Abstract
Intrahepatic cholangiocarcinoma (CHOL) remains a rare malignancy, ranking as the leading lethal primary liver cancer worldwide. However, the biological functions of integrator complex subunit 8 (INTS8) in CHOL remain unknown. Thus, this research aimed to explore the potential role of INTS8 as a novel diagnostic or therapeutic target in CHOL. Differentially expressed genes (DEGs) in two Gene Expression Omnibus (GEO) datasets were obtained by the “RRA” package in R software. The “maftools” package was used to visualize the CHOL mutation data from The Cancer Genome Atlas (TCGA) database. The expression of INTS8 was detected by performing quantitative reverse transcription-PCR (qRT-PCR) and immunohistochemistry in cell lines and human samples. The association between subtypes of tumour-infiltrating immune cells (TIICs) and INTS8 expression in CHOL was determined by using CIBERSORT tools. We evaluated the correlations between INTS8 expression and mismatch repair (MMR) genes and DNA methyltransferases (DNMTs) in pan-cancer analysis. Finally, the pan-cancer prognostic signature of INTS8 was identified by univariate analysis. We obtained the mutation landscapes of an RRA gene set in CHOL. The expression of INTS8 was upregulated in CHOL cell lines and human CHOL samples. Furthermore, INTS8 expression was closely associated with a distinct landscape of TIICs, MMR genes, and DNMTs in CHOL. In addition, the high INTS8 expression group presented significantly poor outcomes, including overall survival (OS), disease-specific survival (DSS) and disease-free interval (DFI) (p < 0.05) in pan-cancer. INTS8 contributes to the tumorigenesis and progression of CHOL. Our study highlights the significant role of INTS8 in CHOL and pan-cancers, providing a valuable molecular target for cancer research.
Similar content being viewed by others
Introduction
As a hepatobiliary malignancy subtype, intrahepatic cholangiocarcinoma (CHOL) has attracted increasing attention due to its increasing global incidence and trends1, with the largest age-standardized incidence rates increasing in China (average annual percent change: 11.1%) from 1993 to 20122. Moreover, the mortality rate of CHOL shows a rising trend globally and is approximately 1–2/100,000 in most countries3. Because of the aggressive and asymptomatic features of CHOL, many patients are diagnosed at an advanced stage. Surgical resection is regarded as the best treatment strategy for achieving a good prognosis and long survival. However, the 5‐year overall survival (OS) rate remains limited to 22–30% after curative hepatectomy due to high recurrence rates4,5. Moreover, a 40–80% recurrence rate was reported for CHOL patients after surgical resection6. The combination of gemcitabine and cisplatin is regarded as the standard chemotherapy regimen, despite showing limited effectiveness for CHOL. Therefore, it is urgent to improve the sensitivity of diagnosis and effectiveness of targeted therapy for CHOL.
Integrators are transcriptional regulatory complexes comprised of at least 14 subunits7. Integrator complex subunit 8 (INTS8) is one of the major components of RNA polymerase II and has been demonstrated to be involved in the cleavage of small nuclear RNAs and transcriptional processes8,9. A recent study found that INTS8 was essential for transcription repression, which was induced by recruiting protein phosphatase 2A to prevent transcription elongation and promote transcription termination10. A previous study revealed that INTS8 was robustly increased in neurodevelopmental diseases11 and numerous tumours12,13. Overexpressed INTS8 could facilitate epithelial-to-mesenchymal transition, which is mediated by the TGF-β signalling pathway in hepatocellular carcinoma (HCC)14. Increasing evidence has demonstrated that mismatch repair (MMR) genes play an important role in maintaining genomic stability, and DNA methylation regulates gene expression. The loss of key gene functions of MMR genes could induce DNA replication errors, resulting in a high level of somatic mutations. It has been reported that the MMR pathway is potently activated during G1/S phase15. DNA methylation is a type of epigenetic modification that can regulate gene expression. As the function of DNA methyltransferases (DNMTs), DNA methylation occurs when the methyl group covalently bonds to the 5′ carbon position of the cytosine in genome CpG dinucleotides16. However, studies focused on the role of INTS8 in CHOL are generally lacking.
In the present study, we used the robust rank aggregation (RRA) method to select differentially expressed genes (DEGs) based on the Gene Expression Omnibus (GEO) database. Then, we explored genes at the intersection between DEGs and gene mutation profiles in the CHOL cohort of The Cancer Genome Atlas (TCGA) and identified INTS8 as a candidate gene. We verified the overexpression of INTS8 in CHOL cell lines and human CHOL samples by quantitative reverse transcription-PCR (qRT-PCR) and immunohistochemistry (IHC). Our study showed that high INTS8 expression is closely correlated with poor prognosis across cancers. Moreover, the underlying mechanism may be attributed to tumour-infiltrating immune cells (TIICs), MMR genes, and DNMTs. Therefore, INTS8 was identified as a therapeutic target in CHOL and pan-cancer series; an association was observed between INTS8 expression and TIICs; MMR genes and DNMTs were suggested to mediate INTS8 effects.
