Thermofisher spectra system HPLC as explained earlier by Ahmed et al., (2019)32.
Genome assembly and comparative genomics
The PA80 whole genome sequence was provided by MicrobesNG (http://www.microbesng.uk) which is supported by the BBSRC (grant number BB/L024209/1). The PA80 gene sequence has been submitted to GenBank (PRJNA675745) and is now publicly available. The PA80 genomic DNA library was prepared using Nextera XT Library Prep Kit (Illumina, San Diego, USA) with slight modifications. Hamilton Microlab STAR automated liquid handling system was used for DNA quantification and library preparation. The pooled libraries were quantified using the Kapa Biosystems Library Quantification Kit for Illumina on a Roche light cycler 96 qPCR machine and sequenced on the Illumina HiSeq using a 250 bp paired end protocol. Reads were adapter trimmed using Trimmomatic 0.3v software and for de novo assembly SPAdes v3.7 was used. The total number of contigs in the PA80 genome assembly was 97. The number of contigs of length ≥ 0 bp and length ≥ 1000 bp were 145 and 90 respectively. The assembled contigs were then annotated and aligned with the reference PAO1 genome (GCF_000006765.1) using BWA-MEM80 and variant calling was performed using VarScan and annotated using Prokka 1.11. Only for MexT and MexS, the PA14 (GCF_006974045.1) genome was used for reference as previously recommended67. From the genome sequences, using NCBI local blast (BLAST v2.10) the specific gene sequences were extracted, and alignments were compared. Using the BAM alignment file generated by BWA-MEM algorithm, variants like SNP, insertion and deletions were identified using Mega-X software.
From the mapping statistics it was found that a large portion (8.74%) of the raw reads remained unmapped. This was performed using BWA-MEM tool with P. aeruginosa PAO1 as the reference. Hence from the alignment files the unmapped reads were extracted and was assembled into contigs using the spades81 tool. The contigs were then aligned against the NCBI nucleotide database using the BLASTN tool and was found to match mostly with the Pseudomonas aeruginosa LESB58 genome. The assembled contigs were annotated using RAST46 and was functionally annotated using the associated SEED viewer82. Later a de novo assembly was generated using the raw reads using spades and it was annotated using RAST and SEED viewer. In addition, the raw reads were also aligned with the publicly available LESB58 (GCF_000026645.1) and PA14 reference genome using bwa80 and samtools83 to generate mapping statistics. Whole genomes were aligned using MAUVE multiple genome alignment. Pseudomonas aeruginosa genomes were downloaded from NCBI assembly. Genome assemblies were annotated with PROKKA and provided as input to Roary. Pangenome analysis was carried out using Roary version 3.12.084. Roary was run using default parameters except for the following: -e -n (to produce alignments with MAFFT) and -i 95. The accessory genome phylogeny was visualised in iTOL using the accessory_binary_genes_fa.newick file output from Roary. Genomic islands and prophages were predicted with IslandViewer85 and PHASTER86 respectively. BLAST Ring Image Generator (BRIG)87 was used to compare P. aeruginosa genomes and visualise mobile genetic elements.
References
Swift, S. et al. Quorum sensing as a population-density-dependent determinant of bacterial physiology. Adv. Microb. Physiol. https://doi.org/10.1016/S0065-2911(01)45005-3 (2001).
Lee, J. et al. A cell-cell communication signal integrates quorum sensing and stress response. Nat. Chem. Biol. 9, 339–343 (2013).
Lee, J. & Zhang, L. The hierarchy quorum sensing network in Pseudomonas aeruginosa. Protein Cell 6, 26–41 (2014).
Smith, R. S. & Iglewski, B. H. Pseudomonas aeruginosa quorum sensing as a potential antimicrobial target. J. Clin. Investig. https://doi.org/10.1172/JCI200320364 (2003).
Pesci, E. C., Pearson, J. P., Seed, P. C. & Iglewski, B. H. Regulation of las and rhl quorum sensing in Pseudomonas aeruginosa. J. Bacteriol. https://doi.org/10.1128/jb.179.10.3127-3132.1997 (1997).
Medina, G., Juárez, K., Valderrama, B. & Soberón-Chávez, G. Mechanism of Pseudomonas aeruginosa RhlR transcriptional regulation of the rhlAB promoter. J. Bacteriol. 185, 5976–5983 (2003).
