Abstract
Downhill folding has been defined as a unique thermodynamic process involving a conformations ensemble that progressively loses structure with the decrease of protein stability. Downhill folders are estimated to be rather rare in nature as they miss an energetically substantial folding barrier that can protect against aggregation and proteolysis. We have previously demonstrated that the prokaryotic zinc finger protein Ros87 shows a bipartite folding/unfolding process in which a metal binding intermediate converts to the native structure through a delicate barrier-less downhill transition. Significant variation in folding scenarios can be detected within protein families with high sequence identity and very similar folds and for the same sequence by varying conditions. For this reason, we here show, by means of DSC, CD and NMR, that also in different pH and ionic strength conditions Ros87 retains its partly downhill folding scenario demonstrating that, at least in metallo-proteins, the downhill mechanism can be found under a much wider range of conditions and coupled to other different transitions. We also show that mutations of Ros87 zinc coordination sphere produces a different folding scenario demonstrating that the organization of the metal ion core is determinant in the folding process of this family of proteins.
Similar content being viewed by others
Introduction
Protein folding is essentially an exploration of conformational space on a funneled energy surface. Several small protein domains fold by two-state equilibrium mechanisms1. Larger domains frequently populate, both transitorily or at equilibrium, intermediate conformations2,3 that can be on-pathway or off-pathway. In this latter case, the intermediate conformation may act as structural traps leading to aggregation phenomena4,5.
Metal-binding proteins represent more than 30% of all folded proteins6. In these proteins the metal cofactor either plays a simply structural role (the most common example is zinc in zinc-finger containing proteins7,8) or plays enzymatic roles (i.e., zinc in metallo-proteases9) to obtain the functional protein. It is also well-known that several proteins involved in degenerative disorders have metal-binding abilities10,11,12,13. Metal-binding happens in cellular environments where the proper metal is either present free in solution or bound to delivery proteins14. However, many in vitro studies have proved that the metal cofactor has clear influences on the structure, stability and conformational dynamics of its cognate protein5,15. Much experimental and theoretical effort has been dedicated to the study of peptides16,17,18 and proteins19,20 folding mechanisms but the comparison between different studies can be limited by differences in the protein constructs and by a variety of experimental conditions and methods employed21.
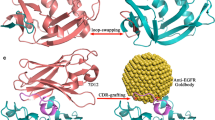
Our group has lately carried out the characterization of the folding mechanism of a metallo-protein belonging to the prokaryotic zinc finger family that we have named Ros875,15 (Fig. 1).
This protein, deletion mutant of Ros from A. tumefaciens22, shares with the eukaryotic domain the tetrahedral coordination of Zn(II) by means of two cysteines and two histidines but differs in the secondary structures topology (Ros87 has a βββαα topology) and in the formation of a large hydrophobic core made by 15 residues23. The study of different members of this protein family24,25,26 allowed us to demonstrate that this protein architecture to functionally fold can substitute the structural ion with a different set of amino acids or can even surrogate the stabilizing role of the metal cofactor with a network of H-bonds and hydrophobic interactions25,27,28,29. These differences among iso-structural proteins led to different mechanisms of folding: the metal lacking Ml452–151 folds/unfolds with a classical two state mechanism while the coordination of the structural zinc confers to Ros87 a partly down-hill unfolding mechanism. As matter of fact, Ros87 shows a bipartite folding/unfolding process in which a metal binding intermediate converts to the native structure through a delicate barrier-less transition. Downhill folders are expected to be quite rare in nature as they miss an energetically significant folding barrier that can protect against aggregation and proteolysis30. Muñoz and co-workers have characterized the downhill folding pathway of the BBL domain31,32,33,34 as a unique thermodynamic state entailing a conformations ensemble that progressively loses structure with the decrease of protein stability35. BBL stability appears particularly sensitive to construct length, the existence of extrinsic fluorophores, ionic strength and pH30,36.
Considering that important variation in folding mechanism can be observed within families of proteins with very similar folds and high sequence identity37 and for the same sequence by varying conditions38, we here describe the characterization of the unfolding mechanism of Ros87 and of one of its mutants in a different buffer with different pH and ionic strength conditions. We have chosen the Tris buffer, a buffer widely used in the characterization of the structure and folding of many proteins. We demonstrate that also in different conditions Ros87 retains the partly downhill folding mechanism demonstrating how the scenario described in the previous work is not simply due to a uniqueness of a particular set of experimental conditions.
