Abstract
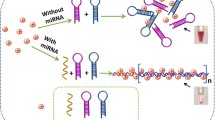
An ultrasensitive optical biosensor for microRNA-155 (miR-155) was developed to diagnose breast cancer at early stages. At first, the probe DNA covalently bind to the negatively charged gold nanoparticles (citrate-capped AuNPs). Then, the target miR-155 electrostatically adsorb onto the positively charged gold nanoparticles (polyethylenimine-capped AuNP) surface. Finally, by mixing citrate-capped AuNP/probe and polyethylenimine-capped AuNP/miR-155, hybridization occurs and the optical signal of the mixture give a measure to quantify the miR-155 content. The proposed biosensor is able to specify 3-base-pair mismatches and genomic DNA from target miR-155. The novelty of this biosensor is in its ability to trap the label-free target by its branched positively charged polyethylenimine. This method increases loading the target on the polyethylenimine-capped AuNPs’ surface. So, proposed sensor enables miR-155 detection at very low concentrations with the detection limit of 100 aM and a wide linear range from 100 aM to 100 fM.
Similar content being viewed by others
Introduction
Breast cancer is one of the main causes of cancer leading to death among women in the world1,2. Mammography and ultrasound scanning are the standard diagnostic techniques which have proven successful for early detection and diagnosis of breast cancer2. However, mammography has limited sensitivity, yielding a high rate of false-positive results. This approach may also bring about an accumulated exposure to radiation, which is considered an extra and crucial risk factor3. Ultrasound, on the other hand, is a non-invasive and safe method, but it cannot be replaced with mammography. Thus, it is thoughtful to develop a non-invasive, simple and low-risk method for screening or diagnosing breast cancer4. Among the various cancer biomarkers, microRNAs (miRs) have recently attracted researchers’ attention. These molecules are short endogenous molecules for noncoding ribonucleic acids that negatively regulate gene expression5. With the regulatory roles in gene expression, the abnormal expression of miRNAs is associated with the occurrence of different cancer types including breast cancer6. More specifically, there are several studies indicating the abnormal expression of miR-155 in breast cancer patients7,8,9. Therefore, overexpression of miR-155 suggests a risk factor for breast cancer10. Currently, methods for detection of miRNA mainly include reverse transcription polymerase chain reaction (RT-PCR)11,12,13, northern blotting14,15, microarray technique16,17, in situ hybridization18,19,20 and so on. Despite the advantages, these methods still suffer some disadvantages such as requiring expensive instruments and reagents, miRNA labeling, large amounts of miRNA and relatively pure miRNA samples21. Therefore, developing a rapid, simple and label free miRNA detection method is very important and highly desirable.
Colorimetric-based detection techniques have attracted great attention due to their simplicity, rapidness and low cost22,23,24. In colorimetric sensing methods, metallic nanoparticles, especially AuNPs because of their high extinction coefficient and strong distance- and size- dependent optical property25 are widely used26,27,28,29,30. The color of small AuNPs (~10–50 nm), when individually dispersed, is red due to the coherent oscillation of AuNP surface electrons (localized surface plasmon resonance)31. Upon aggregation, the solution color changes from red to pinkish/purple32 due to the coupling of AuNP surface plasmons31,33. In colorimetric-based biosensing assays based on AuNPs, there are two kinds of controlled aggregation of AuNPs: Interparticle bonding formation (interparticle crosslinking aggregation) and non-crosslinking aggregation that is produced by the removal of colloidal stabilization effects34,35.
The pioneers of the detection methods based on interparticle crosslinking aggregation of AuNPs were Mirkin and co-workers. They reported a novel method for colorimetric detection of DNA targets based on thiolated oligonucleotide modified GNPs (AuNP/probe) that used the distance-dependent optical properties of aggregated AuNP/probe. They used two sets of AuNP/probe with two different probes. After addition of a linker double-stranded DNA (dsDNA) with overhangs complementary to the two AuNP/probes, aggregation occurred and color of the solution changed from red to blue36. The same group one year later detected single-stranded DNA (ssDNA) with a detection limit of 10 fmol32. Thereinafter several analytes detected by using two kinds of probes that covalently attached to the surface of AuNPs35,37,38,39,40,41.
The biosensing assays based on non-crosslinking aggregation of AuNPs were reported for the first time by Maeda and co-workers30. In using this method several analytes were detected42,43,44. The main purpose of using colorimetric-based sensors being to improve sensitivity of such methods24. Several amplification techniques such as enzymatic signal amplification45,46,47, exonuclease-assisted signal amplification48,49 and rolling circle amplification50,51,52,53 have been developed to improve sensitivity of the colorimetric assays. But, these amplification methods have some disadvantages, such as requiring more time, higher costs and a more complex process52.
