Abstract
Notch signalling activity governs cellular differentiation in higher metazoa, where Notch signals are transduced by the transcription factor CSL, called Suppressor of Hairless [Su(H)] in Drosophila. Su(H) operates as molecular switch on Notch target genes: within activator complexes, including intracellular Notch, or within repressor complexes, including the antagonist Hairless. Mass spectrometry identified phosphorylation on Serine 269 in Su(H), potentially serving as a point of cross-regulation by other signalling pathways. To address the biological significance, we generated phospho-deficient [Su(H)S269A] and phospho-mimetic [Su(H)S269D] variants: the latter displayed reduced transcriptional activity despite unaltered protein interactions with co-activators and -repressors. Based on the Su(H) structure, Ser269 phosphorylation may interfere with DNA-binding, which we confirmed by electro-mobility shift assay and isothermal titration calorimetry. Overexpression of Su(H)S269D during fly development demonstrated reduced transcriptional regulatory activity, similar to the previously reported DNA-binding defective mutant Su(H)R266H. As both are able to bind Hairless and Notch proteins, Su(H)S269D and Su(H)R266H provoked dominant negative effects upon overexpression. Our data imply that Ser269 phosphorylation impacts Notch signalling activity by inhibiting DNA-binding of Su(H), potentially affecting both activation and repression. Ser269 is highly conserved in vertebrate CSL homologues, opening the possibility of a general and novel mechanism of modulating Notch signalling activity.
Similar content being viewed by others
Introduction
Metazoan development depends on a small number of fundamental, conserved signalling pathways that are employed repeatedly to govern cellular differentiation. Not surprisingly, their dysregulation is a major cause of congenital diseases, underlying the pathophysiology of a multitude of different cancers, including solid tumours and leukemias. Together, these fundamental pathways coordinate developmental processes by building up complex signalling networks with extensive molecular cross-talk1,2,3. One such pathway is the Notch signalling pathway, which takes place directly between neighbouring cells, steering them into alternate fates. Appropriate cellular differentiation, therefore, strictly depends on Notch signalling in many instances, explaining its impact on human health (for review4,5,6). The serious consequences of aberrant Notch signalling have triggered large-scale efforts aimed at understanding the regulatory mechanisms underlying Notch activity. One hallmark of these studies is that regulatory mechanisms can occur at multiple steps of signal transduction, ranging from the selection and variation of ligands, crosstalk with other signalling cascades, and various context and tissue specific modification of Notch signalling components5,6,7,8,9. Such investigative studies, however, are complicated by genetic redundancy in vertebrates, e.g. mammals have four Notch receptors and five ligands, highlighting the importance of using model organisms like Drosophila melanogaster to elucidate novel mechanisms of regulation.
The principles of Notch signal transduction are rather simple: the binding of ligands presented on the surface of the signalling cell triggers the cleavage of the Notch receptor in the adjacent signal receiving cell, releasing the intracellular domain NICD (Notch Intracellular Domain). NICD now becomes a transcriptional co-activator: together with co-activators of the Mastermind family (Mam) it assembles a ternary activator complex with the transcription factor CSL (CBF-1/Suppressor of Hairless/Lag-1) on Notch target genes (for review9,10,11). CSL acts as a molecular switch on Notch target genes as it can either assemble activator or repressor complexes depending on its binding partners10. The structure of these complexes has been analysed in several organisms: CSL contains three domains, an N-terminal domain (NTD), a beta-trefoil domain (BTD), and a C-terminal domain (CTD). DNA binding is mediated by NTD and BTD, whereas BTD and CTD are involved in the formation of the activator complex by binding to NICD and Mam, respectively11. In vertebrates, several co-repressors compete with NICD for the binding of BTD10. In Drosophila, repressor complex formation relies on the general Notch antagonist Hairless. Hairless binds with high affinity to the CTD of Su(H), thereby precluding the binding of NICD12,13. In addition, Hairless recruits the two general co-repressors Groucho and C-terminal binding protein to silence Notch target genes14.
As cells are under the influence of several developmental pathways, molecular cross-talk is required for their coordination2,15. Indeed, NICD phosphorylation affects complex formation and stability, which eventually results in either stimulation or inhibition of Notch signalling, depending on the mediating kinase and the targeted region of NICD (for review16,17). As several of these kinases are also an integral part of other signalling pathways they have the potential to act as the mediators of crosstalk. A prime example is the crosstalk between the epidermal growth factor receptor (EGFR) and Notch pathways in Drosophila. These two signalling pathways have a complex relationship, either acting in a cooperative or antagonistic manner, depending on the developmental context8,18. Pivotal to EGFR signalling is the Mitogen-activated protein kinase (MAPK), and two well established substrates of MAPK in Drosophila are the Notch signalling components Groucho (Gro) and Suppressor of Hairless [Su(H)], which are both restricted in their activity by MAPK dependent phosphorylation15,19. Nonetheless, there are likely to be other Notch signalling components that are modified in response to other signalling pathways.
Here, we report the identification of an additional phosphorylation site in Su(H) by a mass spectrometry approach. The identified phospho-serine 269 is located in the beta-trefoil domain (BTD) of Su(H), leaving the possibility of influencing the association of Su(H) with NICD and/or with DNA. Using phospho-site specific mutants we show that the phospho-mimetic Su(H)S269D is impaired in transcriptional regulation. We find that protein complexes with NICD and Hairless form normally; however, we find that DNA binding is affected in Su(H)S269D. Moreover, overexpression analyses during fly development provide evidence for a restricted ability of Su(H)S269D to activate and repress Notch target genes, revealing dominant negative effects at the same time. In contrast, the phospho-deficient mutant Su(H)S269A behaves similarly to the wild type protein. As Ser269 is highly conserved, we propose a new mode of Notch signal regulation at the level of affecting DNA binding by the transcription factor CSL.
Results
Su(H) protein is phosphorylated on Serine 269 in S2 cell culture
The previous finding of a MAPK-site in the CTD of Su(H)19 sparked our interest to search for further phosphorylation sites in Su(H) in order to identify additional cross-talk between Notch and as of yet unknown signalling pathways. To this end, we took a mass spectrometry approach and isolated Su(H) protein from Drosophila Schneider S2 cells. Myc-tagged Su(H) protein and activated RasV12 were ectopically induced in S2 cell culture followed by immunoprecipitation of Su(H) protein with anti-Myc antibodies. The upper of two Su(H) protein bands was excised and in-gel digested with trypsin (Fig. 1a). Resultant peptides were analyzed by nano-LC-ESI-MS/MS with a verified sequence coverage of 38% of the Su(H) protein. A singly phosphorylated peptide (LRpSQTVSTR) corresponding to amino-acids 267–275 of Su(H) was identified by MS/MS analysis (Fig. 1a). The fragmentation spectrum of the phosphopeptide showed a good sequence coverage by y- and b-ions, enabling an unambiguous localization of the phospho-site to Serine 269 (Fig. 1a).