Results
Ros87 thermal unfolding
The investigation of Ros87 unfolding mechanism started with the analysis of its thermal denaturation via far-UV CD spectroscopy in 10 mM Tris buffer, 150 µM TCEP, pH = 6.5 (Fig. 2A). The protein shows a complex reversible unfolding pathway spanning a temperature range of 83 K, between 295 and 372 K, that cannot be fitted using a single cooperative two-state model (Fig. 2A,B and Figure SI 1). Accordingly, the DSC thermogram shows a first broad, reversible endotherm centered at about 324 K followed by a second also reversible broad endotherm centered at 360 K (Fig. 2C) pinpointing a two steps unfolding process separated by a detectable partially folded intermediate state.
Ros87 thermal unfolding. (A) CD spectra recorded in Tris buffer at pH = 6.5 in the temperature range 298 K-372 K. (B) CD melting in Tris buffer followed at 222 nm in the temperature range 298–372 K; the inset shows the melting followed at 222 nm in phosphate buffer at pH = 6.515. (C) DSC thermal unfolding curves and fitting of two different heating cycles in Tris buffer. Baselines are reported with a dashed line. (D) DSC thermal unfolding curves and fitting in phosphate buffer15. Panel C and D were adapted from Ref.15.
In the previously published DSC thermogram of the same protein in phosphate buffer15 (Fig. 2D) this second transition occurs between 370 and 390 K, is quite sharper and irreversible. At first, we have focused our attention on the first transition that in phosphate buffer resulted to be properly described by a downhill scenario. Similar to the former study, also in the new experimental conditions it was not possible to extrapolate from the thermogram the native baseline of this transition. Initially, we fitted the curve by the classic two-state formalism but van’t Hoff to calorimetric enthalpy ratio (ΔHvH∕ΔHcal) resulted far from 1, supporting the above mentioned CD findings which indicate that the two-state model is inadequate to describe Ros87 thermal unfolding. This prompted us to fit the DSC thermogram to a variable-barrier energy model proposed by Muñoz and Sanchez-Ruiz33 obtaining the following parameters: Σα = 109 kJ mol−1, T0 = 324.1 K, and β = 1.5 kJ mol−1 that clearly indicate a downhill scenario. Moreover, this probability distribution is unimodal at all temperatures and varies as normally happens for a one-state transition: with the maximum probability shifts from low-enthalpy values at low temperature to high-enthalpy values at high temperature. Downhill folding for Ros87 has been also confirmed by observing that heat capacity baselines for the native and unfolded protein cross at intermediate temperatures.
We then examined Ros87 thermal unfolding by monitoring temperature-induced chemical shift perturbations using NMR spectroscopy. To this aim, a series of 1H–15N HSQC spectra were analyzed in a temperature range from 298 to 343 K at regular intervals of 5 K (Figure SI 2). Resonances exhibited a continuous chemical shift variation indicating a fast protein folding process in the µs-ms time scale39,40 with most of the residues showing a single sigmoidal behavior and only an exiguous number with a more complex behavior (Figure SI 3).
We obtained different midpoint temperatures (Tm) with an approximately Gaussian distribution spanning the same temperature range covered by the first DSC transition. In agreement with DSC, Tm has a mean value of 324.1 ± 0.2 K. As expected, in agreement with the broad non-cooperative reversible transition, groups of protons with similar Tm do not localize in specific structural regions (Fig. 3A) thus defining the lack of hierarchical behavior in this transition. A three state unfolding mechanism was also ruled out, as it was not possible to fit the NMR data with such a model.
Ros87 NMR unfolding. (A) “Atom-by-atom” unfolding behavior of Ros87 in 278–343 K range. Ros87 ribbon drawing showing the Tm of 24 protons mapped on their corresponding atoms. The inset shows the Tm scale. Each atom color corresponds to the Tm of its sigmoidal transition. (B) 1H–15N HSQC spectrum at 343 K.
Interestingly, consistently with what observed in phosphate15, also in Tris some resonances still preserved a good spectral dispersion and small chemical shift perturbations at the highest measured temperatures (Fig. 3B); their mapping on Ros87 NMR structure indicate the existence at 343 K of a metal binding structural intermediate constituted by the β-hairpin containing the two coordinating cysteines, the last turn of the first helix containing the second coordinating histidine and part of the second helix.
