Abstract
Immune deficiency (IMD) is a death domain-containing protein that is essential for the IMD/NF-κB humoral and epithelial immune responses to Gram-negative bacteria and viruses in insects. In the immune signaling cascade, IMD is recruited together with FADD and the caspase DREDD after the mobilization of PGRP receptors. Activated IMD regulates the expression of effector antimicrobial peptides (AMP) that protect against invading microorganisms. To date, most studies of the IMD pathway, and the IMD gene in particular, have been restricted to Drosophila; few similar studies have been conducted in other model insects. Herein, we cloned and functionally characterized an IMD homolog from the mealworm beetle Tenebrio molitor (TmIMD) and studied its role in host survival in the context of pathogenic infections. Phylogenetic analysis revealed the conserved caspase cleavage site and inhibitor of apoptosis (IAP)-binding motif (IBM). TmIMD expression was high in the hemocytes and Malpighian tubules of Tenebrio late-instar larvae and adults. At 3 and 6 hours’ post-infection with Escherichia coli, Staphylococcus aureus, or Candida albicans, TmIMD expression significantly increased compared with mock-infected controls. Knockdown of the TmIMD transcript by RNAi significantly reduced host resistance to the Gram-negative bacterium E. coli and fungus C. albicans in a survival assay. Strikingly, the expression of nine T. molitor AMPs (TmTenecin1, TmTenecin2, TmTenecin4, TmDefensin2, TmColeoptericin1, TmColeoptericin2, TmAttacin1a, TmAttacin1b, and TmAttacin2) showed significant downregulation in TmIMD knockdown larvae challenged with E. coli. These results suggest that TmIMD is required to confer humoral immunity against the Gram-negative bacteria, E. coli by inducing the expression of critical transcripts that encode AMPs.
Similar content being viewed by others
Introduction
Insects represent the largest group within the arthropod phylum. Over 50% of the described eukaryotes are insects, which inhabit variety of habitats and interact with diverse surrounding environments1. The innate immune system of insects is specialized and is considered as one of the major attributable factors for the robustness and consistency of species across ecosystems. The innate immune system of insects recognizes and removes microbial threats through cellular and systemic immune responses. Cellular responses include phagocytosis mediated by plasmatocytes (these cells can be compared to white blood cells in humans) and lamellocytes2. Systemic responses (including local responses in the epithelia of the skin and intestine) are mediated by antimicrobial peptides (AMPs) produced in the fat body (equivalent to the vertebrate liver) of insects3. The AMPs are transcriptionally regulated by the Toll and immune deficiency (IMD) signaling pathways, in addition to the Janus Kinase/signal transducer and activator of transcription (JAK/STAT) and c-Jun N-terminal kinase (JNK) pathways that are conspicuous and evolutionarily conserved across insect taxa4. The core Toll and IMD signaling pathways act synergistically in insects to activate innate immune responses towards microorganisms via the production of lysozymes, AMPs, and other effector proteins. These synergistic interactions demonstrate cross-regulation of signaling processes that can lead to a broad-spectrum host response. AMPs cloned from many insect models including Drosophila, have revealed the conserved nuclear factor kappaB (NF-κB) binding sites in their promoter elements, suggesting the evolutionary conservation of the AMP production mechanisms and effector actions5. Although both the Toll and IMD pathways are important in controlling the expression of AMPs, the IMD pathway in particular is essential as it regulates the expression of most AMPs in Drosophila6. By contrast the Toll pathway is involved in embryonic axis formation and controls a limited set of AMPs7,8.
The IMD signaling pathway is activated after the recognition of Gram-negative bacteria or viruses9,10. In Drosophila, the IMD signaling pathway is activated through the recruitment of the death domain-containing intracellular protein and Fas-associated protein with death domain (FADD) as well as the caspase death-related ced-3/Nedd2-like protein (DREDD). This occurs as a consequence of bacterial peptidoglycan (PGN)-mediated multimerization and clustering of the transmembrane receptor peptidoglycan recognition protein-LC (PGRP-LC)11,12. DREDD cleavesIMD protein at the caspase cleavage site, creating a novel binding site for inhibitor of apoptosis-2 (IAP-2). The E2-ubiquitin-conjugating enzyme complex (UEV1a, Ubc13, and Ubc5) associates with the IAP-2 binding site and activates the TGF-beta-activated kinase 1 binding protein 2/Mitogen-activated protein kinase kinase kinase 7 (Tab2/Tak1) complex11,13. Subsequently, this complex drives phosphorylation of the IκB kinase (IKK) complex (ird5 and key) to activate Relish. After translocating to the nucleus, Relish activates the transcription of AMP genes that are essential for humoral and epithelial immune responses14,15.
In the context of IMD signaling, the IMD protein acts as an adapter molecule (similar to mammalian receptor interacting proteins) in the antibacterial defense system of insects. This has been established in the Drosophila model in which the overexpression of IMD led to the constitutive expression of AMP genes16. Studies of the IMD pathway, especially of IMD, have been largely restricted to the Drosophila model system, as limited studies have been conducted in other insect models. Although many homologs of the IMD pathway including the IMD gene, have been discovered in other insects through high-throughput RNA sequencing (RNA-seq)17,18, the mechanisms that regulate AMP expression needs to be further explored. In one recent study, the requirement of Bombyx mori IMD (BmIMD) in the induction of Drosomycin, DiptericinA, and DiptericinB was studied via overexpression in Drosophila S2 cells19. In the red flour beetle Tribolium castaneum, the IMD pathway was proposed to confer resistance against the Gram-negative and Gram-positive pathogens Enterobacter cloacae and Bacillus subtilis, respectively, through the robust induction of AMP genes. In T. castaneum pupae, the RNAi mediated knockdown of IMD transcripts led to downregulation of AMPs, such as Attacin1 and Coleoptericin1, at 24 h post-infection with Escherichia coli20.
In the Tenebrio model system, limited information is available about genes in the IMD pathway and the regulation of AMP gene expression. This is in contrast to the elegant studies that have been conducted on the Toll signaling pathway in model insects. For example, the extracellular signaling cascades involving proteases and the Toll ligand Spätzle have been elucidated21,22. Additionally, intracellular signaling cascades emanating from the Toll -Spätzle complex and the antimicrobial responses have been partially explored23,24. In this study, we report for the first time, cloning of the full-length cDNA of the T. molitor IMD gene (TmIMD) and the analysis of its sequence features and phylogenetic relationships at the amino acid level. The temporal and spatial expression profiles of TmIMD transcripts and its response to bacterial (E. coli and S. aureus) and fungal (Candida albicans) challenges were characterized at multiple time-points. We used RNAi to understand the requirement of TmIMD transcripts for survival of T. molitor larvae and AMP gene expression. A set of nine AMP genes was found to be downregulated in the E. coli infection model after the silencing of TmIMD transcripts. Our study advances an understanding of the IMD regulatory pathway in the innate immune response of T. molitor through the regulation of expression of AMP genes.
