Abstract
Polycephalomyces (Ophiocordycipitaceae) species are found in subtropical regions and are parasitic or hyperparasitic on insects. Two new species, P. aurantiacus and P. marginaliradians, parasitic on Ophiocordyceps barnesii and larva of Cossidae respectively, are introduced in this paper. Morphological comparison with extant species and DNA based phylogenies from analyses of a multigene dataset support the establishment of the new taxa. Polycephalomyces aurantiacus, exhibiting a hyperparasitic lifestyle on Ophiocordyceps barnesii, differs from other species in producing orange conidia in mass and have longer β-phialides in culture. Polycephalomyces marginaliradians differs from other Ophiocordyceps species by producing single stromata with a stipe, smaller perithecia and branched α-phialides and catenate α-conidia and is parasitic on Cossidae. A combined nrSSU, nrLSU, ITS, tef-1a, rpb1 and rpb2 sequence data was analysed phylogenetically including Ophiocordyceps and Polycephalomyces taxa. The new species described herein are clearly distinct from other species in Polycephalomyces. We provide a key to the species of Polycephalomyces and discuss relevant interspecies relationships.
Similar content being viewed by others
Introduction
The genus Polycephalomyces was introduced by Kobayasi1 to accommodate P. formosus Kobayasi (1941), based on its asexual characteristics2 and it is presently accommodated in Ophiocordycipitaceae3. Phylogenetic placement of Polycephalomyces has always been a debate within the clavicipitoid fungi as the taxonomic hypotheses based on host substrate and sexual morph affinities were controversial1,4,5. Kepler et al.5 amended the taxonomic circumscription of Polycephalomyces and accepted twelve species (i.e. P. cuboideus, P. cylindrosporus, P. ditmarii, P. formosus, P. kanzashiznus, P. nipponicus, P. paracubiodeus, P. prolificus, P. ramosopulvinatus, P. ramosus, P. ryogamiensis and P. tomentosus) in Ophiocordycipitaceae based on phylogenetic analyses. Later, P. sinensis6, P. lianzhouensis7, P. yunnanensis8, P. agaricus9 and P. onorei10 were introduced as new species within Polycephalomyces based on morphology and DNA sequence data. Then, Liang et al.11 introduced a new species P. ponerae in the genus Polycephalomyces. Based on recent morphological studies and DNA based phylogenetic analyses, Polycephalomyces taxa have been segregated in two sister clades within Ophiocordycipitaceae5,12. Matočec et al.12 considered those two clades as two different genera, viz. Perennicordyceps Matočec & I. Kušan and Polycephalomyces. Perennicordyceps comprises four species (i.e. Pe. cuboidea, Pe. paracuboidea, Pe. prolifica, and Pe. ryogamiensis), which are characterized by superficial perithecia, and hirsutella-like or acremonium-like asexual morphs12,13. Polycephalomyces comprised eight species12. Maharachchikumbura et al.14,15 and Wijayawardene et al.3 maintained Polycephalomyces within the Ophiocordycipitaceae.
The sexual morph of Polycephalomyces has been recorded as fertile, capitulate, globose, tuberiform to pulvinate stromata and immersed, elongated pyriform perithecia12, while the asexual morph has branched or unbranched synnemata, ending up with clavate to spherically flared, hymeniform aggregations of conidiophores, which produce large masses of conidia united in collective globular mucus12. There is only one species (P. lianzhouensis) reported with both sexual and asexual morphs. Ophiocordyceps fungi are ecologically important host species for Polycephalomyces, and to date five Ophiocordyceps species have been reported to be associated with Polycephalomyces6,8,9,16,17,18. At the Engineering Research Center of Southwest Bio-Pharmaceutical Resources (Guizhou University, China) in collaboration with the Center of Excellence in Fungal Research (Mae Fah Luang University, Thailand), we are investigating diversity of microfungi associated with insects in the tropics and clarify their taxonomy based on morphology and multigene phylogeny14,15,19,20,21,22,23,24.
Results
Molecular phylogeny
Table S1 comprises 39 taxa (including the seven newly collected taxa) analysed herein and their accession numbers. DNA sequence data of the new species have been submitted to GenBank. A concatenated sequence data-set was analyzed comprising 5003 characters with gaps (SSU: 971, LSU: 813, ITS: 667, TEF: 875, RPB1: 659, RPB2: 1018).
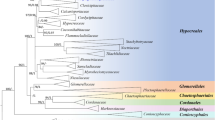
The RAxML analysis of the combined dataset yielded a best scoring tree (Fig. 1) with a final ML optimization likelihood value of −23008.064357. The matrix had 1750 distinct alignment patterns, with 39.76% of undetermined characters or gaps. Parameters for the GTR model of the concatenated data set were as follows: Estimated base frequencies; A = 0.236157, C = 0.276387, G = 0.278052, T = 0.209403; substitution rates AC = 1.217462, AG = 3.345152, AT = 0.776221, CG = 1.574418, CT = 6.177706, GT = 1.000; gamma distribution shape parameter α = 0.251124. The Bayesian analysis resulted in 20001 trees after 2000000 generations. The first 4000 trees, representing the burn-in phase of the analyses, were discarded, while the remaining 16001 trees were used for calculating posterior probabilities in the majority rule consensus tree.
Phylogram of Polycephalomyces aurantiacus, Polycephalomyces marginaliradians and Ophiocordyceps barnesii generated from maximum likelihood analysis of ITS, SSU, LSU, RPB1, RPB2 and TEF1α sequence data. Purpureocillium lilacinum CBS 284.36 and Purpureocillium lilacinum CBS 431.87 were used as outgroup taxon. Maximum likelihood bootstrap values greater than 70% and Bayesian posterior probabilities over 0.9 are indicated above the nodes. The new species were indicated in blue. The host of Polycephalomyces aurantiacus is indicated in bold.