Materials and methods
Data acquisition and processing
We selected 2 CHOL datasets from the GEO database (http://www.ncbi.nlm.nih.gov/geo/): GSE26566 and GSE3222517,18. GSE26566 included 104 CHOL samples and 6 matched surrounding samples. GSE32225 contained 149 CHOL samples and 6 matched surrounding samples. All analyses were undertaken with R version 4.0.4 (https://cran.r-project.org/src/base/R-4/). All expression profiles were downloaded and processed by the “GEOquery” package (www.r-project.org). Considering the differences and batch effects of different platforms, we utilized the “sva” package19 to avoid these effects and remove other unwanted variations. In addition, the transcriptome profiles, mutation and methylation data, and clinical information of tumour samples and corresponding samples were obtained from the TCGA database (https://cancergenome.nih.gov/) and analysed by utilizing the “TCGAbiolinks” package20. Transcripts per million (TPM) values were applied for subsequent analyses21.
DEGs and RRA analysis
The DEGs were determined between CHOL samples and matched surrounding samples using the “limma” package22. RRA was performed for gene list integration23, and a score < 0.05 was used to determine the RRA gene set. These data were visualized by a heatmap and volcano plot.
Identification of INTS8 by mutation analysis and ROC curves
We used the “maftools” package24 to visualize the CHOL mutation data. To assess mutated genes present in the RRA gene set, we obtained the intersection of RRA genes and CHOL mutation genes. To evaluate the diagnostic performance of muted RRA genes, receiver operating characteristic (ROC) curves were generated by using the “pROC” package. Next, we selected the optimal efficacy indicators based on the areas under the curve (AUCs) for further research. The patients were stratified into two groups according to the median expression of INTS8. DEGs between the high and low INTS8 groups were confirmed by using the “limma” package. The cut-offs for the DEGs were as follows: |log2 fold change (FC)|> 1 and false discovery rate (FDR) < 0.05. A heat map constructed by using the “ggplot2” package was used to visualize the DEGs.
Functional enrichment of mutated RRA genes and INTS8-related genes
To identify the possible pathways and biological functions of the 5 mutated RRA genes and differential INTS8-associated genes, we applied the “clusterProfiler” package25 to perform Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses. The protein–protein interaction (PPI) network of the 5 mutated RRA genes was constructed via the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING, http://string-db.org/) online database and then visualized by Cytoscape v.3.7.1 (https://cytoscape.org/). Moreover, we used Molecular Complex Detection (MCODE) to explore functional clusters in the PPI network. To further understand the functions and biological pathways related to INTS8, gene set enrichment analysis (GSEA) was performed on the INTS8 gene by utilizing GSEA software (v.3.0).
CIBERSORT estimation
To characterize the tumour microenvironment, we used “CIBERSORT” (R package) (http://cibersort.stanford.edu/)26 to explore the relative proportions and absolute fraction scores of 22 subtypes of TIICs in CHOL tissues. Moreover, the association between INTS8 expression levels and the infiltration of TIICs was assessed and visualized by heatmaps and violin plots.
Association of INTS8 gene expression with clinical outcome in different tumours
To explore the influence of the expression level of INTS8, we carried out analyses using a publicly available database. We retrieved data from The Cancer Cell Line Encyclopedia (CCLE, https://portals.broadinstitute.org/ccle)27 and the Genotype-Tissue Expression (GTEx) project28 to investigate the gene expression data of INTS8 in a range of tumour tissues and cell lines. Moreover, we downloaded pan-cancer mutation data from the TCGA database and analysed the mutations of INTS8 in samples of 32 different tumour types. Furthermore, we used Cox regression and the Kaplan–Meier method to evaluate the association of INTS8 gene expression with clinical outcome in different cancers. p < 0.05 was regarded as the cut-off to verify the prognostic role of INTS8.
Association of INTS8 expression with MMR genes and DNA methylation
To reveal the role of INTS8 in cancer progression, we evaluated the relationship between the expression level of INTS8 and 5 key DNA MMR genes (including MLH1, MSH2, MSH6, PMS2, and EPCAM). In addition, we performed an integrative analysis of DNA methylation and INTS8 expression to determine its underlying mechanism in pan-cancer.
Real-time PCR
A human normal biliary epithelial cell line (HIBEC) and 3 CHOL cell lines (including HCCC-9810, RBE, and CCLP-1 cells) were used to detect the mRNA expression of INTS8. Total RNA and cDNA synthesis was performed by following the manufacturer’s instructions (Accurate biotechnology, China). Gene expression was measured on an ABI 7500 system by using a SYBR Green kit (Accurate biotechnology, China). The forward primer for INTS8 was 5′-TGCTGGAGGAGTCACTGTTGGAG- 3′, and the reverse primer for INTS8 was 5′-TTATCAGGCGGAGGTTGAACTTGG-3′.
IHC
A total of 155 paired CHOL and 5 peritumoural tissue samples were obtained for experimental validation. Informed consent was obtained from all participants. The study involving human participants was approved by the Ethics Committee of Shanghai Outdo Biotech Company (No. YB M-05-02) and performed following relevant guidelines and regulations. Formalin-fixed paraffin-embedded tissue samples were examined by incubation with primary antibodies (ab18050, Abcam).
Ethical approval
Informed consent was obtained from all participants. The study involving human participants was approved by the Ethics Committee of Shanghai Outdo Biotech Company with NO. YB M-05-02, and performed following relevant guidelines and regulations.
Results
Identification of robust DEGs in GEO
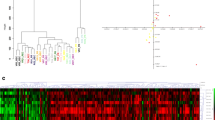
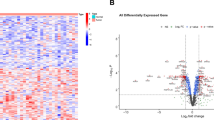
Based on the DEG results, a total of 710 significantly upregulated and 903 significantly downregulated DEGs were confirmed in GSE26566, and 432 significantly upregulated DEGs and 566 significantly downregulated DEGs were identified in GSE32225. The DEGs are shown by heatmaps and volcano plots in Fig. 1A–D. Furthermore, these DEGs were integrated by the RRA method with a score < 0.05. Then, the RRA gene set was visualized by a heatmap, as shown in Fig. 1E. As a result, an RRA gene set was obtained for further investigation.