Results
Gene organization, cDNA sequence analysis, and phylogenetics
We used a standalone local blast server (downloaded from ftp://ftp.ncbi.nlm.nih.gov/blast/executables/blast+/) and the T. molitor RNAseq and expressed sequence tag libraries (unpublished) to retrieve the TmIMD sequence. Local tblastn analysis was conducted using the Tribolium castaneum IMD (TcIMD) sequence as the query (NCBI Reference Sequence XM_008201183.2). We retrieved one full-length TmIMD ORF sequence. This sequence was used as a query against the T. molitor DNAseq database (unpublished) as the subject in a blastn analysis to identify the TmIMD genomic sequence. Analyzing the gene structure of TmIMD, we identified three exons and two introns (Fig. 1A). A detailed study on the TmIMD gene organization is shown in Fig. 1B. The exons are 181 bp (exon 1), 286 bp (exon 2), and 142 bp (exon 3), while the introns 1 and 2 are 52 and 56 bp, respectively. The transcription start site (t) was identified at 751 bp upstream of the translation start codon (ATG) and the poly(A) sequence was found at 448 nucleotides downstream of the translation stop codon (TAA). The ORF sequence contains 609 nucleotides and encodes a 202 amino acid residues (Fig. 2). A death domain (72 amino acids), similar to that of the mammalian receptor-interacting protein (RIP), was predicted at the C-terminus of the protein. This domain is evolutionarily conserved in many multicellular organisms and is critical for the recruitment of the downstream effector molecules involved in apoptotic and immune signaling pathways25. N-glycosylation sites were predicted at N67 and N85, while four polyubiquitination sites were predicted at K26, K28, K86, and K92. No O-glycosylation sites were predicted for TmIMD, in contrast to six O-glycosylation sites in the IMD homolog of B. mori19. The TmIMD nucleotide and translated amino acid sequence was deposited in GenBank (through BankIt submission) under the accession number MK121950.
Genomic organization of the T. molitor IMD (TmIMD) gene. (A) Schematic representation of the TmIMD gene. The genomic structure (open bar) shows three exons (black) interspersed with two introns (grey). The translation start (ATG) and stop (TAA) codons are marked demarcating the ORF region. (B) The sequence of the TmIMD gene with the ORF region (capitalized). Exonic regions are in bold. Transcription start, translation start codon, and translation stop codons are boxed. The poly(A) sequence is indicated in italics. In the mRNA, the poly(A) begins after the last nucleotide in exon 3.
Nucleotide and deduced amino acid sequences of TmIMD. The nucleotides are numbered from the first base of the ORF region represented by the start codon. The death domain extending from position 120 to 191 is shown in an open box. The potential N-glycosylation and polyubiquitination sites are indicated with triangles (N67 and N85) and circles (K26, K28, K86 and K92), respectively.
A blastp search of the TmIMD amino acid sequence performed against the NCBI non-redundant database retrieved several other proteins showing homology to IMD proteins in insects and other organisms. Multiple alignment analysis of TmIMD with other insect homologs showed that the amino acids D and A at the interface of caspase cleavage and the IAP-binding motif (IBM) were conserved (Fig. S1). These form the caspase cleavage site critical for IMD cleavage by DREDD in a FADD-dependent manner as well as the downstream regulation of the transcription factor Relish and induction of the expression of AMPs26. Furthermore, most of lepidopteran insect sequences showed a conserved ‘LKSDA’ motif, whilst in the coleopterans it was ‘LTTDA’ and in dipterans ‘LETDA’ (and was ‘LEKDA’ in Drosophila isoforms).
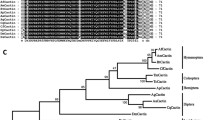
A phylogenetic tree was constructed using IMD amino acid sequences from 16 representative insect species, including Drosophila IMD isoform A and B (DmIMD-A & DmIMD-B) (Fig. 3A). The results revealed the distribution of the IMD sequences into three major groups comprising of Lepidoptera, Coleoptera/Hymenoptera, and Diptera. TmIMD was firstly grouped with its close relative TcIMD protein, and then the members of this coleopteran group was clustered into the branch of hymenopterans comprising of Nasonia vitripennis IMD (NvIMD) and Apis mellifera IMD (AmIMD) proteins. Among the Dipterans, the DmIMD-A and DmIMD-B sequences were identical, while Anopheles gambiae IMD (AgIMD) and Aedes aegypti IMD (AaIMD) were found to be related. The evolutionary divergence matrix also revealed less than 25% identity of TmIMD with IMD proteins from Lepidoptera and Diptera orders (Fig. 3B). Considering the multiple alignment and divergence matrix of IMD sequences from the orthologs, it is evident that there is a selection pressure to maintain the caspase/IBM motifs in TmIMD, whereas the other sequences do not show the same signature of selection.
Phylogenetic tree and percentage identity/distance matrix of TmIMD. (A) The unrooted Neighbor- Joining phylogenetic tree of BmIMD and orthologous proteins from insects based on the JTT matrix model. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1,000 replicates) is shown next to the branches. An arrow head indicates TmIMD. (B) Estimates of evolutionary divergence between sequences of TmIMD and orthologous proteins from insects are illustrated by percentage identity (upper block) and distance matrix (lower block). The abbreviations and GenBank accession numbers (in parenthesis) are: TcIMD, T. castaneum IMD (XP_008199405); DmIMD-A, D. melanogaster IMD, isoform A (NP_573394.1); DmIMD-B, D. melanogaster IMD, isoform B (NP_001286572.1); HgIMD, Hebomoia glaucippe IMD (AFK75936.1); AgIMD, A. gambiae (XP_001688608); PcIMD, Polygonia c-album IMD (AFK75940.1); VcIMD, Vanessa cardui IMD (AFK75942.1); AmIMD, A. mellifera IMD (NP_001157189.1); VaIMD, Vanessa atalanta IMD (AFK75941.1); AaIMD, A. aegypti AAEL010083-PA (EAT37980.1); AlIMD, Appias lyncida IMD (AFK75935.1); SgIMD, Schistocerca gregaria IMD-like protein (AFK75938.1); HsIMD, Heliothis subflexa IMD (AFK75939.1); PxIMD, Plutella xylostella IMD (AFK75937.1); and NvIMD, Nasonia vitripennis IMD (NP_001135910.1).