As already mentioned above, the DSC thermogram shows a second reversible transition that conversely resulted irreversible in phosphate buffer and attributable to the final loss of the metal ion and of the intermediate residual structure.
The effect of a systematic change of ionic strength and pH conditions in Tris buffer on Ros87 unfolding was also examined via far-UV CD spectroscopy (Figure SI 4). In all the cases, it was not possible to fit the obtained data to either a two- or three-state model. The melting profile obtained appear quite similar to the profile reported in Fig. 2, strongly indicating that Ros87 unfolding is certainly downhill over a wider range of conditions.
Apo-Ros87 and Ros87_C27D thermal unfolding
In order to investigate the second transition of the DSC thermogram, we have decided to explore the thermal denaturation behavior of the apo-protein. We have previously shown that the structural metal ion removal causes severe effects on Ros87 structure and stability41: apo-Ros87 misses tertiary structure interactions and adopts flexible conformations that lack ordered secondary structures as well (Figure SI 5A). However, some slight differences with the complete random coil protein were also evidenced. Therefore, we have decided to investigate its thermal behavior via CD, DSC and NMR. The far-UV CD spectrum of apo-Ros87 was characteristic of an unfolded polypeptide with reduced amount of secondary structure content when compared with that of the metal loaded protein (Figure SI 5B). Yet, the depth of the spectrum around the 222 nm wavelength suggests a residual helical structure content. This could result from the formation of transient α-helices at different sites of the unfolded protein and/or from the presence of a small local core that resists the destabilizing effects of metal removal. In agreement with its largely unfolded state, the protein does not exhibit a cooperative unfolding transition when the CD signal at 222 nm was monitored while raising temperature (Fig. 4A), but again the presence of residual structure is suggested by the decrease in amplitude of the 222 nm signal at 316.4 ± 1.8 K (Fig. 4B and SI 5C).
Apo-Ros87 thermal unfolding. (A) CD spectra recorded at increasing temperatures (298–372 K). (B) CD melting followed at 222 nm in the temperature range 298–372 K. (C) Overlay of the 1H–15N HSQC spectra of apo-Ros87 at 298 K (blue) and at 318 K (red). (D) DSC thermal unfolding curves and fitting of two different heating cycles of apo-Ros87.
NMR data appear in substantial agreement with the CD analysis. In fact, in contrast to Ros87, 1H–15N HSQC spectrum of the apo-form (Figure SI 5A) is poorly dispersed in both proton and nitrogen dimensions and lacks any subset of well dispersed signals indicating that the loss of structure is severe and involves all protein regions and that the residual secondary structure shown by the CD spectrum is not due to a core region that remains folded in spite of the destabilizing effect of the metal lacks. Accordingly, the temperature increase results in a change of the distribution pattern of the 1H–15N HSQC resonances above 318 K (Fig. 4C and SI 6). DSC data (Fig. 4D) reconcile with CD and NMR data showings a first small reversible transition centered at 323.1 (± 0.1)K. Interestingly apo-Ros87 DSC thermogram shows, similarly to Ros87, a second broad reversible transition centered at 364.1 (± 0.2)K. We fitted the DSC profile of this second transition according to the variable-barrier model (Figure SI 7). As expected, the fitted energy barrier value β was very high (30 kJ/mol−1) thus confirming that this unfolding transition cannot be considered downhill. This transition fitted perfectly to a two-state model with a 1.010 (± 0.005) van’t Hoff to calorimetric enthalpy ratio (ΔHvH∕ΔHcal).
In order to understand the conformational determinants governing this second transition we investigated also the thermal behaviour of Ros87_C27D, a single point mutant of Ros87 in which the second coordinating cysteine is mutated in aspartate42,43. This mutant has been previously shown to fold around the zinc ion and to overall preserve Ros87 global fold and positions of the secondary structure elements42. Ros87_C27D shows only minimal structural alterations around the mutation site and for few hydrophobic residues, revealing a slight local reorganization of the protein core. However, these small structural changes resulted in a clear change in the unfolding mechanism observed for this mutant via far-UV CD: unlike the wild-type protein this mutant faces a two-state folding mechanism (Fig. 5A).