For the first time, in order to improve the sensitivity of biosensing assays based on crosslinking aggregation, branched polyethylenimine (PEI) capped AuNP (P-AuNP) were used for target trapping. P-AuNPs are highly positively charged because of the presence of nitrogen atoms of branched PEI, which makes them suitable for concentrating negatively charged molecules54,55,56 such as miR-155 resulting in the formation of P-AuNPs/miR-155 complex. On the other hand, thiolated hairpin probe DNA is covalently attached to the citrate-capped AuNPs (C-AuNPs) and the resulting C-AuNPs/probe. By mixing two complexes containing probe and target, DNA/RNA hybrid duplex is formed which cause interparticle cross-linking aggregation. This is quantified as an optical signal for monitoring the target. The significant and interesting point of this study is using branched PEI as concentrating miR-155 on it causes miR-155 detection at very low concentrations without the need for expensive and time-consuming signal amplifications.
Results
Sensing mechanism
Figure 1 illustrates the sensing mechanism of the proposed optical biosensor. In the first step, after C-AuNP synthesis, C-AuNPs are functionalized with the thiolated probes (Fig. 1A). It should be noted that thiolated probe molecules contain a nine-nucleotide thymidine spacer at its 3′ end which provides further optimal immobilization and hybridization efficiency57. In addition, the hairpin structure of our designed probe enhances its specificity more so than the corresponding linear ones58. In parallel, P-AuNPs are prepared and modified with miR-155 (Fig. 1B). Noteworthy, the color of C-AuNP/probe is dark pink while the color of P-AuNPs and P-AuNP/miR-155 are red. In the next step, when C-AuNP/probe add to P-AuNPs/miR-155, the mixture displays interparticle cross-linking aggregates; its color changes from red-pink to pink and the absorption intensity decreases at ~530 nm (Fig. 2). In fact, probe-target hybridization begins by addition of C-AuNP/probe to P-AuNP/miR-155, the distance between nanoparticles is reduced and, consequently, the aggregation process happens.
Since the UV-Vis absorption at ~530 nm indicates the quantity of dispersed nanoparticles and the absorbance at ~750 nm designates the aggregated AuNPs44, the absorbance ratio of 530/750 nm can express molar ratio of dispersed to aggregated AuNPs. Hence, the higher values of A530/A750, the greater stability of AuNPs and; the lower values of A530/A750, the greater aggregation.
Characterization of C-AuNPs and P-AuNPs
Two types of gold nanoparticles were synthesized based on the method reported in the literature59,60,61 by slight modification. C-AuNPs and P-AuNPs were synthesized via reduction of HAuCl4 by using two reductants of trisodiumcitrate and polyethylenimine, respectively. Formation of C-AuNPs and P-AuNPs were confirmed by the UV-Vis spectra of two samples (Fig. 3A). The maxima of the plasmon resonance bands, located at ~530 nm, give evidence of the formation of C-AuNPs and P-AuNPs. The red color of the two samples (insets in Fig. 3A) demonstrates the successful formation of gold nanoparticles. SEM images also confirmed the formation of nanoparticles (Fig. 3B and C). Since citrate ions surround C-AuNPs and PEI cover P-AuNPs, these complexes are negatively and positively charged and their zeta potentials, as might be expected, have different signs i.e. −16.32 ± 0.17 mV and +36.51 ± 1.40 mV, respectively. In addition the hydrodynamic size of C-AuNPs and P-AuNPs were measured by DLS to be 21.2 ± 1.4 nm and 22.6 ± 0.3 nm, respectively (data not shown).
Modification of C-AuNPs by thiolated probe and P-AuNPs by miR-155
As mentioned in the previous section, attachment of thiolated probe molecules to C-AuNPs leads to the coverage of gold nanoparticles with probe molecules on the surface (C-AuNPs/probe) that are used for trapping target molecules. On the other hand, attachment of target molecules (miR-155) to the surface of P-AuNPs results in the formation of P-AuNPs/miR-155 with the ability of hybridization to the probe. To confirm the formation of P-AuNPs/miR-155 and C-AuNPs/probe, UV-Vis spectroscopy was used. As Fig. 4A shows, after coating of C-AuNPs with thiolated probes, the absorption at 530 nm decreases drastically, while that at 550 to 800 nm region increases and the color of nanoparticles changes from red to dark pink. As explained in the literature, the reason for changes in the position and intensity of absorption peak of gold nanoparticles is the changes of plasmon resonance frequency. Several factors influence these changes such as size and morphology of particles, dielectric constant of metal and environment62, shape and size of the charge distribution, electron density, effective electron mass63, electrolyte due to charge screening effects64,65 and agglomeration66 or aggregation of AuNPs. It seems that covering the C-AuNPs by the hairpin structured thiolated probes, possibly results in the vertical placement of probe on the surface of C-AuNPs. This may form a thick insulating layer on the surface of nanoparticles which causes some changes in the aforementioned physical properties of nanoparticles.