The genus Polycephalomyces currently includes 15 species and only 11 species have available DNA sequence data in GenBank (Table S1), excluding the new taxa described in this study. Our multigene phylogenetic analyses herein reveal that our new taxa constitute a strongly supported monophyletic subclade and nested in between other Polycephalomyces species (Fig. 1). In particular it is noted that Polycephalomyces aurantiacus and P. marginaliradians share a close phylogenetic affinity to P. nipponicus and P. kanzashianus (Fig. 1).
In this paper, we illustrate a collection of Ophiocordyceps barnesii, which was parasitized by a Polycephalomyces species. Two new species of Polycephalomyces, one from Ophiocordyceps barnesii, and one from a Cossidae host are also introduced. A phylogenetic tree based on multigene sequence analyses for Ophiocordyceps (11 species) and Polycephalomyces (13 species), is also provided.
Description of Ophiocordyceps barnesii (Thwaites) G.H. Sung, J.M. Sung, Hywel-Jones & Spatafora, in Sung, Hywel-Jones, Sung, Luangsa-ard, Shrestha & Spatafora, Stud. Mycol. 57: 40 (2007)
Index Fungorum number: 504230; Facesoffungi number: FoF 03810 (Fig. 2).
Ophiocordyceps barnesii MFLU 17-1393 (host). (a) Habitat. (b) Overview of the host and stromata. (c) Host. (d) Stroma. (e) Cross section of stroma. (f) Cross sections showing the immersed perithecia. (g) Perithecia. (h–k) Asci. (l–o) Secondary ascospores. Scale Bars: b = 5 cm, c = 5 mm, d = 2 cm, e, f = 500 µm, p = 200 µm, g–k = 100 µm, l–o, s = 20 µm, q = 10 µm, r = 5 µm.
Parasitic on larvae (Coleopteran), buried in the soil. Sexual morph: Host 2–2.5 long × 0.5–1 cm wide, brown to dark brown without hyphae on the surface. Stromata 13–20 long × 0.5–1 cm diam., mostly single, stipitate, unbranched or branched into 2 or 3 fertile head, arising from between the head and thorax of larva (fusiformis), dark-brown, fleshy, cylindrical, often flexuous or angularly crooked. Stipe 1–2 cm long, 2–3 mm diam., brown, with a fertile apex. Fertile head 2–5 cm long, 1–3.5 mm diam., single or branched more than 2, cylindrical, apically tapered, brown with orange mycelium cover on the surface. Perithecia 308–389 × 98–132 μm (\(\bar{x}\) = 349 × 115 µm, n = 60), immersed, brown, elongated pyriform or flask-shaped, thick-walled. Peridium 12–21 µm (\(\bar{x}\) = 16 µm, n = 60) wide, brown, textura angularis to textura globulosa to textura prismatica. Asci 195–229 × 6–9 μm (\(\bar{x}\) = 212 × 7.5 µm, n = 90), 8-spored, hyaline, filiform, with a thin apex. Apical cap 3.1–4.2 × 4.1–5.4 μm (\(\bar{x}\) = 3.6 × 4.7 µm, n = 60), with a small channel in the center. Ascospores 155–200 × 2.2–2.7 μm (\(\bar{x}\) = 178 × 2.5 µm, n = 60), 3-septate, easily breaking into 4 part-spores, filiform, tapered at each end. Secondary ascospores 31.6–41.6 × 2.2–2.7 μm (\(\bar{x}\) = 36.6 × 2.5 µm, n = 90) cylindrical, thickening at each end or tapered at one end, straight, hyaline, smooth-walled. Asexual morph: undetermined.
Notes: We collected Ophiocordyceps barnesii in this study which was colonized by an orange hyperparasite which we introduce below as Polycephalomyces aurantiacus. This species may be important in future industrial production of Cordyceps species, which are increasingly being produced because of their medicinal properties and biopesticides potential25,26. The specimen was deposited in MFLU Herbarium (MFLU 17-1393).
Description of Polycephalomyces aurantiacus Y.P. Xiao, T.C. Wen & K.D. Hyde, sp. nov
Index Fungorum number: IF553936; Facesoffungi number: FoF 03811 (Figs 3, 4).
Etymology: The specific epithet refers to the color of conidia in mass in the specimen and colony.
Holotype: MFLU 17-1393
Hyperparasite on Ophiocordyceps barnesii (Ophiocordycipitaceae), buried in the soil. Sexual morph: undetermined. Asexual morph: Synnemata solitary or not solitary, arising from the fertile head of the stromata, flat-shaped, orange color. Phialides 9.9–14.3 × 0.7–1.4 μm (\(\bar{x}\) = 12.1 × 1.1 µm, n = 90) hyaline. Conidia 2–2.6 × 1.4–2.1 μm (\(\bar{x}\) = 2.3 × 1.8 µm, n = 90), oval to globose shape, hyaline, one-celled, smooth-walled, orange in mass.