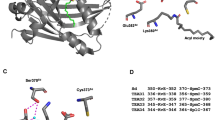
Phosphorylation of Su(H) at Serine 269. (a) Coomassie stained Su(H)myc protein precipitated from S2 cell culture used for mass spectrometry analyses (left, asterisk). Approximate molecular weight is given in kilo Dalton. MS/MS spectrum of the Su(H) phosphopeptide LRpSQTVSTR (precursor ion m/z = 564.2816, z = 2). Identity and sequence of the peptide as well as the phosphorylation site at S269 were confirmed by b- and y-ion series as indicated in blue and red, respectively. Neutral loss reactions of H2O and H3PO4 from the precursor ion are indicated in green. (b) Scheme of the wild type Su(H) protein [Su(H)wt], consisting of three domains: NTD (N-terminal domain; AS 116–252, light blue), BTD (beta-trefoil domain; AS 253–400, purple) and CTD (C-terminal domain; AS 424–516, dark blue). Below, the sequence of a BTD segment overlapping the phospho-peptide isolated by mass spectra is shown [underlined, bold in D. melanogaster Su(H)]; it is completely conserved in H. sapiens CBF-1, M. musculus RBPJ and C. elegans Lag1 proteins. Yellow triangles mark amino acids that contact the DNA backbone, light yellow triangles indicate base contacts via hydrogen bonds22,28. Ser269 is highlighted in red. The phospho-deficient [Su(H)S269A], the phospho-mimetic [Su(H)S269D] and the DNA-binding deficient [Su(H)R266H] mutant isoforms are indicated. Su(H)R266H corresponds to the naturally occurring Su(H) S5 mutant24. (c) Structure of Su(H) bound to DNA [PDB ID: 5E24]. Enlargement of the BTD-DNA contact is shown; Ser269 and Arg266 are highlighted. Figure was assembled using PyMOL, licensed to RAK.
The position of this newly identified phosphorylation site resides within the beta-trefoil domain (BTD) of Su(H), which is known to interact with the RAM domain of the intracellular domain of Notch (NICD) and to mediate contact with DNA (Fig. 1b,c)20,21,22. Therefore, phosphorylation at this site might influence the high-affinity interaction with NICD, interfering with activator complex formation. Similarly, Nemo-like kinase (NLK) inhibits ternary complex formation by phosphorylation of NICD23. Alternatively, phosphorylation might generally affect the binding of Su(H) to target sites on the DNA. Ser269, as well as the sequence surrounding it, is absolutely conserved between Drosophila, C. elegans and mammals, including humans, and lies within a region known to make both specific and nonspecific contacts with the DNA (Fig. 1b,c)20. To address the potential biological significance of Ser269 phosphorylation we generated two mutants by site directed mutagenesis: a phospho-deficient Su(H) isoform by replacing Ser269 with alanine (S269A), and a phospho-mimetic Su(H) isoform by replacing Ser269 with aspartic acid (S269D) (Fig. 1b). In addition, we replaced Arg266 by Histidine (R266H), thereby re-creating a DNA binding-deficient Su(H) protein, also known as Su(H)S5 mutant24 (Fig. 1b). The mutant constructs were shuttled into respective vectors to allow for subsequent analyses in vitro and in vivo.
The phospho-mimetic Su(H)S269D isoform shows reduced activity in Drosophila cells
First, we addressed the transcriptional activity of the phospho-mutant Su(H) isoforms in Drosophila Schneider S2 cells by transient transfection with respective inducible constructs. As a read out, luciferase activity was measured from a Notch response element reporter construct (NRE-luc)25. NRE-reporter gene activity is induced by transfection of NICD, which can be further raised about threefold by co-transfection with wild type Su(H)wt (Fig. 2)12,25,26. Compared to the level of Su(H)wt, the phospho-deficient Su(H)S269A mutant construct resulted in a slightly enhanced activation, whereas Su(H)S269D resulted in a considerably lower reporter activation. As expected, the Su(H)R266H mutant had no impact on the NICD-mediated activation of the NRE-luc reporter (Fig. 2, compare lanes 4 and 5). We conclude that phosphorylation of Su(H) at Ser269 impedes Notch activity, which is not a consequence of a mutation of Ser269 per se, since Su(H)S269A shows an even stronger activity than wild type. Moreover, as our mass spec data shows that Su(H) is phosphorylated in S2 cells, the increased activity of Su(H)S269A suggests that repression of Notch signalling activity by phosphorylation is in fact taking place in these cells even in the absence of activated RasV12.
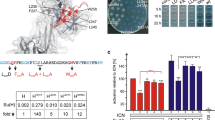
Notch signalling readout is changed in S2 cell culture assays. S2 cells were co-transfected with the NRE luciferase reporter construct and with NICD to induce expression plus the different Su(H) constructs as indicated. Reporter activity in cells co-transfected with Su(H)wt and NICD was taken as 100%. Co-transfection with Su(H)S269A significantly increases reporter activity (about 120%), whereas co-transfection with either Su(H)S269D or Su(H)R266H had a much lower effect on NICD activity. Repressor complexes were tested with a co-transfection of NICD with H and respective Su(H) variants (orange bars). H inhibits ICN-mediated reporter-gene activation down to about 60%. This inhibition is alleviated by the combined co-transfection of NICD and H with either Su(H)wt or Su(H)S269A: apparently the two behave similarly in this context. Co-transfection of NICD and H with either Su(H)S269D or Su(H)R266H show the same low reporter gene activation as NICD and H alone: both seem incapable of interfering with H-mediated repression. Three independent assays each were performed, measurements were in duplicate. Error bars denote standard deviation; ***p < 0.001 highly significant; **p < 0.01 very significant; *p < 0.05 significant; p > 0.05 not significant (ns). Symbol on top of the bars represent comparison to control, to either NICD + Su(H) (blue columns) or NICD + H (orange columns), respectively. Other comparisons are as indicated by horizontal lines.
The Su(H) transcriptional regulator not only activates, but also represses Notch target genes, depending on its bound co-factors. We therefore wanted to determine whether repression activity was likewise affected in the Su(H)S269D mutant. To this end, a co-transfection of the respective constructs with NICD and Hairless (H) was performed. Transfection of H results in a repression of NICD-mediated reporter gene activity by about two-fold26. In combination with either wild type Su(H)wt or Su(H)S269A values are similar to NICD overexpression alone. In contrast neither Su(H)S269D nor Su(H)R266H had a significant effect on H-mediated repression (Fig. 2). Although compensation of H overexpression by Su(H)wt and Su(H)S269A was not significant in this assay, the tendency of differential activity is apparent. Hence, we propose that phosphorylation of Su(H) at Ser269 may mitigate transcriptional activity, affecting both activation and repression of Notch target genes alike.