Latifi, A., Foglino, M., Tanaka, K., Williams, P. & Lazdunski, A. A hierarchical quorum-sensing cascade in Pseudomonas aeruginosa links the transcriptional activators LasR and RhIR (VsmR) to expression of the stationary-phase sigma factor RpoS. Mol. Microbiol. https://doi.org/10.1046/j.1365-2958.1996.00063.x (1996).
Pearson, J. P., Passador, L., Iglewski, B. H. & Greenberg, E. P. A second N-acylhomoserine lactone signal produced by Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA https://doi.org/10.1073/pnas.92.5.1490 (1995).
Déziel, E., Lépine, F., Milot, S. & Villemur, R. rhlA is required for the production of a novel biosurfactant promoting swarming motility in Pseudomonas aeruginosa: 3-(3-hydroxyalkanoyloxy)alkanoic acids (HAAs), the precursors of rhamnolipids. Microbiology https://doi.org/10.1099/mic.0.26154-0 (2003).
Tan, M. W., Rahme, L. G., Sternberg, J. A., Tompkins, R. G. & Ausubel, F. M. Pseudomonas aeruginosa killing of Caenorhabditis elegans used to identify P. aeruginosa virulence factors. Proc. Natl. Acad. Sci. USA https://doi.org/10.1073/pnas.96.5.2408 (1999).
Schuster, M., Lostroh, C. P., Ogi, T. & Greenberg, E. P. Identification, timing, and signal specificity of Pseudomonas aeruginosa quorum-controlled genes: a transcriptome analysis. J. Bacteriol. https://doi.org/10.1128/JB.185.7.2066-2079.2003 (2003).
Venturi, V. Regulation of quorum sensing in Pseudomonas. FEMS Microbiol. Rev. https://doi.org/10.1111/j.1574-6976.2005.00012.x (2006).
Muller, M. Pyocyanin induces oxidative stress in human endothelial cells and modulates the glutathione redox cycle. Free Radic. Biol. Med. https://doi.org/10.1016/S0891-5849(02)01087-0 (2002).
Mariencheck, W. I., Alcorn, J. F., Palmer, S. M. & Wright, J. R. Pseudomonas aeruginosa elastase degrades surfactant proteins A and D. Am. J. Respir. Cell Mol. Biol. https://doi.org/10.1165/rcmb.2002-0141OC (2003).
Schmidtchen, A., Frick, I. M., Andersson, E., Tapper, H. & Björck, L. Proteinases of common pathogenic bacteria degrade and inactivate the antibacterial peptide LL-37. Mol. Microbiol. https://doi.org/10.1046/j.1365-2958.2002.03146.x (2002).
Engel, L. S., Hill, J. M., Caballero, A. R., Green, L. C. & O’Callaghan, R. J. Protease IV, a unique extracellular protease and virulence factor from Pseudomonas aeruginosa. J. Biol. Chem. https://doi.org/10.1074/jbc.273.27.16792 (1998).
Park, P. W. et al. Syndecan-1 shedding is enhanced by LasA, a secreted virulence factor of Pseudomonas aeruginosa. J. Biol. Chem. https://doi.org/10.1074/jbc.275.5.3057 (2000).
Laarman, A. J. et al. Pseudomonas aeruginosa alkaline protease blocks complement activation via the classical and lectin pathways. J. Immunol. https://doi.org/10.4049/jimmunol.1102162 (2012).
Jensen, P. et al. Rapid necrotic killing of polymorphonuclear leukocytes is caused by quorum-sensing-controlled production of rhamnolipid by Pseudomonas aeruginosa. Microbiology https://doi.org/10.1099/mic.0.2006/003863-0 (2007).
Caiazza, N. C., Shanks, R. M. Q. & O’Toole, G. A. Rhamnolipids modulate swarming motility patterns of Pseudomonas aeruginosa. J. Bacteriol. https://doi.org/10.1128/JB.187.21.7351-7361.2005 (2005).
Verstraeten, N. et al. Living on a surface: swarming and biofilm formation. Trends Microbiol. https://doi.org/10.1016/j.tim.2008.07.004 (2008).