Functional enrichment and PPI network analyses of the RRA gene set
GO and KEGG enrichment analyses elucidated the functions of the RRA gene set (Supplementary Fig. 1A,B). The RRA gene set was obviously enriched in biological processes, such as lipid catabolic process, digestion, drug catabolic process, and eicosanoid metabolic process. In addition, the RRA gene set participated in pancreatic secretion, fat digestion and absorption, protein digestion and absorption, and focal adhesion (Supplementary Fig. 1C,D). A PPI network of the RRA gene set, which included 202 interactions, was constructed to identify protein interactions and was visualized by Cytoscape (Supplementary Fig. 2A,C). Two functional clusters in the PPI network were extracted, suggesting their central roles in this network (Supplementary Fig. 2B). Our results showed that the RRA gene set was associated with some metabolic pathways.
Mutation landscape of the RRA gene set in CHOL
To identify the mutational landscape in CHOL patients, the “maftools” package in R software was used. Missense mutations were the predominant type of mutation in patients with CHOL (Fig. 2A). Single nucleotide polymorphisms had a more frequent occurrence than insertions or deletions (Fig. 2B). In particular, C > T remained the most common mutation type of single nucleotide variants in CHOL (Fig. 2C). The mutation types in CHOL are displayed in Fig. 2D,E. The top 10 mutated genes present in CHOL with ranked percentages are as follows: MUC16 (12%), PBRM1 (20%), ARID1A (18%), BAP1 (16%), MUC5B (10%), EPHA2 (14%), IDH1 (12%), LRP1B (10%), CHD7 (10%), and DNAH5 (8%) (Fig. 2F). A total of 5 mutated genes in the RRA gene set were found in mutation profiles, and the mutation information of the RRA gene set was obtained by a waterfall plot (Fig. 2G). BAP1, IDH1 and PBRM1 were the top 3 mutant genes of the RRA gene set (Fig. 2H). The mutant base pair ratio of the RRA gene set showed that C > T was the most common single nucleotide variant in the RRA gene set (Fig. 2I).
Mutation landscape of the RRA gene set in TCGA-CHOL. (A–C) According to different classification categories, the classification of mutation types, including missense mutations, SNPs, and C > T mutations, was performed with statistical calculations. (D) Total mutation number in each sample. (E) Each variant classification in each sample. (F) Top 10 mutated genes in TCGA-CHOL. (G) The mutation information of 5 mutated genes in the RRA gene set was determined by the waterfall plot. (H) The top mutant genes of the RRA gene set are shown by a box plot. (I) Mutant base pair ratio of the RRA gene set.
Identification of DEGs between the high and low INTS8 expression groups
ROC analysis was applied to determine the diagnostic efficacy of the 5 mutated genes of the RRA gene set. INTS8 had the highest AUC value (AUC = 0.852), followed by ATF4 (AUC = 0.836), PPP1CA (AUC = 0.781), PCSK2 (AUC = 0.504) and BUB1B (AUC = 0.5) (Fig. 3A). Considering that INTS8 had the highest AUC, it was selected as the target gene for further analysis. To explore the underlying mechanism of INTS8 in CHOL, the patients were divided into two groups according to the median expression value of INTS8. DEGs between the high and low INTS8 expression groups were identified (Fig. 3B). Furthermore, we found that the mRNA expression of INTS8 was upregulated in 3 CHOL cell lines compared with HIBE in vitro (Fig. 3C). The protein levels of INTS8 via IHC were also verified to be obviously increased in CHOL patient tissue samples compared with normal tissue samples (Fig. 3D). The experimental results were consistent with those of the bioinformatic analysis.
Identification of INTS8 as a candidate gene. (A) ROC curves of 5 genes for diagnostic value. (B) DEGs in the high and low INTS8 expression groups. (C) Expression of INTS8 in HIBEC and 3 CHOL cell lines (including HCCC-9810, RBE, and CCLP-1 cells) by using PCR. (D) Representative images of INTS8 IHC staining in human CHOL and adjacent normal tissues.
Functional enrichment of INTS8 in CHOL
To identify the biological functions and key candidate pathways of the INTS8-related genes, we performed GO and KEGG analyses. The top 10 GO terms are shown in Fig. 4A. Drug metabolism-cytochrome P450 (CYP), retinol metabolism, chemical carcinogenesis, metabolism of xenobiotics by CYP, drug metabolism-other enzymes, and fatty acid degradation were the most significantly enriched in CHOL patients with high INTS8 expression compared with those with low INTS8 expression (Fig. 4B). To elucidate the molecular mechanisms of INTS8, INTS8-related signalling pathways were analysed by GSEA-KEGG and GSEA-GO (Fig. 4C,D). The results suggested that INTS8 might be related to metabolic pathways, such as CYP and retinol metabolism.
