Abstract
Very virulent plus Marek’s disease (MD) virus (vv + MDV) induces tumors in relatively resistant lines of chickens and early mortality in highly susceptible lines of chickens. The vv + MDV also triggers a series of cellular responses in both types of chickens. We challenged birds sampled from a highly inbred chicken line (line 63) that is relatively resistant to MD and from another inbred line (line 72) that is highly susceptible to MD with a vv + MDV. RNA-sequencing analysis was performed with samples extracted from spleen tissues taken at 10-day and 21-day post infection (dpi). A total of 64 and 106 differentially expressed genes was identified in response to the vv + MDV challenge at latent phase in the resistant and susceptible lines of chickens, respectively. Direct comparisons between samples of the two lines identified 90 and 126 differentially expressed genes for control and MDV challenged groups, respectively. The differentially expressed gene profiles illustrated that intensive defense responses were significantly induced by vv + MDV at 10 dpi and 21 dpi but with slight changes in the resistant line. In contrast, vv + MDV induced a measurable suppression of gene expression associated with host defense at 10 dpi but followed by an apparent activation of the defense response at 21 dpi in the susceptible line of chickens. The observed difference in gene expression between the two genetic lines of chickens in response to MDV challenge during the latent phase provided a piece of indirect evidence that time points for MDV reactivation differ between the genetic lines of chickens with different levels of genetic resistance to MD. Early MDV reactivation might be necessary and potent to host defense system readiness for damage control of tumorigenesis and disease progression, which consequently results in measurable differences in phenotypic characteristics including early mortality (8 to 20 dpi) and tumor incidence between the resistant and susceptible lines of chickens. Combining differential gene expression patterns with reported GO function terms and quantitative trait loci, a total of 27 top genes was selected as highly promising candidate genes for genetic resistance to MD. These genes are functionally involved with virus process (F13A1 and HSP90AB1), immunity (ABCB1LB, RGS5, C10ORF58, OSF-2, MMP7, CXCL12, GAL1, GAL2, GAL7, HVCN1, PDE4D, IL4I1, PARP9, EOMES, MPEG1, PDK4, CCLI10, K60 and FST), and tumor suppression (ADAMTS2, LXN, ARRDC3, WNT7A, CLDN1 and HPGD). It is anticipated that these findings will facilitate advancement in the fundamental understanding on mechanisms of genetic resistance to MD. In addition, such advancement may also provide insights on tumor virus-induced tumorigenesis in general and help the research community recognize MD study may serve as a good model for oncology study involving tumor viruses.
Similar content being viewed by others
Introduction
Marek’s disease (MD) is a neoplastic disease of chickens caused by an oncogenic alpha-herpesvirus, commonly known as MD virus (MDV). MD is characterized by a variety of clinical signs, including immune suppression, polyneuritis, and notably, the formation of T-cell lymphomas that manifest as solid tumors1. MD has had being a major concern for the poultry industry worldwide since the 1900s2, which causes an economic loss as high or over $2 billion per year3 up to date resulted from condemnation, vaccination, and extra management measures necessary for the control. Hence, a better understanding on genomic mechanism underlying virus-induced tumorigenesis and progression would not only benefit the poultry industries, but also bring significant merits for human health in the foreseeable future.
Like other herpesviruses, MDV pathogenesis in chicken involves multiple phases including cytolytic and latent phases4. MDV initially enters an early cytolytic infection phase in B cells between 3- and 6-days post infection (dpi)5. The virus replication reaches peak at this stage6. Around 7 dpi, MDV-infected B cells transfer MDV to T cells reportedly through cell to cell contacts and the virus then quickly establishes latency between 7–10 dpi in the T cells7,8. During the latency phase, the MDV genome remains persistent in the host cells4. In susceptible chickens, a second cycle of cytolytic infection takes place at around 14 dpi as a result of virus reactivation, which causes inflammation and lymphoma formation around 21 dpi and onward1. MD regression has been reported and suggested to be affected by virus and host genotypes as well as subsequent immune responses9,10,11. During infection, MDV attempts to control or take advantage of host components to facilitate its replication. At the meantime, host mobilizes its innate immune system to battle against the viral infection and replication processes1,4,12. Therefore, the disease progression is clearly influenced by a complex set of interactions involving, at least, the viral genome and the host’s immune systems. To date, much efforts have been focused on advancing the understanding of the pathogen and its remarkable repertoire of virulence factors. Many viral genes have been extensively studied and documented, such as Meq and vIL-8, which are functionally involved in cell transformation, tumorigenesis and tumor malignance13. On the other hand, what host genes affect MD resistance is relatively far from clearly understood.
It is well documented that following MDV challenge, MD incidence occurs differently among different genetic lines of chickens due to varied susceptibility14,15,16,17,18. Striking examples include the highly inbred chicken lines 63 and 72, which were developed and maintained since 1939 at the USDA, Avian Disease and Oncology Laboratory (ADOL), East Lansing, Michigan, U.S.A. The lines 63 and 72 share a common major histocompatibility complex haplotype (B*2) but differ significantly in MD resistance16,17. When challenged with a partially attenuated very virulent plus strain of MDV (648A passage 40), up to 100% of the line 72 chickens developed MD while over 90% of line 63 chickens remained MD free 8 weeks post MDV inoculation19. These inbred lines of chickens are commonly considered as ideal resources for investigating the mechanisms of genetic resistance to MD and MD tumor progression15,20. Since the establishment of the genetic lines, a variety of differences in response to MDV infection between the two lines of chickens have been investigated, which include the aggregate number of target lymphocytes for virus infection and transformation21,22, expressed antigens on T-cell surface23, the number of infected lymphocytes during the early lytic phase of infection24, viral load during latent phase of infection25,26, the expression of Interferon (INF) genes27 and cytokine genes26. However, little is known about the difference of transcriptomic patterns in response to vv + MDV challenge at the latent and the late cytolytic phases between the resistant and susceptible lines of chickens, despite that extensive efforts have been put in to elucidate the gene expression in response to MDV challenge3,12,28,29,30,31,32,33,34,35,36. Furthermore, most of the studies conducted previously employed microarray analysis, which bares known limitations37.
Establishment and maintenance of latency in host after primary lytic infection is a hallmark of herpesvirus infection38. This study was designed to systematically investigate transcriptomic changes in the line 63 and line 72 chickens in response to a vv + MDV challenge at 10 and 21 dpi by RNA sequencing analysis, which include identification and characterization of differentially expressed genes (DEGs) between the two genetic lines with and without MDV infection during the latent and late cytolytic phases of MDV infection. Public web server tools for high-throughput genomics data analyses were employed to dissect the identified DEGs for functional interpretations and to select candidate DEGs functionally relevant to MD resistance, which was anticipated to provide some insights on the mechanisms of genetic resistance to MD.
