Abstract
Antibodies are proteins working in our immune system with high affinity and specificity for target antigens, making them excellent tools for both biotherapeutic and bioengineering applications. The prediction of antibody affinity changes upon mutations (\({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\)) is important for antibody engineering. Numerous computational methods have been proposed based on different approaches including molecular mechanics and machine learning. However, the accuracy by each individual predictor is not enough for efficient antibody development. In this study, we develop a new prediction method by combining multiple predictors based on machine learning. Our method was tested on the SiPMAB database, evaluating the Pearson’s correlation coefficient between predicted and experimental \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\). Our method achieved higher accuracy (R = 0.69) than previous molecular mechanics or machine-learning based methods (R = 0.59) and the previous method using the average of multiple predictors (R = 0.64). Feature importance analysis indicated that the improved accuracy was obtained by combining predictors with different importance, which have different protocols for calculating energies and for generating mutant and unbound state structures. This study demonstrates that machine learning is a powerful framework for combining different approaches to predict antibody affinity changes.
Similar content being viewed by others
Introduction
Antibodies are proteins working in our immune system that bind to target molecules named antigen such as proteins or chemical ligands with high affinity and specificity. Over the past two decades, antibodies have become popular as biotherapeutics1. Antibodies have important advantages over small-molecule drugs such as antibody dependent cellular cytotoxicity2 and complement dependent cytotoxicity activity3. In addition, antibody–drug conjugates can kill tumor cells with high efficiency4,5. Recently, a single chain fragment variable region of an antibody is used as a receptor for chimeric antigen receptor T-cell therapy6,7, highlighting the adaptability and efficacy of antibodies as biotherapeutics. Antibody engineering is used to improve the properties of antibodies such as affinity, specificity, solubility, and stability. In particular, improving affinity is important for increasing drug efficacy and decreasing the amount of antibody per dose, thereby reducing the drug price. The affinity of an antibody can be improved by introducing mutations in its amino acid sequence while in practice not many mutations increase affinity8. To date, improving affinity requires trial and error, making many mutants and measuring their affinities to identify mutants of interest.
The affinity of an antibody is evaluated by the binding free energy (\({\mathrm{\Delta G}}_{\mathrm{binding}}\)). \({\mathrm{\Delta G}}_{\mathrm{binding}}\) is calculated by the free energy of the bound state minus that of the unbound state. \({\mathrm{\Delta G}}_{\mathrm{binding}}\) is experimentally measured with surface plasmon resonance (SPR), isothermal titration calorimetry (ITC), or enzyme-linked immune-sorbent assay. Although SPR and ITC have high sensitivity, measuring many samples with SPR and ITC requires substantial time and cost. Therefore, it is important for antibody engineering to develop a method for predicting mutants with high affinity prior to experimental evaluation9,10.
A number of software tools have been developed for predicting binding affinity of complexes11,12, some of which are proposed for general protein complexes while others are dedicated specifically to antibody-antigen complexes13,14. These methods are largely divided into two approaches: molecular mechanics and machine learning. The molecular mechanics methods are based on the evaluation of energies calculated from protein structures15,16. Each method utilizes a different scoring function to calculate energies. The typical terms considered in a scoring function include hydrogen bonding, conformational energies, solvation energies, and entropic terms in addition to Coulombic and van der Waals interaction energies17. Normally, the molecular mechanics methods take as input the structure of a wild-type complex only, and mutant structures and structures in the unbound state are computationally generated (i.e. structure regeneration). Therefore, the performance of molecular mechanics methods depends on the choice of scoring functions and structure regeneration methods. Sulea et al.17 have presented a benchmark study to investigate the effect of scoring functions and structure regeneration methods on the prediction accuracy. As an approach different from molecular mechanics, the machine learning methods are proposed based on statistical models that predict affinity changes upon mutations using feature values calculated from protein complex structures13,18. The performance of machine learning methods is determined by the choice of statistical models and feature values.
Sulea et al.17 have also proposed a prediction method in their benchmark study. Their prediction method, termed consensus scoring, is defined as the average of predicted affinity changes calculated by multiple molecular mechanics methods (multiple predictors). In detail, the Z score is calculated for each of predictors for adjusting their difference in mean and standard deviation. Then, the consensus score is calculated as the average of the Z scores of predictors. The consensus scoring method has shown higher prediction accuracy than any of individual molecular mechanics methods (single predictors). However, the consensus scoring method does not consider the different importance of predictors since the method simply takes the average of the Z scores of predictors, assuming all features are equally important. In addition, the predictors used in the consensus scoring method have been selected empirically, thus the best combination of predictors for improving accuracy is unknown.