On the other hand, P-AuNPs possess positive charges. Therefore, they are able to bind to negatively charged phosphate groups of miR-155 through electrostatic interactions which results a slight decrease in the absorption at 600–800 nm (Fig. 4B). Mir-155 molecules are single stranded and may orient parallel to the nanoparticles’ surfaces and consequently form a weak thin insulation-film. This film, in turn, alters the physical surface properties of P-AuNPs including dielectric constant and surface plasmon resonance which causes the absorption change of P-AuNPs/mir-155 being minute relative to that of P-AuNPs alone.
To prove the successful attachment of probe to C-AuNP surface, the zeta potential of C-AuNPs before and after probe addition were compared. Reducing in zeta potential from −16.32 ± 0.17 to −18.35 ± 1.04 mV indicates successful attachment of probe molecules to C-AuNPs surface. The attachment of negatively charged probe molecules to nanoparticles increases the overall negative charges of the latter. On the other hand, after miR-155 addition, zeta potential of P-AuNPs decreases from +36.51 ± 1.40 to +24.34 ± 1.36 mV which implies negative charges of target molecules compensate positive charges of nanoparticles; an observation that confirms the successful attachment of miR-155 to P-AuNPs through electrostatic interactions.
Calibration curve for detection of miR-155
Serial dilutions of miR-155 were prepared and analyzed based on the sensing mechanism (see Methods). To remove the background noise and therefore, better present the sensor response, we used Equation 1, in which \(\delta {{\rm{A}}}_{(\frac{530}{750})}\) is a measure which indicates the amount of aggregated nanoparticles. The more concentration of miR-155, the more hybridization between probe and miR-155. This hybridization results in nanoparticles aggregation and therefore increment in \(\delta {{\rm{A}}}_{(\frac{530}{750})}\) quantity. In Equation 1, \({{\rm{A}}}_{(\frac{530}{750})}\) indicates the absorption ratio of C-AuNP/probe + P-AuNP/miR-155 at 530 nm to that at 750 nm. In addition, \({{\rm{A}}}_{(\frac{530}{750}){\rm{p}}}\) represents the absorption of C-AuNP/probe + P-AuNP at 530 nm to that at 750 nm.
Based on the \(\delta {{\rm{A}}}_{(\frac{530}{750})}\) vs. log miR-155 concentration, a calibration curve was plotted (Fig. 5). The linear segment of the resulting sigmoid from 102 to 105 attomolar (aM) was used as the calibration curve for quantitative detection of target (Fig. 5, inset). Upon increasing the concentration of miR-155, the number of dispersed nanoparticles descended while the number of aggregated nanoparticles ascended. Therefore, as shown in the calibration curve, the \(\delta {{\rm{A}}}_{(\frac{530}{750})}\) value increases by increasing the target concentration. The intersection between two linear functions of maximum and minimum slope at low concentrations was used59,67 to estimate the limit of detection (LOD) of miR-155 to be 100 aM. As compared in Table 1, this LOD is much lower than those reported for other colorimetric methods32,37,68,69,70,71,72,73,74,75. Such a low LOD is probably due to the ability of the branched PEI to increase the target loading by concentrating76 miR-155 molecules on the P-AuNPs surface.