The phospho-mutant Su(H) isoforms are not impaired in protein complex formation
The reduced activation and repression activity of Su(H)S269D in the above luciferase assay could result from a failure to assemble activator complexes together with NICD and Mam, or repressor complexes together with H. To assay protein complex formation of the Su(H) mutant isoforms, yeast two- and three-hybrid assays were performed, addressing the direct interaction of Su(H) with either H or NICD, as well as the assembly of the Notch activator complex, which includes Mam. As shown in Fig. 3, all mutant isoforms, the phospho-specific Su(H)S269A and Su(H)S269D mutants as well as the DNA binding-deficient Su(H)R266H mutant proteins, performed very similar to wild type Su(H) in the binding of either H or NICD, or in the three-hybrid assay together with NICD and Mam. These results imply that phosphorylation of Su(H) at Ser269 does not interfere with the assembly of protein complexes. We next addressed the possibility that the mutations might affect protein expression, stability or nuclear localization of Su(H) protein. To this end, expression of the Su(H) isoforms was induced by a heat pulse and detected 6 hours later on Western blots. We observed little variance in the levels of the induced proteins (Supplementary Fig. S1). Hence the observed differences in Su(H) activity are unlikely to result from differences in expression or stability. Moreover, nuclear localization was observed for the phospho-mutant Su(H) variants when induced in S2 cells (Supplementary Fig. S1). In sum it appears more likely that the DNA-binding activity of Su(H) is altered by the phosphorylation, presumably by adding a negative charge in the vicinity of the residues that are directly contacting DNA (Fig. 1c).
Yeast two- and three hybrid interaction assays. Direct protein-protein interaction between the Su(H) variants (in pJG) and either full length H or the intracellular part of the N receptor (ICN I, in pEG), were tested in a yeast two-hybrid assay. Interaction is seen by the blue colour resulting from the activation of the reporter gene lacZ, whereas white yeast colonies indicate no interaction. Yeast three-hybrid tests12,13 were done with pEG-Mam and pESC-NICD transformed with the different Su(H) variants in pJG vector, respectively. Empty vectors served as mock control.
Altered DNA binding of phospho-mutant Su(H) proteins
To directly address the DNA binding properties of the Su(H) protein isoforms, wild type as well as the two phospho-specific mutants were in vitro transcribed and translated, and subjected to an electro-mobility shift assay (EMSA) using a radiolabelled E(spl)m8-S1 oligo as target DNA27 (Fig. 4a, Supplementary Fig. S2). For comparison, the defined DNA binding deficient Su(H) mutant isoform Su(H)R266H was also used24. Whereas no obvious difference in DNA binding was observed between wild type and the phospho-deficient isoform Su(H)S269A, the phospho-mimetic Su(H)S269D isoform was strongly impaired in DNA binding. As expected, DNA-binding was completely abrogated in the Su(H)R266H mutant (Fig. 4a). However, in contrast to Su(H)R266H, we were able to observe very weak DNA binding by Su(H)S269D (Fig. 4a, red asterisk). In order to quantify the differences we performed isothermal titration calorimetry (ITC) with recombinantly purified Su(H) protein and oligomeric DNA duplexes that contain the consensus binding site of the HES-1 gene28. The ITC experiments revealed little differences between the energetics of either Su(H)wt or Su(H)S269A – DNA binding (Fig. 4b, Supplementary Fig. S2). In contrast to our EMSA results, we could not detect any DNA binding of Su(H)S269D by ITC (Fig. 4b, Supplementary Fig. S2). These differences are likely a result of the differences in detection limits between the two methods. From these results we conclude that phosphorylation at Ser269 interferes with the DNA binding capacity of the transcription factor Su(H). This could mean that, depending on the context of such a phosphorylation, either the activation of Notch target genes or their repression is downregulated by an unknown kinase. For example, if this kinase is activated concurrent with the Notch receptor, Notch target gene activation would be inhibited. If the kinase, however, is active whilst Notch signalling is off, Notch target gene repression would be alleviated.
DNA binding is affected in the phospho-mimetic Su(H)S269D isoform. (a) Electro-mobility shift assay for the binding of the indicated Su(H) isoforms to radiolabelled E(spl)m8-S1 oligo27. In the mock control, no protein was added to the reaction. Su(H)wt and likewise Su(H)S269A bind well, whereas only a very weak binding of Su(H)S269D (red asterisks) and no binding of Su(H)R266H to the oligo was detected. Specificity was tested by adding increasing amounts (1.6 ng, 18 ng, 42 ng) of unlabelled competitor oligo. The arrow points to the shifted complexes; the arrowhead demarks the radiolabelled unbound oligos. (b) Isothermal titration calometry assay of Su(H)-DNA binding. Representative thermograms for Su(H) binding to DNA comprising the consensus HES-1 site are shown (raw heat signal and nonlinear squares fit to the integrated data). Twenty titrations were performed per experiment, consisting of 14 μl injections that were spaced 120 s apart.
In vivo studies of phospho-mutant Su(H) variants during wing imaginal development
With the goal to examine the in vivo consequences of Ser269 phosphorylation, the Su(H) mutant constructs (Fig. 1b) were cloned under UAS-control, enabling us to induce their expression in a tissue specific manner in transgenic flies. Position effects were avoided by using the PhiC31 integrase system that allows insertion of each mutant construct at the identical chromosomal position29. Ectopic expression of the different Su(H) mutant variants and the wild type form was induced with omb-Gal4 in the central wing anlagen. Notch in vivo activity was monitored with the vg BE -lacZ reporter line, which is induced in a stripe along the dorso-ventral boundary of the presumptive wing blade (Fig. 5a)30. In addition, the expression of Cut protein served as readout of Notch activation along the dorso-ventral boundary31 (Supplementary Fig. S3). In this context, overexpression of wild type Su(H) causes a slight expansion of vg BE -lacZ or of Cut expression12,13,26, which was likewise observed upon overexpression of the phospho-deficient Su(H)S269A isoform (Fig. 5a; Supplementary Fig. S3). In accordance with a gain of Notch activity32,33, the wing discs were overgrown, again with little variance between Su(H)wt and Su(H)S269A (Fig. 5a; Supplementary Fig. S3). In contrast, overexpression of Su(H)S269D caused a patchy expression of vg BE -lacZ and a strong repression of Cut, but had little effect on the size of the wing disc (Fig. 5a, Supplementary Fig. S3). Apparently, Su(H)S269D was not only impaired in its ability to activate Notch mediated reporter expression, but instead caused its repression. The latter might be a consequence of Su(H)S269D-NICD protein binding and simultaneous failure of DNA binding, i.e. dysfunctional activator complexes are formed that cannot localize to Notch target genes. Hence, overexpression of Su(H)S269D acts in a dominant negative fashion on activated NICD molecules. Accordingly, overexpression of the Su(H)R266H variant – which completely fails to bind to DNA in vitro – showed an even stronger repression of vg BE -lacZ or Cut, and also no increase in disc size (Fig. 5a, Supplementary Fig. S3).