Morici, L. A. et al. Pseudomonas aeruginosa AlgR represses the Rhl quorum-sensing system in a biofilm-specific manner. J. Bacteriol. https://doi.org/10.1128/JB.01797-06 (2007).
Gutierrez, M. et al. Simultaneous inhibition of Rhamnolipid and polyhydroxyalkanoic acid synthesis and biofilm formation in Pseudomonas aeruginosa by 2-bromoalkanoic acids: effect of inhibitor alkyl-chain-length. PLoS ONE https://doi.org/10.1371/journal.pone.0073986 (2013).
Deligianni, E. et al. Pseudomonas aeruginosa cystic fibrosis isolates of similar RAPD genotype exhibit diversity in biofilm forming ability in vitro. BMC Microbiol. https://doi.org/10.1186/1471-2180-10-38 (2010).
Tramper-Stranders, G. A. et al. Initial Pseudomonas aeruginosa infection in patients with cystic fibrosis: characteristics of eradicated and persistent isolates. Clin. Microbiol. Infect. https://doi.org/10.1111/j.1469-0691.2011.03627.x (2012).
Sousa, A. M. & Pereira, M. O. Pseudomonas aeruginosa diversification during infection development in cystic fibrosis Lungs-A review. Pathogens 3, 680–703 (2014).
Smith, E. E. et al. Genetic adaptation by Pseudomonas aeruginosa to the airways of cystic fibrosis patients. Proc. Natl. Acad. Sci. USA https://doi.org/10.1073/pnas.0602138103 (2006).
Bjarnsholt, T. et al. Quorum sensing and virulence of Pseudomonas aeruginosa during lung infection of cystic fibrosis patients. PLoS ONE https://doi.org/10.1371/journal.pone.0010115 (2010).
Marvig, R. L., Sommer, L. M., Molin, S. & Johansen, H. K. Convergent evolution and adaptation of Pseudomonas aeruginosa within patients with cystic fibrosis. Nat. Genet. https://doi.org/10.1038/ng.3148 (2015).
Salunkhe, P. et al. A cystic fibrosis epidemic strain of Pseudomonas aeruginosa displays enhanced virulence and antimicrobial resistance. J. Bacteriol. https://doi.org/10.1128/JB.187.14.4908-4920.2005 (2005).
Fothergill, J. L., Walshaw, M. J. & Winstanley, C. Transmissible strains of Pseudomonas aeruginosa in cystic fibrosis lung infections. Eur. Respir. J. https://doi.org/10.1183/09031936.00204411 (2012).
Ahmed, S. A. K. S. et al. Natural quorum sensing inhibitors effectively downregulate gene expression of Pseudomonas aeruginosa virulence factors. Appl. Microbiol. Biotechnol. 103, 3521–3535 (2019).
Tan, S. Y. Y. et al. Identification of five structurally unrelated quorum-sensing inhibitors of Pseudomonas aeruginosa from a natural-derivative database. Antimicrob. Agents Chemother. 57, 5629–5641 (2013).
Kim, H. S., Lee, S. H., Byun, Y. & Park, H. D. 6-Gingerol reduces Pseudomonas aeruginosa biofilm formation and virulence via quorum sensing inhibition. Sci. Rep. 5, 1–11 (2015).
Tang, H. B. et al. Contribution of specific Pseudomonas aeruginosa virulence factors to pathogenesis of pneumonia in a neonatal mouse model of infection. Infect. Immun. https://doi.org/10.1128/iai.64.1.37-43.1996 (1996).
Rumbaugh, K. P., Griswold, J. A., Iglewski, B. H. & Hamood, A. N. Contribution of quorum sensing to the virulence of Pseudomonas aeruginosa in burn wound infections. Infect. Immun. 67, 5854–5862 (1999).
Jiricny, N. et al. Loss of social behaviours in populations of Pseudomonas aeruginosa infecting lungs of patients with cystic fibrosis. PLoS ONE https://doi.org/10.1371/journal.pone.0083124 (2014).
LaFayette, S. L. et al. Cystic fibrosis–adapted Pseudomonas aeruginosa quorum sensing lasR mutants cause hyperinflammatory responses. Sci. Adv. https://doi.org/10.1126/sciadv.1500199 (2015).
Feltner, J. B. et al. LasR variant cystic fibrosis isolates reveal an adaptable quorum-sensing hierarchy in Pseudomonas aeruginosa. MBio 7, e01513–e01516 (2016).