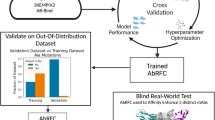
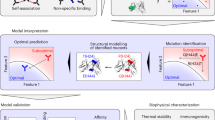
Here, we propose a new computational method for the prediction of antibody affinity changes upon mutations. Our method combines multiple predictors using machine learning. In contrast to the consensus scoring method based on the average of multiple predictors, the use of machine learning enables us to combine multiple predictors with different importance adjusted in model training. The machine learning model takes predictions from multiple methods as feature values (Fig. 1). These predictors include a variety of molecular mechanics predictors with various scoring functions and structure regeneration methods as well as a previous machine-learning-based predictor. In experiments on the SiPMAB database, our method achieves higher prediction accuracy than the best single predictor and the consensus scoring method. We present feature importance analysis to evaluate the contribution of each predictor in our method, showing that the improved accuracy is obtained by combining predictors using different scoring functions and structure regeneration methods. Moreover, we show that the number of combined predictors can be reduced according to the feature importance without compromising the accuracy.
Results
Prediction accuracy improved by combining multiple predictors
We compared our method with the consensus scoring method based on the average of multiple predictors and the 12 kinds of single predictors used as feature values in our method (“Methods” section). As proposed in the previous study17, we used the consensus scoring method with 3 predictors (Cons3 with SIE-Scwrlmut, Rosmut and FoldX-S) and that with 4 predictors (Cons4 with SIE-Scwrlmut, Rosmut, FoldX-S and FoldX-B). Figure 2 shows the Pearson’s correlation coefficient between predicted scores and experimental \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) on the SiPMAB dataset. Our method with GPR and RFR achieved R = 0.69 and R = 0.67, respectively, showing better accuracy than Cons3 (R = 0.63) and Cons4 (R = 0.64). These results demonstrate the effectiveness of machine learning for combining multiple predictors to improve the prediction accuracy.
Comparison of different methods on the SiPMAB dataset. The bar graph shows the Pearson's correlation coefficient between predicted scores and experimental \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) in the SiPMAB dataset. Left: single predictors; Middle: Consensus scoring method; Right: the proposed method. The error bar represents the standard error of the mean (SEM) from 100 calculations using the different splits of subsets in cross validation. P-value was calculated using the Wilcoxon signed rank test.
The best single predictor was Rosmut with R = 0.59. For each molecular mechanics software, R > 0.50 was achieved by using the best choice of scoring functions and structure regeneration methods: SIE-Scwrlmut, Rosmut, FoldX-B, and DS-B. The accuracy of FoldX-S and DS-S was lower than the other methods, which may be because these methods are based on the stability Eq. (3) rather than the binding free energy Eq. (1).
We compared the distribution of predicted scores for each method with experimental \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) (Supplementary Fig. S1). We found that there were outliers in the predictions of Rosiface-sc and RosCDR-loop. Notably, our method with GPR and RFR showed few outliers while it used these features. Such a robustness may be another merit of machine learning for combining multiple predictors.
Analysis of feature importance
An advantage of machine learning is the ability to evaluate the importance of each feature in terms of its contribution to the prediction. We used the feature importance method based on Gini index19 implemented in scikit-learn package (Fig. 3). The most important feature was Rosmut, which also achieved the best accuracy among the single predictors (Fig. 2). Similarly, the feature with the second-highest accuracy, SIE-Scwrlmut, showed the second-highest feature importance whereas the tendency for the rank of accuracy and the rank of feature importance to become equal did not apply to the other features. The importance was above 0.1 for 4 features: Rosmut, SIE-Scwrlmut, mCSM-AB, and DS-B. Interestingly, those predictors were based on different prediction approaches (molecular mechanics or machine learning), and different scoring functions and structure regeneration methods for molecular mechanics. These results suggest that the improved accuracy of our method was obtained by combining predictors based on different principles.