Accordingly, the DSC analysis here reported shows for this mutant a behavior strikingly different from that of the wild-type protein. In particular, the DSC thermogram shows a unique reversible transition centered at 338.8 ± 0.2 K accompanied by an enthalpic change of about 182 kJ/mol in the first heating cycle and 136 kJ/mol in the second heating cycle. Cpexc(T) traces (Fig. 5B) fit perfectly to a two-state model with an average ΔHvH∕ΔHcal of 1.003 ± 0.04, supporting the previous mentioned findings which indicate the lack of intermediate species during the transition. DSC data fitted to the variable-barrier model are reported in Figure SI 8. The fitted energy barrier value β was very high (25 kJ/mol−1) also in this case, thus confirming that Ros87_C27D unfolding cannot be considered downhill.
Interestingly, Ros87_C27D thermogram does not show the second transition seen for the wild type protein. Overall, data collected on apo-Ros87 and on Ros87_C27D allow us to conclude that the second transition seen for apo-Ros87 and Ros87 is likely due to the loss of metal coordination (in Ros87) and the concurrent transition to partially insoluble aggregated forms44 involving the two unbound cysteines.
Discussion
The body of data collected in the present work clearly shows that change of buffer, pH and ionic strength does not influence Ros87partly downhill unfolding mechanism. Differences with what was previously seen in phosphate buffer are found in the second transition and are likely due to the different environment found by the cysteines during metal ion release. This however does not influence the folding/unfolding behavior of the protein described by the first DSC transition indicating that the rate-limiting step in Ros87 folding is represented by zinc recruitment and, once metal ion is recruited, a partially folded intermediate forms and the protein moves toward its native functional state with a non-cooperative downhill mechanism.
Downhill folding mechanisms were originally supposed to occur only for proteins with particularly optimized native interactions, under strongly stabilizing conditions35,45 or when favorable mutations occurred36. We aim at contributing to the general discussion indicating that, at least in metallo-proteins, downhill folding can be commonly found under a much wider range of conditions and coupled to other different transitions. The key event in determining the mechanism of folding is the organization of metal ion center that not only stabilizes the native state but is determinant in the folding process of this proteins family.
The finding that mutations of Ros87 zinc coordination sphere trigger different folding mechanisms resembling a more common two-state process suggests that the complex two-step folding mechanism found in Ros87, including a stable downhill scenario, is a precise evolutionary result. The biological advantage of evolving such bipartite folding process needs further investigations but it could have been driven by the necessity to recruit and retain the metal ion in the most diverse chemical conditions.
Methods
Protein expression and purification
Un-labelled and 15N labelled proteins used for CD, NMR and DSC experiments were over-expressed and purified as formerly reported22. Concisely, transformed E. coli host strain BL21(DE3) were plated onto an LB-agar plate containing ampicillin (100 µg/ml) for selection. For 15N labeling, cells were grown at 37° in a minimal medium that contained 0.5 g/l 15NH4Cl as only nitrogen source. At O.D.600 ~ 0.6 the expression was induced with 1.0 mM isopropyl-β-d-thiogalactopyranoside (IPTG) for 1 h 30 min. Cells were harvested by centrifugation (3750 rpm for 40 min) and the pellet was re-suspended in 20 mM Na2HPO4 (pH 6.8) buffer. The suspension was lysed using a sonicator and centrifuged at 16,500 rpm for 40 min. The supernatant was filtered through a 0.22 µm filter membrane to remove cell debris and applied to a Mono S HR 5/5 cation exchange chromatography column (Amersham Bioscences) equilibrated with phosphate buffer. The fractions that contained the proteins were applied to a HiLoad 26/60 Superdex 75 (Amersham Bioscences) gel filtration chromatography column equilibrated with 20 mM Na2HPO4 (pH 6.8), 0.2 M NaCl. For CD and UV–Vis experiments, unlabeled proteins were expressed in LB-medium and the purification procedure described above was followed. An Amicon ultra-15 centrifugal filter was used to concentrate the proteins after the purification phase to reach the desired final concentration. The zinc ion was removed from obtained samples by acidifying Zn(II)-Ros87 at pH ~ 2.5 adding HCl 0.1 M and dialyzing it against 10 mM Tris, 150 µM TCEP (400 µM for NMR samples) aqueous solution at pH ~ 3.0. The pH was fixed to 6.5 and controlled throughout the experiments. Tris buffer and TCEP were chosen to avoid precipitates formation and to prevent cysteine oxidation. Both Tris and TCEP experience a weak affinity for zinc that does not interfere with the measurements46,47.
