Abstract
Due to rapid advancements in deep learning techniques, the demand for large-volume high-quality datasets grows significantly in chemical research. We developed a quantum-chemistry database that includes 443,106 small organic molecules with sizes up to 10 heavy atoms including C, N, O, and F. Ground-state geometry optimizations and frequency calculations of all compounds were performed at the B3LYP/6-31G* level with the BJD3 dispersion correction, while the excited-state single-point calculations were conducted at the ωB97X-D/6-31G* level. Totally twenty-seven molecular properties, such as geometric, thermodynamic, electronic and energetic properties, were gathered from these calculations. Meanwhile, we also established a comprehensive protocol for the construction of a high-volume quantum-chemistry dataset. Our QCDGE (Quantum Chemistry Dataset with Ground- and Excited-State Properties) dataset contains a substantial volume of data, exhibits high chemical diversity, and most importantly includes excited-state information. This dataset, along with its construction protocol, is expected to have a significant impact on the broad applications of machine learning studies across different fields of chemistry, especially in the area of excited-state research.
Similar content being viewed by others
Background & Summary
In recent decades, the introduction of artificial intelligence (AI) and machine learning (ML) into chemistry research dramatically altered the paradigm of scientific discoveries. With the development of computer science and technology, data played a more and more important role in several areas of chemical research. Several chemical datasets were created from the perspective of cheminformatics, such as PubChem1, GDB2,3,4,5, ZINC6,7, ChEMBL8,9 and so on10,11. These datasets were widely used in various areas of chemistry, which often serve as crucial data sources for in silico drug discovery12,13,14, novel material development15,16,17,18,19,20,21, etc.
Recently, the development of molecular datasets from first-principles quantum chemistry calculations attracted great attention. The incorporation of these electronic-structure calculations largely improves the data consistency in the dataset, eliminates inherent distribution errors, and provides molecular properties based on underlining atomic-level physical insights. Therefore, this types of quantum-chemistry dataset shows the high transferable ability and the unified performance across various applications. Currently, available quantum-chemical datasets can be roughly categorized into two main types: those that focuses the chemical and physical properties of different compounds with high diversities22,23,24,25,26,27,28,29,30,31,32,33, and those that aim at exploring the non-equilibrium structures in the conformational space of specific molecules.
In recent years, the emergence of deep learning algorithms dramatically speed up the growth of the demand for a large-volume, high-quality dataset. In this new era of big data-driven chemical researches, two major limitations of existing molecular-property datasets need to be addressed.
Firstly, the datasets providing information on molecular excited states are very rare24,29,33,34,35,36,37, although the excited-state properties are immensely valuable in practical applications, ranging from photovoltaic devices, organic light-emitting diodes, laser technologies, and photobiological processes. Therefore, there is an urgent need to develop the high-volume datasets that contains high-quality excited-state data of molecules with large chemical diversities.
Secondly, with the development of the deep learning technologies, the quality and quantity of data becomes a new bottleneck. On the one hand, it is well known that very large data volumes can guarantee the correct interpolation ability and enhance the transferable abilities of ML models. Although the simple combination of different datasets seems to be a straightforward solution given their small overlap in chemical spaces38,39, this approach is not always recommended40,41. The main issue is that the data from different datasets do not match to each other due to their different resources. In addition, many available datasets still suffer from the lack of chemical diversity, and this significantly deteriorates the performances of the deep ML models in chemical applications38.
Therefore, our objective is to build a quantum chemistry dataset that includes both ground- and excited-state properties. This dataset must show massive-volume, data-consistency and high-diversity. At the same time, the suitable protocol for the large dataset construction must show a balance between effectiveness, efficiency, and accessibility. To address this, we aim to develop such a comprehensive protocol for general usages, which covers initial geometry selection, quantum chemical calculations, and data quality examinations, along with efficient accessing and retrieval processes. This protocol ensures the robust and efficient construction of a large-volume dataset in chemical research. We wish that the current work provides not only a valuable large dataset but also a useful protocol to meet the increasing tendency to treat large-volume data in future chemistry research.