Correlation between predictors
We also evaluated the Pearson’s correlation coefficient between different predictors (Fig. 4). Among the 4 predictors with high feature importance, the molecular mechanics predictors (Rosmut, SIE-Scwrlmut, and DS-B) were similar to each other (R > 0.66) with Rosmut and DS-B showing the highest correlation. On the other hand, mCSM-AB based on machine learning was distinct from the other predictors (e.g. R = 0.50 between mCSM-AB and Rosmut). These results further support that combining predictors based on different principles may contribute to improving prediction accuracy.
Reduced features
Although our method uses 12 predictors as input, the number of predictors may be reduced, which is desirable for reducing the computational cost. Thus, we developed a prediction method combining only four predictors: Rosmut, SIE-Scwrlmut, mCSM-AB, and DS-B whose feature importance was higher than the others (Fig. 3). Our method using the reduced feature set achieved the accuracy comparable to that using the full feature set (Fig. 5). Using GPR as a machine learning model, the Pearson's correlation coefficient by our method was still higher than that of Cons4 (R = 0.67 compared with R = 0.64, P < 10–15; Wilcoxon signed-rank test). These results indicate that the number of features used for our method can be reduced without compromising prediction accuracy.
Comparison of accuracy using different feature sets. The bar graph shows the Pearson's correlation coefficient between predicted scores and experimental \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) in the SiPMAB dataset. Error bar represents the SEM from 100 calculations using different splits of subsets in cross validation.
Evaluation on independent data
In addition to the cross-validation-based evaluation on SiPMAB database, we performed a benchmark study on independent data not included in SiPMAB database (Methods). We compared our method using the reduced feature set with the 4 kinds of single predictors used as feature values in our method. Figure 6a shows the Pearson’s correlation coefficient between predicted scores and experimental \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) on 34 mutants of an antibody targeting vascular endothelial growth factor (VEGF), called bH1, from AB-Bind database20,21. Our method with GPR and RFR achieved R = 0.54 and R = 0.60, respectively, showing better accuracy than the best single predictor mCSM-AB with R = 0.47. We increased the data size by combining the bH1 data with additional independent data of 12 mutants of an antibody targeting monocyte chemo-attractant protein-1(MCP-1), called 11K2, from Kiyoshi et al.22, and also confirmed that our method achieved better accuracy than the single predictors (Fig. 6b). These results demonstrate the effectiveness of machine learning for combining multiple predictors to improve prediction accuracy, not only for SiPMAB database but also for independent data.
Comparison of different methods on independent data not included in SiPMAB database. The bar graph shows the Pearson's correlation coefficient between predicted scores and experimental \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\). (a) bH1 data (n = 34). (b) bH1 data combined with 11K2 data (n = 46). *P < 0.05, **P < 0.01, ***P < 0.005.
Discussion
Numerous \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) prediction methods have been developed with a variety of scoring functions and structure regeneration methods. However, due to the characteristics of each method, \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) prediction with high accuracy has been difficult. In this study, we demonstrated that the prediction accuracy can be improved by combining multiple predictors using machine learning. Our method with GPR achieved R = 0.69 on the SiPMAB database (Fig. 2), which was more accurate than the best single predictor (Rosmut, R = 0.59) and the consensus scoring method based on the average of multiple predictors (Cons4, R = 0.64). The feature importance analysis suggested that Rosmut, SIE-Scwrlmut, mCSM-AB, and DS-B were particularly important for the improved accuracy (Fig. 3). Our method using these 4 features kept the prediction accuracy comparable to that using the full feature set (Fig. 5). Moreover, our method using these 4 features achieved higher accuracy than single predictors in the benchmark study on the independent data not included in SiPMAB database (Fig. 6). In addition, the feature importance analysis suggested that \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{folding}}\) (DS-S and FoldX-S) was not so important for the improved accuracy (Fig. 3).
The Pearson’s correlation coefficient between predictors ranged from 0.5 to 0.8 (Fig. 4). This result indicates that each predictor has unique information derived from different prediction approaches (molecular mechanics or machine learning), and different scoring functions and structure regeneration methods for molecular mechanics. In particular, the Pearson’s correlation coefficient between mCSM-AB and the other predictors based on molecular mechanics was lower than the other pairs. This result suggests that combining predictors based on molecular mechanics and machine learning is important for accuracy.