Selectivity of the optical biosensor
The selectivity of the optical biosensor was studied using three sequences including perfectly complementary targets (miR-155), three-base mismatched strands and non-complementary strands (genomic DNA). Comparison between the three responses and background are shown in Fig. 6. As seen, the response toward perfectly complementary target is 3.5 times bigger than that of three-base mismatch sequence and about 8 times higher than that of the genomic DNA. The higher sensor response toward miR-155 indicate that the probe-target duplex formation is more probable in comparison to the other duplexes. To confirm this superiority in hybridization, we also calculated the hybridization free energy of probe-target and probe-three base mismatch sequence using “RNA structure” web server. Since, the stability and the probability of hybridization depend on the free energy of hybridization77,78, therefore it has been used for comparison between the stability of duplexes. Results showed that the hybridization energy of probe-target and probe-three base mismatch are −39 and −23/3 kJ/mol, respectively. Since, probe-target hybridization energy is more negative than that of the probe-three base mismatch, therefore, the probe-target duplex is more stable and probable. Indeed, free energy of hybridization is the sum of individual base-pairing reactions involved. Thus, the total number of complementary bases in the two strands determines the duplex stability and probability78. Since, probe-target has more base pair than probe-three base mismatch and probe-genomic DNA, we conclude that duplex stability and probability in probe-target is higher than those for other duplexes. Even there is a possibility of binding of each negatively charged molecule to P-AuNPs, only miR-155 binding causes more cross-linking aggregation.
Discussion
A versatile, sensitive, simple, and cost-effective method for detection of miR-155 was developed for the first time. The sensing signal was based on the hybridization of probe/functionalized C-AuNPs and P-AuNPs modified with miR-155. The mechanism behind the designed sensor lies in the hybridization between probe and target i.e. miR-155 caused interparticle cross-linking aggregation of C-AuNPs and P-AuNPs; which in its macroscopic scale shows a change in color from red-pink to pink. The innovation of this method is the utilization of positively charged AuNPs that are modified with target by electrostatic interaction for interparticle cross-linking aggregation, this reduces time and steps of sensing along with no need to tag target. Moreover, by using the branched P-AuNPs, the absorption of miR-155 on the surface of nanoparticles increases. Of interest, this process dramatically improves the interparticle cross-linking aggregation and, consequently, the detection limit shifts to the lower values. This method can be used to detect any oligonucleotide simply by changing the probe sequence. We expect that sophisticated optimization of the ratio between C-AuNPs, probe, and P-AuNPs might lead to even higher sensitivity.
Methods
Chemicals
The HPLC purified oligonucleotide sequences are as follows: Hsa-miR-155 (5′-UUAAUGCUAAUCGUGAUAGGGGU-3′) and three-base mismatched hsa-miR-155 (5′-UUAAUGCUUAUCGAGAUACGGGU-3′) were purchased from Bioneer Corporation (Republic of South Korea). Thiolated miR-155 probe (5′-AAAAAAAAACCCCTATCACGATTAGCATTAATTTTTTTTT-HS-3′) was synthesized by AnaSpec, Inc. (Canada). HAuCl4, polyethylenimine (50% solution, Mn ~1200, Mw ~1300) and dialysis bag were obtained from Sigma-Aldrich (USA). Trisodiumcitratedihydrate (Na3C6H5O7.2H2O), HCl and Tween-20 were purchased from Merck (Germany).
Preparation of C-AuNPs and C-AuNP/probe
C-AuNPs with the average diameter of 21 nm were synthesized by citrate reduction of HAuCl461. In brief, 1.5 mL of sodium citrate 1% was added to 21 mL of boiling chloroauric acid solution 0.8 mM, while vigorously stirring until its color changed from pale yellow to deep red. The solution, then, was stirred for an additional 15 min and gradually cooled down to room temperature61,79. To prepare C-AuNP/probe, 400 µL C-AuNPs was mixed with 2 µL Tween 20 and 400 µL thiolated probe 1 µM, left for 48 hrs, and centrifuged for 23 min at 10,000 rpm. Finally, the supernatant was removed, and the oily red precipitate redispersed in 200 µL deionized water.
Preparation of P-AuNPs and P-AuNP/miR-155
P-AuNPs were synthesized by thermal reduction with slight modification to the previous reports59,60. In brief, 100 µL of PEI 42 mM was added to 3 mL of 1.5 M HAuCl4 under vigorous stirring, pH adjusted to 7.4 with HCl, and then, the solution was brought to a boil; color change from yellow to red is an indication of the reduction process. The obtained nanoparticles were dialyzed against deionized water with a 3.5-kDa cutoff membrane. The resulting red solution was stored at 4 °C before use. For the preparation of P-AuNPs/miR-155, 5 µL of the miR-155 solution with specific concentration was incubated with 40 µL of synthesized P-AuNPs for 30 min at room temperature.
Characterization
UV-Vis absorption spectra of the particle dispersions were measured using a Varian Cary Bio 100 spectrophotometer. The particles were characterized by dynamic light scattering (DLS) using a 90 Plus Pals (Brookhaven Instruments Corp., USA), and PALS zeta-particle sizing potential analyzer software. All optical sensing experiments were measured by a microplate reader (BioTek, PowerWave XS2, USA). Morphology and size of nanoparticles were further confirmed by scanning electron microscope (SEM) (KYKY, EM3200, China).