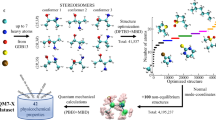
In this work, we reported the QCDGE (Quantum Chemistry Dataset with Ground- and Excited-State Properties) dataset with high chemical diversity, which totally includes 443,106 molecules with up to ten heavy atoms within the carbon (C), nitrogen (N), oxygen (O) and fluorine (F) range. These molecules are collected from the well-known QM925, PubChemQC29 and GDB-112,4 datasets. The ground-state geometry optimization and the frequency analysis were performed at the B3LYP/6-31G* level with the D3 version of Grimme’s dispersion complemented by Becke-Johnson damping (BJD3)42, while the excited-state single-point calculations including the first ten singlet and triplet transition states were performed at the ωB97X-D/6-31G* level. In total, 27 properties are extracted from these calculations, including ground-state energies, thermal properties, transition electric dipole moments, etc. We expect this dataset of small organic molecules to be useful in a wide range of applications in chemistry, especially for excited-state researches.
Methods
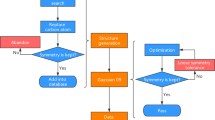
In this work, we tried to construct the QCDGE dataset, which is a quantum chemistry dataset that contains ground-state and excited-state information of ~450k molecules. They are small organic molecules with sizes up to ten heavy atoms in the range of C, O, N and F. The dataset construction procedure is divided into four steps, as detailed in the following subsections. The whole workflow is shown in Fig. 1(a).
Initial geometry collection
Given our objective to build a dataset that reflects chemical diversity, it is imperative to ensure a balanced integration of data sources. The GDB and PubChem datasets commonly serve as molecular sources for the construction of quantum chemistry datasets. The GDB series datasets are constructed through molecular combinatorial enumerations, according to the criteria of chemical stability and synthetic feasibility. In contrast, PubChem data are built from hundreds of data sources, including government agencies, chemical vendors, journal publishers, and others, which offers a broad collection of molecular information. It was observed that the duplication of data between these two datasets may not be significant38,39, which allows us to integrate molecules from these two datasets to define a new one. Therefore, we believe that the GDB and PubChem datasets stand out as original data sources with the optimal balance between chemical diversity and reliability.
In the first stage, our aim was to collect molecules with a size of up to 9 heavy atoms. To save computational resources, we first consider quantum chemistry datasets derived from GDB and PubChem, in which the three-dimensional optimized molecular structures were already given. Specifically, we took molecules from two primary datasets: QM9 and PubChemQC, which are derived from GDB-17 (also built from GDB) and PubChem, respectively. The QM9 dataset contains 132,177 molecules with sizes up to nine heavy atoms in the C, O, N and F range. Because of its significance and reliability, it is generally considered one of the golden standard datasets in the field of ML chemistry. Therefore, we chose compounds from the QM9 dataset according to the above selection rule. At the same time, we selected 122,785 molecules from PubChemQC, by using the same selection rule of QM9, in terms of the same limitation on heavy atoms.
In the second step, we broadened our selection criteria to include molecules with up to ten heavy atoms, in order to cover a wider chemical space. As the PubChemQC dataset contains many large compounds, we simply extracted a subset of 105,085 molecules that meet this new standard. As contrast, no such molecules are found in the QM9 dataset. Therefore, we selected additional molecules from GDB-11, a dataset generated from the original GDB. It is necessary to mention that many datasets were derived from GDB, such as GDB-11, GDB-17, QM9, etc. In principle, the GDB-17 dataset might be a better choice since the QM9 dataset is derived from it. However, in practice, the large size of the GDB-17 dataset brings numerical challenges. Because both GDB-11 and GDB-17 are generated on the basis of the same approach, we chose the smaller GDB-11 dataset here.
Here GDB-11 still contains over 3,000,000 molecules characterized by 10 heavy atoms (C, N, O and F elements). The direct inclusion of such large amounts of molecules would disrupt the balance of data distributions. To manage this, we used the mini-batch K-Means clustering algorithm43 to divide these molecules into 10,000 clusters. We then randomly selected molecules from these clusters, ensuring that the number of molecules chosen from each cluster was proportional to ratio between the cluster size and the total number of molecules. In this way, we selected 134,681 molecules, achieving a balanced representation across the chemical space.