We note that our method has the limitations summarized below. First, although our method achieved higher accuracy than single predictors and the previous method using the average of multiple predictors, our methods requires a relatively high computational cost. Second, although our method achieved higher accuracy, it requires training data. On the other hand, the consensus scoring method and single predictors do not require training data. Third, our method, like other existing methods, requires the three-dimensional structure of the antigen–antibody complex. However, antibody-antigen complexes are easier to crystalize than monomers because the complexes are normally stable23. In addition, complex structures can be predicted using homology modeling24,25 and docking simulation26. In this study, we focused on affinity changes upon single point mutations as in previous studies. Nonetheless, our method can be easily extended to multiple point mutations by using scores of multiple point mutants as feature values.
In conclusion, our method performs the best for predicting affinity changes upon mutations of antibody-antigen complexes (\({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\)). The method is more accurate than the single predictors and the consensus scoring method using the average of multiple predictors. The improved accuracy is obtained by combining multiple predictors with different importance using machine learning. Our method can contribute to the design of antibodies for therapeutics and diagnostics by improving speed and reducing the associated costs.
Methods
Overview of the proposed method
The idea of our method is to combine multiple predictors for antibody affinity changes using machine learning (Fig. 1). The machine learning model takes as input predictions from multiple methods as feature values, and outputs the \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) as the final prediction. These predictors (feature values) included those based on molecular mechanics with different scoring functions and structure regeneration methods (Table S1). In addition, we also employed a previous machine-learning-based predictor as a feature value in our method (Table S1). We used two different machine learning models and compared their performance: gaussian process regression (GPR)27 and random forest regressor (RFR)28. GPR and RFR are one of the most popular machine learning models, which have been used for study such as antibody engineering field29,30. As an advantage of the use of machine learning, our method can evaluate the importance of each feature in terms of its contribution to the prediction. Specifically, we evaluated the feature importance based on the Gini index in RFR19. Our method was implemented in Python using scikit-learn package31.
Predictors based on molecular mechanics
\({\mathrm{\Delta G}}_{\mathrm{binding}}\) of an antigen–antibody complex is calculated with Eq. (1). \({\mathrm{G}}_{\mathrm{Ag}+\mathrm{Ab}}\) is the Gibbs free energy of the antigen–antibody complex. \({\mathrm{G}}_{\mathrm{Ag}}\) and \({\mathrm{G}}_{\mathrm{Ab}}\) are the Gibbs free energies of the unbound state of the antigen and the antibody, respectively.
The change in the binding energy after mutagenesis (\({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\)) is calculated with Eq. (2). \({\mathrm{\Delta G}}_{\mathrm{binding }}^{\mathrm{mutant}}\) and \({\mathrm{\Delta G}}_{\mathrm{binding }}^{\mathrm{wild}-\mathrm{type}}\) are the \({\mathrm{\Delta G}}_{\mathrm{binding}}\) of the mutant and the wild-type complexes, respectively.
The stability of an antigen–antibody complex is also calculated because it is related to binding free energy17. The stability (\({\mathrm{\Delta G}}_{\mathrm{folding}}\)) of an antigen–antibody complex is calculated with Eq. (3).
\({\mathrm{G}}_{\mathrm{fold}}\) and \({\mathrm{G}}_{\mathrm{unfold}}\) are the Gibbs free energies of the folded state and the unfolded state, respectively. The change in the structure stability after mutagenesis (\({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{folding}}\)) is calculated with Eq. (4) \({\mathrm{\Delta G}}_{\mathrm{folding }}^{\mathrm{mutant}}\) and \({\mathrm{\Delta G}}_{\mathrm{folding }}^{\mathrm{wild}-\mathrm{type}}\) are the \({\mathrm{\Delta G}}_{\mathrm{folding}}\) of the mutant and the wild-type complexes, respectively.
In this study, we used 11 molecular mechanics predictors as feature values in our method. Among them, 9 predictors have been evaluated in the previous benchmark study17, while 2 predictors were newly employed in this study. Each predictor was different in the choice of a scoring function and a structure regeneration method, in addition to whether it used the binding energy Eq. (1) or the stability Eq. (3). The scoring functions included SIE32, Talaris201315, Talaris-interface33, CHARMm Polar H16 and FOLDEF34. For regenerating mutant structures from the wild-type complex structure, only the side chain at the mutated site was repacked with the other residues fixed, or the side chains around the mutated site were also repacked (see the details below). Structures in unbound state were refined by separating the antibody and the antigen as rigid bodies, or by refining their structures after the separation. Below, for clarity, we divide the 11 predictors into 4 groups: Discovery Studio (2 predictors), FoldX (2 predictors), Rosetta (6 predictors), SIE-Scwrlmut (1 predictor).