Optical sensing of miR-155
For miR-155 detection, 40 µL of P-AuNPs and 5 µL of miR-155 solution were mixed and incubated for ∼30 min at room temperature, followed by the addition of 5 µL of C-AuNP/probe to it. After 15 min, UV-Vis absorption spectra of the aggregated particles were recorded by a microplate reader.
References
Jemal, A. et al. Cancer statistics, 2009. CA: a cancer journal for clinicians 59, 225–249 (2009).
Schrauder, M. G. et al. Circulating micro-RNAs as potential blood-based markers for early stage breast cancer detection. Plos One 7, e29770 (2012).
Yahalom, G. et al. An antibody-based blood test utilizing a panel of biomarkers as a new method for improved breast cancer diagnosis. Biomarkers in cancer 5, 71 (2013).
Cardoso, A. R., Moreira, F. T., Fernandes, R. & Sales, M. G. F. Novel and simple electrochemical biosensor monitoring attomolar levels of miRNA-155 in breast cancer. Biosensors and Bioelectronics 80, 621–630 (2016).
Wang, Q., Li, R.-D., Yin, B.-C. & Ye, B.-C. Colorimetric detection of sequence-specific microRNA based on duplex-specific nuclease-assisted nanoparticle amplification. Analyst 140, 6306–6312 (2015).
Zhao, H., Qu, Y., Yuan, F. & Quan, X. A visible and label-free colorimetric sensor for miRNA-21 detection based on peroxidase-like activity of graphene/gold-nanoparticle hybrids. Analytical Methods 8, 2005–2012 (2016).
Mattiske, S., Suetani, R. J., Neilsen, P. M. & Callen, D. F. The oncogenic role of miR-155 in breast cancer. Cancer Epidemiology and Prevention Biomarkers 21, 1236–1243 (2012).
Sun, Y. et al. Serum microRNA-155 as a potential biomarker to track disease in breast cancer. PloS one 7, e47003 (2012).
Jiang, S. et al. MicroRNA-155 functions as an OncomiR in breast cancer by targeting the suppressor of cytokine signaling 1 gene. Cancer research 70, 3119–3127 (2010).
Zeng, H., Fang, C., Nam, S., Cai, Q. & Long, X. The clinicopathological significance of microRNA-155 in breast cancer: a meta-analysis. Biomed Res Int 2014 (2014).
Varkonyi-Gasic, E., Wu, R., Wood, M., Walton, E. F. & Hellens, R. P. Protocol: a highly sensitive RT-PCR method for detection and quantification of microRNAs. Plant methods 3, 12 (2007).
Wang, Z. & Yang, B. In MicroRNA Expression Detection Methods 131–140 (Springer, 2010).
Ho, Y. K., Xu, W. T. & Too, H. P. Direct quantification of mRNA and miRNA from cell lysates using reverse transcription real time PCR: a multidimensional analysis of the performance of reagents and workflows. PloS one 8, e72463 (2013).
Várallyay, É., Burgyán, J. & Havelda, Z. MicroRNA detection by northern blotting using locked nucleic acid probes. Nat Protoc 3, 190–196 (2008).
Rio, D. C. Northern blots for small RNAs and microRNAs. Cold Spring Harbor Protocols 2014, pdb. prot080838 (2014).
Zhao, B. et al. A simple and fast method for profiling microRNA expression from low‐input total RNA by microarray. IUBMB life 64, 612–616 (2012).
Liang, R.-Q. et al. An oligonucleotide microarray for microRNA expression analysis based on labeling RNA with quantum dot and nanogold probe. Nucleic Acids Res 33, e17–e17 (2005).
Song, R., Ro, S. & Yan, W. In situ hybridization detection of microRNAs. RNA Therapeutics: Function, Design, and Delivery, 285–292 (2010).
Doné, S. C. & Beltcheva, O. In Situ Hybridization Detection of miRNA Using LNA™ Oligonucleotides. RNA Mapping: Methods and Protocols, 57–71 (2014).
Zhang, X., Lu, X., Lopez-Berestein, G., Sood, A. & Calin, G. In situ hybridization-based detection of microRNAs in human diseases. microRNA Diagn Ther 1, 12–23 (2013).
Zhou, Y., Wang, M., Meng, X., Yin, H. & Ai, S. Amplified electrochemical microRNA biosensor using a hemin-G-quadruplex complex as the sensing element. RSC Advances 2, 7140, https://doi.org/10.1039/c2ra20487h (2012).