Temporal and spatial expression of TmIMD transcripts
We analyzed the expression patterns of TmIMD mRNA transcripts in different days of larval, pupal, and adult development using qRT-PCR. The expression of TmIMD mRNA was observed in all the developmental stages of the insect, but was more pronounced in the late larval, prepupal, and very early pupal stages (P0 and P1) (Fig. 4A). Expression data in Fig. 4 refer to the late larval stage (set to 1.0). The expression levels of TmIMD mRNA transcripts peaked at stage P1 (close to the larval-pupal transition). In the mid- and late-pupal stages, the mRNA transcript levels of TmIMD were found to be lower compared to the other stages, with the lowest levels at stage P5. In the A1 and A2 stages, the expression of TmIMD mRNA was high, but non-significant compared to other stages, except for the P1 and P5 stages.
Developmental and tissue distribution of TmIMD. (A) The mRNA transcript levels of TmIMD in different developmental stages of the insect were analyzed using qPCR. LL: late-instar larva; PP: pre-pupa; P0–P7: zero to seven-day old pupa; A1-A2: one to two-day old adult. (B) The mRNA transcript levels of TmIMD in larval and adult tissues of the insect were analyzed by qPCR. IT: integument; MT: Malpighian tube; GT: gut; HC: hemocytes; FB: fat body; OV: ovary; TE: testis. T. molitor Ribosomal protein L27a (TmL27a) was used as an internal control. Vertical bars represent standard errors (n = 3). One-way ANOVA and Tukey’s multiple range tests at 95% confidence level (P < 0.05) were performed and used to determine the level of significance of differences.
To gain further insights into the function of TmIMD in different tissues of T. molitor larvae and adults, we examined the expression patterns of TmIMD mRNA by qRT-PCR analysis. Here, TmIMD expression referred to the expression in the integument (set to 1.0). We found that TmIMD was expressed in all of the tested tissues of T. molitor larva and adult. The expression levels of TmIMD mRNA were the highest in the hemocytes compared to the expression levels in all other larval tissues analyzed (Fig. 4B). In other tissues, such as the fat body and gut that are involved in systemic and local immune responses, respectively, the expression levels of TmIMD mRNA was also found to be higher. In the adult tissues of T. molitor, the expression levels of TmIMD mRNA were found to be the highest in the hemocytes, followed by the expression in Malpighian tubules and fat body, respectively (Fig. 4C).
Microbial challenge-induced expression of TmIMD transcripts
The changes in expression of TmIMD mRNA were analyzed in response to immune challenges with the Gram-negative bacteria E. coli, Gram-positive bacteria S. aureus, and fungus C. albicans (Fig. 5). TmIMD mRNA increased more than 8-fold 6 h after the injection of E. coli. In S. aureus-injected T. molitor larvae, TmIMD mRNA was increased 6-fold at 6 h. These results were found to be significant at 95% confidence levels. Even after the injection of the fungus, there was a significant expression of TmIMD transcripts. In our time-course, TmIMD transcripts decreased by 12 h after microbial challenge. These early induction patterns in the microbial challenge studies suggests a putative role of TmIMD in systemic immune signaling.
The mRNA transcript levels of TmIMD in whole larvae (10–12th instar) after microbial challenge. After challenge injections of E. coli, S. aureus, or C. albicans, the mRNA transcript levels of TmIMD were analyzed by qPCR. TmL27a was used as an internal control. Results of triplicate experiments are shown with standard errors (n = 3). Asterisks indicate significant differences at P < 0.05 between control (PBS) and microbial challenge groups for each time point.
TmIMD in vivo knockdown assay
To determine whether TmIMD is involved in conferring immune response against bacterial and fungal challenge, we generated double-stranded RNA (dsRNA) against TmIMD and knocked it down in systemic circulation in the T. molitor whole-larvae. After confirmation of efficient knockdown (generally 2 days after dsRNA treatment), we investigated the lethality rates of TmIMD knockdown larvae following E. coli, S. aureus, and C. albicans challenge using larval mortality assays. We constructed a dsRNA corresponding to a 363 bp portion of the TmIMD gene. A dsRNA for the Enhanced Green Fluorescent Protein (EGFP) sequence (derived from plasmid EGFP-C1) was also synthesized to act as a negative control in the survival analysis. The dsRNA for the EGFP sequence corresponded to a 546 bp portion of the ORF sequence. We confirmed efficient knockdown (95%) of TmIMD mRNA by qRT-PCR (Fig. 6A). We find that the knockdown efficiency of TmIMD mRNA was stable at least for a duration of 10 days from day 2 after dsTmIMD injection. Next, we challenged TmIMD knockdown larvae with E. coli, S. aureus, or C. albicans, and subsequently analyzed the survival of larvae for the duration of 10 days. The survival analysis was not continued beyond 10 days as TmIMD knockdown was not active. The survival of the infected dsEGFP-injected T. molitor was considered for comparison. Of note, there was a 40% reduction in survival compared to controls in IMD knockdown beetles subjected to E. coli infection (Fig. 6B). A non-significant 15–20% reduction in the survival of larvae was observed in the case of S. aureus infection (Fig. 6B), and a significant 40% reduction was observed in the case of C. albicans infection (Fig. 6C). Considering the survival analysis, TmIMD was confirmed as a positive regulator because its knockdown resulted in increased susceptibility against the Gram-negative bacteria E. coli and the fungus C. albicans. However, TmIMD is not directly involved in immune response against S. aureus.
RNAi knockdown of TmIMD enhanced the susceptibility of T. molitor larvae to E. coli and C. albicans infection. (A) T. molitor larvae (10–12th instar) were injected with dsTmIMD (2 µg per insect) and transcript silencing was measured by qPCR. The EGFP dsRNA-treated larvae served as a negative control. Data represent means ± standard error of three independent biological replicates. E. coli (B), S. aureus (C), and C. albicans (D) were inoculated at 2-days post RNAi, and larval survival was recorded daily for 10 days in each treatment group. The experiment was repeated three times with similar results. Statistical analysis of survival analysis was carried out based on Kaplan-Meier plots (log-rank chi-square test; *p < 0.05).