Stewart, L. et al. Draft genomes of 12 host-adapted and environmental isolates of Pseudomonas aeruginosa and their positions in the core genome phylogeny. Pathog. Dis. 71, 20–25 (2014).
Kos, V. N. et al. The resistome of Pseudomonas seudomonas aeruginosa in relationship to phenotypic susceptibility. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.03954-14 (2015).
Freschi, L. et al. Clinical utilization of genomics data produced by the international Pseudomonas aeruginosa consortium. Front. Microbiol. https://doi.org/10.3389/fmicb.2015.01036 (2015).
Subedi, D., Vijay, A. K., Kohli, G. S., Rice, S. A. & Willcox, M. Comparative genomics of clinical strains of Pseudomonas aeruginosa strains isolated from different geographic sites. Sci. Rep. https://doi.org/10.1038/s41598-018-34020-7 (2018).
Cramer, N. et al. Microevolution of the major common Pseudomonas aeruginosa clones C and PA14 in cystic fibrosis lungs. Environ. Microbiol. 13, 1690–1704 (2011).
Klockgether, J., Cramer, N., Wiehlmann, L., Davenport, C. F. & Tümmler, B. Pseudomonas aeruginosa genomic structure and diversity. Front. Microbiol. https://doi.org/10.3389/fmicb.2011.00150 (2011).
Aziz, R. K. et al. The RAST server: rapid annotations using subsystems technology. BMC Genom. 9, 75 (2008).
Heurlier, K. et al. Quorum-sensing-negative (lasR) mutants of Pseudomonas aeruginosa avoid cell lysis and death. J. Bacteriol. 187, 4875–4883 (2005).
Cruz, R. L. et al. RhlR-regulated acyl-homoserine lactone quorum sensing in a cystic fibrosis isolate of Pseudomonas aeruginosa. MBio 11, e00532 (2020).
Chen, R., Déziel, E., Groleau, M. C., Schaefer, A. L. & Greenberg, E. P. Social cheating in a Pseudomonas aeruginosa quorum-sensing variant. Proc. Natl. Acad. Sci. USA 116, 7021–7026 (2019).
Kostylev, M. et al. Evolution of the Pseudomonas aeruginosa quorum-sensing hierarchy. Proc. Natl. Acad. Sci. USA 116, 7027–7032 (2019).
Meng, X., Ahator, S. D. & Zhang, L.-H. Molecular mechanisms of phosphate stress activation of Pseudomonas aeruginosa quorum sensing systems. mSphere https://doi.org/10.1128/msphere.00119-20 (2020).
Zulianello, L. et al. Rhamnolipids are virulence factors that promote early infiltration of primary human airway epithelia by Pseudomonas aeruginosa. Infect. Immun. https://doi.org/10.1128/IAI.01772-05 (2006).
Reis, R. S., Pereira, A. G., Neves, B. C. & Freire, D. M. G. Gene regulation of rhamnolipid production in Pseudomonas aeruginosa—a review. Bioresour. Technol. 102, 6377–6384 (2011).
Toder, D. S., Gambello, M. J. & Iglewski, B. H. Pseudomonas aeruginosa LasA: a second elastase under the transcriptional control of lasR. Mol. Microbiol. 5, 2003–2010 (1991).
Perfumo, A. et al. Rhamnolipids are conserved biosurfactants molecules: implications for their biotechnological potential. Appl. Microbiol. Biotechnol. 97, 7297–7306 (2013).
Winstanley, C., O’Brien, S. & Brockhurst, M. A. Pseudomonas aeruginosa evolutionary adaptation and diversification in cystic fibrosis chronic lung infections. Trends Microbiol. 24, 327–337 (2016).
Hoffman, L. R. et al. Pseudomonas aeruginosa lasR mutants are associated with cystic fibrosis lung disease progression. J. Cyst. Fibros. 8, 66–70 (2009).
Wilder, C. N., Allada, G. & Schuster, M. Instantaneous within-patient diversity of Pseudomonas aeruginosa quorum-sensing populations from cystic fibrosis lung infections. Infect. Immun. 77, 5631–5639 (2009).
Lorè, N. I. et al. Cystic fibrosis-niche adaptation of Pseudomonas aeruginosa reduces virulence in multiple infection hosts. PLoS ONE 7, e35648 (2012).