Colonies on PDA medium, growing slowly, attaining 4 cm in 17 days at 25 °C, white, reverse yellow. Synnemata emerging after 30 days, solitary or not solitary, branched or unbranched, 1.3–2.2 μm long (\(\bar{x}\) = 60), showing 1–2 radiating ring-like distributions. Conidial masses generating from the apex of the synnemata or covering the surface of the colony (Fig. 4). Hyphae hyaline, branched, smooth-walled, 0.3–50 mm (\(\bar{x}\) = 20) wide. Conidiophores undetermined, not clear. Phialides existing in two types: α-and β-phialides. α-phialides 10.4–18.3 × 0.8–1.8 μm (\(\bar{x}\) = 14.4 × 1.3 µm, n = 90) hyaline, narrow slender, smooth. β-phialides 22.9–64.2 × 1–1.5 μm (\(\bar{x}\) = 43.6 × 1.3 µm, n = 90) solitary, growing from hyphae, narrow slender, catenateblasto conidia, smooth. α-conidia 1.8–2.2 × 1.4–1.9 μm (\(\bar{x}\) = 2 × 1.7 µm, n = 90) globose to subglobose, occurring in the conidial mass on the agar or on the final portion of synnemata, one-celled, smooth-walled, yellow slimy in mass. β-conidia 3.2–3.9 × 1.4–1.8 μm (\(\bar{x}\) = 3.5 × 1.6 µm, n = 90) fusiform, and produced on the surface mycelium of colony or on the top of the synnemata, one-celled, smooth-walled, hyaline, usually in chains on a phialide.
Polycephalomyces aurantiacus MFLUCC 17-2113. (a–c) Upper side of the culture. (d–f) Reverse side of the culture. (g,o) β-phialides. (h–l) Synnemata growing on PDA medium. (m) β-phialides with hyphae. (n) α-conidia. (p) β-conidia. Scale Bars: h = 1000 µm, i = 2000 µm, j = 200 µm, l = 500 µm, g, k, m = 50 µm, o = 20 µm, n, p = 5 µm.
Material examined: THAILAND, Prachuap Khiri Khan. On dead larvae (Coleopteran), 29 July 2015, YuanPin Xiao, BK15072907 (MFLU 17-1393, holotype); BK15072902, BK15072906 (MFLU 17-1394, HKAS100693, paratypes); ex-type living cultures, MFLUCC 17-2113, MFLUCC 17-2114, MFLUCC 17-2115, KUMCC 17-0256, KUMCC 17-0257.
Description of Polycephalomyces marginaliradians Y.P. Xiao, T.C. Wen & K.D. Hyde, sp. nov
Index Fungorum number: IF553937; Facesoffungi number: FoF 03812 (Figs 5, 6).
Polycephalomyces marginaliradians MFLU 17-1582. (a) Habitat. (b,d) Overview of the host and stromata. (c) Part of the stroma. (e,f) Stroma. (g) Cross sections showing the immersed perithecia. (h) Perithecia. (j–l) Asci. (m) Part of the ascospores. (n) Apical cap. (o) Secondary ascospores. (p,q) Synnemata. (r) Phialide. (s) Conidia. Scale Bars: b–d = 1 cm, e, f = 2000 µm, g = 1000 µm, h = 500 µm, p, q = 200 µm, j–l = 100 µm, i = 50 µm, m, r = 10 µm, n, o, s = 5 µm.
Polycephalomyces marginaliradians MFLUCC 17-2276. (a) Upper side of the culture. (b) Reverse side of the culture. (c,d) Synnemata growing on PDA medium. (e) Synnemata. (f,h) α-phialides. (g,j) β-phialides with hyphae. (i) α-conidia. (k) β-conidia. Scale Bars: c, d = 5000 µm, e = 1000 µm, f, g, j = 20 µm, h = 10 µm, i, k = 5 µm.
Etymology: The specific epithet refers to the feature of the colonies on the culture.
Holotype: MFLU 17-1582
Parasitic on a Cossidae larva (Lepidoptera), buried in the soil. Sexual morph: Thallus within host. Host 3.2–3.5 long × 0.4–0.6 cm wide, yellow to brown, without hyphae on the surface. Stromata 3–3.5 long × 0.2–0.45 cm diam., mostly single, stipitate, cylindrical, unbranched or branched, arising from the head of larva, brown to yellow. Stipe 1–2 cm long, 2–3 mm diam., cylindrical, yellow to brown, with one or two lateral fertile head. Fertile head 0.40–0.42 cm long, 0.3–0.45 mm diam., capitate, lateral, globose to subglobose, pale yellow to yellow, with protruding ostiolar necks. Ascomata 676–803 × 246–328 μm (\(\bar{x}\) = 739 × 287 µm, n = 60), immersed, yellow, flask-shaped, thick-walled. Peridium 11–19 µm (\(\bar{x}\) = 15 µm, n = 60) wide, brown, textura angularis to textura globulosa to textura prismatica. Asci 459–556 × 3.1–4.3μm (\(\bar{x}\) = 508 × 3.7 µm, n = 90), 8-spored, hyaline, filiform, with a thin apex. Apical cap 1.4–2.5 × 2.2–3.2 μm (\(\bar{x}\) = 2 × 2.7 µm, n = 60), with a small channel in the center. Ascospores as long as the asci, easily breaking into part-spores, filiform. Secondary ascospores 3.2–4.2 × 1.3–1.7 μm (\(\bar{x}\) = 3.8 × 1.5 µm, n = 90) cylindrical, straight, hyaline, smooth. Asexual morph: Synnemata solitary or not solitary, arising from the fertile head of the host, cylindrical, pale yellow. Phialides 11–14.4 × 1.2–1.8 μm (\(\bar{x}\) = 12.7 × 1.5 µm, n = 90) hyaline. Conidia 3.6–4.9 × 1.8–2.5 μm (\(\bar{x}\) = 4.2 × 2.1 µm, n = 90), fusiform, hyaline, one-celled, smooth-walled.