AMP gene expression analysis
To investigate the involvement of TmIMD in humoral immunity, we measured the transcript levels of 14 Tenebrio AMPs after microbial challenge in in vivo TmIMD silenced whole-larvae by qRT-PCR. The level of expression of AMPs were compared between TmIMD knockdown and EGFP control groups. The T. molitor AMPs such as TmTenecin, TmDefensin, TmCecropin, TmThaumatin-like protein, TmColeoptericin, and TmAttacin were considered for the studies. After confirmation of in vivo knockdown (2 days after dsRNA treatment), we injected larvae with E. coli, S. aureus, C. albicans, or phosphate buffered saline (PBS). After one day of microbial infection, AMP gene expression was analyzed. We found that from among the 14 AMPs analyzed, the expression levels of 9 AMPs were significantly upregulated (i.e., greater than 2000-fold) in dsEGFP-injected larvae. Except for TmTenecin3, TmDefensin1, TmCecropin2, TmThaumatin-likeprotein1, and TmThaumatin-like protein2, other AMPs showed very high transcriptional upregulation. TmColeoptericin1 (Fig. 7E) and TmTenecin4 (Fig. 7C) showed the maximum expression followed by TmTenecin2 (Fig. 7B), TmDefensin2 (Fig. 7D), TmAttacin1b (Fig. 7H), TmAttacin2 (Fig. 7I), TmAttacin1a (Fig. 7G), TmColeoptericin2 (Fig. 7F), and TmTenecin1 (Fig. 7A). In the dsTmIMD- injected larvae, expression levels of all of these 9 AMPs were significantly reduced after E. coli infection, explaining the larval susceptibility to the Gram-negative bacterial infection. Hence, it is confirmed that TmIMD is required for AMP induction and humoral immunity functions after Gram-negative bacterial infections. Further, AMPs such as TmTenecin3 (Fig. S2A), TmThaumatin-like protein1 (Fig. S2D), TmDefensin1 (Fig. S2B), TmThaumatin-like protein2 Fig. S2E), and TmCecropin2 (Fig. S2C) were not detected or were slightly induced in larvae treated with dsEGFP. In contrast, knock-down of IMD increased TmDefensin1 expression by more than 8,500-fold in E. coli-challenged conditions. In the C. albicans infection group, only TmDefensin2 and TmAttacin2 showed a significant reduction in expression under TmIMD knockdown conditions.
AMP expression levels in TmIMD-knockdown T. molitor larvae upon microorganism challenge. E. coli (Ec), S. aureus (Sa), or C. albicans (Ca) were injected into dsTmIMD-treated T. molitor larvae. The mRNA transcript levels of TmTenecin1 (A), TmTenecin2 (B) TmTenecin4 (C), TmDefensin2 (D), TmColeoptericin1 (E), TmColeoptericin2 (F), TmAttacin1a (G), TmAttacin1b (H), and TmAttacin2 (I) were analyzed by qPCR. EGFP dsRNA was used as a silencing control and TmL27a was used as an internal control. TmIMD knockdown showed impaired expression of AMPs upon E. coli infection in T. molitor larvae. Results show an average of three independent biological replicates with standard errors. All data were analyzed for significance using Student’s t-test (*p < 0.05).
Discussion
In this study, we identified a full-length ORF of the TmIMD gene in the coleopteran beetle T. molitor using bioinformatics analysis. The full-length ORF of TmIMD was then queried against the T. molitor genome database to retrieve the TmIMD genomic sequence. All IMD homologs in different insect species are 240–280 amino acids long. The TmIMD amino acid sequence was comparatively shorter to IMD-like proteins from most other Lepidopteran and Dipteran species. The coleopteran IMD homologs lack sequences that correspond to the non-conserved regions in IMD proteins but showed a completely conserved structure at the caspase cleavage site and the IBM motif. In TmIMD, the putative cleavage site occurs at 10LTTD/A15. In the Drosophila IMD protein, the caspase cleavage site (27LEKD/A31) was found to be critical for DREDD- mediated IMD cleavage in an FADD- dependent manner, which then drives further downstream signaling events12. Moreover, the cleavage is necessary to expose a neo N-terminus with a highly conserved IBM motif that binds the Death-associated inhibitor of apoptosis 2 (E3 ligase DIAP2) via its baculovirus IAP repeat 2/3 domains27. For the IMD pathway in Drosophila, the DIAP2-IMD interaction is rapidly ubiquitinated at the K63 position, and the polyubiquitin chains serve to recruit the TAB2/TAK1 complex28. Unlike the K63 polyubiquitination site in Drosophila, a potential polyubiquitination site at K92 occurs in TmIMD and BmIMD. Hence, it can be speculated that TmDREDD could cleave TmIMD in a TmFADD-dependent manner leading to the recruitment of the TmTAB2/TmTAK1 complex. This could subsequently activate TmRelish for downstream induction of AMPs. Similar studies have been attempted in the red flour beetle T. castaneum to understand the host defense mechanism against bacteria20,29, taking the advantage of the T. castaneum genome annotation (BioProjects: PRJNA15718, PRJNA12540)30. Taken together, we have identified the IMD gene from T. molitor and characterized the protein in detail. Because limited studies have been conducted on the functional characterization of IMD homologs and activation of the IMD-Relish-AMPs cascade in insects, our study should help to advance our understanding of innate immunity among insects.