Association between TIICs and INTS8 expression in CHOL
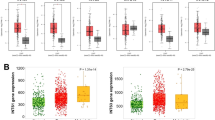
TIICs significantly impact the development and progression of many types of cancers, including CHOL. By applying CIBERSORT tools, we observed a high level of M0 macrophages, M2 macrophages, monocytes, and resting CD4+ memory T cells and a lower level of activated dendritic cells, eosinophils, neutrophils and activated CD4+ memory T cells in CHOL (Fig. 5A,B). Moreover, we assessed the relationship between TIICs and INTS8 expression in CHOL. We found that the high INTS8 expression group presented a unique TIIC landscape, including a significantly high level of M0 macrophages but a low level of M2 macrophages, an elevated level of resting CD4+ memory T cells but a low level of CD4 naive T cells, and an increased level of resting mast cells but a low level of activated mast cells. In addition, low expression of gamma delta T cells and monocytes was also found in the high INTS8 expression group (Fig. 5C,D).
Identification of complex associations between 22 TIIC subsets and INTS8 expression in CHOL. (A) Relative proportions of 22 subtypes of tumour-infiltrating immune cells for each sample in CHOL. (B) Relative proportions of 22 subtypes of tumour-infiltrating immune cells for each sample. (C,D) Comparison of the immune cell fraction difference between the low and high INTS8 expression groups. Note: Blue refers to low INTS8 expression, and brown refers to high INTS8 expression.
INTS8 expression in multiple dimensions
Considering the extensive mutational heterogeneity of cancers, we systematically performed large-scale profiling of INTS8 expression in 21 cell lines and 31 related tissues based on CCLE and GTEx. As shown in Fig. 6A,B, the expression levels of INTS8 in diverse cancer tissues, including the biliary tract, liver, and bone marrow, and cell lines were elevated to differing degrees. In addition, we found that INTS8 harboured the most prevalent mutations, such as missense, truncating and fusion mutations, in different tumours (Fig. 6C).
Associations between INTS8 and clinicopathologic characteristics and survival information
As shown in Table 1, increased INTS8 expression was directly associated with age and grade. INTS8 expression gradually increased from stages I/II to stage IV CHOL. To assess the prognostic capacity of INTS8, we constructed Kaplan–Meier curves for OS, disease-specific survival (DSS), and disease-free interval (DFI) by using multivariate Cox regression analysis. Regarding prognostic outcomes, patients in the high INTS8 group exhibited poor OS, DSS and DFI (p < 0.05) in a pan-cancer analysis (Supplementary Figs. 3–5). These findings suggested that INTS8 expression is a potent potential prognostic biomarker for various cancers.
MMR genes and DNA methylation genes involved in CHOL
To explore the underlying DNA repair mechanism associated with INTS8 mutation, we investigated the association between INTS8 and MMR genes (including MLH1, MSH2, MSH6, PMS2, and EPCAM). We found that INTS8 was positively correlated with the expression of MSH2, MSH6, and PMS2 but showed no association with MLH1 and EPCAM. Due to the extensive function of MMR genes in cancers, we performed a pan-cancer analysis to analyse the relationship between INTS8 and MMR genes. Interestingly, a positive association between INTS8 and MMR genes was present in numerous cancers, such as brain lower-grade glioma, liver HCC, and pancreatic cancer (Fig. 7A).
MMR genes and DNA methylation genes involved in CHOL. (A) A total of 3 MMR genes were detected in CHOL. The right-angled triangle in the lower left corner represents the Pearson’s correlation between INTS8 and MMR gene expression. The right-angled triangle in the upper right corner represents the p value; *p < 0.05, **p < 0.01 and ***p < 0.001 show significant differences. (B) Circle diagram showing the association between INTS8 and DNMTs. The first circle from outer to inner is the abbreviation of cancers. The second circle is the specific DNMTs, with DNMT1 (red), DNMT2 (blue), DNMT3A (green), and DNMT3B (purple). The third circle is the correlation coefficient. Green represents negative correlations; Red represents positive correlations. The fourth circle is the p value. And the fifth circle is the specific value of the correlation coefficient and p.
As shown in Fig. 7B, an epigenetic signature was discovered and showed a high correlation between INTS8 and DNMTs (DNMT1: r = 0.31, p < 0.05; DNMT2: r = 0.53, p < 0.05; DNMT3A: r = 0.53, p < 0.05; DNMT3B: r = 0.42, p < 0.05). Furthermore, a pan-cancer analysis of DNMTs was performed and showed that INTS8 was positively related to the expression profiles of 4 DNMTs in most cancers except testicular germ cell tumours. All these results indicated that MMR genes and certain DNMTs may play an important role in INTS8 mutations in CHOL.
Discussion
CHOL is an extremely aggressive biliary neoplasm with increasing incidence and poor prognosis worldwide29. Currently, prognostic model in biliary tract cancers has reached interesting results. For example, the PECS index was identified as a replicable and promising tool to assess the prognosis of biliary tract cancer patients in future clinical practice; it is based on a real-life population and has robust numerosity, with C-indexes of 0.73–0.83 and survival curves showing clear separation. With an integration with clinicopathological model, the potential value of molecular data could contribute to the clinical practice30. In this study, the TCGA and GEO databases were applied to systematically analyse the mutational status of RRA genes in CHOL, and 5 mutant genes were found by intersection analysis. Based on the diagnostic efficacy of the 5 mutant genes, we selected INTS8, which had the largest AUC value, for follow-up research, which showed that INTS8 played a significant role in CHOL and even across all cancers.
Various studies have suggested that the integrator complex plays an essential role in RNA processing and transcription regulation. Previous studies have shown that INTS8 mutation can induce severe neurodevelopmental syndrome11 and pan-cancer31. In this study, we found that INTS8 was significantly overexpressed in CHOL compared to normal samples, which was consistent with the results of IHC and PCR. Our results showed that INTS8 overexpression was positively related to poor prognosis in many tumour types.