Effect of phospho-mutant Su(H) variants on Notch target gene vestigial expression. (a) vg BE-lacZ reporter activity along the dorso-ventral boundary of the wing imaginal disc was detected with anti-beta-Galactosidase antibodies (red). Su(H) isoforms as indicated (shown in green) were induced in the central part of the wing disc anlagen with the omb-Gal4 driver. As control UAS-GFP (green) was overexpressed. Overexpression of UAS-Su(H)wt or UAS-Su(H)S269A caused overproliferation in the affected tissue (double headed arrow) and enforced vg BE -lacZ activity at the dorso-ventral boundary (arrows). In contrast, overexpression of UAS-Su(H)S269D reduced vg BE -lacZ expression (repressive beam) and had little effect on tissue size. A similar phenotype was seen after overexpression of UAS-Su(H)R266H. (b) Overexpression of UAS-H (shown in blue) using omb-Gal4 represses vg BE -lacZ reporter activity (red) at the dorso-ventral boundary (repressive bar). Co-overexpression with the indicated UAS-Su(H) constructs (shown in green; appears turquoise in the combination with UAS-H): wild type and phospho-deficient Su(H) enhances H-mediated repression, leading to a strong reduction of vg expression (repressive bars) and a further decrease in size of affected tissue. The phospho-mimetic and the DNA-binding deficient Su(H) isoforms had little effect on H-overexpression phenotypes. This is especially apparent when comparing tissue size (double headed arrow). Size bar represents 100 μm in all panels.
Due to its role as a molecular switch, Su(H) is required for both Notch target gene activation and repression. With the goal to analyse the ability of the different phospho-specific Su(H) variants during repression, co-expression experiments with H were performed. Overexpression of H interferes with many Notch-dependent processes during larval development, for example the formation of the peripheral nervous system like mechano-sensory organs or during wing development (for review14). As H directly competes with Notch for interacting with Su(H) protein, gain of H equalizes loss of Notch activity. Accordingly, overexpression of H results in repression of Notch target gene expression, which can be visualized with either vg BE -lacZ or Cut, and is accompanied by a reduction of tissue size32 (Fig. 5b; Supplementary Fig. S4)26. A combined overexpression of H and Su(H) results in an extreme inhibition of Notch activity as a result of excessive repressor complex formation12,13,34. As a consequence many of the affected cells are lost presumably by apoptosis35,36,37, and only a small remainder of the expression domain is observed accompanied by a total repression of Notch target gene expression (Fig. 5b; Supplementary Fig. S4). A very similar result was seen when H is overexpressed together with Su(H)S269A, indicating that a mutation of Ser269 per se does not interfere with Su(H) activity. A considerably weaker phenotype was seen upon combined overexpression of H with Su(H)S269D, and even more so with Su(H)R266H (Fig. 5b; Supplementary Fig. S4). These results show that repressor complex assembly itself is not affected, but rather these complexes are less active, which can be attributed to the impairment of DNA binding. Dominant negative effects are not apparent in the combined overexpression, as one might have expected a size increase of tissue relative the H overexpression alone (Supplemental Fig. S4). This might be explained by the fact, the Su(H) mutant proteins interfere with activated Notch at the same time, contributing to a loss of Notch activity in addition to impeding H. Overall, these in vivo data support the results from our cell culture studies, indicating that phosphorylation of Ser269 is equivalent to a deficit of Su(H) activity as a consequence of lowered DNA binding.
In vivo studies of phospho-mutant Su(H) variants in the adult fly
Next we wanted to visualize the consequences of an overexpression of the phospho-mutant Su(H) variants on fly development. This is hampered by the fact that a manipulation of Notch signalling activity during larval development most often results in larval or pupal lethality, limiting the processes and tissues to be analysed. Lateral inhibition is one of the best-studied processes mediated by the Notch pathway, a process in which a few cells are selected from a group of equipotent cells to assume the primary cell fate, whereas the others are forced into a secondary fate (for review5,6,7). For example, during wing vein development Notch signalling is required to restrict vein competence, thereby limiting the width of the vein to its normal size38. We used this process to address the activity of the various Su(H) isoforms. Using vg QE-Gal4, the respective UAS-constructs were overexpressed exclusively within the presumptive wing blade, sparing the morphogenetic antero-posterior and dorso-ventral boundaries19,30. Both, Su(H)wt and Su(H)S269A overexpression caused a loss of vein material, in agreement with an enhancement of vein fate restriction due to increased Notch activity. Notably, the distal parts of the longitudinal L4 and L5 veins were affected (Fig. 6a–c), a phenotype indistinguishable from a heterozygous H null mutant fly, which also has an increase in Notch activity39,40. Notch gain of function phenotypes resulting from Su(H) overexpression can be largely explained by a Su(H)-H titration effect. In fact, overexpression of just the Su(H) CTD, which is capable of binding to H but is itself inactive in Notch signal transduction, is sufficient to induce Notch gain of function phenotypes41. One might have expected a similar increase in Notch activity when overexpressing Su(H)S269D or Su(H)R266H since both are able to titrate H. This was not observed. In contrast, overexpression of Su(H)S269D and Su(H)R266H caused predominantly thickened veins and deltas at the margin (Fig. 6d,e), a typical consequence of a Notch loss of function, resulting from a failure to restrict vein width38. As both two Su(H) isoforms are hampered in DNA-binding, any activator complex formed together with NICD will be non-functional, i.e. both isoforms also titrate active Notch, providing an explanation for the observed phenotypes. In addition to thickened veins, overexpression of Su(H)S269D also caused distinct vein gaps in the longitudinal vein 4 (Fig. 6d), indicative of a gain of Notch signalling activity. This mixed phenotype, i.e. increased vein thickness and vein loss, may be explained by the observation that Su(H)S269D has somewhat residual DNA binding capacity, allowing little signal transduction and hence, some of the expected gain of Notch phenotypes. In addition to vein thickening, we also noted a conspicuous change in the wing size: an enlargement when overexpressing either Su(H)wt or Su(H)S269A, and a decrease when inducing Su(H)S269D or Su(H)R266H, reflecting primarily gain of Notch activity in the former two and loss in the latter (Fig. 6f).