CD spectroscopy
Ros87 and apo-Ros87 in 10 mM Tris, 150 µM TCEP, at pH 6.5 were thermally denaturated by using a JASCO J-815 CD spectropolarimeter equipped with Peltier temperature control. Experiments were repeated in the same buffer also at different pH values (6, 7.5 and 8.5) and at different ionic strengths (100, 300 and 400 mM NaCl). CD spectra were measured at 5 K intervals in the temperature range of 278–373 K (3 K for Zn(II)-Ros87). At the end, samples were cooled back to 298 K and a final set of spectra recorded. Data were collected using a quartz cuvette with a 1 cm path-length in the 200–260 nm wavelength range with 1 nm data pitch and 50 nm/min scanning speed. Data, normalized against reference spectra to remove buffer background contribution, were fitted by two-state or three-state model.
NMR spectroscopy
NMR samples were made of ~ 250 μM protein solution, 10 mM Tris, 400 μM TCEP adjusted to pH 6.5 in 550 μL of 90% H2O/10% 2H2O. The NMR spectra were recorded on a Bruker Avance III HD 600 MHz, equipped with a cryoprobe, at the Department of Environmental, Biological and Pharmaceutical Sciences and Technologies, University of Campania “L. Vanvitelli” (Caserta, Italy). A series of 1H–15N HSQC spectra were acquired increasing temperatures at regular intervals of 5 K from 298 to 343 K with: 256 complex points for 15N (F1), 1024 for 1H (F2). The 1H and 15N chemical shifts were indirectly calibrated using Tetramethylsilane as external reference. Data were processed using TopSpin 3.5 (Bruker) and NMRPIPE48, analyzed with CARA49 and SPARKY50 software. The structures were visualized and evaluated using the software CHIMERA51 and PyMol52.
Differential scanning calorimetry (DSC)
Protein samples were prepared after extensive dialysis against the buffer (10 mM Tris, 150 µM TCEP, pH 6.5). Next, after vacuum degassing, they were heated at 1 K/min scan rate in the temperature range 278–373 K. A 3 atm extra external nitrogen pressure was applied to samples to prevent formation of air bubbles during heating. In all measurements, the reference cell of the calorimeters was filled by the buffer from the last dialysis step. To ensure an accurate equilibration of the calorimeter, several buffer–buffer heating scans were routinely performed before DSC measurement. To obtain the molar heat capacity curves Cp(T), buffer–buffer/metal baselines were recorded at the same scanning rate, subtracted from raw DSC curves and normalized by the protein concentration. Excess molar heat capacities curves (Cpexc) were obtained from Cp(T), by subtracting a baseline obtained by a fifth-order polynomial fit of pre- and post-transition Cp trends as described elsewhere53. As previously observed for other metallo-proteins54,55, the heat capacity curve of the unfolded protein is significantly distorted due to the concomitant occurrence of phenomena linked to the presence of metal ions. For this reason, we estimated the baseline by using a fifth-order polynomial fit. For all experiments, two consecutive heating–cooling cycles were performed to determine the process reversibility. This value was calculated as the enthalpy change ratio of the second scan (reheating) vs. the first one. Cpexc curves were deconvoluted by NanoAnalyze software using the Gaussians model. The number of DSC components to be adopted in peak deconvolution procedure was selected in order to minimize fitting errors. DSC experiments were run by a NanoDSC instrument (TA Instruments).
References
Olsson, U. & Wolf-Watz, M. Overlap between folding and functional energy landscapes for adenylate kinase conformational change. Nat. Commun. 1, 111. https://doi.org/10.1038/ncomms1106 (2010).
Roder, H. & Colón, W. Kinetic role of early intermediates in protein folding. Curr. Opin. Struct. Biol. 7, 15–28. https://doi.org/10.1016/s0959-440x(97)80004-8 (1997).
Matthews, C. R. Pathways of protein folding. Annu. Rev. Biochem. 62, 653–683. https://doi.org/10.1146/annurev.bi.62.070193.003253 (1993).