Results
Global transcriptomic profiles of the lines 63 and 72
Total RNA was extracted from spleen samples of the inbred line 63 and 72 birds of both control and vv + MDV challenged groups at 10 and 21 dpi, which was subjected to RNA-seq analyses. A total of 198.28 million clean reads was generated with an average of 24.78 million reads per treatment group. Mapping rates of the clean reads to the chicken reference genome ranged from 86.4% to 88.9% (Supplementary Table S1). Expression level (reads) for each of the identified genes was normalized as the fragments per kilobase of transcript per million mapped reads (FPKM). A total of 11,426 and 11,227 genes for line 63, 11,575 and 11,083 genes for line 72 was identified in the control and MDV challenged groups, respectively, with FPKM values ≥1 at 10 dpi. A total of 11,343 and 10,922 genes for line 63, 10,955 and 10,927 genes for line 72 was identified in the control and MDV challenged groups, respectively, at 21 dpi. The total numbers of genes identified at 10 and 21 dpi were 12,132 and 11,793 with FPKM values ≥1, respectively, from both control and MDV challenged groups of the two lines.
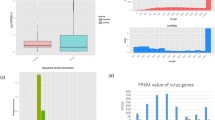
Overall gene expression levels were evidently clustered by treatment groups and by the genetic lines of chickens in a Principle Component Analysis (PCA)-space along the PC1 and PC2 coordinates, respectively, with one relatively notable deviation of the line 63 MDV challenged group at 10 dpi, which was relatively less distanced from its control group and a bit more distanced from the group at 21 dpi (Fig. 1A). The average FPKM across all treatment groups was 87.0. Distributions of the FPKM were very comparable among the treatment groups (Fig. 1B), which suggested a general uniformity of the transcriptomes and basically free of global shift biases in transcriptomic levels among all the treatment groups. Therefore, it is highly likely that the differences detected in transcriptomic expression levels between the treatment groups and between the genetic lines were indeed due to the MDV challenge and the host genetics of different chicken lines, respectively.
RNA-Seq overview. (A) A PCA score plot of 10,257 gene expression data, illustrating the clusters along the PC1 and PC2 coordinates by the treatment groups (MDV challenged vs. control) and genetic lines (63 and 72) of chickens; (B) A plot of FPKM distribution post normalization by chicken line and treatment groups, illustrating the overall uniformity of the expressional data across the treatment groups and the genetic lines of birds.
Differentially expressed genes in response to MDV challenge
To improve the reliability and comparability of differential expression analysis, a total of 10,257 identified genes with FPKM values ≥1 in all treatment groups were included in this and subsequent analyses. At 10 dpi, 61 and 79 DEGs in response to MDV challenge were identified in the line 63 and 72 birds, respectively. Of which, 54 (88.52%) and 46 (58.22%) DEGs were significantly up-regulated in the line 63 and 72 birds, respectively (Fig. S1A,C). At 21 dpi, 11 and 60 DEGs in response to MDV challenge were identified in the line 63 and 72 birds, respectively. Interestingly, all the 11 DEGs of line 63 were upregulated while 51 (85%) of the 60 DEGs in line 72 were upregulated (Fig. 2A,B, and Fig. S1B,D). Detailed lists of all DEGs induced by MDV challenge in the two genetic lines at 10 and 21 dpi are given in Supplementary Tables S2–S5. As shown in the Venn diagrams (Fig. 2C,D), there were dozens of mutually exclusive DEGs between the lines 63 and 72 at 10 and 21 dpi except the line 63 at 21 dpi with only a single mutually exclusive DEG, in addition to 26 and 10 DEGs identified in common between the lines of birds at 10 and 21 dpi, respectively. Within each of the lines between the two time points, 10 and 21 dpi, there were a few (2 in line 63 at 21 dpi) up to dozens of DEGs identified mutually exclusive between the two time points, in addition to 9 and 33 DEGs in common between the two time points in line 63 and line 72, respectively (Fig. 2E,F).
Graphical summaries of DEGs of line 63 and 72 birds in response to MDV challenge at 10 and 21 dpi by RNA-Seq. (A) The number of DEGs identified in each of the lines and in both lines at each of the time points. The length of the segments is proportional to the number of identified DEGs. (B) Illustrating the relative proportion of up- and down-regulated DEGs (red and blue portions of each bar, respectively) for each line at each time point. (C) A Venn diagram showing mutually exclusive DEGs and DEGs in common in response to MDV challenge at 10 dpi between the lines 63 and 72 groups. (D) A Venn showing mutually exclusive DEGs and DEGs in common in response to MDV challenge at 21 dpi between the lines. (E,F) Venn diagrams showing mutually exclusive and in common DEGs between the 10 and 21 dpi groups in response to MDV challenge for the line 63 and line 72 groups, respectively.
Deferentially expressed genes between the genetic lines with and without MDV challenge
In contrast to line 72 control groups, 36 and 32 DEGs were significantly expressed at higher levels, 38 and 10 DEGs were expressed at lower levels in the line 63 control groups at 10 and 21 dpi, respectively (Fig. 3A, Fig. S2A,B, and Tables S6, S7). Following MDV challenge, 32 and 59 DEGs were expressed at significantly higher levels, 16 and 37 DEGs were expressed significantly at lower levels in the line 63 compared to the line 72 at 10 and 21 dpi, respectively (Fig. 3B, Supplementary Fig. S2C,D, Supplementary Tables S8, S9). There were dozens of genes expressed differentially between the two lines regardless of MDV-challenge, and there were 19 and 18 genes differentially expressed in common between the two lines as well at 10 (Fig. 3C) and 21 dpi (Fig. 3D), respectively. In MDV-challenged birds, there were certain degrees of overlaps of genes differentially expressed between the two lines at 10 (Fig. 3E) and 21 dpi (Fig. 3F). A close examination of 5 and 12 genes at 10 and 21 dpi, respectively, between the lines 63 and 72 for both the control groups and the MDV challenged groups is depicted in Fig. 3G,H. The expression levels of those genes were significantly differentiated between the two lines, but none of the expression levels of those genes was between the control groups.