Consequences of ectopic expression of phospho-mutant Su(H) variants on adult wings. Overexpression of the indicated UAS-constructs was induced in the distal wing anlagen with the vg QE-Gal4 driver; (a–f) shows Su(H) variants alone and (g–k) in combination with full length H. Representative wings from female flies are shown. (a) UAS-lacZ was used as control. Longitudinal veins L1-L5 are indicated. (b) Overexpression of Su(H) wt and (c) Su(H)S269A causes gaps in the longitudinal veins L4 and L5 (black arrows). In contrast, overexpression of (d) Su(H) S269D or (e) Su(H) R266H results in thickened veins (red arrows) and deltas at the wing margin (open arrowheads) indicative of a reduced Notch signalling activity. In some cases also gaps can be detected within the same wing (arrow; example is shown in (d). (f) Measurements of the area from 8 female wings overexpressing the given UAS-Su(H) construct. Su(H) wt and Su(H) S269A overexpression increased wing size, whereas that of Su(H) S269D or Su(H) R266H reduced it significantly relative to control. Error bars represent standard deviation (n = 8); ***p < 0.001 highly significant; p > 0.5 not significant (ns). (g) Wings derived from a combination of UAS-H plus UAS-lacZ served for comparison with UAS-H plus any of the different UAS-Su(H) variants, to account for the titration effect of Gal4. Crosses were kept at 18 °C to circumvent early lethality. Compared to the control UAS-lacZ (a), UAS-H (g) causes thickened veins (red arrows) and blistered wings (arrowhead) when induced with vg QE-Gal4. This effect is strongly enhanced by a concomitant overexpression of H with either (h) Su(H) wt or (i) Su(H) S269A resulting in tiny wings where large parts are completely transformed into vein material, recognized by more densely spaced cells and ectopic bristles (red asterisks). This enhanced phenotype was not observed when H was co-overexpressed with either (j) Su(H) S269D or (k) Su(H) R266H. Yet, distal veins were conspicuously thicker (red arrows) in these combinations [compare (j) and (k) with (g)].
To follow the effects of the phospho-mutant Su(H) isoforms exclusively on the repression of Notch signalling activity, we simultaneously overexpressed H within the wing anlagen. As expected, sole overexpression of H with vg QE caused a broadening of the longitudinal veins (Fig. 6g), a phenotype which was strongly enhanced when Su(H) or Su(H)S269A were induced at the same time: the wings were extremely small and much of the tissue was directed to vein fate (Fig. 6h,i). In contrast, the wings obtained after co-overexpression of H with either Su(H)S269D or Su(H)R266H resembled those resulting from the sole H overexpression with regard to size and residual intervein material, although the distal portions of the longitudinal veins were clearly thicker (compare Fig. 6j,k with g).
Discussion
In this study we identified phosphorylation of Su(H) at Serine 269 by nano-LC-ESI-MS/MS and provide evidence for the function of this modification in vivo. A phospho-mimetic mutation of this site [Su(H)S269D] results in strongly reduced DNA binding, which is accompanied by a mitigation of Notch signalling readout in several different developmental contexts tested. As Su(H) acts as a switch in the Notch pathway, being the nuclear effector for both activator and repressor complexes, it follows that Su(H)S269D could either affect the transcriptional activation or the repression of Notch target genes, depending on the cellular context. If Su(H) phosphorylation occurred simultaneously with Notch receptor activation, transcriptional activation would be lessened by the reduced DNA binding affinity of Su(H). At other times, however, Su(H) phosphorylation would result in a release of target gene inhibition, since binding of the Su(H)-H repressor complex would be mitigated. Consistent with this premise, our overexpression studies demonstrate that both, transcriptional activation and repression are affected in vivo. Bearing the limitations of overexpression studies in mind, the respective contribution of Ser 269 phosphorylation during normal development and signalling situations, however, remains to be determined.
Structure analyses revealed that CSL transcription factors recognize their cognate DNA binding site [C/tGTGGGAA] through extensive sequence-specific and non-specific DNA contacts mediated by the BTD and NTD20. The NTD makes major-groove sequence specific contacts to the second half of the binding site (5′-GGGA-3′) in a manner similar to other Rel-DNA complexes. The BTD also makes specific DNA contacts via a beta-hairpin loop that inserts into the minor-groove sequence, thereby selecting for the 5′ pyrimidine followed by a G (5′-C/t G)20. CSL-DNA crystal structures have been determined from human, mouse, C. elegans and Drosophila proteins with DNA corresponding to the consensus HES-1 binding site, and remarkably, the structures are very similar despite the huge evolutionary distance of the studied species (for review11). Interestingly, the beta-hairpin loop of the BTD involved in DNA contacts appears to be flexible as it can adopt alternate conformations28,42: in the preferred conformation the backbone carbonyl of Ser269 and the side chain of Gln270 make sequence specific contacts [numbering is from Drosophila Su(H)]. In alternative conformations, however, Ser269 can move into the place of Gln270 to make side chain-hydrogen bonds28. Alanine replacement of Ser269 should likewise allow for the backbone carbonyl contacts but not for side chain-hydrogen bonds, which would likely have little impact on Su(H)-DNA complex stability, but may affect specificity. Whereas aspartate at this position is expected to interfere with DNA binding altogether due to its negative charge. Perhaps the flexibility of the beta hairpin-loop is an important aspect of phospho-Ser269 regulation, since it is conceivable that its conformation influences the accessibility of Ser269 by protein kinases. This flexibility may also underlie the molecular basis for why Su(H)S269D retains some DNA binding in our EMSA data, whereas Su(H)R266H does not. Moreover, there are indications that the conformation of the BTD beta-hairpin loop may correlate with target sequences bound by CSL28. In this case, Ser269 phosphorylation of Su(H) might be target gene specific and would result in a specific rather than a general downregulation of Notch signalling activity within a cell.
Our new results uncover extensive crosstalk between the Notch signalling pathway and other signalling cascades by means of phosphorylation. They reveal a second example of CSL phosphorylation. Earlier, we have shown that Su(H) is targeted within its C-terminus at Thr426 by MAPK, resulting in an attenuation of Notch signalling readout19. Also the MAPK-target site is absolutely conserved between fly and vertebrates. In mouse, phosphorylation of RBPJ at the corresponding Thr339 by the stress kinase p38 MAPK results in a destabilization of RBPJ43. In Drosophila we have, however, no indications to date that phosphorylation regulates Su(H) protein stability, despite the fact that Su(H) protein stability is an important means of regulating its availability44. The relative protein levels of either Su(H)wt, Su(H)S269D or Su(H)S269A were very similar after overexpression. If phosphorylation at this site resulted in a destabilisation or even degradation of the protein, we might have expected to see differences in stability. There is ample evidence for the influence of phosphorylation on the stability and the half-life of a protein. A prime example is the Notch receptor itself, as NICD is phosphorylated and degraded upon signalling45.