Privalov, P. L. Intermediate states in protein folding. J. Mol. Biol. 258, 707–725. https://doi.org/10.1006/jmbi.1996.0280 (1996).
Malgieri, G. et al. Folding mechanisms steer the amyloid fibril formation propensity of highly homologous proteins. Chem. Sci. 9, 3290–3298. https://doi.org/10.1039/c8sc00166a (2018).
Gray, H. B. Biological inorganic chemistry at the beginning of the 21st century. Proc. Natl. Acad. Sci. USA 100, 3563–3568. https://doi.org/10.1073/pnas.0730378100 (2003).
Malgieri, G. et al. The prokaryotic zinc-finger: structure, function and comparison with the eukaryotic counterpart. FEBS J. 282, 4480–4496. https://doi.org/10.1111/febs.13503 (2015).
Isernia, C. et al. Zinc fingers. Met. Ions Life Sci. https://doi.org/10.1515/9783110589757-018 (2020).
Malgieri, G. & Grasso, G. The clearance of misfolded proteins in neurodegenerative diseases by zinc metalloproteases: an inorganic perspective. Coord. Chem. Rev. 260, 139–155 (2014).
Gaeta, A. & Hider, R. C. The crucial role of metal ions in neurodegeneration: the basis for a promising therapeutic strategy. Br. J. Pharmacol. 146, 1041–1059. https://doi.org/10.1038/sj.bjp.0706416 (2005).
Travaglia, A. et al. Zinc(II) interactions with brain-derived neurotrophic factor N-terminal peptide fragments: inorganic features and biological perspectives. Inorg. Chem. 52, 11075–11083. https://doi.org/10.1021/ic401318t (2013).
Travaglia, A. et al. The inorganic perspective of nerve growth factor: interactions of Cu2+ and Zn2+ with the N-terminus fragment of nerve growth factor encompassing the recognition domain of the TrkA receptor. Chemistry 17, 3726–3738. https://doi.org/10.1002/chem.201002294 (2011).
Quintanar, L. & Lim, M. H. Metal ions and degenerative diseases. J. Biol. Inorg. Chem. 24, 1137–1139. https://doi.org/10.1007/s00775-019-01744-4 (2019).
Palm-Espling, M. E., Niemiec, M. S. & Wittung-Stafshede, P. Role of metal in folding and stability of copper proteins in vitro. Biochim. Biophys. Acta 1823, 1594–1603. https://doi.org/10.1016/j.bbamcr.2012.01.013 (2012).
Palmieri, M. et al. Structural Zn(II) implies a switch from fully cooperative to partly downhill folding in highly homologous proteins. J. Am. Chem. Soc. 135, 5220–5228. https://doi.org/10.1021/ja4009562 (2013).
Diana, D. et al. Structural analysis of a helical peptide unfolding pathway. Chemistry 16, 5400–5407. https://doi.org/10.1002/chem.200903428 (2010).
Blasie, C. A. & Berg, J. M. Entropy-enthalpy compensation in ionic interactions probed in a zinc finger peptide. Biochemistry 43, 10600–10604. https://doi.org/10.1021/bi0363230 (2004).
Rich, A. M. et al. Thermodynamics of Zn2+ binding to Cys2His2 and Cys2HisCys zinc fingers and a Cys4 transcription factor site. J. Am. Chem. Soc. 134, 10405–10418. https://doi.org/10.1021/ja211417g (2012).
Dobson, C. M. Protein folding and misfolding. Nature 426, 884–890. https://doi.org/10.1038/nature02261 (2003).
Bhowmik, D., Gao, S., Young, M. T. & Ramanathan, A. Deep clustering of protein folding simulations. BMC Bioinform. 19, 484. https://doi.org/10.1186/s12859-018-2507-5 (2018).
Maxwell, K. L. et al. Protein folding: defining a “standard” set of experimental conditions and a preliminary kinetic data set of two-state proteins. Protein Sci. 14, 602–616. https://doi.org/10.1110/ps.041205405 (2005).
Esposito, S. et al. A novel type of zinc finger DNA binding domain in the Agrobacterium tumefaciens transcriptional regulator Ros. Biochemistry 45, 10394–10405. https://doi.org/10.1021/bi060697m (2006).