Depicting the numbers of DEGs identified between the resistant line 63 and susceptible line 72 birds without and with MDV challenge. (A) Number of DEGs significantly expressed higher (red bars) or lower (blue bars) in line 63 birds in contrast to the line 72 birds of the control groups at the ages matching the counterpart groups at 10 and 21 dpi. (B) Number of DEGs significantly expressed higher or lower in the line 63 birds than in line 72 birds in response to MDV challenge at 10 and 21 dpi. (C) A Venn diagram showing the number of DEGs in common (19) and mutually exclusive (55 and 29) between the line 63 and 72 control groups’ contrast and the MDV challenged groups’ contrast, respectively at 10 dpi. (D) A Venn diagram showing the number of DEGs in common (18) and mutually exclusive (24 and 78) between the line 63 and 72 control groups’ contrast and the MDV challenged groups’ contrast, respectively, at 21 dpi. (E) A Venn diagram showing the numbers of DEGs in common and mutually exclusive between the line 63 challenged/control, line 72 challenge/control, and the challenged groups of lines 63 and 72 at 10 dpi. (F) A Venn diagram showing the numbers of DEGs in common and mutually exclusive between the line 63 challenged/control, line 72 challenge/control, and the challenged groups of lines 63 and 72 at 21 dpi. (G) Depicting both the expression levels of 5 genes at 10 dpi and (H) 12 genes at 21 dpi without (control) and with MDV challenge between the line 63 and line 72 groups. A red star indicates a gene that was differentially expressed (FDR <0.05 and FC >2) in response to MDV challenge in the line 72 birds, whereas a green star indicates that gene was differentially expressed (FDR <0.05 and FC >2) in response to MDV challenge in the line 63 birds. Both groups of charts in G and H demonstrated that the difference in gene expression in the control groups did not alter the differential expression status of genes in response to MDV challenge, at least in this study.
ddPCR validation of the RNA-Seq data
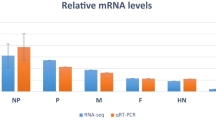
To assess the validity of RNA-Seq data, a total of 12 pairs of primers (Table S10) was designed targeting 12 selected DEGs that were identified in a combination of 23 comparisons of this study by time point (dpi), MDV treatment and chicken lines. These primers were used in droplet digitalTM PCR (QX200TM ddPCR system; Bio-Rad Laboratories, Inc., Hercules, CA, USA) analysis for the validation. Three biological replicates were used for each treatment group. A scatterplot was generated by comparing the log2FC determined by RNA-Seq analysis and ddPCR analysis. The result showed there was a fairly high correlation between the two groups of log2FCs determined by the two methods (Fig. S3, R2 = 0.496, P value < 0.001). The ddPCR data validated the RNA-Seq data.
ddPCR analysis of MDV microRNAs’ expressions associated with MDV latency
The expressions of two MDV microRNAs, miR-M3-3p and miR-M12-3p, in each of the total RNA samples extracted from MDV challenged birds and included in the current RNA_Seq analysis were quantitatively assessed by ddPCR to ensure that each of those total RNA sample of this study was from a challenged bird that had entered the latent phase. The miR-M3-3p and miR-M12-3p have been characterized with significantly higher expression in latent phases in contrast to that at the early cytolytic phase39. The ddPCR analysis was conducted with technical replicates using customer primers (Table S11). Splenic total RNA samples extracted from MDV challenged birds at 5 DPI were also used to contrast the difference of the microRNA expressions. The 5-DPI total RNA samples were aliquoted from a sister project conducted simultaneously along with this project using the same hatch of line 63 and line 72 birds under the same exact conditions. In a 20 μL reaction, an average of 7,020 ± 653.9 and 4,587 ± 610.0 miR-M3-3p copies at 10 DPI and 5,123 ± 866.6 and 3,750 ± 1,092.7 copies at 21 DPI was detected for lines 63 and 72, respectively; in contrast, an average of 422 ± 30.6 and 356 ± 35.9 copies was detected in the 5 DPI samples of line 63 and 73, respectively. The average of miR-M3-3p expression copies among all the 10 and 21 DPI individual total RNA samples ranged from 2,760 ± 180 to 8,120 ± 40 for line 63, and 1,860 ± 20 to 6,470 ± 470 for line 72.
The ddPCR quantification of the miR-M12-3p microRNA resulted in an average of 218 ± 17.7 and 197 ± 13.0 copies at 10 DPI, and 247 ± 18.3 and 195 ± 3.7 copies at 21 DPI for the line 63 and 72 samples, respectively. In contrast, an average of 63 ± 4.1 and 70 ± 8.4 copies at 5 DPI was detected for the lines 63 and 72, respectively. The average of miR-M12-3p copies expressed among all the individual birds at both 10 and 21 DPI ranged from 174 ± 12.0 to 300 ± 4.0, and 170 ± 6.0 to 216 ± 36.0 for lines 63 and 72, respectively.
The results of Analysis of Variance showed both MDV microRNAs’ expressions at 10 and 21 DPI significantly differed from those at 5 DPI (P < 0.01; leverage plots of the MDV microRNA expression data are given in Figs. S4 and S5 graphically illustrating the differences in expression at 5, 10, and 21 DPI). This in combination with the ranges of individual bird MDV microRNA expression evidently suggested that none of the MDV challenged birds under this study was remained at the early cytolytic phase.
Functional analysis of the DEGs
To better understand the MDV-induced DEGs, functional enrichment analysis was conducted for four separate sets of the DEGs, which included two sets of MDV-induced DEGs in lines 63 and 72 at 10 dpi and two sets of MDV-induced DEGs in lines 63 and 72 at 21 dpi. At 10 dpi, the up-regulated genes in response to MDV challenge in line 63 were primarily enriched in 13 GO terms, which are primarily associated with the immune system including immune effector process, immune system process, immune response, defense response, and regulation of immune response. The down-regulated genes in line 63 at 10 dpi were associated with cell communication (FAM132 A, FN1 and PROKR2) and immune system process (AQP3). The up-regulated genes of line 72 at 10 dpi were also significantly enriched in two GO terms of immune response and immune effector process. Some up-regulated DEGs of both lines 63 and 72 were also involved with the Influenza A pathway. In contrast, the line 72 down-regulated genes were enriched in four GO terms and four KEGG pathways. Notably, these functional categories included several terms involved in fatty acid metabolism, such as fatty acid biosynthetic process, biosynthesis of unsaturated fatty acids and fatty acid metabolism (Table 1). Of the 11 significantly up-regulated genes in line 63 at 21 dpi (Table S4), two were granzyme (GZMA and GZMK) genes, three were immune response genes (MX, OAS*A and CCLI10), one was an Interleukin Four Induced gene 1 (IL4l1) and one was avidin gene (AVD). For line 72, the up-regulated DEGs were significantly enriched in GO terms primarily associated with immune response, including defense response, innate immune response, and response to interferon-gamma, in addition to influenza A pathway (Table 2).
Gene function analysis showed that DEGs that were relatively expressed at higher levels in the unchallenged line 63 birds than those of the unchallenged line 72 birds were significantly enriched in extracellular structure organization and extracellular matrix organization, as well as pathways of Focal adhesion and ECM-receptor interaction (Table 3). DEGs that were relatively expressed higher in the unchallenged line 72 birds than those of the unchallenged line 63 birds were mainly involved in metabolic pathways, such as pathways of Butanoate metabolism, Fatty acid metabolism, Biosynthesis of amino acids. Following infection at 10 dpi, six of the DEGs were immune genes (GAL1, GAL2, GAL7, ABCB1, LEPR and SNED1) and two were involved in NOD-like receptor signaling pathway (K60 and HSP90B1), all were upregulated in expression in line 63 birds. In contrast, four DEGs associated with response to stimulus (HSP90AB1, GNG4, ANXA1, CD180) were upregulated in expression in line 72 birds. At 21 dpi, upregulated DEGs identified in line 63 were significantly enriched in GO terms involved in system development and two pathways including Focal adhesion and ECM-receptor interaction, while upregulated DEGs identified in line 72 were mainly associated in immunity, as indicated in enriched functional categories such as antigen processing and presentation, adaptive immune response, cell killing and KEGG pathway (Table 4).