According to computational predictions based on the phosphorylation site database algorithm (PHOSIDA) several kinases may be considered to recognize Ser269 as target site46. Although we have identified the phosphorylation of Ser269 in cultured S2 cells in the presence of activated RasV12, the sequence surrounding Ser269 does not have a typical MAPK signature. Perhaps phosphorylation of Ser269 is mediated by a kinase, which itself is activated upon high levels of Ras signalling. As active Ras controls key aspects of growth, cell death, metabolism and mediates inter-cellular stress responses47 it might also pilot Notch signalling activity via Su(H) phosphorylation by an as yet unknown kinase. Alternatively, Ser269 phosphorylation may be independent of Ras activity. To our knowledge, no murine or human homologue phosphorylated at the corresponding Ser has been identified in high throughput screens (http://www.phosphosite.org/). It is conceivable that Ser269 in Su(H) is phosphorylated constitutively, and dephosphorylated specifically by an unknown phosphatase in a context specific manner. As this phosphorylation target site is absolutely conserved in all mammalian CSL orthologues it is tempting to speculate that – as in Drosophila - this site is used for crosstalk in mammals as well, opening the possibility for a new strategy to achieve tissue homeostasis and to consequently coordinate and react on cellular stress signals.
Methods
Plasmid construction, generation of transgenic flies and fly work
Su(H) cDNA in Bluescript vector was used for QuickChange site-directed mutagenesis (Agilent, La Jolla CA, USA) to generate Su(H)S269A, Su(H)S269D and Su(H)R266H. Sequence verified constructs were cloned as Acc65I/XbaI fragments into the pUAST-attB vector. Transgenic flies were established with the PhiC31 integrase-based system29 using the ΦX-58A or ΦX-96E strain harbouring the landing site at 58A and 96E, respectively. Correct integration was tested by PCR. Other fly stocks used: omb md65-Gal4/FM748, vg QE-Gal419, vg BE-lacZ30, Sco/CyO hs-Gal4 UAS-GFP (BL5702), UAS-GFP (BL4776), UAS-lacZ (BL8529), UAS-H at 68E12. For co-overexpression experiments UAS-Su(H) wild type12 or mutant constructs at 96E were recombined with UAS-H at 68E. Flies were raised on standard cornmeal at 18 °C and crosses kept at 18 °C or 25 °C as indicated.
Mass spectrometry
Myc-tagged Su(H) protein was derived from stably transfected Schneider S2 cells by immuno-precipitation as described before19. The coomassie stained Su(H) protein was in-gel-digested with trypsin and mass spectrometry analysis was performed on a Thermo LTQ-Orbitrap instrument (Thermo Fisher Scientific, Germany) equipped with a nanoelectrospray ion source as described before49. Survey spectra (m/z = 300–1800) were detected in the Orbitrap at a resolution of 60,000 at m/z = 400. Database searches were performed with MASCOT search algorithm to identify the Su(H)-phosphopeptide, verified by manual inspection of the MS/MS spectra49.
Yeast two-hybrid constructs and assays
Yeast two- and three-hybrid experiments were performed as described before12,13,35. The following constructs were used: pEG-ICN I (codons 1827–2259)50, pEG-Mam (codons 118–194), pESC-NICD (codons 1762–2176)13, pEG-H (full length Hairless, corresponding to pEG-HFL)51. The Su(H)S269A, Su(H)S269D and Su(H)R266H mutant variants were cloned as full length constructs into pJG vector52 and were sequence verified.
S2 cell culture and transfection assays
Schneider S2 cell culture reporter assays were performed at least in triplicate according to ref.12. Cells were transiently transfected with 1 μg NRE-luciferase reporter25 and 0.2 μg of control Renilla plasmid for normalization. cDNA of Su(H)wt, Su(H)R266H, Su(H)S269A and Su(H)S269D was cloned as 1.8 kb Eco RI/Kpn I fragment into pRmHa3 (pMT) vector53. In addition, the constructs were tagged C-terminally with myc. Protein expression was induced with CuSO4 6 h after transfection and luciferase activity was measured 18 h later using dual-luciferase reporter assay (Promega, Heidelberg, Germany) with Lumat LB 9507 (EG and Salem, MA, USA). Su(H) myc-tagged protein was precipitated from about 108 Schneider S2 cells stably transfected with pMT-rasV12, pMT-Gal4 and UAS-Su(H)-myc19. The expression was induced with 0.7 mM CuSO4 and extracts were prepared 18-22 hours after induction.
DNA binding studies
Electro mobility shift assays were performed as described before19 with a double stranded DNA-oligomer harbouring the E(spl)m8-S1 Su(H) binding site [5′-TTGGGTGGCTCGTGGCGTGGGAACCGAGCTGAAAG-3′, 5′-GGTTCTTTCAGCTCGGTTCCCACGCCACGAGCCAC-3′]27. Su(H) protein isoforms were produced by coupled in vitro transcription / translation with TNT-Coupled Reticulocyte Lysate System (Promega, Mannheim, Germany) and 5 µl used for the reaction. The binding buffer contained 10 mM Tris-HCl pH 7.5, 50 mM NaCl, 1 mM EDTA, 0.0275 µg/µl salmon sperm DNA.
For isothermal titration calometry (ITC), recombinant Su(H) (98–523) was overexpressed and purified from bacteria, as previously described13. ITC experiments were carried out using a MicroCal VP-ITC microcalorimeter exactly as outlined earlier using double stranded DNA with the Hes-1 consensus [5′-GTTACTGTGGGAAAGAAAG-3′, 5′-GGAAGTTTCCCACAGGCCG-3′]28. All Su(H)-DNA experiments were performed at 10 °C in 50 mM sodium phosphate pH 6.5 and 150 mM NaCl. Su(H) was degassed and buffer-matched using dialysis and size exclusion chromatography. A typical Su(H)-DNA binding experiment contained 10 μM Su(H) in the cell and 100 μM DNA in the syringe. The data were analyzed using ORIGIN software and fit to a one-site binding model.
Immunochemistry and confocal imaging
Overexpression in wing anlagen was induced with omb md65-Gal448. Wing discs were isolated from late third instar larvae and stained as described before54. Expression of vestigial (vg) was determined using the vg BE-lacZ reporter line30 and anti-β-galactosidase antibodies (DSHB, University of Iowa, Iowa City, IA, USA). For Western blots, hs-Gal4 was crossed to the respective UAS-Su(H) lines, and expression induced in embryos with one hour heat shock at 37 °C as described earlier51. Other antisera used were: anti-Cut (from DSHB, University of Iowa, Iowa City, IA, USA), anti-H-A55 or anti-Su(H) (Santa Cruz Biotechnology, Santa Cruz, USA). S2 cells transiently transfected with myc-tagged pMT-Su(H) constructs were stained as outlined before56 with anti-myc (Santa Cruz Biotechnology, Santa Cruz, USA), anti-Su(H)39 and anti-Pzg57 as nuclear marker. Secondary antibodies with minimal cross-reactivity coupled to DTAF, Cy3 and Cy5 generated in donkey were purchased from Jackson Immuno-Research Laboratories (Dianova, Hamburg, Germany). Discs were mounted in VectaShield (Vector Laboratories, California, USA) and documented with a Bio-Rad MRC1024 confocal system using LaserSharp 2000 imaging software coupled to a Zeiss Axiophot microscope (Carl Zeiss AG, Oberkochen, Germany). Pictures were assembled with Corel-PhotoPaint and CorelDRAW Version 9.0 software.