Ciofu, O., Mandsberg, L. F., Bjarnsholt, T., Wassermann, T. & Høiby, N. Genetic adaptation of Pseudomonas aeruginosa during chronic lung infection of patients with cystic fibrosis: strong and weak mutators with heterogeneous genetic backgrounds emerge in mucA and/or lasR mutants. Microbiology 156, 1108–1119 (2010).
Martínez, O. F., Cardoso, M. H., Ribeiro, S. M. & Franco, O. L. Recent advances in anti-virulence therapeutic strategies with a focus on dismantling bacterial membrane microdomains, toxin neutralization, quorum-sensing interference and biofilm inhibition. Front. Cell. Infect. Microbiol. https://doi.org/10.3389/fcimb.2019.00074 (2019).
Davies, E. V., James, C. E., Brockhurst, M. A. & Winstanley, C. Evolutionary diversification of Pseudomonas aeruginosa in an artificial sputum model. BMC Microbiol. https://doi.org/10.1186/s12866-016-0916-z (2017).
Winstanley, C. et al. Newly introduced genomic prophage islands are critical determinants of in vivo competitiveness in the liverpool epidemic strain of Pseudomonas aeruginosa. Genome Res. 19, 12–23 (2009).
Kukavica-Ibrulj, I. et al. In vivo growth of Pseudomonas aeruginosa strains PAO1 and PA14 and the hypervirulent strain LESB58 in a rat model of chronic lung infection. J. Bacteriol. https://doi.org/10.1128/JB.01572-07 (2008).
Brockhurst, M. A., Buckling, A. & Rainey, P. B. The effect of a bacteriophage on diversification of the opportunistic bacterial pathogen, Pseudomonas aeruginosa. Proc. R. Soc. B Biol. Sci. https://doi.org/10.1098/rspb.2005.3086 (2005).
LoVullo, E. D. & Schweizer, H. P. Pseudomonas aeruginosa mexT is an indicator of PAO1 strain integrity. J. Med. Microbiol. 69, 139–145 (2020).
Richardot, C. et al. Amino acid substitutions account for most mexS alterations in clinical nfxC mutants of Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 60, 2302–2310 (2016).
Boursier, M. E. et al. Structure-function analyses of the N-Butanoyl l-Homoserine lactone quorum-sensing signal define features critical to activity in RhlR. ACS Chem. Biol. 13, 2655–2662 (2018).
O’loughlin, C. T. et al. A quorum-sensing inhibitor blocks Pseudomonas aeruginosa virulence and biofilm formation. Proc. Natl. Acad. Sci. https://doi.org/10.1073/pnas.1316981110 (2013).
Rust, L., Pesci, E. C. & Iglewski, B. H. Analysis of the Pseudomonas aeruginosa elastase (lasB) regulatory region. J. Bacteriol. 178, 1134–1140 (1996).
Schuster, M., Urbanowski, M. L. & Greenberg, E. P. Promoter specificity in Pseudomonas aeruginosa quorum sensing revealed by DNA binding of purified LasR. Proc. Natl. Acad. Sci. 101, 15833–15839 (2004).
Oshri, R. D., Zrihen, K. S., Shner, I., Omer Bendori, S. & Eldar, A. Selection for increased quorum-sensing cooperation in Pseudomonas aeruginosa through the shut-down of a drug resistance pump. ISME J. 12, 2458–2469 (2018).
Yan, H. et al. A metabolic trade-offmodulates policing of social cheaters in populations of Pseudomonas aeruginosa. Front. Microbiol. https://doi.org/10.3389/fmicb.2018.00337 (2018).
Faure, E., Kwong, K. & Nguyen, D. Pseudomonas aeruginosa in chronic lung infections: how to adapt within the host?. Front. Immunol. 9, 2416 (2018).
Jacobs, M. A. et al. Comprehensive transposon mutant library of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA https://doi.org/10.1073/pnas.2036282100 (2003).
Zhang, Y. & Miller, R. M. Enhanced octadecane dispersion and biodegradation by a Pseudomonas rhamnolipid surfactant (biosurfactant). Appl. Environ. Microbiol. 58, 3276–3282 (1992).