Key genes for MD resistance
To further identify key genes that may confer genetic resistance to MD from all the unique DEGs, we focused on the followings: (1) DEGs located within reported QTL regions of MD resistance (http://www.animalgenome.org/cgi-bin/QTLdb/GG/index); (2) DEGs with large fold change (FC >2) in one line but not in the other (FC < 0.5); (3) DEGs differentially expressed between the two lines regardless of MDV challenge; (4) MDV induced DEGs with differential expression between infected line 63 and line 72 birds (Fig. 3E–H). These comparisons resulted in a total of 104 unique DEGs. Each of these DEGs was subjected to a separate Gene Ontology (GO) analysis and a comprehensive literature review. Jointly considering the expression pattern of each gene in different chicken lines and the potential functionality, a short list of 27 top DEGs was selected as the likely candidate genes identified from this study, which may potentially play key roles in conferring genetic resistance to MD (Fig. 4). These genes were broadly grouped into three functional groups, including virus process, immunity and tumor suppression.
Discussion
Several studies have documented the global gene expression responses to MDV infection in chicken3,12,28,29,30,31,32,33,34,35,36. These analyses, while providing valuable and important mechanistic clues to host responses and virus-host interactions, are of limited scope due to coverage of gene arrays used and lack information of comparisons between chickens exhibiting different resistance to MD. By this study, we further expanded the current knowledge with regard to host transcriptomic response upon MDV challenge through comparing the whole transcriptome changes between a highly inbred MD-resistant and -susceptible lines of chicken using RNA-Seq technology. This study was primarily focused on the latent stages post MDV inoculation. Our results revealed quite different transcriptome patterns, which may provide new insights potentially conferring genetic resistance to MD.
Close examinations of the transcriptomic changes in response to MDV challenge showed that many genes associated with immunity and anti-tumor functions were activated in both lines of chickens. For example, two granzyme genes, GZMA and GZMK, that are associated with apoptosis, and the MX1 gene that is a well-known IFNs-induced gene40, were up-regulated in both lines at 10 and 21 dpi, suggesting on going interactions that took place between host and the MDV pathogen. In contrast, more different features in response to MDV infection between the two inbred lines were observed. More DEGs were identified in the MD susceptible line 72 than those in the resistant line 63 at both 10 and 21 dpi (Fig. 2A), indicating that MDV induced much stronger response at the transcriptional level in line 72 than in line 63 birds. However, it is interesting to note that the proportion of up-regulated DEGs was much higher in line 63 than that in line 72 (Fig. 2B), which suggested that insufficient immune response was probably activated against tumorigenesis regardless more immune-related gene expressions were altered in response to MDV challenge. Specifically, in MD resistant line 63, a notable immune response was observed at 10 dpi, which was characterized by the overrepresentation of DEGs enriched in many immune response-related GO terms, such as immune response, defense response to virus, positive regulation of immune response, and so on. While at 21 dpi, only 11 genes were identified to exhibit a statistically significant change in expression, indicative of a minimal change compared to normal tissues. In susceptible line 72, a totally different transcription pattern was revealed across the two-time points. At 10 dpi, 33 out of 79 DEGs (41.77%) were down-regulated in expression (Fig. 2B, Fig. S2C). These down-regulated genes were significantly enriched in multiple GO terms and pathways, including several terms associated with fatty acid metabolism, such as fatty acid biosynthetic process, pathways of fatty acid metabolism, and biosynthesis of unsaturated fatty acids (Table 2). Genes involved in these terms included SCD, FADS1, FADS2 and HPGDS. Previous studies showed that Hepatitis C virus (HCV) could induce imbalance in lipid homeostasis in host cells41,42. However, it seemed that inhibition of fatty acid biosynthesis can suppress virus replication43,44 and control cancer cell proliferation45. Therefore, whether the down-regulation of these genes in susceptible line 72 playing a role in establishment of latency or representing a host response that may contribute to repressing the activities of MDV infection needs to be further investigated. At 21 dpi, 60 DEGs were identified being differentially expressed compared to the control birds and 51 of those were up-regulated. Gene function analysis showed that the up-regulated genes were significantly enriched in immune response-related terms such as innate immune response, interferon-gamma-mediated signaling pathway, and so on. In particular, the IRF1 (Interferon regulatory factor-1) gene expression was upregulated by 4.2 folds (from an FPKM of 1278.1 to 5415.7). IRF1 is a member of the IRF gene family of transcription factors that binds to the virus-inducible cis-elements of IFN-α and IFN-β gene promoter as well as to the interferon response sequence of IFN-inducible gene promoters46. Accordingly, the 51 identified up-regulated DEGs were significantly enriched in IRF-1 (P value = 5.66E-5) by a motif enrichment analysis using the g:Profilier (http://biit.cs.ut.ee/gprofiler/). A total of 15 up-regulated genes containing potential IRF-1 binding site was revealed, including IFIH1, BCL2L14, GZMK, ISG12(2), SERPINB10, IFI27L2, AVD, TAP1, CCL19, CMPK2, ENSGALG00000026152, ENSGALG00000001629, ENSGALG00000006384, ENSGALG00000013057 and ENSGALG00000019141. These gene functions implied strong defense responses might have been activated in susceptible line 72 at 21 dpi, which is in good agreement with earlier observations that MDV were reactivated in susceptible line of birds at late latency stage1. Furthermore, we observed some genes showed extreme difference in response to MDV challenge between the two genetic lines of birds at each time points (Fig. S2C,D). These genes might also play key roles modulating genetic resistance to MD, therefore, follow-up investigations are warranted. One of those DEGs is IL4I1. Reportedly it is an inhibitor of the CD8+ antitumor T-cell response and may facilitate tumor growth. Our data showed that the expression of IL4I1 was significantly increased in the susceptible line 72 birds at 10 dpi, while it remained unchanged in the resistant line 63 birds at the same time point, which indicated that genes like IL4I1 may contribute to immunosuppression and facilitate tumorigenesis through inhibiting the CD8+ antitumor T-cell response in MD susceptible chickens. If so, it is then in good agreement with the functional findings of this gene in mice47.