Malgieri, G. et al. The prokaryotic Cys2His2 zinc-finger adopts a novel fold as revealed by the NMR structure of Agrobacterium tumefaciens Ros DNA-binding domain. Proc. Natl. Acad. Sci. USA 104, 17341–17346. https://doi.org/10.1073/pnas.0706659104 (2007).
Netti, F. et al. An experimentally tested scenario for the structural evolution of eukaryotic Cys2His2 zinc fingers from eubacterial ros homologs. Mol. Biol. Evol. 30, 1504–1513. https://doi.org/10.1093/molbev/mst068 (2013).
Baglivo, I. et al. The structural role of the zinc ion can be dispensable in prokaryotic zinc-finger domains. Proc. Natl. Acad. Sci. USA 106, 6933–6938. https://doi.org/10.1073/pnas.0810003106 (2009).
D’Abrosca, G. et al. Structural insight of the full-length Ros protein: a prototype of the prokaryotic zinc-finger family. Sci. Rep. 10, 9283. https://doi.org/10.1038/s41598-020-66204-5 (2020).
Baglivo, I. et al. Molecular strategies to replace the structural metal site in the prokaryotic zinc finger domain. Biochim. Biophys. Acta 1844, 497–504. https://doi.org/10.1016/j.bbapap.2013.12.019 (2014).
Pirone, L. et al. Identifying the region responsible for Brucella abortus MucR higher-order oligomer formation and examining its role in gene regulation. Sci. Rep. 8, 17238. https://doi.org/10.1038/s41598-018-35432-1 (2018).
Palmieri, M. et al. Deciphering the zinc coordination properties of the prokaryotic zinc finger domain: the solution structure characterization of Ros87 H42A functional mutant. J. Inorg. Biochem. 131, 30–36. https://doi.org/10.1016/j.jinorgbio.2013.10.016 (2014).
Cho, S. S., Weinkam, P. & Wolynes, P. G. Origins of barriers and barrierless folding in BBL. Proc. Natl. Acad. Sci. USA 105, 118–123. https://doi.org/10.1073/pnas.0709376104 (2008).
Sadqi, M., Fushman, D. & Muñoz, V. Atom-by-atom analysis of global downhill protein folding. Nature 442, 317–321. https://doi.org/10.1038/nature04859 (2006).
Garcia-Mira, M. M., Sadqi, M., Fischer, N., Sanchez-Ruiz, J. M. & Muñoz, V. Experimental identification of downhill protein folding. Science 298, 2191–2195. https://doi.org/10.1126/science.1077809 (2002).
Muñoz, V. & Sanchez-Ruiz, J. M. Exploring protein-folding ensembles: a variable-barrier model for the analysis of equilibrium unfolding experiments. Proc. Natl. Acad. Sci. USA 101, 17646–17651. https://doi.org/10.1073/pnas.0405829101 (2004).
Muñoz, V., Sadqi, M., Naganathan, A. N. & de Sancho, D. Exploiting the downhill folding regime via experiment. HFSP J. 2, 342–353. https://doi.org/10.2976/1.2988030 (2008).
Bryngelson, J. D., Onuchic, J. N., Socci, N. D. & Wolynes, P. G. Funnels, pathways, and the energy landscape of protein folding: a synthesis. Proteins 21, 167–195. https://doi.org/10.1002/prot.340210302 (1995).
Johnson, C. M. & Sharpe, T. D. Protein Folding, Energy Landscapes and Downhill Protein Folding Scenarios. In Encyclopedia of Biophysics (eds Roberts G. & Watts A.) 1–19 (Springer, Berlin, 2018).
Ivarsson, Y., Travaglini-Allocatelli, C., Brunori, M. & Gianni, S. Mechanisms of protein folding. Eur. Biophys. J. 37, 721–728. https://doi.org/10.1007/s00249-007-0256-x (2008).
Oliveberg, M. & Wolynes, P. G. The experimental survey of protein-folding energy landscapes. Q. Rev. Biophys. 38, 245–288. https://doi.org/10.1017/S0033583506004185 (2005).
Fung, A., Li, P., Godoy-Ruiz, R., Sanchez-Ruiz, J. M. & Muñoz, V. Expanding the realm of ultrafast protein folding: gpW, a midsize natural single-domain with alpha+beta topology that folds downhill. J. Am. Chem. Soc. 130, 7489–7495. https://doi.org/10.1021/ja801401a (2008).