Essar, D. W., Eberly, L., Hadero, A. & Crawford, I. P. Identification and characterization of genes for a second anthranilate synthase in Pseudomonas aeruginosa: interchangeability of the two anthranilate synthase and evolutionary implications. J. Bacteriol. 172, 884–900 (1990).
Du Noüy, P. L. An interfacial tensiometer for universal use. J. Gen. Physiol. https://doi.org/10.1085/jgp.7.5.625 (1925).
Smyth, T. J. P., Perfumo, A., Marchant, R. & Banat, I. M. Isolation and analysis of low molecular weight microbial glycolipids. In Handbook of Hydrocarbon and Lipid Microbiology (eds McGenity, T. et al.) 3706–3723 (Springer, 2010). https://doi.org/10.1007/978-3-540-77587-4.
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics https://doi.org/10.1093/bioinformatics/btp698 (2010).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. https://doi.org/10.1089/cmb.2012.0021 (2012).
Overbeek, R. et al. The SEED and the rapid annotation of microbial genomes using Subsystems Technology (RAST). Nucl. Acids Res. https://doi.org/10.1093/nar/gkt1226 (2014).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Page, A. J. et al. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics 31, 3691–3693 (2015).
Bertelli, C. et al. IslandViewer 4: expanded prediction of genomic islands for larger-scale datasets. Nucl. Acids Res. 45, W30–W35 (2017).
Arndt, D. et al. PHASTER: a better, faster version of the PHAST phage search tool. Nucl. Acids Res. 44, W16–W21 (2016).
Alikhan, N.-F., Petty, N. K., Ben Zakour, N. L. & Beatson, S. A. BLAST ring image generator (BRIG): simple prokaryote genome comparisons. BMC Genom. 12, 402 (2011).
Jani, M., Mathee, K. & Azad, R. K. Identification of novel genomic islands in liverpool epidemic strain of Pseudomonas aeruginosa using segmentation and clustering. Front. Microbiol. 7, 1210 (2016).
Kiratisin, P., Tucker, K. D. & Passador, L. LasR, a transcriptional activator of Pseudomonas aeruginosa virulence genes, functions as a multimer. J. Bacteriol. https://doi.org/10.1128/JB.184.17.4912-4919.2002 (2002).
Smith, R. S., Harris, S. G., Phipps, R. & Iglewski, B. The Pseudomonas aeruginosa quorum-sensing molecule N-(3-oxododecanoyl)homoserine lactone contributes to virulence and induces inflammation in vivo. J. Bacteriol. https://doi.org/10.1128/jb.184.4.1132-1139.2002 (2002).
Parsek, M. R., Schaefer, A. L. & Greenberg, E. P. Analysis of random and site-directed mutations in rhII, a Pseudomonas aeruginosa gene encoding an acylhomoserine lactone synthesis. Mol. Microbiol. https://doi.org/10.1046/j.1365-2958.1997.5741935.x (1997).
Déziel, E. et al. The contribution of MvfR to Pseudomonas aeruginosa pathogenesis and quorum sensing circuitry regulation: multiple quorum sensing-regulated genes are modulated without affecting IasRI, rhIRI or the production of N-acyl-L-homoserine lactones. Mol. Microbiol. https://doi.org/10.1111/j.1365-2958.2004.04448.x (2005).
Rampioni, G. et al. RsaL provides quorum sensing homeostasis and functions as a global regulator of gene expression in Pseudomonas aeruginosa. Mol. Microbiol. 66, 1557–1565 (2007).
Fuchs, E. L. et al. The Pseudomonas aeruginosa Vfr regulator controls global virulence factor expression through cyclic AMP-dependent and -independent mechanisms. J. Bacteriol. https://doi.org/10.1128/JB.00363-10 (2010).
Kong, K. F. et al. Pseudomonas aeruginosa AmpR is a global transcriptional factor that regulates expression of AmpC and PoxB β-lactamases, proteases, quorum sensing, and other virulence factors. Antimicrob. Agents Chemother. https://doi.org/10.1128/AAC.49.11.4567-4575.2005 (2005).
Branny, P. et al. Inhibition of quorum sensing by a Pseudomonas aeruginosa dksA homologue. J. Bacteriol. https://doi.org/10.1128/JB.183.5.1531-1539.2001 (2001).