Differences in gene expression levels were also observed between the two lines at both time points for birds without and with MDV challenge. Notably, genes associated with Focal adhesion and ECM-receptor interaction were observed with higher expression in the resistant line 63 control birds (Table 3). Interestingly, following MDV challenge, genes involved with these two pathways exhibited consistently elevated expression in the resistant line 63 birds than that in the susceptible line 72 birds at 21 dpi (Table 3), indicative of potential roles of these two pathways in MD resistance. The Focal adhesion and ECM-receptor interaction pathways have a profound influence on major cell programs including growth, differentiation, migration, and survival48, which suggests differences in functional properties of cells between the resistant and the susceptible lines of chicken. Furthermore, earlier studies have documented that these two pathways play a major role in immunity49,50. More interestingly, these pathways are also reportedly linked to tumor progression51,52. Therefore, the higher expression of these genes involved in Focal adhesion and ECM-receptor interaction may be contributing to tumor formation and development. Notably, the majority of the significantly enriched functional categories for DEGs with higher levels of transcriptional expression were observed in the susceptible line 72 birds other than in the resistant line 63 birds in response to MDV challenge, which are related to immune response, including positive regulation of T cell mediated immunity, adaptive immune response, and positive regulation of cell killing. This, again, implicated the strong reactivation of the susceptible line 72 birds to MDV infection at these points of time.
Merging the DEGs identified between the infected and uninfected birds as well as between the two lines resulted to a total of 284 unique genes. Among them, 189 DEGs (66.6%) have been previously reported in studies of host response to MDV challenge3,12,28,29,30,31,32,33,34,35,36, including genes associated with chemokine (CCL1, CXCL13L2, CXCL12 and CCL19), cytokine (IL21R, IL2RA, LEPR, IL12RB2, FLT3), innate immune response (IRF1, STAT1, CD36, IFIH1, SLC11A1, IGJ, PAPR9, GCH1, LAG3, RSAD2 and HPX), and adaptive immune response (B2M, P2RX7, SLC11A1, ANXA1, RSAD2, HPX, and BFIV21). This overlap confirms the roles of these candidate genes that may play important roles in response to MDV challenge. On the other hand, due to other differences including genetic background of chickens, MD virus strains, tissue samples and sampling time, novel DEGs were identified in this study, which expanded the candidate gene pool conferring genetic resistance to MD. Jointly considering the gene expression pattern in two lines, gene function, and comparison with QTL regions, 27 promising top candidate genes were proposed based on this study, which may highly likely play key roles in MD including conferring genetic resistance to MD (Fig. 4). These genes showed large differences either between resistant and susceptible lines, or in response to MDV challenge within each of the genetic lines, or both, or are located at reported MD-related QTL regions. Functional analysis classified these genes into three broadly functional categories, including viral process, immunity, and tumor development.
Earlier studies have shown that viral load was different between the line 63 and line 72 at latency phase of infection25,26. Consistent with these observations, we found two genes, F13A1 and HSP90AB1, with known functions in promoting virus replication53,54, that were significantly induced in expression in response to MDV challenge in the susceptible line 72 birds, but with little change in the resistant line 63 birds at 10 dpi. Presumably, these two genes might be hijacked by MDV to facilitate viral replication in the susceptible line 72 birds. Additionally, there were 19 genes that may potentially play roles in immunity. Among them, ABCB1LB, RGS5, C10ORF58, OSF-2, MMP7 and CXCL12 were observed with higher expression levels in the resistant line 63 birds regardless of MDV challenge treatment. Our results also revealed that some immune response-related genes, including HVCN1 and PDE4D, were down-regulated in the susceptible line 72 birds in response to MDV challenge. Together with the identification of up-regulation of IL4I1 and down-regulation of defensing genes, GAL1, GAL2 and GAL3, in the susceptible line 72, it is postulated here that the susceptible line 72 birds may suffer from immunosuppression at latent stage of MDV infection, which is in good agreement with a previous report29. Other promising candidates involved in immunity were uniquely induced by MDV in the resistant line 63 (PARP9, EOMES, MPEG1, PDK4, CCLI10, K60 and FST) compared to the susceptible line 72. Presumably, these genes may belong to the group that contribute to genetic resistance to MD. MD is a virus-induced tumorous disease of chicken and has been proposed to be an invaluable model for investigation of virus-induced cancers7,55,56. Our results showed that six putative tumor suppressor genes, including ADAMTS257, LXN58, ARRDC359, WNT7A60, CLDN161 and HPGD62,63,64, exhibited higher expression levels in the resistant line 63 than in the susceptible line 72 birds, which should confer genetic resistance to MD. The findings from this study underscored the value of MD study serving as a model for better understanding tumorigenesis and raised the possibility that more novel genes involved in tumorigenesis and development need to be further explored.
Conclusion
In summary, we have carried out a comprehensive gene expression study using RNA-Seq analysis and identified hundreds of differentially expressed genes in response to a vv + MDV challenge in two highly inbred lines of chickens at 10 and 21 days post MDV infection. It is well-documented that one of the lines, the line 63, is relatively resistant to MD, while the other, the line 72, is highly susceptible to MD. We identified a total of 284 unique coding genes that likely affect the resistance to MD, which provided a sizable and valuable addition to the current candidate gene pool that is reportedly involved with genetic resistance to MD. We showed that the response pattern of gene expression at 21 dpi supported a reactivation of MDV in the susceptible line. We further proposed 27 promising candidate genes that may play key roles conferring genetic resistance to MD. Notably, most of these promising candidate genes identified in this study are reportedly associated with immunity and tumor suppression, which may have an important implication on virus-induced tumorigenesis in general and highlighted the value of MD model for tumor virus-induced cancer study. Further directions should include work on the detailed function of these candidate genes and regulatory control of the expression of those genes upon explosion to tumor viruses like MDV.
Materials and Methods
Experimental design
One-day old chickens from two highly inbred lines were sampled for an MDV challenge trial in this study. One genetic line is known as line 63 and the other, as line 72. The two genetic lines were developed and have been maintained at the USDA, Agricultural Research Service, Avian Disease and Oncology Laboratory (ADOL) in East Lansing, Michigan. The lines 63 and 72 share a common major histocompatibility complex (B*2) haplotype but are resistant and highly susceptible to MD, respectively15. On the day of hatch, chickens from each line were randomly divided into MDV challenge group and control group. Each of the chicks in the MDV challenge groups of both lines was inoculated intraabdominally with 500 plaque-forming units of 648A passage 10 MDV at day 5 post hatch. No inoculation was implemented in the control groups. Three chickens from each group were randomly euthanized at 10 (latency period) and 21 dpi (reactivation period), respectively. Spleen samples were individually collected, immediately placed into RNAlater solution (Qiagen, Valencia, CA, USA), and stored at −20 °C until total RNA extraction.
All chickens used in this study were housed in a BSL-2 experimental facility during the trial. Feed and water were supplied ad libitum. The chickens were observed daily throughout the entire duration of the experiment. The animal challenge experiment was approved by the USDA, Avian Disease and Oncology Laboratory Institutional Animal Care and Use Committee (IACUC). The IACUC guidelines established and approved by the ADOL IACUC (April 2005) and the Guide for the care and use of Laboratory Animals by Institute for Laboratory Animal Research (2011) were closely followed throughout the experiment.