Documentation of adult phenotypes
Adult wings were dehydrated in ethanol and mounted in Euparal (Roth, Karlsruhe, Germany), pictures of wings were taken with an ES120 camera (Optronics, Goleta, USA) using Pixera viewfinder software version 2.0. Wing size was measured with ImageJ programme.
Statistical evaluation
Statistical significance of probes was evaluated by ANOVA using a two-tailed Tukey-Kramer approach for multiple comparisons (highly significant ***p < 0.001; very significant **p < 0.01; significant *p < 0.05; not significant (ns) p > 0.05).
Data availability statement
All relevant data are within the manuscript and its supplementary information files.
References
Pires-daSilva, A. & Sommer, R. J. The evolution of signalling pathways in animal development. Nat Rev Genet. 4, 39–49 (2003).
Hurlbut, G. D., Kankel, M. W., Lake, R. J. & Artavanis-Tsakonas, S. Crossing paths with Notch in the hyper-network. Curr Opin Cell Biol. 19, 166–175 (2007).
Borggrefe, T. et al. The Notch intracellular domain integrates signal from Wnt, hedgehog, TGFß/BMP and hypoxia pathways. Biochim Biophys Acta 1863, 301–313 (2015).
Louvi, A. & Artavanis-Tsakonas, S. Notch and disease: A growing field. Semin Cell Dev Biol. 23, 473–480 (2012).
Hori, K., Sen, A. & Artavanis-Tsakonas, S. Notch signaling at a glance. J Cell Sci. 126, 2135–2140 (2013).
Bray, S. J. Notch signalling in context. Nat Rev Mol Cell Biol. 17, 722–735 (2016).
Fortini, M. E. Notch signaling: the core pathway and its posttranslational regulation. Dev. Cell 16, 633–647 (2009).
Hurlbut, G. D., Kankel, M. W. & Artavanis-Tsakonas, S. Nodal points and complexity of Notch-Ras signal integration. Proc Natl Acad Sci USA 106, 2218–2223 (2009).
Kovall, R. A., Gebelein, B., Sprinzak, D. & Kopan, R. The Canonical Notch Signaling Pathway: Structural and Biochemical Insights into Shape, Sugar, and Force. Dev Cell 41, 228–241 (2017).
Borggrefe, T. & Oswald, F. The Notch signaling pathway: transcriptional regulation at Notch target genes. Cell Mol Life Sci 66, 1631–1646 (2009).
Kovall, R. A. & Blacklow, S. C. Mechanistic insights into Notch receptor signaling from structural and biochemical studies. Curr Top Dev Biol. 92, 31–71 (2010).
Maier, D. et al. Structural and functional analysis of the repressor complex in the Notch signaling pathway of Drosophila melanogaster. Mol Biol Cell 22, 3242–3252 (2011).
Yuan, Z. et al. Structure and function of the Su(H)-Hairless repressor complex, the major antagonist of Notch signaling in Drosophila melanogaster. PLoS Biol 14(7), e1002509 (2016).
Maier, D. Hairless: the ignored antagonist of the Notch signalling pathway. Hereditas 143, 212–221 (2006).
Hasson, P. & Paroush, Z. Crosstalk between the EGFR and other signalling pathways at the level of the global transcriptional corepressor Groucho/TLE. Br J Cancer 94, 771–775 (2006).
Lee, H. J., Kim, M. Y. & Park, H. S. Phosphorylation-dependent regulation of Notch1 signalling. BMB Rep. 48, 431–437 (2015).
Carrieri, F. A. & Dale, J. K. Turn it down a Notch. Front Cell Dev Biol. 18(4), 151 (2017).
Doroquez, D. B. & Rebay, I. Signal integration during development: mechanisms of EGFR and Notch pathway function and cross-talk. Crit Rev Biochem Mol Biol. 41, 339–385 (2006).
Auer, J. S., Nagel, A. C., Schulz, A., Wahl, V. & Preiss, A. MAPK-dependent phosphorylation modulates the activity of Suppressor of Hairless in Drosophila. Cell Signal. 27, 115–124 (2015).
Kovall, R. A. & Hendrickson, W. A. Crystal structure of the nuclear effector of Notch signalling, CSL. EMBO J. 23, 3441–3451 (2004).
Nam, Y., Sliz, P., Song, L., Aster, J. C. & Blacklow, S. C. Structural basis for cooperativity in recruitment of MAML coactivators to Notch transcription complexes. Cell 124, 973–983 (2006).
Wilson, J. J. & Kovall, R. A. Crystal structure of the CSL-Notch-Mastermind ternary complex bound to DNA. Cell 124, 985–996 (2006).
Ishitani, T. et al. Nemo-like kinase suppresses Notch signalling by interfering with formation of the Notch active transcriptional complex. Nat Cell Biol. 12, 278–285 (2010).
Furukawa, T. et al. Suppressor of Hairless, the Drosophila homologue of RBP-Jkappa, transactivates the neurogenic gene E(spl)m8. Jpn J Genet. 70, 505–524 (1995).
Bray, S. J., Musisi, H. & Bienz, M. Bre1 is required for Notch signaling and histone modification. Dev Cell 8, 279–286 (2005).
Nagel, A. C. et al. Hairless mediated repression of Notch target genes requires the combined activity of Groucho and CtBP corepressors. Mol Cell Biol. 25, 10433–10441 (2005).
Bailey, A. M. & Posakony, J. W. Suppressor of Hairless directly activates transcription of Enhancer of split complex genes in response to Notch receptor activity. Genes Dev. 9, 2609–2622 (1995).
Friedmann, D. R. & Kovall, R. A. Thermodynamic and structural insights into CSL-DNA complexes. Protein Sci. 19, 34–46 (2010).
Bischof, J., Maeda, R. K., Hediger, M., Karch, F. & Basler, K. An optimized transgenesis system for Drosophila using germ-line-specific φC31 integrase. Proc Natl Acad Sci. 104, 3312–3317 (2007).