Parent structure preparation
Predictors based on molecular mechanics require a parent structure that is prepared from an experimental structure for computational analyses. In this study, we used the parent structures provided by SiPMAB database for antibodies included in SiPMAB database. For other antibodies not included in SiPMAB database, we prepared the parent structures using the same procedure as SiPMAB database according to Sulea et al.17 Briefly, the starting structure was retrieved from the protein data bank (PDB ID: 3BDY for the anti-VEGF antibody; 2BDN for the anti-MCP-1-antibody), and we removed non-protein compounds including waters and ions, and deleted non-variable domains in the antibody. Protons were added with neutral pH condition. The structure was energy-minimized using Amber force field35,36.
Discovery studio
Discovery Studio37 is biomolecular simulation software where CHARMm Polar H force field16 is used as a scoring function. Two types of protocols were used by Discovery Studio 2018: DS-B and DS-S. \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) (DS-B) was calculated by the “Calculate Mutation Energy (Binding)” protocol, and \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{folding}}\)(DS-S) was calculated by the “Calculate Mutation Energy (Stability)” protocol. The structure of the mutant was refined with repacking and energy minimization of the side chain at the mutated site. The structures in unbound state were refined by rigid separation. All runs were performed with default parameters.
FoldX, SIE-Scwrlmut, and Rosetta
Predictions for SiPMAB by FoldX34 (FoldX-B and FoldX-S), SIE-Scwrlmut17, and Rosetta38 (SIE-Rosmut, SIE-Rosiface-sc, SIE-RosCDR-loop , Rosmut, Rosiface-sc, and RosCDR-loop) were obtained from the previous benchmark study17. Predictions for the complex of anti-VEGF antibody and VEGF were calculated according to the previous benchmark study17. The descriptions of these methods were shown in Table S1. Briefly, FoldX is protein free energy calculation software using FOLDEF as a scoring function. Two types of protocols were used by FoldX: FoldX-B and FoldX-S using \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) and \({{\Delta \Delta {\mathrm{G}}}}_{{{\text{folding}}}}\), respectively. SIE-Scwrlmut uses 2 software and a scoring function: SCWRL is software for regenerating protein structures based on empirical side chain rotamers. Amber is software for molecular dynamics simulation and SIE is a scoring function32,39. In this protocol, mutant structures after refined by SCWRL with repacking and energy minimization of mutated side chains were further energy-minimized around mutated sites using Amber, and then \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) was calculated using SIE. Rosetta suite40 is a protein design and structure prediction tool based on Monte Carlo simulation. It is capable of predicting scores and generating a mutant structure with backrub sampling of the backbone and repacking of side chains. Six types of protocols were used by Rosetta: SIE-Rosmut, SIE-Rosiface-sc, SIE-RosCDR-loop, Rosmut, Rosiface-sc, and RosCDR-loop. Rosetta employed 3 scoring functions: Talaris201315 (Rosiface-sc and RosCDR-loop), Talaris-interface33 (Rosmut), and SIE32 (SIE-Rosmut, SIE-Rosiface-sc, and SIE-RosCDR-loop).
Machine-learning-based predictor (mCSM-AB)
In addition to molecular mechanics predictors described above, we also used a previous machine-learning-based predictor, mCSM-AB13, as a feature value in our method. mCSM-AB is a machine learning model that predicts antibody affinity changes using the graph-based signatures of protein structures. In the previous study, the model has been trained using experimental \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) from the AB-Bind database20. To use mCSM-AB in our method, we took care to prevent the leakage of training data into the performance evaluation (see the “Performance evaluation” section for details).