RNA sequencing
Total RNA was extracted using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions. RNA concentration and quality were assessed using an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). Equal amount of RNA samples from three biological replicates within each line each treatment group were pooled in preparation to construct standard cDNA libraries using Illumina TruSeq kits and reagents following the manufacturer’s protocol for deep sequencing. The libraries were sequenced on an Illumina HiSeq2000 sequencer for single end 50 base sequencing run. The post sequencing processes, including image analysis, base calling, and Q-Score calculation, were carried out using Real Time Analysis (v1.13.48); read demultiplexing and conversion to final FASTQ files, using CASAVA (v1.8.2) software tools (Illumina Inc., San Diego, CA, USA). The library preparation, RNA sequence read extraction, and preliminary read quality control were performed at the Research Technology Support Facility, Michigan State University.
Mapping and gene expression quantitation
Sequence adaptors were removed in the first quality control process using Trimmomatic (version 0.32) software65 to obtain the pass-filter (PF) reads. Low quality bases were further trimmed from the PF reads using custom Python scripts eliminating the first 15 nucleotides. Sickle (v1.33)66 was used with a sliding window average score of 30 in removing reads with “N”s, and minimum read length of 30 bps, and ended with clean reads. The clean reads were then used to map to the chicken genome reference (galGal4) using TopHat2 (v2.0.12)67 and Bowtie2 (v2.2.3)68 with default parameters. Transcript abundance and differential expression of genes were estimated with Cufflink (v2.2.1)69. FPKM values were obtained to quantify relative expression of transcripts.
Analyses of DEGs between treatment and chicken line groups
The number of reads per gene for each sample were counted using HTSeq70. In each of the pairwise comparisons (between infected birds and control in each line, and between the two lines with and without MDV infection), DEGs were identified by using the DESeq R package (2.1.0) and selected using a filter criterion of FDR <0.05 and FC >2. To better understand the functional involvements of these DEGs, g:Proflier (http://biit.cs.ut.ee/gprofiler/index.cgi)71 were used for the gene annotation, GO term and pathway enrichment analyses.
Droplet DigitalTM PCR validation of gene expression
To validate sequencing data, three genes from each of the treatment groups were selected and re-evaluated on a Droplet DigitalTM PCR (QX200TM ddPCR system; Bio-Rad Laboratories, Inc., Hercules, CA, USA). The primers for ddPCR were designed with Primer3Plus (http://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi/) and are listed in Table S10. A total of seven genes was selected. The cDNA samples used in ddPCR validation were reversely transcribed from individual RNA samples using the iScriptTM RT Supermix Kit (Cat No. 170–8841) and following the manufacturer’s instructions (Rio-Rad). Those total RNA samples were the same samples used in cDNA library preparation for the RNA-Seq analysis of this study. A ddPCR reaction mixture of 25 μL in final volume was initially prepared per gene per biological sample including 2 μL of cDNA, 12.5 μL of EvaGreen Supermix (Cat No. 1864034), 0.5 μL of each of the forward and reverse primers (200 nM; synthesized by Eurofins Genomics, Huntsville, AL), and 9.5 μL of nuclease-free water. Of which, 20 μL were loaded into one of 8 sample channels of a DG8TM cartridge (Cat No. 1864008, Bio-Rad). Each oil well was loaded with 70 μL of droplet generating oil (Cat No. 1864006, Bio-Rad). The loaded DG8TM cartridges were placed on a QX200TM droplet generator (Bio-Rad) to generate the digital droplets. Forty μL of the generated droplet emulsion for each sample were transferred to a well in a 96-well PCR plate followed by polymerase chain reaction with EvaGreen on a C1000TM Thermal Cycler (Bio-Rad). The cycling conditions were 95 °C for 5 min, followed by 40 cycles of 95 °C for 15 s, 58 °C for 60 s, and a final extension step of 98 °C for 10 min. The droplets post PCR were read well by well on a QX200TM droplet reader (Bio-Rad). PCR-positive and PCR-negative droplets in each of the wells were counted and analyzed with the QuantaSoftTM Software (Version 1.7, Bio-Rad).
Following similar procedures, two MDV microRNA expression levels in each of the total RNA samples included in this study were determined by ddPCR using customer primers (Table S11) to ensure that all the MDV challenged birds included in the samplings had entered the latent phase.
References
Calnek, B. W. Pathogenesis of Marek’s disease virus infection. Curr. Top. Microbiol. Immunol 255, 25–55 (2001).
Marek, J. Multiple Nervenentzündung (Polyneuritis) bei Hühnern. Dtsch. Tierärztl. Wochenschr 15, 417–421 (1907).
Smith, J. et al. Systems analysis of immune responses in Marek’s disease virus-infected chickens identifies a gene involved in susceptibility and highlights a possible novel pathogenicity mechanism. Journal of virology 85, 11146–11158 (2011).
Baigent, S. J. & Davison, F. In Marek’s Disease - An Evolving Problem (eds Davison, F. & Nair, V.) Ch. 6, 62–77 (ELSEVIER Academic Press, 2004).
Mitra, A. et al. Histone modifications induced by MDV infection at early cytolytic and latency phases. BMC Genomics 16, 311, https://doi.org/10.1186/s12864-015-1492-6 (2015).
Bumstead, J. M., Howes, K. & Payne, L. N. Effect of cyclosporin a on normal, mitogen-stimulated and Marek’s disease virus-exposed and transformed chicken lymphoid cells. Avian Pathol 14, 99–113, https://doi.org/10.1080/03079458508436211 (1985).
Calnek, B. W. Marek’s disease–a model for herpesvirus oncology. Crit Rev. Microbiol 12, 293–320 (1986).
Schermuly, J. et al. In vitro model for lytic replication, latency, and transformation of an oncogenic alphaherpesvirus. Proc Natl Acad Sci USA 112, 7279–7284, https://doi.org/10.1073/pnas.1424420112 (2015).
Lawn, A. M. & Payne, L. N. Chronological study of ultrastructural changes in the peripheral nerves in Marek’s disease. Neuropathol. Appl. Neurobiol 5, 485–497 (1979).
Payne, L. N. & Biggs, P. M. Studies on Marek’s disease. II. Pathogenesis. J. Natl. Cancer Inst 39, 281–302 (1967).
Sharma, J. M., Witter, R. L. & Burmester, B. R. Pathogenesis of Marek’s disease in old chickens: lesion regression as the basis for age-related resistance. Infect. Immun 8, 715–724 (1973).
Abdul-Careem, M. F. et al. Induction of innate host responses in the lungs of chickens following infection with a very virulent strain of Marek’s disease virus. Virology 393, 250–257, https://doi.org/10.1016/j.virol.2009.08.001 (2009).