Li, K. et al. SuhB is a regulator of multiple virulence genes and essential for pathogenesis of Pseudomonas aeruginosa. MBio https://doi.org/10.1128/mBio.00419-13 (2013).
Jin, S., Ishimoto, K. S. & Lory, S. PilR, a transcriptional regulator of piliation in Pseudomonas aeruginosa, binds to a cis-acting sequence upstream of the pilin gene promoter. Mol. Microbiol. https://doi.org/10.1111/j.1365-2958.1994.tb01338.x (1994).
Köhler, T., Van Delden, C., Curty, L. K., Hamzehpour, M. M. & Pechere, J. C. Overexpression of the MexEF-OprN multidrug efflux system affects cell-to-cell signaling in Pseudomonas aeruginosa. J. Bacteriol. https://doi.org/10.1128/JB.183.18.5213-5222.2001 (2001).
Juhas, M. et al. Global regulation of quorum sensing and virulence by VqsR in Pseudomonas aeruginosa. Microbiology https://doi.org/10.1099/mic.0.26906-0 (2004).
Gupta, R. & Schuster, M. Negative regulation of bacterial quorum sensing tunes public goods cooperation. ISME J. https://doi.org/10.1038/ismej.2013.109 (2013).
Kay, E. et al. Two GacA-dependent small RNAs modulate the quorum-sensing response in Pseudomonas aeruginosa. J. Bacteriol. https://doi.org/10.1128/JB.00409-06 (2006).
Moskowitz, S. M., Ernst, R. K. & Miller, S. I. PmrAB, a two-component regulatory system of Pseudomonas aeruginosa that modulates resistance to cationic antimicrobial peptides and addition of aminoarabinose to lipid A. J. Bacteriol. https://doi.org/10.1128/JB.186.2.575-579.2004 (2004).
de Bentzmann, S. et al. Unique biofilm signature, drug susceptibility and decreased virulence in drosophila through the Pseudomonas aeruginosa two-component system PprAB. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1003052 (2012).
Cai, Z. et al. RpoN regulates virulence factors of Pseudomonas aeruginosa via modulating the PqsR quorum sensing regulator. Int. J. Mol. Sci. https://doi.org/10.3390/ijms161226103 (2015).
Suh, S. J. et al. Effect of rpoS mutation on the stress response and expression of virulence factors in Pseudomonas aeruginosa. J. Bacteriol. https://doi.org/10.1128/jb.181.13.3890-3897.1999 (1999).
Tanaka, K. & Takahashi, H. Cloning and analysis of the gene (rpoDA) for the principal σ factor of Pseudomonas aeruginosa. BBA Gene Struct. Expr. https://doi.org/10.1016/0167-4781(91)90092-Z (1991).
Leoni, L., Orsi, N., De Lorenzo, V. & Visca, P. Functional analysis of PvdS, an iron starvation sigma factor of Pseudomonas aeruginosa. J. Bacteriol. https://doi.org/10.1128/JB.182.6.1481-1491.2000 (2000).
Acknowledgements
This work was supported by Ulster University, Northern Ireland through a Vice Chancellor’s Research Scholarship studentship to S.A.K.S Ahmed. The authors would also like to thank the Centre for Cognitive and Skill Enhancement at IUB for computational support to the bioinformatic analysis.
Funding
This work was supported by the Northern Ireland Research & Development Office, HPSS(NI) Grant RRG9.3. Professor Banat and group would like to acknowledge support of the European Union Framework Programme for Research and Innovation, Horizon 2020 under Grant agreement No. 635340 MARISURF.
Author information
Authors and Affiliations
Contributions
J.S.G.D. and S.A.K.S.A. conceived and designed the study. S.A.K.S.A. performed all experiments. S.M.E. performed WGS analysis of PA80. MR carried out all comparative genomic analyses. T.J.S. performed all HPLC-MS analyses. M.R. and S.A.K.S.A. prepared the figures and wrote the manuscript. J.S.G.D., I.B. and R.M. reviewed the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ahmed, S.A.K.S., Rudden, M., Elias, S.M. et al. Pseudomonas aeruginosa PA80 is a cystic fibrosis isolate deficient in RhlRI quorum sensing. Sci Rep 11, 5729 (2021). https://doi.org/10.1038/s41598-021-85100-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-85100-0
- Springer Nature Limited