This study showed that the mRNA expression levels of TmIMD are upregulated at 3–6 h after E. coli, S. aureus, or C. albicans challenge compared to a healthy wounding control. The expression of BmIMD is activated at the midgut level during oral infection with E. coli or S. aureus19. Irrespective of the mode of infection by microorganisms, IMD is implicated as a regulator of innate immune signaling in insect models. To understand how this occurs in T. molitor, we designed TmIMD-knockdown T. molitor larvae and studied larval mortality after infection with microorganisms. The RNAi technique is quite popular for characterizing cell signaling pathways triggered by a specific class of pathogen. In the Drosophila model, the components of the IMD pathway were analyzed after the synthesis of nearly 7,000 dsRNAs31. Furthermore, the technique has been employed to understand the molecular interactions and effector functions of extracellular and intracellular Toll signaling pathways in T. molitor23,32. In the present study, we found that T. molitor larvae are susceptible to E. coli infection with a mortality percentage of >60% in the TmIMD- knockdown cohort. A 40% mortality of larvae were also observed in the context of fungal infection. But in case of S. aureus infection, the mortality % in the TmIMD knockdown larvae was non-significant compared with the dsEGFP treated group. This suggests that TmIMD contributes to the innate immune response against the Gram-negative bacteria E. coli and the fungus C. albicans. Further in the intestinal infection model of D. melanogaster, IMD pathway is indispensable to the innate immunity process33. In a previous study, we reported the involvement of TmMyD88 in the positive regulation of host immune responses against S. aureus using RNAi23. MyD88 is an adapter protein in the Toll signaling pathway that mediates immune responses against most Gram-positive bacteria, including S. aureus34. Hence, we speculate that the induction of TmIMD after S. aureus challenge is not necessarily related with the immune responses and overall survival of T. molitor larvae. In systemically infected insect models, Gram-positive bacterial infections are mostly regulated by the Toll signaling pathway, but in the gut-specific immune responses in Drosophila, the IMD pathway is implicated for the clearance of Gram-positive bacteria S. aureus35. The IMD knockdown pupae of T. castaneum failed to mount an appropriate immune response against E. coli or B. subtilis infection36. Furthermore, in the Drosophila model, IMD pathway components were found to be proactively involved in promoting host resistance against the bacterium S. marcescens37. Understanding the preferential regulation of AMP genes under the influence of the Toll or IMD pathway is important in the context of infection models because these pathways are suggested to act synergistically and show cross-regulation20,38. A study has attributed the presence of GGGGATTYYY as the kB consensus motif for IMD-responsive genes and GGGAAADYCC for Toll-responsive genes, explaining the differential regulation of AMPs39. In one of our earlier studies on the T. molitor Toll signaling pathway, we confirmed the role of the Cactin protein in regulating the expression of seven AMP genes in response to E. coli and S aureus32. This study for the first time explains the E. coli-induced AMP gene expression under a TmIMD knockdown condition in T. molitor larvae. The significantly strong and robust expression of nine Tenebrio AMPs (TmColeoptericin1, TmTenecin4, TmTenecin2, TmTenecin1, TmDefensin2, TmAttacin1b, TmAttacin2, TmAttacin1a, and TmColeoptericin2) in an E. coli- challenged condition indicates that these AMPs are required to elicit a host defense response against infection. Further, in the TmIMD-knockdown insects, these AMPs showed a significant reduction in expression, suggesting that in T. molitor, TmIMD is required to regulate the expression of these AMPs and mount a host-protective response against the Gram-negative bacteria E. coli (Fig. 8). Nevertheless, it will be interesting to determine whether all upregulated AMPs show activity against such infective agents. The moderate expression of 2 AMPs (TmDefensin2 and TmAttacin2) was also observed in a C. albicans- challenged condition, indicating the involvement of these AMPs in the immune response of T. molitor against the fungi. Moreover, a downregulation in the expression of these AMPs after TmIMD knockdown suggests a possible requirement of this gene in regulating the AMPs and defense against fungal infection. AMPs, such as TmTenecin3, TmDefensin1, TmCecropin2, and TmThaumatin-like protein1/2-, showed low to very low expression in T. molitor larvae under infective conditions. This is surprising because TmTenecin3 contributes to host defense against fungal infections40. In T. castaneum, IMD knockdown reduced the expression levels of T. castaneum Attacin1, Defensin2, Defensin3, and Coleoptericin1 after E. coli challenge. However, that study was conducted in the context of understanding the preferential induction of AMP genes in response to microbial challenges by using RNAi to target either MyD88 (a Toll pathway adaptor protein) or IMD20. Irrespective of the dominant role of the IMD pathway in contributing to defense against E. coli, we also speculate the promiscuous role of the Toll pathway in directing the expression of AMPs via Dif- Relish heterodimer formation because the DAP-type peptidoglycan in Gram-negative bacteria is known to activate the Toll ligand Spätzle and induce defensin-like AMPs in Tenebrio larvae22.
A putative model for TmIMD-mediated regulation of AMPs in response to the Gram-negative bacteria E. coli. (A) In Tenebrio, PGRP-LC senses bacterial and/or fungal pathogens and processes the signal via the cleavage of the cytoplasmic adapter protein IMD. Subsequently, IMD drives the phosphorylation of the NF-κB homolog in the IMD pathway, Relish. Activated Relish translocates to the nucleus and controls the transcriptional activation of specific AMP genes. However, the nexus between the pathogen types that regulate the expression of specific sets of AMP genes remains obscure. (B) We propose that the knockdown of IMD transcripts prevents the activation and subsequent translocation of Relish to the nucleus, thereby downregulating the expression of nine AMP genes in the Tenebrio model system.
Materials and Methods
T. molitor and microbial strains
Larvae of the mealworm beetle, T. molitor, were reared in our laboratory with an artificial diet (4.4 g NeoVita, 0.5 g chloramphenicol, 0.4 g L-ascorbic acid, 0.5 g sorbic acid, 0.5 ml propionic acid, 2.2 g yeast extract, 2.2 g bean powder, 7.6 g agar, 4.4 g wheat powder, and 73 g wheat bran in 200 ml deionized water; autoclaved at 121 °C for 15 min) at 26 ± 1 °C, and 60% ± 5% relative humidity under dark conditions. In all experiments, only healthy 10–12th instar larvae were taken. Gram-positive bacteria, Staphylococcus aureus, strain RN4220 (also referred to as NRS144) were purchased from BEI Resources, NIAID, NIH, USA. Escherichia coli K12; Strain SMG 123 (PTA-7555) were procured from the American Type Culture Collection (ATCC). C. albicans was cultured in Sabouraud Dextrose broth, while overnight cultures of E. coli and S. aureus were cultured in Luria- Bertani (LB) broth at 37 °C. For the injection experiments, microorganisms were harvested and washed in phosphate-buffered saline (PBS; pH 7.0) by centrifugation at 3,500 g for 10 min. The washed samples were resuspended in PBS and concentrations at OD600 were measured. The final concentrations were adjusted to 106 cells/µl for E. coli and S. aureus and 5 × 104 cells/µl for C. albicans. The concentrations of the microorganisms have been validated in T. molitor larvae for immune challenge experiments as reported in previous studies32.
Identification of TmIMD and in silico analysis
The full-length TmIMD gene was identified by taking the T. castaneum IMD (TcIMD) amino acid sequence as the query (XP_008199405) and the T. molitor nucleotide database derived from the RNAseq/EST (unpublished) as the subject using local tblastn analysis. To further understand the genomic organization of TmIMD, we conducted local-blastn analysis using the obtained nucleotide sequence of TmIMD as the query and the T. molitor DNAseq database (unpublished) as the subject. Subsequently, the TmIMD ORF sequence was mapped in the genomic sequences using the eukaryotic gene-finding program FGENESH41. The conserved domains/motifs were predicted using InterProScan 5.042 and blastp programs. N-glycosylation and potential ubiquitination sites were predicted using the NetPhos 3.1 Server43 and UbPred44, respectively. The multiple alignment, percentage identity, and distance analysis of TmIMD amino acid and orthologous sequences from different orders were conducted using the ClustalX2 program45 and visualized using Genedoc software (http://iubio.bio.indiana.edu/soft/molbio/ibmpc/genedoc-readme.html). The phylogenetic tree was constructed based on the maximum likelihood tree method using the MEGA 7.0 program46.