The GO enrichment analyses showed that high INTS8 expression was mainly associated with organic anion transport, organic acid transport, carboxylic acid transport and acute inflammatory response. In addition, retinol metabolism, chemical carcinogenesis, drug metabolism-CYP, metabolism of xenobiotics, drug metabolism-other enzymes, and fatty acid degradation were most significantly enriched in CHOL patients with high INTS8 expression compared with those with low INTS8 expression. Retinol is a fat-soluble nutrient that is essential for maintaining physiological functions in many tissues32. Retinol metabolism abnormalities caused by genetic or environmental factors could induce developmental pathologies, including mammalian placental and embryonic development33, ovary disease32 and fatty liver disease34. A previous study showed that the administration of retinol facilitated hepatocarcinogenesis development during its early stages35. Drug metabolism-CYP was related to DNA methylation-driven genes in prostate adenocarcinoma36. In addition, previous data showed that hepatic CYP family enzymes, especially increased CYP2A6 and diminished CYP2E1, might participate in the progression of CHOL37. Lipid metabolism is newly recognized as a hallmark of cancer, and inhibiting fatty acid availability could control the development of malignancy38,39. Li et al. found that CHOL tumorigenesis was insensitive to fatty acid synthase deprivation, which contributed to high fatty acid uptake and resulted in rapid tumour growth. Therefore, promoting fatty acid degradation may be a novel therapeutic approach for CHOL40.
DNA damage and repair provide protection for mutation avoidance, which plays central roles in maintaining genome stability41,42. To date, it has been reported that 4 major DNA repair pathways are involved in maintaining gene expression, including nucleotide excision repair, base excision repair, MMR, and double-strand break repair43. The expression of INTS8 was positively correlated with MSH2, MSH6, and PMS2 but not associated with MLH1 and EPCAM. The IHC analysis44 results showed that there was no loss of the expression of DNA repair enzymes/MMR proteins (MLH1, MSH2, PMS2, and MSH6) in either occupational CHOL45 or cohorts with CHOL46.
MMR gene mutations and tumour MLH1 promoter methylation are the main causes of microsatellite instability (MSI) in patients with colorectal cancer (CRC)47. Although the overall number of MSI-high (MSI-H) CHOL cases is low (1.3%), MSI testing of cholangiocarcinoma exhibited an atypical histomorphology, especially in younger patients48. EPCAM, a stemness-related marker, is positively correlated with poor prognosis in CHOL and HCC49,50. However, we did not observe an association between INTS8 and EPCAM in CHOL.
Recently, epigenetic alterations have been characterized by any heritable modification of chromatin DNA or histone proteins but without changes in the DNA sequence51,52; they can be observed in many human cancers and cooperate with genetic alterations to dominate the formation of cancers53. DNA methylation is one of the main epigenetic changes and is specifically mediated by the DNMT family (including DNMT1, DNMT1, DNMT3A and DNMT3B)54. DNMTs could establish and maintain DNA methylation patterns, which induce gene silencing, transcriptional activation and posttranscriptional regulation mediated by DNMT2-dependent RNA methylation. Here, we found that INTS8 is positively associated with DNMTs in CHOL, suggesting that the effect of INTS8 on CHOL development may be caused by mutations in DNMT genes. Thus, we hypothesized that functional impairment of INTS8, which is associated with MMR genes and DNMTs, promotes malignancy across cancers, suggesting the potential of INTS8 for cancer research.
A significant increase in macrophages was shown in locally advanced CHOL patients compared to metastatic CHOL patients55. In our study, the immune landscape showed a distinctly high expression of macrophages in CHOL, which was consistent with the findings of other studies. However, the results showed that low M2 macrophage levels but no significant alteration in M1 macrophages appeared in the high INTS8 expression group, suggesting that remarkable cellular heterogeneity exists in macrophage subtypes. M1 macrophages are currently known to promote inflammation, while M2 macrophages are characterized by anti-inflammatory functions. Thus, whether INTS8 is involved in locally advanced CHOL remains to be experimentally validated. In particular, a case study discovered that gamma delta T cell-based immunotherapy showed no adverse effects and could positively regulate peripheral immune functions in patients with CHOL56. We found that a low level of gamma delta T cells was present in the high INTS8 expression group in CHOL. Based on this promising finding, INTS8 could be considered in the development of promising therapies for CHOL. Previous studies proved that CD4 regulatory T cell infiltration is a prominent immunosuppressive characteristic in CHOL57. Although Tregs were relatively high in CHOL, they had no association with INTS8.
Although our study provided insights into the relationship between INTS8 and CHOL, there were still some limitations. Due to the smaller occurrence of cases compared with other common cancers, the sample size for IHC involved in cohort validation (especially for the peritumoural tissue samples) was relatively small. In addition, there were relatively few samples with complete clinical information from the TCGA. However, to avoid bias caused by the small sample size, we used data from both the GEO and TCGA databases to confirm the findings, extended the potential functions and mechanisms of INTS8 to pan-cancer research, and discussed the functions of INTS8 in depth. Although the associations between INTS8 expression and MMR genes and DNMTs was shown, the direct mechanisms require further exploration and solid experimental evidence.