Given that GDB-11 solely offers SMILES representations for its molecules, we needed to perform the initial geometry optimization. For this purpose, Cartesian coordinates of these molecules were generated using the in-house Python interface to Open Babel (version 2.8.1)44,45. Subsequently, these geometries were optimized using the semi-empirical method GFN2-xTB46 in the xtb program47. The whole process of collecting initial geometries from GDB-11 is shown in Fig. 1(b). For the in-depth information ranging from molecular descriptors, clustering methods, to the data selection strategies, please refer to the Supplementary Information.
Finally, we collected 494,701 pre-optimized molecular structures with balanced data sources as shown in Table 1, in which all small organic molecules at most contain ten heavy atoms of C, N, O, and F. It is noted that only molecules selected from GDB-11 were newly generated from SMILES strings in this step. This choice is sufficient because our goal is to build a quantum chemistry dataset with high diversity. Moreover, our approach not only extends the existing dataset but also establishes a unified method for merging different datasets. Therefore, we aim to ensure that all molecules are optimized and calculated at the same computational level. Certainly, it is essential to incorporate more new compounds and develop a more comprehensive dataset, which will be the focus of our future research efforts.
Ground-state calculations
Once the initial geometries were collected, we moved to the step of the quantum chemistry calculations, which is the most time-consuming one. Here, ground-state geometry optimizations and frequency analyses were conducted for all molecules. These calculations were performed at the DFT level using the B3LYP functional and the 6-31 G(d) basis set, which employ the Gaussian 16 package (B.01 version)48. To enhance accuracy, we incorporated the BJD3 dispersion corrections. Among all optimization jobs, about 0.3% (1,534) calculations failed to converge. This indicates that most initial structures are reasonable.
Optimized geometry check
In order to check the optimized geometries and to streamline the dataset by removing duplicated compounds, we proposed an examination process as shown in Fig. 1(c).
The first goal of this step is to confirm the consistency between optimized geometries and their initial counterparts. We generate InChI (IUPAC International Chemical Identifier) strings49 of initial and optimized geometries with the Python scripts interfaced with Open Babel (version 2.8.1). In most situations, the initial and optimized geometries give consistent InChI representations, indicating the reliability of the optimization tasks. Occasionally, some pairs show obvious discrepancies. This may refer to situations where the optimized geometry and the initial geometry are significantly different, implying that the optimization may not obtain a consistent result. However, such discrepancies may simply be due to the fact that the definition of InChI codes is too rigorous, and this very tight rule largely exaggerates stereoisomeric differences, even for minor ones. Therefore, when initial and optimized molecular structures show different InChI representations, additional examinations of geometrical details should be performed to avoid misjudgments. In practice, the redundant internal coordinates50,51 of the initial and optimized geometries were extracted using Gaussian 16 software. The direct comparison of them gave us solid answers to address whether the optimization task brings significant changes in molecular structures. When the optimization task gives the consistent structure with respect to the initial one, the molecule was retained in the dataset.
The second target is to eliminate duplicate geometries from our dataset. To achieve this, we simultaneously compare the structures using both SMILES and InChI strings generated via Open Babel. Given that these two representations highlight different aspects of the molecular geometries, it is enough to use them to classify duplicated molecules. In addition, this detection methodology shows the effective balance between computational accuracy and efficiency.
After removing 51,595 geometries that failed in optimization (1,534) and discard in duplicated checking process (50,061), finally the refined dataset contains 443,106 geometries. The relatively low proportion of duplicates further confirms that two original datasets cover different areas of the chemical space.
Excited-state calculations
All excited-state calculations were performed using the TDDFT method with the ωB97X-D functional and the 6-31 G(d) basis set. The reason to choose the ωB97X-D functional is mainly due to the fact that it gives reasonable descriptions of the charge transfer states, while the employment of the B3LYP level here may significantly underestimate the excitation energies of the charge transfer states. In the TDDFT calculations, the first ten singlet and triplet excited states were included. These calculations were carried out with Gaussian 16 software, employing the molecular geometries optimized at the B3LYP/6-31G(d)/BJD3 level.
Data Records
All data, including optimized molecular structures and important molecular properties, were extracted from the results of the quantum chemistry calculations. They are organized in a standard manner, which are accessible either in the figshare repository52 and or on the website of this data-driven excited-state information project (langroup.site/QCDGE). To ensure the integrity of all data in the further applications, we also provide the corresponding 512-bit cryptographic hash generated by the Secure Hash Algorithm 512 (SHA-512) for verification.