Gene expression analysis and immune challenge studies
Reverse transcription quantitative PCR (RT-qPCR) was performed using the ExicyclerTM real-time Quantitative Thermal Block (Bioneer co., Daejon, South Korea) to analyze developmental and tissue distributions of TmIMD at the mRNA transcript level. In brief, total RNA was isolated (using 6 M Guanidine hydrochloride, 20 mM each of 0.5 M EDTA and 1 M 2-(N-morpholino) ethanesulfonic acid, and 40 µM phenol red) from different developmental stages, such as late larvae (10th–12th instar larvae; lengths of approximately 3 cm), pre-pupae, pupae (1–7 days old), and adults (1–2 days old). For tissue-specific expression profiles, total RNA was isolated from the larval tissues, such as integument, gut, fat body, Malpighian tubules, and hemocytes, and adult tissues, such as integument, gut, fat body, Malpighian tubules, hemocytes, ovary, and testes after disrupting the tissues with Precellys Evolution Homogenizer (Bertin Instruments, Montigny-le-Bretonneux, France) Notably, hemocytes were collected as pellets after centrifugation of the hemolymph at 5,000 × g for 4 min. For gene expression analysis, 15 insects were used (five insects were pooled together as one group). From each of the groups, tissue samples were collected to make three samples from each tissue.
For challenge experiments, healthy T. molitor larvae were microinjected intra-abdominally with 106 cells/µl of E. coli and 5 × 104 cells/µl of S. aureus, or C. albicans, respectively. PBS was used as a wounding control. Larvae (n = 3) were collected at 3, 6, 12, and 24 h postinjection (p.i) and total RNA was extracted at each time point using FavorPrepTM Tri-RNA reagent (Favorgen biotech corp., Ping-Tung, Taiwan). For cDNA synthesis, 2 µg total RNA was used with AccuPower RT Pre-Mix (Bioneer) and Oligo (dT)12–18 primer. The real-time PCR assay was performed at an initial denaturation of 95°C for 20 s, followed by 45 cycles of 95°C for 5 s, and 60°C for 20 s using AccuPower 2X GreenStar qPCR Master Mix (Bioneer). Ribosomal protein L27a (RpL27a) was used as an internal control for normalizing the expression results obtained in the experimental groups. Experiments were performed in triplicate and are presented as mean ± standard error of three biological replications.
TmIMD gene silencing
For TmIMD RNAi, a 363 bp PCR product was amplified by Ex TaqTM Polymerase (Takara, Japan) using specific primers containing the T7 promoter sequences at the 5′-ends. As a negative control, a 546 bp PCR product of the EGFP gene (derived from plasmid EGFP-C1) was used as a template for EGFP RNA synthesis. The primers for RNAi (Table 1) were designed using the SnapDragon-Long dsRNA design software available at DRSC/TRiP functional genomics resources (https://www.flyrnai.org/cgi-bin/RNAi_find_primers.pl) to minimize the off-target effects, if any. PCR was conducted at an initial denaturation of 98°C for 5 min, followed by 30 cycles of denaturation at 98°C for 10 s, annealing at 55°C for 30 s and extension at 72°C for 1 min. The purified PCR product was used to synthesize TmIMD double-strand RNA (dsRNA) using an AmpliScribe T7-Flash Transcription Kit (Epicentre Biotechnologies, Madison, WI, USA) following the manufacturer’s instructions. Briefly, the 20 µl reaction mixture was incubated at 42°C for 16 h, followed by 75°C for 5 min, and 25°C for 30 min. The dsRNA was purified using PCI (Phenol: Chloroform: Isoamyl alcohol), precipitated using 5 M ammonium acetate and ethanol, and resuspended in molecular biology grade water. The sample was measured spectrophotometrically at 260 nm and the integrity was determined by agarose gel electrophoresis. A total of 2 µg (in 1 µl) synthesized TmIMD dsRNA was injected intra-abdominally into the 10th–12th stage larvae of T. molitor. The dsEGFP was injected into the host using a disposable needle mounted onto a Picospritzer III micro-dispense system (Parker). The EGFP dsRNA was injected as a negative control in RNAi studies. For each gene, at least 30 last-instar larvae (10th–12th instar) were injected. The experiment was conducted in triplicate to confirm the silencing of TmIMD transcripts.
Survival analysis
As a measure of the mortality under TmIMD knockdown conditions, 106 cells/µl of E. coli and 5 × 104 cells/µl of S. aureus and C. albicans were injected to separate sets of larvae. The survival analysis was conducted with 10 insects for each set and results represented an average of three biological replicates. To monitor mortality, larvae that died during the first 3 h of microorganism challenge were discarded. Surviving larvae were incubated at 26 °C and scored daily for a period of 10 days.
AMP expression analysis
To understand the role of TmIMD transcripts in regulating the expression of 14 AMP genes, including TmTenecin1, TmTenecin2, TmTenecin3, TmTenecin4, TmDefensin1, TmDefensin2, TmColeoptericin1, TmColeoptericin-2, TmAttacin1a, TmAttacin1b, TmAttacin2, TmCecropin2, TmThaumatin-like protein1, and TmThaumatin-like protein2, microorganisms were injected into TmIMD-knockdown T. molitor larvae. A EGFP dsRNA was used as a negative control and PBS was used as an injection control. Samples at 12 h post injection of microorganisms were processed for total RNA analysis, cDNA synthesis, and qPCR analysis using AMP primers (Table 1).
Statistical analysis
For qPCR-based expression analysis of TmIMD transcripts, the 2−ΔΔCt method47 was used. Data were subjected to one-way analysis of variance (ANOVA) and Tukey’s multiple range tests to evaluate differences between groups (P < 0.05). For the survival analysis, a Kaplan-Meier plot (log-rank chi-square test) was generated using Real Statistics using Excel (http://www.real-statistics.com/survival-analysis/kaplan-meier-procedure/real-statistics-kaplan-meier/).