Colonies on PDA medium, circular, attaining 4 cm in 10 days at 25 °C, white, reverse yellow. Synnemata emerging after 14 days in the margin of the colony, single or branched into 2 or 3 branched, 3200.8–4566.3 × 142.9–661.8 μm (\(\bar{x}\) = 3883.5 × 402.3 µm, n = 30), showing 1–2 radiating ring-like distributions. Conidial masses generating from the middle of the synnemata or covering the surface of the colony, pale yellow to yellow, with hyaline to pale yellow exucate. Hyphae hyaline, branched, smooth-walled, 1.8–2.7 μm (\(\bar{x}\) = 2.2) wide. Conidiophores undetermined, not clear. Phialides existing in two types: α-and β-phialides. α-phialides 11–14.4 × 1.2–1.8 μm (\(\bar{x}\) = 12.7 × 1.5 µm, n = 90), hyaline, smooth, elongated lageniform,caespitose, palisade-like, crowed, monoverticillate, mostly branched into 2 phialides, 3 branched on one metula. β-phialides 12.8–23.9 × 1.8–2.7 μm (\(\bar{x}\) = 18.3 × 2.2 µm, n = 90), hyaline, smooth, solitary, growing from hyphae, narrow slender to narrow lageniform, with or without metula at the base. α-conidia 1.9–2.6 μm (\(\bar{x}\) = 2.3 µm, n = 90) diam, globose, catenate, occurring in the conidial mass on the middle of synnemata, one-celled, smooth-walled, pale yellow slimy in mass. β-conidia 3.1–3.9 × 1.6–2.1 μm (\(\bar{x}\) = 3.5 × 1.8 µm, n = 90) fusiform, and produced on the surface mycelium of colony or on the branch of the synnemata, one-celled, smooth-walled, hyaline.
Material examined: THAILAND, Chiang Mai, The Mushroom Research Center. On dead Cossidae larvae (Lepidoptera), 11 June 2017, Yuan Pin Xiao, MRC170611 (MFLU 17-1582, holotype); CM48 (MFLU 17-1583 MFLU 17-1584, HKAS100694, paratypes); ex-type living cultures, MFLUCC 17-2276, MFLUCC 17-2277, MFLUCC 17-2278, KUMCC 17-0258, KUMCC 17-0259.
Discussion
Studies based on morphology and DNA sequence analyses have provided insights into the phylogeny of Polycephalomyces to resolve generic delimitation. Species of this genus are commonly known to exhibit a parasitic mode of life on insects and other fungi2,5,12. Our fungal diversity studies on entomophagous fungi have led to the discovery of two species, new to science, which we accommodate in Polycephalomyces. Molecular data also reveals that our new genus belongs to the family Ophiocordycipitaceae as circumscribed by Matočec et al.12. Species which exist in their sexual state display morphs such as fertile, capitulate, globose, tuberiform to pulvinate stromata and immersed, elongated pyriform perithecia while the asexual morphs occur as branched or unbranched synnemata, ending up with clavate to spherically flared, hymeniform aggregations of conidiophores, and produce large masses of conidia united in collective globular mucus12. To date, six species, including P. ramosus16,17, P. sinensis6,18, and P. agaricus9 are considered as parasites of entomogenous fungi, while six species are recorded as entomogenous6,7,9,11. Some species such as P. lianzhouensis and P. yunnanensis colonise both entomogenous fungi and insects7,8. Because of their economic importance, species of this genus have been the subject for various research. The most recent new species introduced is P. yunnanensis and multigene phylogeny reveals a close relationship to P. formosus, P. ramosopulvinatus and P. sinensis based on 5-loci (nrSSU, nrLSU, tef-1α, rpb1 and rpb2) phylogenetic analyses8.
Our taxonomic investigations herein reveal two new species of Polycephalomyces, P. aurantiacus and P. marginaliradians. Our morphological examination suggests that both species fit clearly within the generic concept of Polycephalomyces and both species produce two types of conidia. However, they exhibit different mode of life and there are sufficent morphological differences that can justify their segregation into two species. These two new species similar to P. agaricus, P. formosus, P. ponerae, P. sinensis, P. ramosus, and P. yunnanensis have produce two types of conidia, while P. ditmarii, P. lianzhouensis, P. paludosus and P. tomentosus have only one type of conidia. Polycephalomyces aurantiacus and P. marginaliradians have two types of phialides, while P. formosus and P. sinensis have only one type of phialide. Polycephalomyces ponerae also differs from P. aurantiacus and P. marginaliradians by producing Akanthomyces-like β-phialides and parasitic on ant (Ponera Latreille). Polycephalomyces agaricus differs from P. aurantiacus and P. marginaliradians by producting agaric shaped synnemata and parasitic on Ophiocordyceps sp. Polycephalomyces yunnanensis is distinct from P. aurantiacus and P. marginaliradians as it produces longer α-conidia fusiform, catenate or clump together β-conidia and parasitic on O. nutans. Polycephalomyces ramosus differs from P. aurantiacus and P. marginaliradians in having longer synnemata, shorter β-phialides and parasitic on Hirsutella guignardii. Polycephalomyces aurantiacus is distinct from P. marginaliradians as it is parasitic on the fungus Ophiocordyceps barnesii and produces longer synnemata, shorter β-phialides and catenate α-conidia, wheras P. marginaliradians is parasitic on insect and produces shorter synnemata, longer β-phialides and catenate β-conidia (further morphological differences are outlined in Table S2).