In the uploaded files, the final_all. csv summarizes the basic information of all molecules, such as their features (SMILES and InChI strings), the number of the heavy atoms, the number of ring moieties and so on. Within this file, the string in the first column serves as the unique identifier for each molecule. All data obtained from ground- and excited-state quantum chemistry calculations are saved in a binary file with the compressed version of the Hierarchical Data Format version 5 (HDF5)53 format. The HDF5 format is specifically designed to handle large volumes of numerical data, offering more efficient disk space utilization compared to other file formats such as text, JSON, and YAML. It supports reading data in chunks, enhancing efficiency in complex data analysis. The compressed version of HDF5 further reduces the record space significantly. In the current compressed version of the HDF5 file, the information of each molecule is organized as a dataset named after its identifier. Several versions of the SMILES and InChI strings are assigned as attributes for the dataset. Within each molecular dataset, 14 ground-state and 13 excited-state properties were recorded, as described in Table 2.
Technical Validation
For the current dataset, technical validation includes two main parts. First, the basic qualities of the data must be examined, such as their source, consistency, and correctness. Second, the chemical diversity of the dataset is crucial for its further applications.
We consider the following checks to ensure basic data quality. During the dataset construction process, the identifier for each molecule is created on the basis of its original number in the source dataset, as detailed in the Supporting Information. This allows us to easily track each molecule throughout the construction process and find the initial information in the source dataset. The primary technical validation of data quality is carried out by the checking process shown in Fig. 1(c). Since this is a crucial step in the dataset construction process, we described all details in the Methods section.
Our primary objective is to construct a dataset with high chemical diversity, making chemical diversity a crucial indicator of data quality. Therefore, we validated the chemical diversity in QCDGE in various ways to ensure data quality meets this high-diversity goal. The technical validation of chemical diversity is presented in six aspects, primarily performed on the basis of the Python interface of RDKit (version 2023.3.1, https://doi.org/10.5281/zenodo.591637).
Here, given that the current dataset does not contain duplicated structures, we simply divided all data into two subsets, namely data_A and data_B according to their original resources, the GDB series datasets and the PubChemQC dataset, respectively. These labels are used mainly for better illustrations in the following discussion. The differences in the chemical spaces of data_A and data_B were discussed in the Supplementary Information. We wish to emphasize that the conclusions drawn from the forthcoming analysis of data_A and data_B can not be generalized to describe the properties of the GDB and PubChemQC datasets themselves.
Element composition
First, the chemical diversity was examined by the element composition. Fifteen elemental compositions were identified according to different combinations of four heavy atoms (C, N, O and F). The numbers of compounds in several leading composition groups are given in Table 3. Among them, the group composed of molecules containing three heavy elements (C, N, and O) at the same time is the largest one, accounting for slightly less than the half of the total. The molecules including both C and O atoms together, as well as C and N, define the second and third largest groups, their total contributions accounting for about 3/4 of the total.
The same analysis was conducted on two molecular subsets, i.e., data_A and data_B, as shown in Tables S2 and S3. The results clearly demonstrate that the data from two different resources in fact do not overlap with each other in the chemical space. Interestingly, data_A has a high proportion of molecules that contain other heavy atoms except C, whereas data_B exhibits a greater diversity of carbon skeletons. However, molecules containing only N atoms, only O atoms, those containing both O and F, both N and F, and those containing N, O and F simultaneously, only appear in data_B.
Topology
To ensure high-diversity in topology of molecules, we show the distribution of molecules by the number of rings in Fig. 2(a). More than 3/4 of molecular structures contain ring moieties and almost half of all molecules include only one ring from a topological perspective. Among them, the proportions of acyclic molecules in data_A and data_B are approximately ~15% and ~35%, respectively (Fig. S2). On average, molecules in data_A possess 1.51 rings, whereas in data_B, the average is 0.81 rings. This finding prompts us to make a more comprehensive investigation of the existence of various ring units, as the ring moieties, particularly the aromatic rings, play important roles in determining excited-state properties.