Datasets
To assess prediction accuracy, a dataset from the SiPMAB database17 was used. This dataset is comprised of 212 single point mutant antibodies in their CDRs, across 7 different antibody-antigen complexes. The wild-type structures of the antibody-antigen complexes are available, which are solved by high resolution X-ray crystallography. The majority of experimental binding free energies are measured by SPR and ITC. The \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) values range between − 0.65 and 7.32 kcal/mol. When the multiple \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) measurements are recorded for the same mutant, which originates from different publications, we selected one \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) value as previously described17. Briefly, the \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) value was selected considering the reliability of assay methods, and the scale of the assay in the original publication. Although AB-Bind20 and SKEMPI41 are another database for affinity changes upon mutations, we used the SiPMAB database since it collects mutants on antibodies excluding mutants on antigens, and thus is more suitable for the purpose of our study.
For evaluation on independent data, 34 mutants of the complex of anti-VEGF antibody, called bH1, and VEGF were collected from the AB-Bind. These mutants are not included in SiPMAB database, and have mutations in the antibody side. We also used the data of 12 mutants of the complex of anti-MCP-1 antibody, called 11K2, and MCP-1 from Kiyoshi et al.22.
Performance evaluation
We evaluated the Pearson's correlation coefficient between predicted and experimental \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) as a measure of prediction accuracy. To compare the performance of our method with previous methods, we conducted the following procedures to ensure a fair comparison avoiding potential overfitting.
For RFR, we used fourfold nested cross-validation for optimizing a hyperparameter in our method (the number of trees). In the outer loop of the nested cross-validation procedure, the dataset was split into 4 subsets where each subset was used as an independent test dataset named Test and the remaining data were used for the inner loop. In the inner loop of the nested cross-validation procedure, the dataset was split into 4 subsets where each subset was used as a validation dataset named Validation, and the remaining data were named Training. A model was trained using Training dataset with various hyperparameter values, and the performance was measured using Validation dataset. The performance of our method with the best hyperparameter value was evaluated using the Test dataset. The Pearson's correlation coefficient between predicted and experimental \({{\Delta \Delta {\mathrm{G}}}}_{\mathrm{binding}}\) was calculated for each Test dataset, and the average value was used as a final evaluation measure. For GPR, we used the radial basis function (RBF) kernel with a constant kernel and a white kernel. The hyperparameters of GPR can be optimized during model training without looking at validation or test datasets by maximizing log-marginal-likelihood as implemented in scikit-learn package. Thus, we conducted a fourfold cross-validation rather than a fourfold nested cross-validation.
One of the predictors used in our method, mCSM-AB, was itself based on machine learning. Therefore, we took care to ensure that the data used for training mCSM-AB were always separated from the training data of our method. The mCSM-AB implemented in the public web server (https://biosig.unimelb.edu.au/mcsm_ab/prediction) has been trained using the AB-Bind database. Thus, we checked the overlap of data between the AB-Bind and SiPMAB databases. For each mutant in the SiPMAB database, when the mutant did not exist in the AB-Bind, we simply used the mCSM-AB web server for calculating the feature value in our method. When the mutant existed in the AB-Bind, we used the mCSM-AB predictions in the "Predictions on cross validation" provided by the developers of mCSM-AB (https://biosig.unimelb.edu.au/mcsm_ab/data) rather than the web server. These mCSM-AB predictions were obtained from the tenfold cross-validation where the mutant was separated from the training data.
In addition to cross-validation-based evaluation above, we performed a benchmark study where our method was trained on SiPMAB database, and evaluated on independent data not included in SiPMAB database. For this purpose, we used the bH1 data of the anti-VEGF antibody and the 11K2 data from Kiyoshi et al.22.
Data availability
Our method was implemented in Python using scikit-learn package. The codes and datasets for reproducing the results in this study are available at the authors' GitHub website: https://github.com/ykurumida/ab-predictor.
References
Kaplon, H. & Reichert, J. M. Antibodies to watch in 2019. Mabs-Austin 11, 219–238 (2019).
Wang, W., Erbe, A. K., Hank, J. A., Morris, Z. S. & Sondel, P. M. NK cell-mediated antibody-dependent cellular cytotoxicity in cancer immunotherapy. Front. Immunol. 6, 368 (2015).
Pawluczkowycz, A. W. et al. Binding of submaximal C1q promotes complement-dependent cytotoxicity (CDC) of B cells opsonized with anti-CD20 mAbs Ofatumumab (OFA) or rituximab (RTX): considerably higher levels of CDC are induced by OFA than by RTX. J. Immunol. 183, 749–758 (2009).