Hunt, E. A., Goulding, A. M. & Deo, S. K. Direct detection and quantification of microRNAs. Analytical Biochemistry 387, 1–12, https://doi.org/10.1016/j.ab.2009.01.011 (2009).
Dykman, L. & Khlebtsov, N. Gold nanoparticles in biomedical applications: recent advances and perspectives. Chem Soc Rev 41, 2256–2282 (2012).
Miao, X., Ning, X., Li, Z. & Cheng, Z. Sensitive detection of miRNA by using hybridization chain reaction coupled with positively charged gold nanoparticles. Sci Rep-Uk 6, 32358 (2016).
Lee, J. H., Wang, Z., Liu, J. & Lu, Y. Highly sensitive and selective colorimetric sensors for uranyl (UO22+): Development and comparison of labeled and label-free DNAzyme-gold nanoparticle systems. J Am Chem Soc 130, 14217–14226 (2008).
Liu, J. & Lu, Y. Preparation of aptamer-linked gold nanoparticle purple aggregates for colorimetric sensing of analytes. Nat Protoc 1, 246 (2006).
Zhao, W., Gonzaga, F., Li, Y. & Brook, M. A. Highly Stabilized Nucleotide‐Capped Small Gold Nanoparticles with Tunable Size. Adv Mater 19, 1766–1771 (2007).
Storhoff, J. J. et al. What controls the optical properties of DNA-linked gold nanoparticle assemblies? J Am Chem Soc 122, 4640–4650 (2000).
Mirkin, C. A., Letsinger, R. L., Mucic, R. C. & Storhoff, J. J. A DNA-based method for rationally assembling nanoparticles into macroscopic materials. Nature 382, 607–609 (1996).
Sato, K., Hosokawa, K. & Maeda, M. Rapid aggregation of gold nanoparticles induced by non-cross-linking DNA hybridization. Journal of the American Chemical Society 125, 8102–8103 (2003).
Zhao, W., Brook, M. A. & Li, Y. Design of gold nanoparticle‐based colorimetric biosensing assays. Chembiochem 9, 2363–2371 (2008).
Elghanian, R., Storhoff, J. J., Mucic, R. C., Letsinger, R. L. & Mirkin, C. A. Selective colorimetric detection of polynucleotides based on the distance-dependent optical properties of gold nanoparticles. Science 277, 1078–1081 (1997).
Broude, N. E. Molecular beacons and other hairpin probes. Encyclopedia of Diagnostic Genomics and Proteomics, 846–850 (2005).
Zhao, W., Brook, M. A. & Li, Y. F. Design of Gold Nanoparticle-Based Colorimetric Biosensing Assays. Chembiochem 9, 2363–2371, https://doi.org/10.1002/cbic.200800282 (2008).
Wong, K. F. J. Ultra-stable Oligonucleotide-nanoparticle Conjugates Prepared by Silica Reinforcement Method: Properties and Applications for Colorimetric DNA Detection. (Interdisciplinary Division of Biomedical Engineering, The Hong Kong Polytechnic University, 2012).
Mirkin, C. A., Letsinger, R. L., Mucic, R. C. & Storhoff, J. J. A DNA-based method for rationally assembling nanoparticles into macroscopic materials. Nature 382, 607–609, https://doi.org/10.1038/382607a0 (1996).
Li, J. et al. A colorimetric method for point mutation detection using high-fidelity DNA ligase. Nucleic Acids Res 33, e168–e168 (2005).
Xue, X., Wang, F. & Liu, X. One-step, room temperature, colorimetric detection of mercury (Hg2+) using DNA/nanoparticle conjugates. J Am Chem Soc 130, 3244–3245 (2008).
Han, M. S., Lytton-Jean, A. K. & Mirkin, C. A. A gold nanoparticle based approach for screening triplex DNA binders. J Am Chem Soc 128, 4954–4955 (2006).
Han, M. S., Lytton‐Jean, A. K., Oh, B. K., Heo, J. & Mirkin, C. A. Colorimetric Screening of DNA‐Binding Molecules with Gold Nanoparticle Probes. Angewandte Chemie International Edition 45, 1807–1810 (2006).
Liu, J. & Lu, Y. Fast colorimetric sensing of adenosine and cocaine based on a general sensor design involving aptamers and nanoparticles. Angewandte Chemie 118, 96–100 (2006).
Conde, J., de la Fuente, J. M. & Baptista, P. V. RNA quantification using gold nanoprobes-application to cancer diagnostics. J Nanobiotechnol 8, 5 (2010).