Analysis of molecular ring distribution and category diversity in the QCDGE dataset. (a) Distribution of molecules by the number of rings. The histogram illustrates the counts of molecules categorized by their corresponding ring numbers. (b) Diversity of molecular categories. The horizontal axis quantifies the number of molecules examined, while the vertical axis lists several types of molecules discovered. Each bar represents the frequency of a particular molecular type.
Compound type
Compound types were counted to determine if they exhibited a high diversity in this aspect. According to the descending order of the number of molecules, all composition groups were sorted as follows: heterocycles (24.6%), fused heterocycles (22.1%), heteroacyclic (15.3%), heteroaromatics (11. 9%), carbocycles (11. 9%), carboacyclic compounds (7.9%), fused carbocycles (5.4%), and aromatics with carbon rings (0.9%). This distribution is consistent with the chemical intuitive notion, as the introduction of heteroatoms should largely extend the chemical space with respect to the situations with only carbon atoms.
Significant differences appear in the distributions of compound types in data_A and data_B, as illustrated in Fig. S3. This divergence could be attributed to the following reasons. More ring structures appears in data_A, and thus the fused heterocycles are popular. In contrast, data_B contains more acyclic compounds, including both heteroacyclic and carboacyclic ones. As a consequence, data_A features a greater complexity of ring moieties, whereas data_B is characterized by a relative abundance of linear or non-ring structures.
Scaffold analysis
Using the RDKit (version 2023.3.1) toolkit, the Murcko scaffold analysis was conducted to explore the diversity of molecular backbone in the QCDGE dataset. Aside from acyclic molecules (~23.2%), totally 59,898 distinct scaffolds were identified among the remaining molecules. Among all identified scaffolds, the most dominant one is three-membered carbon ring (C1CC1) moieties, as shown in Table 4. This may be attributed to the following fact. As our selection rule only chose molecules limited to ten heavy atoms, both small and large molecules may easily include stable three-membered rings.
To facilitate a more comprehensive analysis, we can also make Murcko scaffolds generic as illustrated in Table 5, by converting all types of atom to carbon and treating all bonds as single bonds. In such analysis, 3,258 Murcko scaffolds were identified, while five-membered rings predominated.
The scaffold analysis were also carried out in data_A and data_B, and results are detailed in Tables S4 to S7 of Supplementary Information. In the standard scaffold analysis, 46,234 scaffolds were identified in data_A and 17,290 in data_B. In contrast, the generic scaffold analysis yielded 2,161 and 1,934 scaffolds for data_A and data_B, respectively. This observation suggests that data_A show higher chemical diversity in the ring part than data_B, consistent with their individual features.
Functional group analysis
The diversity of functional groups was explored using the Ertl algorithm54,55, achieved with the RDKit (version 2023.3.1) toolkit. Initially, the original RDKit version only recognizes a limited range of generic functional groups composed of C, N, O, and F. To enhance the analysis ability, we expanded its functionality to identify 109 functional groups (as shown in Fig. S4), according to their definitions in Checkmol software56. The current in-house expansion mainly improves the analysis protocol in two ways, (i) making the distinction of substituents such as dialkylether and alkylarylether; (ii) including some larger functional groups such as hemiaminal.
Across the dataset, 102 functional groups were detected and the number of functional groups on average is 2.4 per molecule. Top twenty functional groups are shown in Table 6. Importantly, the absence of certain functional groups in our dataset does not suggest a lack of chemical diversity, while it may be attributed to the limitation on the number of atoms due to our selection rule.
The analysis of functional groups in molecules from data_A and data_B showed different distributions, as detailed in Tables S8 and S9. 102 and 98 types of functional groups were identified within data_A and data_B, respectively. The similar numbers here suggest that both datasets display very high degrees of chemical diversity. However, this does not imply that the distribution of chemicals in two datasets is similar. For the same functional group, it is clear that its proportion is different in two subgroups. These results highlight the differences in chemical diversity between two subgroups, and further confirm the importance of merging two data sources.