Kim, J. et al. Integration of positional signals and regulation of wing formation and identity by Drosophila vestigial gene. Nature 382, 133–138 (1996).
Micchelli, C. A., Rulifson, E. J. & Blair, S. S. The function and regulation of cut expression on the wing margin of Drosophila: Notch, Wingless and a dominant negative role for Delta and Serrate. Development 124, 1485–1495 (1997).
Go, M. J., Eastman, D. S. & Artavanis-Tsakonas, S. Cell proliferation control by Notch signaling in Drosophila development. Development 127, 3553–3566 (1998).
Djiane, A. et al. Dissecting the mechanisms of Notch induced hyperplasia. EMBO J. 32, 60–71 (2013).
Morel, V. et al. Transcriptional repression by Suppressor of Hairless involves the binding of a Hairless-dCTBP complex in Drosophila. Curr Biol. 11, 789–792 (2001).
Müller, D., Kugler, S. J., Preiss, A., Maier, D. & Nagel, A. C. Genetic modifier screens on Hairless gain-of-function phenotypes reveal genes involved in cell differentiation, cell growth and apoptosis in Drosophila melanogaster. Genetics 171, 1–16 (2005).
Protzer, C. E., Wech, I. & Nagel, A. C. Hairless induces cell death by downregulation of EGFR signalling activity. J Cell Sci. 121, 3167–3176 (2008).
Kurth, P., Preiss, A., Kovall, R. A. & Maier, D. Molecular analysis of the Notch repressor-complex in Drosophila: characterization of potential Hairless binding sites on Supressor of Hairless. PLoS One 6(11), e27986 (2011).
de Celis, J. F. Pattern formation in the Drosophila wing: the development of the veins. BioEssays 25, 443–451 (2003).
Bridges, C. B. & Morgan, T. H. Third-chromosome group of mutant characters of Drosophila melanogaster. Carnegie Inst Wash Publ. 327, 1–251 (1923).
Praxenthaler, H., Smylla, T. K., Nagel, A. C., Preiss, A. & Maier, D. Generation of new Hairless alleles by genomic engineering at the Hairless locus in Drosophila melanogaster. PLoS One 10(10), e0140007 (2015).
Maier, D., Praxenthaler, H., Schulz, A. & Preiss, A. Gain of function Notch phenotypes associated with ectopic expression of the Su(H) C-terminal domain illustrate separability of Notch and Hairless-mediated activities. PLoS One 8(11), e81578 (2013).
Friedmann, D. R., Wilson, J. J. & Kovall, R. A. RAM-induced allostery facilitates assembly of a Notch pathway active transcription complex. J Biol Chem. 283, 14781–14791 (2008).
Kim et al. Presenilin-2 regulates the degradation of RBP-Jk protein through p38 mitogen-activated protein kinase. J Cell Sci. 125, 1296–1308 (2012).
Praxenthaler, H. et al. Hairless-binding deficient Suppressor of Hairless alleles reveal Su(H) protein levels are dependent on complex formation with Hairless. PLoS Genetics 13(5), e1006774 (2017).
Fryer, C. J., White, J. B. & Jones, K. A. Mastermind recruits CycC:CDK8 to phosphorylate the Notch ICD and coordinate activation with turnover. Mol Cell 16, 509–520 (2004).
Gnad, F. et al. PHOSIDA (phosphorylation site database): management, structural and evolutionary investigation, and prediction of phosphosites. Genome Biol. 8, R250 (2007).
Jindal, G. A., Goyal, Y., Burdine, R. D., Rauen, K. A. & Shvartsman, S. Y. RASopathies: unraveling mechanisms with animal models. Dis Model Mech. 8, 769–782 (2015).
Lecuit, T. et al. Two distinct mechanisms for long-range patterning by Decapentaplegic in the Drosophila wing. Nature 381, 387–393 (1996).
Voolstra, O., Beck, K., Oberegelsbacher, C., Pfannstiel, J. & Huber, A. Light-dependent phosphorylation of the Drosophila transient receptor potential ion channel. J Biol Chem. 285, 14275–14284 (2010).
Matsuno, K., Diederich, R. J., Go, M. J., Blaumueller, C. M. & Artavanis-Tsakonas, S. Deltex acts as a positive regulator of Notch signaling through interactions with the Notch ankyrin repeats. Development 121, 2633–2644 (1995).
Maier, D. et al. In vivo structure-function analysis of Drosophila Hairless. Mech Dev. 67, 97–106 (1997).
Gyuris, J., Golemis, E., Chertkov, H. & Brent, R. Cdi1, a human G1 and S phase protein phosphatase that associates with cdk2. Cell 75, 791–803 (1993).
Bunch, T. A., Grinblat, Y. & Goldstein, L. S. Characterization and use of the Drosophila metallothionein promotor in cultured Drosophila melanogaster cells. Nucleic Acids Res. 16, 1043–1061 (1988).
Zimmermann, M., Kugler, S. J., Schulz, A. & Nagel, A. C. Loss of putzig activity results in apoptosis during wing imaginal development in Drosophila. PLoS One 10(4), e0124562 (2015).
Maier, D., Nagel, A. C., Johannes, B. & Preiss, A. Subcellular localization of Hairless protein shows major focus of activity within the nucleus. Mech Dev. 89, 195–199 (1999).
Fehon, R. G. et al. Molecular interactions between the protein products of the neurogenic loci Notch and Delta, tow EGF-homologous genes in Drosophila. Cell 61, 523–534 (1990).
Kugler, S. J. & Nagel, A. C. Putzig is required for cell proliferation and regulates Notch activity in Drosophila. Mol Biol Cell 18, 3733–3740 (2007).
Acknowledgements
We thank A.-C. Krahl, I. Steiner and M. Ziegler for performing initial site-direct mutagenesis, as well as T. Stößer and E. Kuprijanow for technical support. We acknowledge H. Schmid and M. Meier for help in overexpression studies. We thank the Bloomington Drosophila Stock Center, funded by the National Science foundation, for sending fly stocks. Some of the antisera were received from the Developmental Studies Hybridoma Bank (DSHB), developed under the auspices of the National Institute of Child Health and Human Development and maintained by the University of Iowa, Dept. Biology (Iowa 52242). In particular, we are indebted to D. Maier for discussion, his constant support and for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
A.N., R.K. and A.P. conceived and designed the experiments; A.N., J.A., A.S., J.P., Z.Y. and C.C. conducted the experiments; A.N. performed the statistical analysis; A.N., J.P., R.K. and A.P. collected and analyzed the data and prepared the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nagel, A.C., Auer, J.S., Schulz, A. et al. Phosphorylation of Suppressor of Hairless impedes its DNA-binding activity. Sci Rep 7, 11820 (2017). https://doi.org/10.1038/s41598-017-11952-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-11952-0
- Springer Nature Limited