Jarosinski, K. W., Tischer, B. K., Trapp, S. & Osterrieder, N. Marek’s disease virus: lytic replication, oncogenesis and control. Expert. Rev. Vaccines 5, 761–772, https://doi.org/10.1586/14760584.5.6.761 (2006).
Bacon, L. D. Influence of the major histocompatibility complex on disease resistance and productivity. Poult Sci 66, 802–811, https://doi.org/10.3382/ps.0660802 (1987).
Bacon, L. D., Hunt, H. D. & Cheng, H. H. A review of the development of chicken lines to resolve genes determining resistance to diseases. Poult. Sci 79, 1082–1093 (2000).
Bacon, L. D., Hunt, H. D. & Cheng, H. H. Genetic resistance to Marek’s disease. Curr. Top. Microbiol. Immunol 255, 121–141 (2001).
Chang, S. et al. Host genetic resistance to Marek’s disease sustains protective efficacy of herpesvirus of turkey in both experimental and commercial lines of chickens. Vaccine 32, 1820–1827, https://doi.org/10.1016/j.vaccine.2014.01.092 (2014).
Chang, S. et al. Genetic susceptibility to and presence of endogenous avian leukosis viruses impose no significant impact on survival days of chickens challenged with very virulent plus Marek’s disease virus. Annals of Viorology and Research 1, 1007 (2015).
Xie, Q. M. et al. Genomic Variation between Genetic Lines of White Leghorns Differed in Resistance to Marek’s Disease. Journal of Clinical Epigenetics 3, 7, https://doi.org/10.21767/2472-1158.100063 (2017).
Stone, H. A. In Technical Bulletin Vol. No. 1514 (ed. Agriculture Research Service USDA) (USDA-ARS, Washington, DC, 1975).
Gallatin, W. M. & Longenecker, B. M. Genetic resistance to herpesvirus-induced lymphoma at the level of the target cell determined by the thymic microenvironment. Int. J. Cancer 27, 373–380 (1981).
Lee, L. F., Powell, P. C., Rennie, M., Ross, L. J. & Payne, L. N. Nature of genetic resistance to Marek’s disease in chickens. J. Natl. Cancer Inst 66, 789–796 (1981).
Fredericksen, T. L., Longenecker, B. M., Pazderka, F., Gilmour, D. G. & Ruth, R. F. A T-cell antigen system of chickens: Ly-4 and Marek’s disease. Immunogenetics 5, 535–552 (1977).
Baigent, S. J., Ross, L. J. & Davison, T. F. Differential susceptibility to Marek’s disease is associated with differences in number, but not phenotype or location, of pp38+ lymphocytes. J. Gen. Virol 79(Pt 11), 2795–2802, https://doi.org/10.1099/0022-1317-79-11-2795 (1998).
Bumstead, N., Sillibourne, J., Rennie, M., Ross, N. & Davison, F. Quantification of Marek’s disease virus in chicken lymphocytes using the polymerase chain reaction with fluorescence detection. J Virol. Methods 65, 75–81, S0166093496021726 (1997).
Kaiser, P., Underwood, G. & Davison, F. Differential cytokine responses following Marek’s disease virus infection of chickens differing in resistance to Marek’s disease. J Virol 77, 762–768 (2003).
Quere, P., Rivas, C., Ester, K., Novak, R. & Ragland, W. L. Abundance of IFN-alpha and IFN-gamma mRNA in blood of resistant and susceptible chickens infected with Marek’s disease virus (MDV) or vaccinated with turkey herpesvirus; and MDV inhibition of subsequent induction of IFN gene transcription. Arch. Virol 150, 507–519, https://doi.org/10.1007/s00705-004-0435-3 (2005).
Abdul-Careem, M. F. et al. Host responses in the bursa of Fabricius of chickens infected with virulent Marek’s disease virus. Virology 379, 256–265 (2008).
Heidari, M. et al. Marek’s disease virus-induced immunosuppression: array analysis of chicken immune response gene expression profiling. Viral immunology 23, 309–319 (2010).
Hu, X. et al. Transcriptional analysis of host responses to Marek’s disease virus infection in chicken thymus. Intervirology 58, 95–105, https://doi.org/10.1159/000370069 (2015).
Liu, H., Cheng, H. H., Tirunagaru, V., Sofer, L. & Burnside, J. A strategy to identify positional candidate genes conferring Marek’s disease resistance by integrating DNA microarrays and genetic mapping. Animal genetics 32, 351–359 (2001).
Morgan, R. W. et al. Induction of host gene expression following infection of chicken embryo fibroblasts with oncogenic Marek’s disease virus. Journal of virology 75, 533–539 (2001).
Perumbakkam, S., Muir, W. M., Black-Pyrkosz, A., Okimoto, R. & Cheng, H. H. Comparison and contrast of genes and biological pathways responding to Marek’s disease virus infection using allele-specific expression and differential expression in broiler and layer chickens. BMC genomics 14, 64, https://doi.org/10.1186/1471-2164-14-64 (2013).
Sarson, A. J., Abdul-Careem, M. F., Zhou, H. & Sharif, S. Transcriptional analysis of host responses to Marek’s disease viral infection. Viral immunology 19, 747–758 (2006).
Sarson, A. J., Parvizi, P., Lepp, D., Quinton, M. & Sharif, S. Transcriptional analysis of host responses to Marek’s disease virus infection in genetically resistant and susceptible chickens. Anim Genet 39, 232–240, https://doi.org/10.1111/j.1365-2052.2008.01710.x (2008).
Yu, Y. et al. Temporal transcriptome changes induced by MDV in marek’s disease-resistant and-susceptible inbred chickens. BMC genomics 12, 501 (2011).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet 10, 57–63, https://doi.org/10.1038/nrg2484 (2009).
Speck, S. H. & Ganem, D. Viral latency and its regulation: lessons from the gamma-herpesviruses. Cell Host Microbe 8, 100–115, https://doi.org/10.1016/j.chom.2010.06.014 (2010).
Zhao, P. et al. In vivo expression patterns of microRNAs of Gallid herpesvirus 2 (GaHV-2) during the virus life cycle and development of Marek’s disease lymphomas. Virus Genes 50, 245–252, https://doi.org/10.1007/s11262-015-1167-z (2015).
Haller, O., Staeheli, P. & Kochs, G. Interferon-induced Mx proteins in antiviral host defense. Biochimie 89, 812–818, https://doi.org/10.1016/j.biochi.2007.04.015 (2007).
Sheikh, M. Y., Choi, J., Qadri, I., Friedman, J. E. & Sanyal, A. J. Hepatitis C virus infection: molecular pathways to metabolic syndrome. Hepatology 47, 2127–2133, https://doi.org/10.1002/hep.22269 (2008).
Syed, G. H., Amako, Y. & Siddiqui, A. Hepatitis C virus hijacks host lipid metabolism. Trends Endocrinol. Metab 21, 33–40, https://doi.org/10.1016/j.tem.2009.07.005 (2010).