Excitation energy
The high diversity of the QCDGE dataset can also be examined via the analysis of excited state properties. Considering that functional groups containing double or triple bonds are typically responsible for the photoexcitation to the low-lying excited states of molecular systems, we mainly focus on molecules containing such bonds. In the QCDGE dataset, 346,312 molecules, representing over 78% of total molecules, contain double or triple bonds. The excitation energies of the lowest singlet states of them are shown in Fig. 3(a). The vast majority of these molecules display the lowest singlet state excitation energies distributed between 2 and 8 eV. Since the excitation energy is closely relevant to the type of compound, the corresponding distributions are given in Fig. 3. Among all compound types, aromatic compounds have the lowest average singlet state excitation energies, while carboacyclic compounds have the highest values. This observation is highly consistent with chemical intuition. Additionally, the distribution of the lowest singlet-state excitation energies across all molecules in the QCDGE dataset is shown in Fig. S5.
Distribution of the lowest singlet state excitation energy across various molecular categories. (a) All selected molecules with double or triple bonds. (b) Heterocycles. (c) Fused heterocycles. (d) Heteroacyclic. (e) Heteroaromatics., (f) Carbocycles. (g) Carboacyclic compounds. (h) Fused carbocycles. (i) Aromatics with carbon rings.
Usage Notes
We offer a Python script named extract_data.py, designed to extract relevant data from HDF5 files. This script allows for extracting molecular properties from the QCDGE dataset, in which many options are supported as well. It can process the full list of all molecules in the dataset, a predefined list of molecules, or a chosen set of molecules filtered by the number of heavy atoms and their elemental compositions. It is also possible to import the extractData() class from this script, providing the seamless integration with other Python codes. All script and data files are available in the figshare repository52 and the project website (http://langroup.site/QCDGE).
Code availability
All research was supported by the Python programming language (version 3.8.5), while several important Python libraries and their respective versions are outlined below. Open Babel (version 2.8.1) and RDKit (version 2023.3.1) Python libraries were used to generate cheminformatic representations and to perform analysis. The management of HDF5 files was facilitated by h5py (version 2.10.0, https://doi.org/10.5281/zenodo.3401726), while pandas (version 1.1.3, https://doi.org/10.5281/zenodo.4067057) were used to implement CSV files and perform relevant data analysis. All related scripts are also available on GitHub (https://github.com/Yifei-Zhu/Database_codes.git). All scripts fall into three categories: calculation, check, analysis, while a separate Python script extract_data.py is also given to extract data information.
References
Kim, S. et al. Pubchem 2023 update. Nucleic Acids Res. 51, D1373–D1380 (2023).
Fink, T., Bruggesser, H. & Reymond, J.-L. Virtual exploration of the small-molecule chemical universe below 160 daltons. Angewandte Chemie International Edition 44, 1504–1508 (2005).
Blum, L. C. & Reymond, J.-L. 970 million druglike small molecules for virtual screening in the chemical universe database gdb-13. Journal of the American Chemical Society 131, 8732–8733 (2009).
Fink, T. & Reymond, J.-L. Virtual exploration of the chemical universe up to 11 atoms of C, N, O, F: Assembly of 26.4 million structures (110.9 million stereoisomers) and analysis for new ring systems, stereochemistry, physicochemical properties, compound classes, and drug discovery. Journal of Chemical Information and Modeling 47, 342–353 (2007).
Ruddigkeit, L., Van Deursen, R., Blum, L. C. & Reymond, J.-L. Enumeration of 166 billion organic small molecules in the chemical universe database GDB-17. Journal of Chemical Information and Modeling 52, 2864–2875 (2012).
Sterling, T. & Irwin, J. J. Zinc 15–ligand discovery for everyone. J. Chem. Inf. Model. 55, 2324–2337 (2015).
Tingle, B. I. et al. Zinc-22– a free multi-billion-scale database of tangible compounds for ligand discovery. J. Chem. Inf. Model. 63, 1166–1176 (2023).
Zdrazil, B. et al. The chembl database in 2023: a drug discovery platform spanning multiple bioactivity data types and time periods. Nucleic Acids Res. 52, D1180–D1192 (2024).
Davies, M. et al. Chembl web services: streamlining access to drug discovery data and utilities. Nucleic Acids Res. 43, W612–W620 (2015).
Pence, H. & Williams, A. Chemspider: An online chemical information resource. Journal of Chemical Education 87 (2010).
Wishart, D. S. et al. Drugbank 5.0: a major update to the drugbank database for 2018. Nucleic Acids Res. 46, D1074–D1082 (2018).