Beck, A., Goetsch, L., Dumontet, C. & Corvaia, N. Strategies and challenges for the next generation of antibody drug conjugates. Nat. Rev. Drug Discov. 16, 315–337 (2017).
Polakis, P. Antibody drug conjugates for cancer therapy. Pharmacol. Rev. 68, 3–19 (2016).
Neelapu, S. S. et al. Axicabtagene ciloleucel CAR T-cell therapy in refractory large B-cell lymphoma. N. Engl. J. Med. 377, 2531–2544 (2017).
Kochenderfer, J. N. et al. Construction and preclinical evaluation of an anti-CD19 chimeric antigen receptor. J. Immunother. 32, 689–702 (2009).
Daugherty, P. S., Chen, G., Iverson, B. L. & Georgiou, G. Quantitative analysis of the effect of the mutation frequency on the affinity maturation of single chain Fv antibodies. Proc. Natl. Acad. Sci. USA 97, 2029–2034 (2000).
Vivcharuk, V. et al. Assisted design of antibody and protein therapeutics (ADAPT). PLoS ONE 12, e0181490 (2017).
Gromiha, M. M. & Yugandhar, K. Integrating computational methods and experimental data for understanding the recognition mechanism and binding affinity of protein-protein complexes. Prog. Biophys. Mol. Biol. 128, 33–38 (2017).
Gromiha, M. M., Yugandhar, K. & Jemimah, S. Protein–protein interactions: scoring schemes and binding affinity. Curr. Opin. Struct. Biol. 44, 31–38 (2017).
Geng, C. L., Xue, L. C., Roel-Touris, J. & Bonvin, A. M. J. J. Finding the ΔΔG spot: are predictors of binding affinity changes upon mutations in protein-protein interactions ready for it?. Wires Comput. Mol. Sci. 9, e1410 (2019).
Pires, D. E. & Ascher, D. B. mCSM-AB: a web server for predicting antibody-antigen affinity changes upon mutation with graph-based signatures. Nucleic Acids Res. 44, W469-473 (2016).
Myung, Y., Rodrigues, C. H. M., Ascher, D. B. & Pires, D. E. V. mCSM-AB2: guiding rational antibody design using graph-based signatures. Bioinformatics 36, 1453–1459 (2020).
Leaver-Fay, A. et al. Scientific benchmarks for guiding macromolecular energy function improvement. Method Enzymol. 523, 109–143 (2013).
Neria, E., Fischer, S. & Karplus, M. Simulation of activation free energies in molecular systems. J. Chem. Phys. 105, 1902–1921 (1996).
Sulea, T., Vivcharuk, V., Corbeil, C. R., Deprez, C. & Purisima, E. O. Assessment of solvated interaction energy function for ranking antibody-antigen binding affinities. J. Chem. Inf. Model. 56, 1292–1303 (2016).
Pires, D. E. V., Ascher, D. B. & Blundell, T. L. mCSM: predicting the effects of mutations in proteins using graph-based signatures. Bioinformatics 30, 335–342 (2014).
Archer, K. J. & Kirnes, R. V. Empirical characterization of random forest variable importance measures. Comput. Stat. Data Anal. 52, 2249–2260 (2008).
Sirin, S., Apgar, J. R., Bennett, E. M. & Keating, A. E. AB-Bind: Antibody binding mutational database for computational affinity predictions. Protein Sci. 25, 393–409 (2016).
Bostrom, J. et al. Variants of the antibody herceptin that interact with HER2 and VEGF at the antigen binding site. Science 323, 1610–1614 (2009).
Kiyoshi, M. et al. Affinity improvement of a therapeutic antibody by structure-based computational design: generation of electrostatic interactions in the transition state stabilizes the antibody-antigen complex. PLoS ONE 9, e87099 (2014).
Arimori, T. et al. Fv-clasp: an artificially designed small antibody fragment with improved production compatibility, stability, and crystallizability. Structure 25, 1611–1622 (2017).
Sali, A., Potterton, L., Yuan, F., Vanvlijmen, H. & Karplus, M. Evaluation of comparative protein modeling by modeler. Proteins Struct. Funct. Genet. 23, 318–326 (1995).