Conclusion
In summary, we observed that increased INTS8 expression can contribute to malignancies and was directly associated with age, grade, and sex in CHOL. Moreover, the pan-cancer analysis revealed that the altered expression of INTS8, which may be mediated by MMR genes and DNA methylation status, might participate in the development of multiple cancer types. In addition, the high INTS8 group displayed an obvious poor prognosis in terms of OS, DSS, and DFI in multiple cancer types. Our results showed the potential of INTS8 as a therapeutic target for CHOL.
Abbreviations
- AUCs:
-
Areas under the curve
- BER:
-
Base excision repair
- CCLE:
-
The Cancer Cell Line Encyclopedia
- CHOL:
-
Intrahepatic cholangiocarcinoma
- CRC:
-
Colorectal cancer
- CYP:
-
Cytochrome P450
- DEGs:
-
Differentially expressed genes
- DNMTs:
-
DNA methyltransferases
- DFI:
-
Disease-free interval
- DSBR:
-
Double strand break repair
- DSS:
-
Disease-specific survival
- FDR:
-
False discovery rate
- GEO:
-
Gene Expression Omni
- GO:
-
Gene Ontology
- GSEA:
-
Gene set enrichment analysis
- GTEx:
-
The Genotype-Tissue Expression
- HCC:
-
Hepatocellular carcinoma
- HIBEC:
-
Human normal biliary epithelial cell line
- IHC:
-
Immunohistochemistry
- INTS8:
-
Integrator complex subunit 8
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- MCODE:
-
Molecular Complex Detection
- MMRs:
-
Mismatch repair genes
- OS:
-
Overall survival
- RRA:
-
Robust rank aggregation
- STRING:
-
Search Tool for the Retrieval of Interacting Genes
- TCGA:
-
The cancer genome atlas
- TIICs:
-
Tumor-infiltrating immune cells
References
Rizvi, S., Khan, S. A., Hallemeier, C. L., Kelley, R. K. & Gores, G. J. Cholangiocarcinoma—Evolving concepts and therapeutic strategies. Nat. Rev. Clin. Oncol. 15, 95–111. https://doi.org/10.1038/nrclinonc.2017.157 (2018).
Florio, A. A. et al. Global trends in intrahepatic and extrahepatic cholangiocarcinoma incidence from 1993 to 2012. Cancer 126, 2666–2678. https://doi.org/10.1002/cncr.32803 (2020).
Bertuccio, P. et al. Global trends in mortality from intrahepatic and extrahepatic cholangiocarcinoma. J. Hepatol. 71, 104–114. https://doi.org/10.1016/j.jhep.2019.03.013 (2019).
Chun, Y. S. & Javle, M. Systemic and adjuvant therapies for intrahepatic cholangiocarcinoma. Cancer Control 24, 1073274817729241. https://doi.org/10.1177/1073274817729241 (2017).
Miyata, T. et al. CXCL12 expression in intrahepatic cholangiocarcinoma is associated with metastasis and poor prognosis. Cancer Sci. 110, 3197–3203. https://doi.org/10.1111/cas.14151 (2019).
Sirica, A. E. et al. Intrahepatic cholangiocarcinoma: Continuing challenges and translational advances. Hepatology 69, 1803–1815. https://doi.org/10.1002/hep.30289 (2019).
Albrecht, T. R. et al. Integrator subunit 4 is a “Symplekin-like” scaffold that associates with INTS9/11 to form the Integrator cleavage module. Nucleic Acids Res. 46, 4241–4255. https://doi.org/10.1093/nar/gky100 (2018).
Kirstein, N., Gomes Dos Santos, H., Blumenthal, E. & Shiekhattar, R. The Integrator complex at the crossroad of coding and noncoding RNA. Curr. Opin. Cell Biol. 70, 37–43. https://doi.org/10.1016/j.ceb.2020.11.003 (2020).
Lai, F., Gardini, A., Zhang, A. & Shiekhattar, R. Integrator mediates the biogenesis of enhancer RNAs. Nature 525, 399–403. https://doi.org/10.1038/nature14906 (2015).
Huang, K. L. et al. Integrator recruits protein phosphatase 2A to prevent pause release and facilitate transcription termination. Mol. Cell. 80, 345–58.e9. https://doi.org/10.1016/j.molcel.2020.08.016 (2020).
Zhang, Y. et al. The integrator complex prevents dedifferentiation of intermediate neural progenitors back into neural stem cells. Cell Rep. 27, 987–96.e3. https://doi.org/10.1016/j.celrep.2019.03.089 (2019).
Cheng, L. et al. A 4-gene panel as a marker at chromosome 8q in Asian gastric cancer patients. Genomics 102, 323–330. https://doi.org/10.1016/j.ygeno.2013.05.004 (2013).
Wee, Y., Wang, T., Liu, Y., Li, X. & Zhao, M. A pan-cancer study of copy number gain and up-regulation in human oncogenes. Life Sci. 211, 206–214. https://doi.org/10.1016/j.lfs.2018.09.032 (2018).
Tong, H. et al. INTS8 accelerates the epithelial-to-mesenchymal transition in hepatocellular carcinoma by upregulating the TGF-β signaling pathway. Cancer Manag. Res. 11, 1869–1879. https://doi.org/10.2147/cmar.S184392 (2019).
Jia, D. et al. Integrative analysis of DNA methylation and gene expression to identify key epigenetic genes in glioblastoma. Aging (Albany NY). 11, 5579–5592. https://doi.org/10.18632/aging.102139 (2019).