Phylogeny based on our concatenated dataset recovered also support that our two new species belong to Polycephalomyces and are distinct from each other (Fig. 1). A close relationship is observed between the two species, but both constitute independent and strongly supported monophyletic subclades indicative of two phylogenetically distinct speces. To further compare our two species, we delved in pairwise nucleotide sequence comparison and noted sufficient differences to justify them as independent taxa27. ITS pairwise nucleotide sequence comparison between P. aurantiacus and P. marginaliradians revealed striking differences in 15 base pairs that justifies that both are different from each other and hence can be considered as two distinct species. There are also 6, 21, 4, 15 and 13 differences in the nrSSU, nrLSU, tef-1α, rpb1, and rpb2 DNA sequence data respectively. Two species not considered in our phylogenetic sampling are P. ditmarii and P. paludosus due to the unavailability of sequence data. However these two are different from our new species with respect to one type of conidia occurring on its natural substrate and under cultural conditions28,29. The hosts from which our new species have been recovered are also different. Polycephalomyces ponerae was not considered in our phylogenetic sampling as the DNA (ITS) sequence is too short, ambiguous and did not align well with other species. However, Polycephalomyces ponerae is morphologically different from our new species with respect to Akanthomyces-like β-phialides and parasitic on ant (Ponera Latreille). Further morphological differences among species are detailed in Tables S2 and S3.
Our multigene phylogeny derived herein also provides robust and well-resolved intergeneric relationships between Polycephalomyces and Ophiocordyceps. Members of both genera are clearly distinct from each other and we managed to successfully identify and sequence Ophiocordyceps barnesii, the host from which P. aurantiacus was isolated. Further interspecies taxonomic relationships are also elucidated in our molecular phylogeny. All Polycephalomyces species currently analysed constitute a strongly supported monophyletic lineage (Fig. 1), which corroborates with previous taxonomic schemes5,12. In particular, a robust relationship in observed between P. onorei and P. agaricus sharing P. yunnanensis as sister taxa. These three species are also markedly different in terms of morphological characters. Polycephalomyces yunnanensis is clearly distinct from P. onorei and P. agaricus in terms of being parasitic on Ophiocordyceps nutans (Pat.) G.H. Sung, J.M. Sung, Hywel-Jones & Spatafora, longer synnemata, cylindrical to subulate α-phialides and subglobose or ellipsoidal α-conidia8,10. Polycephalomyces ramosopulvinatus is closely related to P. lianzhouensis, but each species is positioned in different well-supported subclades and hence merit species status. Polycephalomyces ramosopulvinatus is also different from P. lianzhouensis with respect to being parasitic on nymph of Cicada and characterised by a long stipe and pseudo-immersed, pyriform perithecia. While phylogeny resolves our new species into well-segregated subclades, we note that relationships of P. formosus, P. tomentosus, P. ramosus and P. sinensis are still obscure and the concatenated dataset used herein did not provide adequate species resolution. A similar phylogenetic scenario is observed for P. nipponicus and P. kazanshianus. Whether these species are conspecific warrants further taxonomic investigations. The latter two species do share some morphological resemblances to P. marginaliradians especially with respect to the yellow cylindrical stipe with capitate lateral fertile part (known from their sexual morph). However, P. marginaliradians differs in having a capitate stromata with stipe, smaller perithecia and parasitic on Cossidae, while P. nipponicus and P. kanzashianus have polycephalous stromata and parasitic on Cicadidae. Meanwhile, P. onorei and P. ramosopulvinatus are distinct from P. marginaliradians by producing bigger perithecia and parasitic on caterpillar (Arctinae) and nymph of Cicada respectively. Phylogenies retrieved herein also support them as separate taxonomic entities.
Key to the species of the genus Polycephalomyces
1. Synnemata arising from fungi or insect or culture…………………………2
1. Stromata arising from insect…………………………13
2. Two types of conidia absent in nature or culture…………………………3
2. One type of conidia absent in nature and culture…………………………10
3. Synnemata agaric-shaped…………………………P. agaricus
3. Synnemata other shaped…………………………4
4. Two types of phialides absent…………………………5
4. Only one type of phialides absent…………………………9
5. α-conidia globose to subglobose (1.4–3.2 × 1.2–2.2) μm…………………………6
5. α-conidia ovoid (2.4–3.2 × 1.6–2.4) μm…………………………P. ramosus
6. β-phialides Akanthomyces-like, inflated at base, slenderneck at top…………………………P. ponerae
6. β-phialides lanceolate or narrowly lageniform or subulate…………………………7
7. β-phialides lanceolate or narrowly lageniform, 22.9–64.2 μm length…………………………P. aurantiacus
7. β-phialides narrowly lageniform or subulate, 7–30 μm length…………………………8
8.α-conidia subglobose, not catenate, β-conidia fusiform, catenate…………………………P. marginaliradians
8. α-conidia globose, catenate, β-conidia fusiform, not catenate…………………………P. yunnanensis
9. Phialides lanceolate or narrowly lageniform, 12.5–66 μm length…………………………P. sinensis
9. Phialides cylindrical, subulate, 10–15 μm length…………………………P. formosus
10. Host is insect…………………………11
10. Host is myxomycetes…………………………P. tomentosus
11. Conidia globose to subglobose or cylindrical…………………………12
11. Conidia obovoid, covered by a mucus, agglutinating…………………………P. paludosus
12. Conidia globose to subglobose, 2.2–3.4 × 1.3–1.6um…………………………P. ditmarii
12. Conidia subglobose to cylindrical, 5–7 × 1.3–1.6um…………………………P. lianzhouensis
13. Host is Cicada or nymph of Cicada…………………………14
13. Host is neither Cicada nor nymph of Cicada…………………………16
14. Stipe less than 80 mm…………………………15
14. Stipe more than 90 mm…………………………P. ramosopulvinatus
15. Perithecia flask-shaped, 900–1050 × 270–300 um.…………………………P. kanzashianus
15. Perithecia flask-shaped or ovioid, 800–950 × 300–370 um…………………………P. nipponicus
16. Stromata numerous…………………………17
16. Stromata single…………………………P. marginaliradians
17. Fertile part narrowly ovoid 355–473 × 158–197 μm…………………………P. lianzhouensis
17. Fertile part pyriform, 854–950 × 330–395 um…………………………P. onorei
Materials and Methods
Collection, isolation, and morphology study
Four fresh specimens were collected from southern Thailand (Prachuap Khiri Khan Province), and two from northern Thailand in the soil. The specimens were noted and photographed in the field and transported to the laboratory individually in plastic boxes and stored at 4 °C until examined. Strains were isolated from single spore isolation from both stomata and synnemata following the protocol described in Chomnunti et al.30.Cultures were incubated at 18 °C for 14–25 days on potato extract agar (PDA) as outlined by Vijaykrishna et al.31. Herbarium material is deposited at MFLU herbarium and HKAS herbarium and Facesoffungi numbers and Index Fungorum numbers are provided as in Jayasiri et al.32 and Index Fungorum33. New species are based on recommendations outlined by Jeewon & Hyde27.