Godoy-Ruiz, R. et al. Estimating free-energy barrier heights for an ultrafast folding protein from calorimetric and kinetic data. J. Phys. Chem. B 112, 5938–5949. https://doi.org/10.1021/jp0757715 (2008).
Sivo, V. et al. Ni(II), Hg(II), and Pb(II) coordination in the prokaryotic zinc-finger Ros87. Inorg. Chem. 58, 1067–1080. https://doi.org/10.1021/acs.inorgchem.8b02201 (2019).
D’Abrosca, G. et al. The (unusual) aspartic acid in the metal coordination sphere of the prokaryotic zinc finger domain. J. Inorg. Biochem. 161, 91–98. https://doi.org/10.1016/j.jinorgbio.2016.05.006 (2016).
Sivo, V. et al. Co(II) Coordination in prokaryotic zinc finger domains as revealed by UV–Vis spectroscopy. Bioinorg. Chem. Appl. 2017, 1527247. https://doi.org/10.1155/2017/1527247 (2017).
Vassall, K. A. et al. Decreased stability and increased formation of soluble aggregates by immature superoxide dismutase do not account for disease severity in ALS. Proc. Natl. Acad. Sci. USA 108, 2210–2215. https://doi.org/10.1073/pnas.0913021108 (2011).
Gruebele, M. Downhill protein folding: evolution meets physics. C. R. Biol. 328, 701–712. https://doi.org/10.1016/j.crvi.2005.02.007 (2005).
Magyar, J. S. & Godwin, H. A. Spectropotentiometric analysis of metal binding to structural zinc-binding sites: accounting quantitatively for pH and metal ion buffering effects. Anal. Biochem. 320, 39–54 (2003).
Krezel, A., Latajka, R., Bujacz, G. D. & Bal, W. Coordination properties of tris(2-carboxyethyl)phosphine, a newly introduced thiol reductant, and its oxide. Inorg. Chem. 42, 1994–2003. https://doi.org/10.1021/ic025969y (2003).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Keller, R. The Computer Aided Resonance Assignment Tutorial (Cantina-Verlag, Switzerland, 2004).
Lee, W., Tonelli, M. & Markley, J. L. NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31, 1325–1327. https://doi.org/10.1093/bioinformatics/btu830 (2015).
Pettersen, E. F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612. https://doi.org/10.1002/jcc.20084 (2004).
PyMOL: An Open-Source Molecular Graphics Tool (DeLano Scientific, San Carlos, California, USA, 2002).
Grasso, D., La Rosa, C., Milardi, D. & Fasone, S. The effects of scan rate and protein concentration on DSC thermograms of bovine superoxide dismutase. Thermochim. Acta 265, 163–175. https://doi.org/10.1016/0040-6031(95)02402-N (1995).
Roe, J. A. et al. Differential scanning calorimetry of Cu, Zn-superoxide dismutase, the apoprotein, and its zinc-substituted derivatives. Biochemistry 27, 950–958. https://doi.org/10.1021/bi00403a017 (1988).
La Rosa, C., Milardi, D., Grasso, D., Guzzi, R. & Sportelli, L. Thermodynamics of the thermal unfolding of azurin. J. Phys. Chem. 99, 14864–14870 (1995).
Acknowledgements
Financial support was provided by Italian MIUR (Grant Number 2017WBZFHL). S.G. acknowledges the European Union Horizon 2020 research and innovation program for funding her Ph.D. fellowship under the Marie Sklodowska-Curie Grant Agreement INCIPIT No. 665403.
Author information
Authors and Affiliations
Contributions
R.G., S.G. and G.D.: acquired and analyzed all the NMR and DSC experiments. I.B.: expressed and purified the protein samples. P.V.P., D.M., R.F. and C.I.: designed the experiments and supervised the study. L.R. and G.M.: conceived and designed the study and wrote the manuscript. All the authors revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Grazioso, R., García-Viñuales, S., D’Abrosca, G. et al. The change of conditions does not affect Ros87 downhill folding mechanism. Sci Rep 10, 21067 (2020). https://doi.org/10.1038/s41598-020-78008-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-78008-8
- Springer Nature Limited