Conclusions
This study is the first to identify and functionally characterize the IMD homologue from the coleopteran pest T. molitor. We found that TmIMD is required for innate immune-mediated host defense of T. molitor larva against the Gram-negative bacteria E. coli and the fungus C. albicans. Furthermore, systemic E. coli infection in T. molitor larvae with TmIMD knockdown downregulates the production of a specific set of AMP genes that includes TmTenecin1, TmTenecin2, TmTenecin4, TmDefensin2, TmColeoptericin1, TmColeoptericin2, TmAttacin1a, TmAttain1b, and TmAttacin2. In future studies, it will be interesting to note the functional roles of downstream regulatory molecules in IMD signaling pathway, such as FADD, DREDD, IKKγ, TAB2/TAK1, and Relish, in the context of innate immune-mediated host defense against microorganisms, and to study the conservation of IMD pathways across insect species.
References
Stork, N. E., McBroom, J., Gely, C. & Hamilton, A. J. New approaches narrow global species estimates for beetles, insects, and terrestrial arthropods. Proc Natl Acad Sci USA 112, 7519–7523, https://doi.org/10.1073/pnas.1502408112 (2015).
Strand, M. R. The insect cellular immune response. Insect Sci 15, 1–14, https://doi.org/10.1111/j.1744-7917.2008.00183.x (2008).
Lee, K.-A. & Lee, W.-J. Immune-metabolic interactions during systemic and enteric infection in Drosophila. Curr Opin Insect Sci 29, 21–26, https://doi.org/10.1016/j.cois.2018.05.014 (2018).
Viljakainen, L. Evolutionary genetics of insect innate immunity. Brief Funct Genomics 14, 407–412, https://doi.org/10.1093/bfgp/elv002 (2015).
Engstrom, Y. et al. kappa B-like motifs regulate the induction of immune genes in Drosophila. J Mol Biol 232, 327–333, https://doi.org/10.1006/jmbi.1993.1392 (1993).
Dushay, M. S., Asling, B. & Hultmark, D. Origins of immunity: Relish, a compound Rel-like gene in the antibacterial defense of Drosophila. Proc Natl Acad Sci USA 93, 10343–10347, https://doi.org/10.1073/pnas.93.19.10343 (1996).
Belvin, M. P. & Anderson, K. V. A conserved signaling pathway: the Drosophila toll-dorsal pathway. Ann Rev Cell Dev Biol 12, 393–416, https://doi.org/10.1146/annurev.cellbio.12.1.393 (1996).
Valanne, S., Wang, J.-H. & Ramet, M. The Drosophila Toll Signaling Pathway. J Immunol 186, 649–656, https://doi.org/10.4049/jimmunol.1002302 (2011).
Kingsolver, M. B., Huang, Z. & Hardy, R. W. Insect antiviral innate immunity: pathways, effectors, and connections. J Mol Biol 425, 4921–4936, https://doi.org/10.1016/j.jmb.2013.10.006 (2013).
Myllmaki, H., Valanne, S. & Ramet, M. The Drosophila Imd Signaling Pathway. J Immunol 192, 3455–3462, https://doi.org/10.4049/jimmunol.1303309 (2014).
Naitza, S. et al. The Drosophila immune defense against Gram-negative infection requires the death protein dFADD. Immunity 17, 575–581, https://doi.org/10.1016/S1074-7613(02)00454-5 (2002).
Paquette, N. et al. Caspase-mediated cleavage, IAP-binding, and ubiquitination: linking three mechanisms crucial for Drosophila NF-kappaB signaling. Mol Cell 37, 172–182, https://doi.org/10.1016/j.molcel.2009.12.036 (2010).
Meinander, A. et al. Ubiquitylation of the initiator caspase DREDD is required for innate immune signaling. EMBO J 31, 2770–2783, https://doi.org/10.1038/emboj.2012.121 (2012).
Onfelt Tingvall, T., Roos, E. & Engstrom, Y. The imd gene is required for local Cecropin expression in Drosophila barrier epithelia. EMBO Rep 2, 239–243, https://doi.org/10.1093/embo-reports/kve048 (2001).
Ganesan, S., Aggarwal, K., Paquette, N. & Silverman, N. NF-Kb/Rel proteins and the humoral immune responses of Drosophila melanogaster. Curr Topics Microbiol Immunol 349, 25–60, 10.1007%2F82_2010_107 (2011).
Georgel, P. et al. Drosophila immune deficiency (IMD) is a death domain protein that activates antibacterial defense and can promote apoptosis. Dev Cell 1, 503–514, https://doi.org/10.1016/S1534-5807(01)00059-4 (2001).
Bang, K., Hwang, S., Lee, J. & Cho, S. Identification of immunity-related genes in the larvae of Protaetia brevitarsis seulensis (Coleoptera: Cetoniidae) by a Next-Generation Sequencing-Based Transcriptome Analysis. J Insect Sci 15, 142, https://doi.org/10.1093/jisesa/iev120 (2015).
Xia, X. et al. Genome-wide characterization and expression profiling of immune genes in the diamondback moth, Plutella xylostella (L.). Scientific Rep 5, 9877, https://doi.org/10.1038/srep09877 (2015).
Zhan, M. Y., Yang, P. J. & Rao, X. J. Molecular cloning and analysis of PGRP-L1 and IMD from silkworm Bombyx mori. Comp Biochem Physiol Part-B Biochem Mol Biol 215, 19–30, https://doi.org/10.1016/j.cbpb.2017.10.002 (2018).
Yokoi, K., Koyama, H., Minakuchi, C., Tanaka, T. & Miura, K. Antimicrobial peptide gene induction, involvement of Toll and IMD pathways and defense against bacteria in the red flour beetle, Tribolium castaneum. Results Immunol 2, 72–82, https://doi.org/10.1016/j.rinim.2012.03.002 (2012a).
Roh, K. B. et al. Proteolytic cascade for the activation of the insect Toll pathway induced by the fungal cell wall component. J Biol Chem 284, 19474–19481, https://doi.org/10.1074/jbc.M109.007419 (2009).
Yu, Y. et al. Diversity of innate immune recognition mechanism for bacterial polymeric meso-diaminopimelic acid-type peptidoglycan in insects. J Biol Chem 285, 32937–32945, https://doi.org/10.1074/jbc.M110.144014 (2010).
Patnaik, B. B. et al. Gene structure, cDNA characterization and RNAi-based functional analysis of a myeloid differentiation factor 88 homolog in Tenebrio molitor larvae exposed to Staphylococcus aureus infection. Dev Comp Immunol 46, 208–221, https://doi.org/10.1016/j.dci.2014.04.009 (2014).
Johnston, P. R., Makarova, O. & Rolff, J. Inducible defenses stay up late: Temporal patterns of immune gene expression in Tenebrio molitor. G3 (Bethesda) 4, 947–955, https://doi.org/10.1534/g3.113.008516 (2014).