DNA extraction, PCR amplification and determination of DNA sequences
DNA was extracted from both dried specimens and cultures by using E.Z.N.A.TM Fungal DNA MiniKit (Omega Biotech, CA, USA) according to the manufacturer’s protocols. The primers used in PCR amplification were (Table S4); ITS4/ITS5 for internal transcribed spacer gene region (ITS)34, NS1/NS4 for partial small subunit ribosomal RNA gene region (SSU)34, LROR/LR5 for partial large subunit rDNA gene region (LSU)35. 983 F/2218 R for partial translation elongation factor 1-alpha gene region (TEF-1α)36, CRPB1A/RPB1Cr for partial RNA polymerase II largest subunit gene region (RPB1)37. RPB2-5F/RPB2-5R for partial RNA polymerase II second largest subunit gene region (RPB2)37. PCR amplifications were conducted as outlined by Jeewon et al.38,39 and PCR products were sequenced by GenScript Biotechnology Co., Nanjing, China.
Phylogenetic analyses
All reference sequences were obtained from GenBank based on previously published data (Table S1). MAFFT v.740 (http://mafft.cbrc.jp/alignment/server/) was used to align combined datasets of ITS, SSU, LSU, TEF1α and RPB1. BioEdit41 was used to check alignment manually. Gaps were treated as missing data. Two strains of Perennicordyceps prolifica (Kobayasi) Matočec & I. Kušan, in Matočec et al.12 were selected as outgroup taxa.
ML trees were estimated by using the software RAxML 7.2.8 Black Box42,43 in the CIPRES Science Gateway platform44. MrModeltest v.2.345 was used to determine the best-fit model of evolution for Bayesian analyses. MrBayes v.3.1.246 was used to evaluate posterior probabilities (PP)47,48 by Markov Chain Monte Carlo sampling (BMCMC). Six simultaneous Markov chains were run for 2,000,000 generations and trees were sampled every 100th generation and 20,000 trees were obtained. The first 20% of trees were discarded, which representing the burn-in phase of the analyses, while the remaining trees were used for calculation posterior probabilities in the majority rule consensus tree (critical values for the topological convergence diagnostic is 0.01). Pylogenetic trees were also constructed based on parsimony analyses as detailed by Cai et al.49 and Jeewon et al.50,51. Trees were figured in FigTree v1.4.0 program52. Bayesian Posterior Probabilities (BYPP) equal to or great than 0.95 were given53,54,55,56,57,58,59,60,61,62,63 below each node (Fig. 1).
References
Kobayasi, Y. The genus Cordyceps and its allies. Sci Rep Tokyo BunrikaDaigaku, Sec B. 84, 53–260 (1941).
Wijayawardene et al. Notes for genera: Ascomycota. Fungal Diversity. 86, 1–594 (2017).
Wijayawardene et al. Outline of Ascomycota – 2017. Fungal Diversity. 88, 167–263 (2018).
Massee, G. A revision of the genus Cordyceps. Annals bot-london. 9, 1–44 (1895).
Kepler, R. et al. The phylogenetic placement of hypocrealean insect pathogens in the genus Polycephalomyces: an application of One Fungus One Name. Fungal biol-uk. 117, 611–622 (2013).
Wang, W. J. et al. Molecular and morphological studies of Paecilomyces sinensis reveal a new clade in clavicipitaceous fungi and its new systematic position. Syst Biodivers. 10, 221–232 (2012).
Wang, L., Li, H. H., Chen, Y. Q., Zhang, W. M. & Qu, L. H. Polycephalomyces lianzhouensis sp. nov., a new species, co-occurs with Ophiocordyceps crinalis. Mycol prog. 13, 1089–1096 (2014).
Wang, Y. B. et al. Polycephalomyces yunnanensis (Hypocreales), a new species of Polycephalomyces parasitizing Ophiocordyceps nutans and stink bugs (hemipteran adults). Phytotaxa. 208, 34–44 (2015).
Wang, Y. B. et al. Polycephalomyces agaricus, a new hyperparasite of Ophiocordyceps sp. infecting melolonthid larvae in southwestern China. Mycol Prog. 14, 70 (2015).