Sivasubramanian, A., Sircar, A., Chaudhury, S. & Gray, J. J. Toward high-resolution homology modeling of antibody Fv regions and application to antibody-antigen docking. Proteins 74, 497–514 (2009).
Pierce, B. G. et al. ZDOCK server: interactive docking prediction of protein-protein complexes and symmetric multimers. Bioinformatics 30, 1771–1773 (2014).
Quinonero-Candela, J. Q. & Rasmussen, C. E. A unifying view of sparse approximate Gaussian process regression. J. Mach. Learn. Res. 6, 1939–1959 (2005).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Obrezanova, O. et al. Aggregation risk prediction for antibodies and its application to biotherapeutic development. Mabs-Austin 7, 352–363 (2015).
Sankar, K. et al. Prediction of methionine oxidation risk in monoclonal antibodies using a machine learning method. Mabs-Austin 10, 1281–1290 (2018).
Pedregosa, F. et al. Scikit-learn: machine learning in python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Naim, M. et al. Solvated interaction energy (SIE) for scoring protein-ligand binding affinities. 1. Exploring the parameter space. J. Chem. Inf. Model. 47, 122–133 (2007).
Conchuir, S. O. et al. A web resource for standardized benchmark datasets, metrics, and rosetta protocols for macromolecular modeling and design. PLoS ONE 10, e0130433 (2015).
Guerois, R., Nielsen, J. E. & Serrano, L. Predicting changes in the stability of proteins and protein complexes: a study of more than 1000 mutations. J. Mol. Biol. 320, 369–387 (2002).
Hornak, V. et al. Comparison of multiple amber force fields and development of improved protein backbone parameters. Proteins Struct. Funct. Bioinform. 65, 712–725 (2006).
Cornell, W. D. et al. A second generation force field for the simulation of proteins, nucleic acids, and organic molecules (vol 117, pg 5179, 1995). J. Am. Chem. Soc. 118, 2309–2309 (1996).
Spassov, V. Z. & Yan, L. pH-selective mutagenesis of protein-protein interfaces: in silico design of therapeutic antibodies with prolonged half-life. Proteins Struct. Funct. Bioinform. 81, 704–714 (2013).
Leaver-Fay, A. et al. Rosetta3: an object-oriented software suite for the simulation and design of macromolecules. Methods Enzymol. 487, 545–574 (2011).
Cui, Q. Z. et al. Molecular dynamics-solvated interaction energy studies of protein-protein interactions: the MP1-p14 scaffolding complex. J. Mol. Biol. 379, 787–802 (2008).
Rohl, C. A., Strauss, C. E. M., Misura, K. M. S. & Baker, D. Protein structure prediction using rosetta. Methods Enzymol. 383, 66–93 (2004).
Jankauskaite, J., Jimenez-Garcia, B., Dapkunas, J., Fernandez-Recio, J. & Moal, I. H. SKEMPI 2.0: an updated benchmark of changes in protein-protein binding energy, kinetics and thermodynamics upon mutation. Bioinformatics 35, 462–469 (2019).
Acknowledgements
Computations were partially performed on the NIG supercomputer at ROIS National Institute of Genetics, AI Bridging Cloud Infrastructure (ABCI) at National Institute of Advanced Industrial Science and Technology (AIST), and the supercomputer of ACCMS at Kyoto University.
Funding
This work was supported by Japan Science and Technology Agency Advanced Integrated Intelligence- Public/Private R&D Investment Strategic Expansion Program [JPMJCR18Y3]; Ministry of Education, Culture, Sports, Science and Technology/Japan Society for the Promotion of Science KAKENHI [17H06410 to Y.S., 19K20409 to Y.S., 19K06502 to Y.S., 19K06077 to Y.S.]; and Japan Agency for Medical Research and Development [JP19ak0101122 to Y.S., JP19am0401023 to Y.S.].
Author information
Authors and Affiliations
Contributions
Y.K. designed and performed the experiments with the support of Y.S. Y.K. and Y.S. wrote the manuscript. T.K. conceived and supervised this work. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kurumida, Y., Saito, Y. & Kameda, T. Predicting antibody affinity changes upon mutations by combining multiple predictors. Sci Rep 10, 19533 (2020). https://doi.org/10.1038/s41598-020-76369-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-76369-8
- Springer Nature Limited