Bigger, C. B. et al. Intrahepatic gene expression during chronic hepatitis C virus infection in chimpanzees. J. Virol 78, 13779–13792, https://doi.org/10.1128/JVI.78.24.13779-13792.2004 (2004).
Su, A. I. et al. Genomic analysis of the host response to hepatitis C virus infection. Proc. Natl. Acad. Sci. USA 99, 15669–15674, https://doi.org/10.1073/pnas.202608199 202608199 (2002).
Currie, E., Schulze, A., Zechner, R., Walther, T. C. & Farese, R. V. Jr. Cellular fatty acid metabolism and cancer. Cell Metab 18, 153–161, https://doi.org/10.1016/j.cmet.2013.05.017 (2013).
Miyamoto, M. et al. Regulated expression of a gene encoding a nuclear factor, IRF-1, that specifically binds to IFN-beta gene regulatory elements. Cell 54, 903–913, S0092-8674(88)91307-4 (1988).
Lasoudris, F. et al. IL4I1: an inhibitor of the CD8(+) antitumor T-cell response in vivo. Eur. J Immunol 41, 1629–1638, https://doi.org/10.1002/eji.201041119 (2011).
Jones, F. S. & Jones, P. L. The tenascin family of ECM glycoproteins: structure, function, and regulation during embryonic development and tissue remodeling. Dev. Dyn 218, 235–259, doi:10.1002/(SICI)1097-0177(200006)218:2<235::AID-DVDY2>3.0.CO;2-G;10.1002/(SICI)1097-0177(200006)218:2<235::AID-DVDY2>3.0.CO;2-G [doi] (2000).
Bandyopadhyay, S. et al. Hepatitis C virus infection and hepatic stellate cell activation downregulate miR-29: miR-29 overexpression reduces hepatitis C viral abundance in culture. J. Infect. Dis 203, 1753–1762, https://doi.org/10.1093/infdis/jir186 (2011).
Bozym, R. A. et al. Focal adhesion kinase is a component of antiviral RIG-I-like receptor signaling. Cell Host. Microbe 11, 153–166, https://doi.org/10.1016/j.chom.2012.01.008 (2012).
Krupp, M. et al. The functional cancer map: a systems-level synopsis of genetic deregulation in cancer. BMC Med. Genomics 4, 53, https://doi.org/10.1186/1755-8794-4-53 (2011).
Kuphal, S., Bauer, R. & Bosserhoff, A. K. Integrin signaling in malignant melanoma. Cancer Metastasis Rev 24, 195–222, https://doi.org/10.1007/s10555-005-1572-1 (2005).
Konig, R. et al. Human host factors required for influenza virus replication. Nature 463, 813–817, https://doi.org/10.1038/nature08699 (2010).
Okamoto, T. et al. Hepatitis C virus RNA replication is regulated by FKBP8 and Hsp90. EMBO J 25, 5015–5025, https://doi.org/10.1038/sj.emboj.7601367 (2006).
Osterrieder, N., Kamil, J. P., Schumacher, D., Tischer, B. K. & Trapp, S. Marek’s disease virus: from miasma to model. Nat. Rev. Microbiol 4, 283–294, https://doi.org/10.1038/nrmicro1382 (2006).
Schat, K. A. Marek’s disease: a model for protection against herpesvirus-induced tumours. Cancer Surv 6, 1–37 (1987).
Dubail, J. et al. ADAMTS-2 functions as anti-angiogenic and anti-tumoral molecule independently of its catalytic activity. Cell Mol. Life Sci 67, 4213–4232, https://doi.org/10.1007/s00018-010-0431-6 (2010).
Muthusamy, V. et al. Epigenetic silencing of novel tumor suppressors in malignant melanoma. Cancer Res 66, 11187–11193, https://doi.org/10.1158/0008-5472.CAN-06-1274 (2006).
Draheim, K. M. et al. ARRDC3 suppresses breast cancer progression by negatively regulating integrin beta4. Oncogene 29, 5032–5047, https://doi.org/10.1038/onc.2010.250 (2010).
Lindberg, D., Akerstrom, G. & Westin, G. Mutational analyses of WNT7A and HDAC11 as candidate tumour suppressor genes in sporadic malignant pancreatic endocrine tumours. Clin. Endocrinol. (Oxf) 66, 110–114, https://doi.org/10.1111/j.1365-2265.2006.02694.x (2007).
Hoevel, T., Macek, R., Swisshelm, K. & Kubbies, M. Reexpression of the TJ protein CLDN1 induces apoptosis in breast tumor spheroids. Int. J Cancer 108, 374–383, https://doi.org/10.1002/ijc.11571 (2004).
Huang, G. et al. 15-Hydroxyprostaglandin dehydrogenase is a target of hepatocyte nuclear factor 3beta and a tumor suppressor in lung cancer. Cancer Res 68, 5040–5048, https://doi.org/10.1158/0008-5472.CAN-07-6575 (2008).
Myung, S. J. et al. 15-Hydroxyprostaglandin dehydrogenase is an in vivo suppressor of colon tumorigenesis. Proc. Natl. Acad. Sci. USA 103, 12098–12102, https://doi.org/10.1073/pnas.0603235103 (2006).
Wolf, I. et al. 15-hydroxyprostaglandin dehydrogenase is a tumor suppressor of human breast cancer. Cancer Res 66, 7818–7823, https://doi.org/10.1158/0008-5472.CAN-05-4368 (2006).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120, https://doi.org/10.1093/bioinformatics/btu170 (2014).
A sliding-window, adaptive, quality-based trimming tool for fastQ files v. 1.33 (2011).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14, R36, https://doi.org/10.1186/gb-2013-14-4-r36 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359, https://doi.org/10.1038/nmeth.1923 (2012).
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat Biotechnol 31, 46–53, https://doi.org/10.1038/nbt.2450 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169, https://doi.org/10.1093/bioinformatics/btu638 (2015).
Reimand, J., Kull, M., Peterson, H., Hansen, J. & Vilo, J. g:Profiler–a web-based toolset for functional profiling of gene lists from large-scale experiments. Nucleic Acids Res 35, W193–200, https://doi.org/10.1093/nar/gkm226 (2007).
Author information
Authors and Affiliations
Contributions
H.Z., S.C. and Q.X. jointly conceived the study and conducted the challenge trial; K.D. analyzed the data, wrote the main manuscript text, and prepared the figures. P.Z. revised the manuscript. All authors contributed in the final revision and review of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dong, K., Chang, S., Xie, Q. et al. RNA Sequencing revealed differentially expressed genes functionally associated with immunity and tumor suppression during latent phase infection of a vv + MDV in chickens. Sci Rep 9, 14182 (2019). https://doi.org/10.1038/s41598-019-50561-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-50561-x
- Springer Nature Limited