Edwards, J. R., Yarychkivska, O., Boulard, M. & Bestor, T. H. DNA methylation and DNA methyltransferases. Epigenet. Chromatin. 10, 23. https://doi.org/10.1186/s13072-017-0130-8 (2017).
Andersen, J. B. et al. Genomic and genetic characterization of cholangiocarcinoma identifies therapeutic targets for tyrosine kinase inhibitors. Gastroenterology 142, 1021–31.e15. https://doi.org/10.1053/j.gastro.2011.12.005 (2012).
Sia, D. et al. Integrative molecular analysis of intrahepatic cholangiocarcinoma reveals 2 classes that have different outcomes. Gastroenterology 144, 829–840. https://doi.org/10.1053/j.gastro.2013.01.001 (2013).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883. https://doi.org/10.1093/bioinformatics/bts034 (2012).
Colaprico, A. et al. TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 44, e71. https://doi.org/10.1093/nar/gkv1507 (2016).
Wagner, G. P., Kin, K. & Lynch, V. J. Measurement of mRNA abundance using RNA-seq data: RPKM measure is inconsistent among samples. Theory Biosci. 131, 281–285. https://doi.org/10.1007/s12064-012-0162-3 (2012).
Law, C. W. et al. RNA-seq analysis is easy as 1-2-3 with limma, Glimma and edgeR. F1000Res 5, ISCB. https://doi.org/10.12688/f1000research.9005.3 (2016).
Kolde, R., Laur, S., Adler, P. & Vilo, J. Robust rank aggregation for gene list integration and meta-analysis. Bioinformatics 28, 573–580. https://doi.org/10.1093/bioinformatics/btr709 (2012).
Mayakonda, A., Lin, D. C., Assenov, Y., Plass, C. & Koeffler, H. P. Maftools: Efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 28, 1747–1756. https://doi.org/10.1101/gr.239244.118 (2018).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16, 284–287. https://doi.org/10.1089/omi.2011.0118 (2012).
Becht, E. et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 17, 218. https://doi.org/10.1186/s13059-016-1070-5 (2016).
Nusinow, D. P. et al. Quantitative proteomics of the cancer cell line encyclopedia. Cell 180, 387-402.e16. https://doi.org/10.1016/j.cell.2019.12.023 (2020).
Bahcall, O. G. Human genetics: GTEx pilot quantifies eQTL variation across tissues and individuals. Nat. Rev. Genet. 16, 375. https://doi.org/10.1038/nrg3969 (2015).
Kelley, R. K., Bridgewater, J., Gores, G. J. & Zhu, A. X. Systemic therapies for intrahepatic cholangiocarcinoma. J. Hepatol. 72, 353–363. https://doi.org/10.1016/j.jhep.2019.10.009 (2020).
Rovesti, G. et al. Prognostic role of a new index tested in European and Korean advanced biliary tract cancer patients: The PECS Index. J. Gastrointest. Cancer. https://doi.org/10.1007/s12029-021-00596-z (2021).
Federico, A. et al. Pan-cancer mutational and transcriptional analysis of the integrator complex. Int. J. Mol. Sci. 5, 936. https://doi.org/10.3390/ijms18050936 (2017).
Liu, Z. et al. Follicle-stimulating hormone (FSH) promotes retinol uptake and metabolism in the mouse ovary. Reprod. Biol. Endocrinol. 16, 52. https://doi.org/10.1186/s12958-018-0371-9 (2018).
Marceau, G., Gallot, D., Lemery, D. & Sapin, V. Metabolism of retinol during mammalian placental and embryonic development. Vitam. Horm. 75, 97–115. https://doi.org/10.1016/s0083-6729(06)75004-x (2007).
Romeo, S. & Valenti, L. Regulation of retinol-binding protein 4 and retinol metabolism in fatty liver disease. Hepatology 64, 1414–1416. https://doi.org/10.1002/hep.28722 (2016).
Ohkawa, K. et al. The facilitated effect of retinol on rat hepatocarcinogenesis induced by 3’-methyl-4-dimethylaminoazobenzene. Carcinogenesis 12, 2357–2360. https://doi.org/10.1093/carcin/12.12.2357 (1991).
Xu, N. et al. Identification of key DNA methylation-driven genes in prostate adenocarcinoma: An integrative analysis of TCGA methylation data. J. Transl. Med. 17, 311. https://doi.org/10.1186/s12967-019-2065-2 (2019).
Yongvanit, P. et al. Hepatic cytochrome P450 2A6 and 2E1 status in peri-tumor tissues of patients with Opisthorchis viverrini-associated cholangiocarcinoma. Parasitol. Int. 61, 162–166. https://doi.org/10.1016/j.parint.2011.06.026 (2012).
Currie, E., Schulze, A., Zechner, R., Walther, T. C. & Farese, R. V. Jr. Cellular fatty acid metabolism and cancer. Cell Metab. 18, 153–161. https://doi.org/10.1016/j.cmet.2013.05.017 (2013).
Cheng, C., Geng, F., Cheng, X. & Guo, D. Lipid metabolism reprogramming and its potential targets in cancer. Cancer Commun. (Lond). 38, 27. https://doi.org/10.1186/s40880-018-0301-4 (2018).
Li, L. et al. Differential requirement for de novo lipogenesis in cholangiocarcinoma and hepatocellular carcinoma of mice and humans. Hepatology 63, 1900–1913. https://doi.org/10.1002/hep.28508 (2016).