Crous, P. W. et al. Fungal Planet description sheets: 558–624. Persoonia. 38, 240–384 (2017).
Liang, Z., Chen, W., Liang, J., Han, Y. & Zou, X. Phenotypic polymorphism of the synnematous entomogenous fungi in an ant nest of Ponera I. Mycosystema 35, 906–917 (2016).
Matočec, N., Kušan, I. & Ozimec, R. The genus Polycephalomyces (Hypocreales) in the frame of monitoring Veternica cave (Croatia) with a new segregate genus Perennicordyceps. Ascomycete. org. 6, 125–133 (2014).
Ban, S., Sakane, T., Toyama, K. & Nakagiri, A. Teleomorph–anamorph relationships and reclassification of Cordyceps cuboidea and its allied species. Mycoscience 50, 261–272 (2009).
Maharachchikumbura, S. S. N. et al. Towards a natural classification and backbone tree for Sordariomycetes. Fungal Divers. 72, 199–301 (2015).
Maharachchikumbura, S. S. N. et al. Families of Sordariomycetes. Fungal Divers. 79, 1–317 (2016).
Seifert, K. A monograph of Stilbella and some allied Hyphomycetes. Stud in Mycol. 27, 1–224 (1985).
Bischoff, J. F., Sullivan, R. F., Hywel-Jones, N. L. & White, J. F. Resurrection of Blistum tomentosum and its exclusion from Polycephalomyces (Hyphomycetes, Deuteromycota) based on 28S rDNA sequence data. Mycotaxon. 86, 433–444 (2003).
Chen, Q.T., Xiao, S.R. & Shi, Z.Y. Paecilomyces sinensis sp. nov. and its connection with Cordyceps sinensis. Acta Mycologica Sinica. 24–28 (1984).
Wen, T. C., Xiao, Y. P., Li, W. J., Kang, J. C. & Hyde, K. D. Systematic analyses of Ophiocordyceps ramosissimum sp. nov., a new species from a larvae of Hepialidae in China. Phytotaxa. 161, 227–234 (2014).
Wen, T. et al. Metacordyceps shibinensis sp. nov. from larvae of Lepidoptera in Guizhou Province, southwest China. Phytotaxa. 226, 51–62 (2015).
Wen, T. C., Xiao, Y. P., Zha, L. S., Hyde, K. D. & Kang, J. C. Multigene phylogeny and morphology reveal a new species, Ophiocordyceps tettigonia, from Guizhou Province, China. Phytotaxa. 280, 141–151 (2016).
Wen, T. C. et al. Multigene phylogeny and morphology reveal that the Chinese medicinal mushroom ‘Cordyceps gunnii’is Metacordyceps neogunnii sp. nov. Phytotaxa. 302, 27–39 (2017).
Li, G. J. et al. Fungal diversity notes 253–366: taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers. 78, 1–237 (2016).
Tibpromma, S. et al. Fungal diversity notes 491–602: taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers. 83, 1–261 (2017).
Cock, M. J. & Allard, G. B. Observations on white grubs affecting sugar cane at the Juba Sugar Project, South-Western Somalia, in the 1980s, and implications for their management. Insects. 4, (241–272 (2013).
Han, H., Du, L. & An, T. Main Wild Medical Fungi and Their Functions in Eastern Forest Region of Zhongtiao Mountain in Shanxi Province. Agricultural Science & Technology. 16, 247–250 (2015).
Jeewon, R. & Hyde, K. D. Establishing species boundaries and new taxa among fungi: recommendations to resolve taxonomic ambiguities. Mycosphere. 7, 1669–1677 (2016).
Van Vooren, N. & Audibert, C. Révision du complexe «Cordyceps sphecocephala». 1re partie: les guêpes végétales. B mens Soc linn Lyon. 74, 221–254 (2005).
Mains, E. B. Entomogenous fungi. Mycologia. 40, 402–416 (1948).
Chomnunti, P. et al. The sooty moulds. Fungal Divers. 66, 1–36 (2014).
Vijaykrishna, D. et al. Pleurostomophora, an anamorph of Pleurostoma (Calosphaeriales), a new anamorph genus morphologically similar to Phialophora. Stud Mycol 50, 387–395 (2004).
Jayasiri, S. C. et al. The Faces of Fungi database: fungal names linked with morphology, phylogeny and human impacts. Fungal Divers. 74, 3–18 (2015).
Index Fungorum. www.indexfungorum.org 2017 (2017).
White, T. J., Bruns, T., Lee, S. & Taylor, J. W. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols: a guide to methods and applications. 18, 315–322 (1990).
Vilgalys, R. & Hester, M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol. 172, 4238–4246 (1990).
Sung, G. H., Sung, J. M., Hywel-Jones, N. L. & Spatafora, J. W. A multi-gene phylogeny of Clavicipitaceae (Ascomycota, Fungi): identification of localized incongruence using a combinational bootstrap approach. Mol Phylogenet Evol. 44, 1204–1223 (2007b).
Castlebury, L. A., Rossman, A. Y., Sung, G. H., Hyten, A. S. & Spatafora, J. W. Multigene phylogeny reveals new lineage for Stachybotrys chartarum, the indoor air fungus. Mycol Res 108, 864–872 (2004).
Jeewon, R., Cai, L., Liew, E. C. Y., Zhang, K. & Hyde, K. D. Dyrithiopsis lakefuxianensis gen et sp. nov. from Fuxian Lake, Yunnan, China and notes on the taxonomic confusion surrounding Dyrithium. Mycologia 95, 911–920 (2003).