Zu, Y., Ting, A. L. & Gao, Z. Visualizing Low‐Level Point Mutations: Enzyme‐like Selectivity Offered by Nanoparticle Probes. Small 7, 306–310 (2011).
Chen, S.-J. et al. Colorimetric determination of urinary adenosine using aptamer-modified gold nanoparticles. Biosensors and Bioelectronics 23, 1749–1753, https://doi.org/10.1016/j.bios.2008.02.008 (2008).
Zhang, Y., Guo, Y., Quirke, P. & Zhou, D. Ultrasensitive single-nucleotide polymorphism detection using target-recycled ligation, strand displacement and enzymatic amplification. Nanoscale 5, 5027–5035 (2013).
Chen, D. M., Gao, Z. F., Jia, J., Li, N. B. & Luo, H. Q. A sensitive and selective electrochemical biosensor for detection of mercury (II) ions based on nicking endonuclease-assisted signal amplification. Sensors and Actuators B: Chemical 210, 290–296 (2015).
Li, L., Wang, Q., Feng, J., Tong, L. & Tang, B. Highly sensitive and homogeneous detection of membrane protein on a single living cell by aptamer and nicking enzyme assisted signal amplification based on microfluidic droplets. Anal Chem 86, 5101–5107 (2014).
Bi, S., Li, L. & Cui, Y. Exonuclease-assisted cascaded recycling amplification for label-free detection of DNA. Chem Commun 48, 1018–1020 (2012).
Wang, M. et al. One-step, ultrasensitive, and electrochemical assay of microRNAs based on T7 exonuclease assisted cyclic enzymatic amplification. Anal Chem 86, 5606–5610 (2014).
Deng, R. et al. Toehold‐initiated Rolling Circle Amplification for Visualizing Individual MicroRNAs In Situ in Single Cells. Angewandte Chemie International Edition 53, 2389–2393 (2014).
Gao, A. et al. Signal-to-noise ratio enhancement of silicon nanowires biosensor with rolling circle amplification. Nano Lett 13, 4123–4130 (2013).
Russell, C. et al. Gold nanowire based electrical DNA detection using rolling circle amplification. Acs Nano 8, 1147–1153 (2014).
Li, D. et al. A colorimetric biosensor for detection of attomolar microRNA with a functional nucleic acid-based amplification machine. Talanta 146, 470–476 (2016).
Dunlap, D. D., Maggi, A., Soria, M. R. & Monaco, L. Nanoscopic structure of DNA condensed for gene delivery. Nucleic Acids Res 25, 3095–3101 (1997).
Benjaminsen, R. V., Mattebjerg, M. A., Henriksen, J. R., Moghimi, S. M. & Andresen, T. L. The possible “proton sponge” effect of polyethylenimine (PEI) does not include change in lysosomal pH. Molecular Therapy 21, 149–157 (2013).
Yang, T. et al. Positively charged polyethylenimines enhance nasal absorption of the negatively charged drug, low molecular weight heparin. Journal of controlled release 115, 289–297 (2006).
Pöhlmann, C. & Sprinzl, M. Electrochemical detection of microRNAs via gap hybridization assay. Anal Chem 82, 4434–4440 (2010).
Khudyakov, Y. E. & Fields, H. A. Artificial DNA: Methods and applications. (CRC Press, 2002).
Mohammed, F. S., Cole, S. R. & Kitchens, C. L. Synthesis and enhanced colloidal stability of cationic gold nanoparticles using polyethyleneimine and carbon dioxide. ACS Sustainable Chemistry & Engineering 1, 826–832 (2013).
Sun, X., Jiang, X., Dong, S. & Wang, E. One‐Step Synthesis and Size Control of Dendrimer‐Protected Gold Nanoparticles: A Heat‐Treatment‐Based Strategy. Macromolecular rapid communications 24, 1024–1028 (2003).
Ji, X. et al. Size control of gold nanocrystals in citrate reduction: the third role of citrate. Journal of the American Chemical Society 129, 13939–13948 (2007).
Reinhard, B. M., Siu, M., Agarwal, H., Alivisatos, A. P. & Liphardt, J. Calibration of dynamic molecular rulers based on plasmon coupling between gold nanoparticles. Nano Lett 5, 2246–2252 (2005).
Kelly, K. L., Coronado, E., Zhao, L. L. & Schatz, G. C. (ACS Publications, 2003).
Mulvaney, P. Surface plasmon spectroscopy of nanosized metal particles. Langmuir 12, 788–800 (1996).