Cheng, T., Pan, Y., Hao, M., Wang, Y. & Bryant, S. H. Pubchem applications in drug discovery: a bibliometric analysis. Drug Discovery Today 19, 1751–1756 (2014).
Miller, M. A. Chemical database techniques in drug discovery. Nature Reviews Drug Discovery 1, 220–227 (2002).
Bohacek, R. S., McMartin, C. & Guida, W. C. The art and practice of structure-based drug design: A molecular modeling perspective. Medicinal Research Reviews 16, 3–50 (1996).
Himanen, L., Geurts, A., Foster, A. S. & Rinke, P. Data-driven materials science: status, challenges, and perspectives. Advanced Science 6, 1900808 (2019).
Tripathi, M. K., Kumar, R. & Tripathi, R. Big-data driven approaches in materials science: A survey. Materials Today: Proceedings 26, 1245–1249 (2020). 10th International Conference of Materials Processing and Characterization.
Cai, J., Chu, X., Xu, K., Li, H. & Wei, J. Machine learning-driven new material discovery. Nanoscale Adv. 2, 3115–3130 (2020).
Zou, S.-J. et al. Recent advances in organic light-emitting diodes: toward smart lighting and displays. Mater. Chem. Front. 4, 788–820 (2020).
Salehi, A., Fu, X., Shin, D.-H. & So, F. Recent advances in oled optical design. Advanced Functional Materials 29, 1808803 (2019).
Zhao, Q., Stalin, S., Zhao, C.-Z. & Archer, L. A. Designing solid-state electrolytes for safe, energy-dense batteries. Nature Reviews Materials 5, 229–252 (2020).
Bruno, I. J. & Groom, C. R. Crystallographic perspective on sharing data and knowledge. Journal of Computer-Aided Molecular Design 28, 1015–1022 (2014).
Montavon, G. et al. Machine learning of molecular electronic properties in chemical compound space. New Journal of Physics 15, 095003 (2013).
Kim, H., Park, J. Y. & Choi, S. Energy refinement and analysis of structures in the QM9 database via a highly accurate quantum chemical method. Scientific Data 6, 109 (2019).
Ramakrishnan, R., Hartmann, M., Tapavicza, E. & Von Lilienfeld, O. A. Electronic spectra from TDDFT and machine learning in chemical space. J. Chem. Phys. 143, 084111 (2015).
Ramakrishnan, R., Dral, P. O., Rupp, M. & Von Lilienfeld, O. A. Quantum chemistry structures and properties of 134 kilo molecules. Scientific Data 1, 140022 (2014).
Rupp, M., Tkatchenko, A., Müller, K.-R. & von Lilienfeld, O. A. Fast and accurate modeling of molecular atomization energies with machine learning. Physical Review Letters 108, 58301 (2012).
Nakata, M. & Maeda, T. PubChemQC B3LYP/6-31G*//PM6 data set: The electronic structures of 86 million molecules using B3LYP/6-31G* calculations. J. Chem. Inf. Model. 63, 5734–5754 (2023).
Nakata, M., Shimazaki, T., Hashimoto, M. & Maeda, T. PubChemQC PM6: A dataset of 221 million molecules with optimized molecular geometries and electronic properties. Journal of Chemical Information and Modeling 60, 5891–5899 (2020).
Nakata, M. & Shimazaki, T. PubChemQC Project: A large-Scale first-principles electronic structure database for data-driven chemistry. Journal of Chemical Information and Modeling 57, 1300–1308 (2017).
Chen, G. et al. Alchemy: A quantum chemistry dataset for benchmarking ai models. arXiv arXiv:1906.09427 (2019).
Pereira, F. et al. Machine learning methods to predict density functional theory b3lyp energies of HOMO and LUMO orbitals. Journal of Chemical Information and Modeling 57, 11–21 (2017).
Liang, J., Xu, Y., Liu, R. & Zhu, X. QM-sym, a symmetrized quantum chemistry database of 135 kilo molecules. Scientific Data 6, 213 (2019).
Liang, J. et al. QM-symex, update of the QM-sym database with excited state information for 173 kilo molecules. Scientific Data 7, 400 (2020).