Williams, A. B. & Schumacher, B. p53 in the DNA-damage-repair process. Cold Spring Harb. Perspect. Med. 5, a026070. https://doi.org/10.1101/cshperspect.a026070 (2016).
Phillips, D. H. Mutational spectra and mutational signatures: Insights into cancer aetiology and mechanisms of DNA damage and repair. DNA Repair (Amst). 71, 6–11. https://doi.org/10.1016/j.dnarep.2018.08.003 (2018).
Yang, Y. et al. The diagnostic value of DNA repair gene in breast cancer metastasis. Sci. Rep. 10, 19626. https://doi.org/10.1038/s41598-020-76577-2 (2020).
Liengswangwong, U. et al. Immunohistochemical expression of mismatch repair genes: A screening tool for predicting mutator phenotype in liver fluke infection-associated intrahepatic cholangiocarcinoma. World J. Gastroenterol. 12, 3740–3745. https://doi.org/10.3748/wjg.v12.i23.3740 (2006).
Sato, Y. et al. Immunosuppressive tumor microenvironment in occupational cholangiocarcinoma: Supportive evidence for the efficacy of immune checkpoint inhibitor therapy. J. Hepatobiliary Pancreat. Sci. 27, 860–869. https://doi.org/10.1002/jhbp.788 (2020).
Winkelmann, R. et al. Microsatellite instability occurs rarely in patients with cholangiocarcinoma: A retrospective study from a German Tertiary Care Hospital. Int. J. Mol. Sci. 5, 1421. https://doi.org/10.3390/ijms19051421 (2018).
Parsons, M. T., Buchanan, D. D., Thompson, B., Young, J. P. & Spurdle, A. B. Correlation of tumour BRAF mutations and MLH1 methylation with germline mismatch repair (MMR) gene mutation status: A literature review assessing utility of tumour features for MMR variant classification. J. Med. Genet. 49, 151–157. https://doi.org/10.1136/jmedgenet-2011-100714 (2012).
Goeppert, B. et al. Mismatch repair deficiency is a rare but putative therapeutically relevant finding in non-liver fluke associated cholangiocarcinoma. Br. J. Cancer. 120, 109–114. https://doi.org/10.1038/s41416-018-0199-2 (2019).
Kim, G. J., Kim, H. & Park, Y. N. Increased expression of Yes-associated protein 1 in hepatocellular carcinoma with stemness and combined hepatocellular-cholangiocarcinoma. PLoS One 8, e75449. https://doi.org/10.1371/journal.pone.0075449 (2013).
Wakizaka, K. et al. CD133 and epithelial cell adhesion molecule expressions in the cholangiocarcinoma component are prognostic factors for combined hepatocellular cholangiocarcinoma. Hepatol. Res. 50, 258–267. https://doi.org/10.1111/hepr.13443 (2020).
Jones, P. A. & Baylin, S. B. The epigenomics of cancer. Cell 128, 683–692. https://doi.org/10.1016/j.cell.2007.01.029 (2007).
Ginder, G. D. & Williams, D. C. Jr. Readers of DNA methylation, the MBD family as potential therapeutic targets. Pharmacol. Ther. 184, 98–111. https://doi.org/10.1016/j.pharmthera.2017.11.002 (2018).
Baylin, S. B. & Jones, P. A. Epigenetic determinants of cancer. Cold Spring Harb. Perspect. Biol. 9, a019505. https://doi.org/10.1101/cshperspect.a019505 (2016).
Lyko, F. The DNA methyltransferase family: A versatile toolkit for epigenetic regulation. Nat. Rev. Genet. 19, 81–92. https://doi.org/10.1038/nrg.2017.80 (2018).
Tamma, R. et al. Inflammatory cells infiltrate and angiogenesis in locally advanced and metastatic cholangiocarcinoma. Eur. J. Clin. Investig. 49, e13087. https://doi.org/10.1111/eci.13087 (2019).
Alnaggar, M. et al. Allogenic Vγ9Vδ2 T cell as new potential immunotherapy drug for solid tumor: A case study for cholangiocarcinoma. J. Immunother. Cancer. 7, 36. https://doi.org/10.1186/s40425-019-0501-8 (2019).
Zhang, M. et al. Single-cell transcriptomic architecture and intercellular crosstalk of human intrahepatic cholangiocarcinoma. J. Hepatol. 73, 1118–1130. https://doi.org/10.1016/j.jhep.2020.05.039 (2020).
Acknowledgements
We sincerely thank the TCGA and GEO database for providing their platforms and contributors for their valuable datasets.
Funding
This research was supported by the Leading Talent of Hundred, Thousand and Ten Thousand Project of Xingliao Gifted Person Program of Liaoning Province (NO. XLYC1905013), the Leading Talent Team of Support Program for High-Level Talent's Innovation of Dalian in 2019, and the Type A Project of Leading Talent's Innovative Research of Dalian.
Author information
Authors and Affiliations
Contributions
D.S. and S.X. contributed to designing the study and revising the manuscript. Q.Z., L.J. and X.S. performed the experiments and drafted the manuscript. D.D., F.G. and Z.W. provided the method support and coordinated the study. W.L. and J.Z. contributed to the figures and tables. All the authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhou, Q., Ji, L., Shi, X. et al. INTS8 is a therapeutic target for intrahepatic cholangiocarcinoma via the integration of bioinformatics analysis and experimental validation. Sci Rep 11, 23649 (2021). https://doi.org/10.1038/s41598-021-03017-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-03017-0
- Springer Nature Limited