Kersse, K., Verspurten, J., Vanden Berghe, T. & Vandenabeele, P. The death-fold superfamily of homotypic interaction motifs. Trend Biochem Sci 36, 541–552, https://doi.org/10.1016/j.tibs.2011.06.006 (2011).
Stoven, S. et al. Caspase-mediated processing of the Drosophila NF-kB factor Relish. Proc Natl Acad Sci USA 100, 5991–5996, https://doi.org/10.1073/pnas.1035902100 (2003).
Ribeiro, P. S. et al. DIAP2 functions as a mechanism-based regulator of drICE that contributes to the caspase activity threshold in living cells. J Cell Biol 179, 1467–1480, https://doi.org/10.1083/jcb.200706027 (2007).
Huh, J. R. et al. The Drosophila inhibitor of apoptosis (IAP) DIAP2 is dispensable for cell survival, required for the innate immune response to gram-negative bacterial infection, and can be negatively regulated by the reaper/hid/grim family of IAP-binding apoptosis inducers. J Biol Chem 282, 2056–2068, https://doi.org/10.1074/jbc.M608051200 (2007).
Shrestha, S. & Kim, Y. Activation of immune-associated phospholipase A2 is functionally linked to Toll/Imd signal pathways in the red flour beetle, Tribolium castaneum. Dev Comp Immunol 34, 530–537, https://doi.org/10.1016/j.dci.2009.12.013 (2010).
Zou, Z. et al. Comparative genomic analysis of the Tribolium immune system. Genome Biol 8, R177, https://doi.org/10.1186/gb-2007-8-8-r177 (2007).
Foley, E. & O’Farrell, P. H. Functional dissection of an innate immune response by a genome-wide RNAi screen. PLoS Biol 2, e203, https://doi.org/10.1371/journal.pbio.0020203 (2004).
Jo, Y. H. et al. TmCactin plays an important role in Gram-negative and –positive bacterial infection by regulating the expression of 7 AMP genes in Tenebrio molitor. Scientific Rep 7, 46459, https://doi.org/10.1038/srep46459 (2017).
Nehme, N. T. et al. A model of bacterial intestinal infections in Drosophila melanogaster. PLoS Pathog 3, e173, 10.1371/journal.ppat.0030173.
Ryu, J.-H., Kim, C.-H. & Yoon, J.-H. Innate immune responses of the airway epithelium. Molecules Cells 30, 173–183, https://doi.org/10.1007/s10059-010-0146-4 (2010).
Hori, A., Kurata, S. & Kuraishi, T. Unexpected role of the IMD pathway in Drosophila gut defense against Staphylococcus aureus. Biochem Biophys Res Commun 495, 395–400, https://doi.org/10.1016/j.bbrc.2017.11.004 (2018).
Yokoi, K. et al. Involvement of NF-Kb transcription factors in antimicrobial peptide gene induction in the red flour beetle, Tribolium castaneum. Dev Comp Immunol 38, 342–351, https://doi.org/10.1016/j.dci.2012.06.008 (2012b).
Cronin, S. J. et al. Genome-wide RNAi screen identifies genes involved in intestinal pathogenic bacterial infection. Science 325, 340–343, https://doi.org/10.1126/science.1173164 (2009).
Lemaitre, B. & Hoffmann, J. The host defense of Drosophila melanogaster. Ann. Rev Immunol 25, 697–743, https://doi.org/10.1146/annurev.immunol.25.022106.141615 (2007).
Busse, M. S., Arnold, C. P., Towb, P., Katrivesis, J. & Wasserman, S. A. A kB sequence code for pathway-specific innate immune responses. EMBO J 26, 3826–3835, https://doi.org/10.1038/sj.emboj.7601798 (2007).
Maistrou, S. et al. A constitutively expressed antifungal peptide protects Tenebrio molitor during a natural infection by the entomopathogenic fungus Beauveria bassiana. Dev Comp Immunol 86, 26–33, https://doi.org/10.1016/j.dci.2018.04.015 (2018).
Solovyev, V. et al. Automatic annotation of eukaryotic genes, pseudogenes and promoters. Genome Biol 7, S10, https://doi.org/10.1186/gb-2006-7-s1-s10 (2006).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240, https://doi.org/10.1093/bioinformatics/btu031 (2015).
Blom, N., Gammeltoft, S. & Brunak, S. Sequence and structure-based prediction of eukaryotic protein phosphorylation sites. J Mol Biol 294, 1351–1362, https://doi.org/10.1006/jmbi.1999.3310 (1999).
Radivojac, P. et al. Identification, analysis, and prediction of protein ubiquitination sites. Proteins 78, 365–380, https://doi.org/10.1002/prot.22555 (2010).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948, https://doi.org/10.1093/bioinformatics/btm404 (2007).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol Biol Evol 30, 2725–2729, https://doi.org/10.1093/molbev/mst197 (2013).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C (T)) Method. Methods 25, 402–408, https://doi.org/10.1006/meth.2001.1262 (2001).
Acknowledgements
Financial support was provided by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT and Future Planning (Grant No. 2018R1A2A2A05023367).
Author information
Authors and Affiliations
Contributions
Conception and experimental design: Yeon Soo Han and Yong Hun Jo. Performance of experiments: Yong Hun Jo, Ki Beom Park, Jihun Hwang. Bharat Bhusan Patnaik, Hye Jin Ko, Chang Eun Kim. Data analysis: Yong Hun Jo, Bharat Bhusan Patnaik, Woo Jin Jung, Young Min Bae. Manuscript writing: Bharat Bhusan Patnaik, Yong Hun Jo. English language editing and data analysis: Yong Seok Lee.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jo, Y.H., Patnaik, B.B., Hwang, J. et al. Regulation of the expression of nine antimicrobial peptide genes by TmIMD confers resistance against Gram-negative bacteria. Sci Rep 9, 10138 (2019). https://doi.org/10.1038/s41598-019-46222-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-46222-8
- Springer Nature Limited
This article is cited by
-
Transcriptome analysis of Corvus splendens reveals a repertoire of antimicrobial peptides
Scientific Reports (2023)
-
Transcriptome analysis of the endangered dung beetle Copris tripartitus (Coleoptera: Scarabaeidae) and characterization of genes associated to immunity, growth, and reproduction
BMC Genomics (2023)
-
The holobiont transcriptome of teneral tsetse fly species of varying vector competence
BMC Genomics (2021)
-
TmRelish is required for regulating the antimicrobial responses to Escherichia coli and Staphylococcus aureus in Tenebrio molitor
Scientific Reports (2020)