Jeewon, R. et al. Nomenclatural and identification pitfalls of endophytic mycota based on DNA sequence analyses of ribosomal and protein genes phylogenetic markers: A taxonomic dead end? Mycosphere 8, 1802–1817 (2017).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 30, 772–780, https://doi.org/10.1093/molbev/mst010 (2013).
Hall, T. BioEdit: an important software for molecular biology. GERF Bull. Biosci. 2, 60–61 (2011).
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics. 22, 2688–2690 (2006).
Stamatakis, A., Hoover, P. & Rougemont, J. A rapid bootstrap algorithm for the RAxML web servers. Syst Biol. 57, 758–771 (2008).
Miller, M.A., Pfeiffer, W. & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. Gateway Computing Environments Workshop (GCE). 1–8 (2010).
Nylander, J. A. MrAIC [Internet] Available from: http://www.abc.se/∼nylander/. program distributed by the author (2004).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 19, 1572–1574 (2003).
Rannala, B. & Yang, Z. Probability distribution of molecular evolutionary trees: a new method of phylogenetic inference. J Mol Evol. 43, 304–311 (1996).
Zhaxybayeva, O. & Gogarten, J. P. Bootstrap, Bayesian probability and maximum likelihood mapping: exploring new tools for comparative genome analyses. BMC Genomics. 3, 4 (2002).
Cai, L., Jeewon, R. & Hyde, K. D. Phylogenetic evaluation and taxonomic revision of Schizothecium based on ribosomal DNA and protein coding genes. Fungal Divers 19, 1–21 (2005).
Jeewon, R., Liew, E. C. Y. & Hyde, K. D. Phylogenetic relationships of Pestalotiopsis and allied genera inferred from ribosomal DNA sequences and morphological characters. Mol Phylogenet Evol 25, 378–392 (2002).
Jeewon, R., Liew, E. C. Y., Simpson, J. A., Hodgkiss, I. J. & Hyde, K. D. Phylogenetic significance of morphological characters in the taxonomy of Pestalotiopsis species. Mol Phylogenet Evol 27, 372–383 (2003).
Rambaut, A. Figtree 1.4.0. http://tree.bio.ed.ac.uk/software/figtree/ (18 August 2014, date last accessed) (2012).
Kepler, R. M. et al. New teleomorph combinations in the entomopathogenic genus Metacordyceps. Mycologia. 104, 182–197 (2012).
Luangsa-Ard, J. J., Ridkaew, R., Mongkolsamrit, S., Tasanathai, K. & Hywel-Jones, N. L. Ophiocordyceps barnesii and its relationship to other melolonthid pathogens with dark stromata. Fungal Biol. 114, 739–745 (2010).
Spatafora, J. W., Sung, G. H., Sung, J. M., Hywel-Jones, N. L. & White, J. F. Phylogenetic evidence for an animal pathogen origin of ergot and the grass endophytes. Mol Ecol. 16, 1701–1711 (2007).
Nikoh, N. & Fukatsu, T. Interkingdom host jumping underground: phylogenetic analysis of entomoparasitic fungi of the genus Cordyceps. Mol Biol Evol. 17, 629–638 (2000).
Zhang, W. M., Wang, L., Tao, M. H., Chen, Y. Q. & Qu, L. H. Two species of Cordyceps simultaneously parasitic on a larva of Lepidoptera. Mycosystema. 26, 7–21 (2007).
Schoch, C. L. et al. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. P Natl Acad Sci USA 109, 6241–6246 (2012).
Kobayasi, Y. & Shimizu, D. Cordyceps species from Japan 4. Bulletin of the National Science Museum Tokyo. 8, 1–21 (1982).
Kobayasi, Y. On the genus Cordyceps and its allies on cicadae from Japan. Bulletin of the Biogeographical Society of Japan. 9, 145–176 (1939).
Sung, G. H., Spatafora, J. W., Zare, R., Hodge, K. T. & Gams, W. A revision of Verticillium sect. Prostrata. II. Phylogenetic analyses of SSU and LSU nuclear rDNA sequences from anamorphs and teleomorphs of the Clavicipitaceae. Nova Hedwigia. 72, 311–328 (2001).
Luangsa-ard, J. J., Hywel-Jones, N. L. & Samson, R. A. The polyphyletic nature of Paecilomyces sensu lato based on 18S-generated rDNA phylogeny. Mycologia 96, 773–780 (2004).
Luangsa-ard, J. et al. Purpureocillium, a new genus for the medically important Paecilomyces lilacinus. FEMS Microbiol Lett 321, 141–149 (2011).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 31460012) and the Science and Technology Foundation of Guizhou Province (No. [2016]2863). Yuanpin Xiao also thanks the Mushroom Research Foundation, Chiang Rai, Thailand for supporting this research. R.Jeewon thanks Mae Fah Luang University and University of Mauritius for research support.
Author information
Authors and Affiliations
Contributions
Y.P.X. and T.C.W. designed the study. Y.P.X. and F.Y.L. conducted all the experiments. Y.P.X., S.H., J.J.L., S.B., D.N., R.J. and K.D.H. analysed the result. Y.P.X., S.H., R.J., J.J.L., S.B., D.N., T.C.W. and K.D.H. edited the manuscript. All authors reviewed the manuscript and approved the manuscript for publication.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xiao, YP., Wen, TC., Hongsanan, S. et al. Multigene phylogenetics of Polycephalomyces (Ophiocordycipitaceae, Hypocreales), with two new species from Thailand. Sci Rep 8, 18087 (2018). https://doi.org/10.1038/s41598-018-36792-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-36792-4
- Springer Nature Limited