Storhoff, J. J., Elghanian, R., Mucic, R. C., Mirkin, C. A. & Letsinger, R. L. One-pot colorimetric differentiation of polynucleotides with single base imperfections using gold nanoparticle probes. J. Am. Chem. Soc 120, 1959–1964 (1998).
Tan, J. et al. A Gold Nanoparticle-enhanced Surface Plasmon Resonance Aptasensor for the Detection of 2, 4, 6-trinitrotoluene. Journal of Forensic Science and Medicine 2, 195 (2016).
Ivanova, O. S. & Zamborini, F. P. Size-dependent electrochemical oxidation of silver nanoparticles. J Am Chem Soc 132, 70–72 (2009).
Sato, K., Hosokawa, K. & Maeda, M. Non-cross-linking gold nanoparticle aggregation as a detection method for single-base substitutions. Nucleic acids research 33, e4–e4 (2005).
Lee, H. et al. Colorimetric genotyping of single nucleotide polymorphism based on selective aggregation of unmodified gold nanoparticles. Biosensors and Bioelectronics 26, 730–735 (2010).
Ali, M. et al. Nanoparticle sensor for label free detection of swine DNA in mixed biological samples. Nanotechnology 22, 195503 (2011).
Lam, M. K. et al. Tuning toehold length and temperature to achieve rapid, colorimetric detection of DNA from the disassembly of DNA–gold nanoparticle aggregates. Langmuir 32, 1585–1590 (2016).
Xu, W., Xue, X., Li, T., Zeng, H. & Liu, X. Ultrasensitive and selective colorimetric DNA detection by nicking endonuclease assisted nanoparticle amplification. Angewandte Chemie International Edition 48, 6849–6852 (2009).
Li, H. & Rothberg, L. Colorimetric detection of DNA sequences based on electrostatic interactions with unmodified gold nanoparticles. P Natl Acad Sci USA 101, 14036–14039 (2004).
Xia, F. et al. Colorimetric detection of DNA, small molecules, proteins, and ions using unmodified gold nanoparticles and conjugated polyelectrolytes. Proceedings of the National Academy of Sciences 107, 10837–10841 (2010).
Li, J. et al. Enzyme‐Based Multi‐Component Optical Nanoprobes for Sequence‐Specific Detection of DNA Hybridization. Advanced Materials 20, 497–500 (2008).
Cuk, K. et al. Circulating microRNAs in plasma as early detection markers for breast cancer. International journal of cancer. Journal international du cancer 132, 1602–1612, https://doi.org/10.1002/ijc.27799 (2013).
Schmitt, T. J. & Knotts, T. A. IV Thermodynamics of DNA hybridization on surfaces. The Journal of chemical physics 134, 05B614 (2011).
Hemmings, H. C. & Hopkins, P. M. Foundations of anesthesia: basic sciences for clinical practice. (Elsevier Health Sciences, 2006).
Gu, P. et al. A novel visible detection strategy for lysozyme based on gold nanoparticles and conjugated polymer brush. Sensors and Actuators B: Chemical 246, 78–84 (2017).
Acknowledgements
Financial supports provided by the Research Council of the University of Tehran are gratefully appreciated. The authors also thank to Mr. Jamal Hakimian for buying the DNA samples.
Author information
Authors and Affiliations
Contributions
Fatemeh Hakimian: Doing experiments and writing the manuscript. Hedayatollah Ghourchian: Supervisor and editor. Azam sadat Hashemi: Providing RNA samples, advisor. Mohammad Reza Arastoo: RNA handling, helping in writing the manuscript. Mohammad Behnam Rad: Helping in doing some experiments, helping in writing the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hakimian, F., Ghourchian, H., Hashemi, A.s. et al. Ultrasensitive optical biosensor for detection of miRNA-155 using positively charged Au nanoparticles. Sci Rep 8, 2943 (2018). https://doi.org/10.1038/s41598-018-20229-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-20229-z
- Springer Nature Limited
This article is cited by
-
Ag nanorod@PEI-Ag nanohybrid as an excellent signal label for sensitive and rapid detection of serum HER2
Scientific Reports (2023)
-
Recent Advances in Electrochemical and Optical Biosensors for Cancer Biomarker Detection
BioChip Journal (2023)
-
Advanced graphene oxide-based paper sensor for colorimetric detection of miRNA
Microchimica Acta (2022)
-
GST-Based Plasmonic Biosensor for Hemoglobin and Urine Detection
Plasmonics (2022)
-
Role of Sensor Technology in Detection of the Breast Cancer
BioNanoScience (2022)