Zou, Z. et al. A deep learning model for predicting selected organic molecular spectra. Nature Computational Science 3, 957–964 (2023).
Kayastha, P., Chakraborty, S. & Ramakrishnan, R. The resolution- vs. -accuracy dilemma in machine learning modeling of electronic excitation spectra. Digital Discovery 1, 689–702 (2022).
Pengmei, Z., Liu, J. & Shu, Y. Beyond MD17: The Reactive xxMD Dataset. Scientific Data 11, 1 (2024).
Vinod, V. & Zaspel, P. CheMFi: A Multifidelity Dataset of Quantum Chemical Properties of Diverse Molecules. arXiv. http://www.arxiv.org/abs/2406.14149 (2024).
Glavatskikh, M., Leguy, J., Hunault, G., Cauchy, T. & Da Mota, B. Dataset’s chemical diversity limits the generalizability of machine learning predictions. J. Cheminformatics 11, 69 (2019).
Isert, C., Atz, K., Jiménez-Luna, J. & Schneider, G. QMugs, quantum mechanical properties of drug-like molecules. Scientific Data 9, 273 (2022).
Kokkinos, I. Ubernet: Training a universal convolutional neural network for low-, mid-, and high-level vision using diverse datasets and limited memory. In Proceedings of the IEEE conference on computer vision and pattern recognition, 6129–6138 (2017).
Zhang, D. et al. Dpa-2: Towards a universal large atomic model for molecular and material simulation. arXiv arXiv:2312.15492 (2023).
Grimme, S., Ehrlich, S. & Goerigk, L. Effect of the damping function in dispersion corrected density functional theory. J. Comput. Chem. 32, 1456–1465 (2011).
Sculley, D. Web-scale k-means clustering. In Proceedings of the 19th International Conference on World Wide Web, WWW ’10, 1177–1178 (Association for Computing Machinery, New York, NY, USA, 2010).
O’Boyle, N. M., Morley, C. & Hutchison, G. R. Pybel: a python wrapper for the openbabel cheminformatics toolkit. Chemistry Central Journal 2, 1–7 (2008).
O’Boyle, N. M. et al. Open babel: An open chemical toolbox. J. Cheminformatics 3, 1–14 (2011).
Bannwarth, C., Ehlert, S. & Grimme, S. Gfn2-xtb—an accurate and broadly parametrized self-consistent tight-binding quantum chemical method with multipole electrostatics and density-dependent dispersion contributions. J. Chem. Theory Comput 15, 1652–1671 (2019).
Bannwarth, C. et al. Extended tight-binding quantum chemistry methods. Wiley Interdisciplinary Reviews: Computational Molecular Science 11, e1493 (2021).
Frisch, M. J. et al. Gaussian 16 Revision C.01 (2016). Gaussian Inc. Wallingford CT.
Heller, S. R., McNaught, A., Pletnev, I., Stein, S. & Tchekhovskoi, D. Inchi, the iupac international chemical identifier. J. Cheminformatics 7, 1–34 (2015).
Pulay, P. & Fogarasi, G. Geometry optimization in redundant internal coordinates. J. Chem. Phys. 96, 2856–2860 (1992).
Peng, C., Ayala, P. Y., Schlegel, H. B. & Frisch, M. J. Using redundant internal coordinates to optimize equilibrium geometries and transition states. J. Comput. Chem. 17, 49–56 (1996).
Zhu, Y., Li, M., Xu, C. & Lan, Z. QCDGE dataset. Figshare https://doi.org/10.6084/m9.figshare.c.7259125.v1 (2024).
The HDF Group, N., Koziol, Q. & of Science, U. O. HDF5-version 1.12.0, https://doi.org/10.11578/dc.20180330.1 (2020).
Ertl, P. An algorithm to identify functional groups in organic molecules. J. Cheminformatics 9, 36 (2017).
Schaub, J.Development and implementation of in silico molecule fragmentation algorithms for the cheminformatics analysis of natural product spaces. Ph.D. thesis, Friedrich-Schiller-Universität, Jena https://doi.org/10.22032/dbt.59051 (2023).
Haider, N. Functionality pattern matching as an efficient complementary structure/reaction search tool: an open-source approach. Molecules 15, 5079–5092 (2010).
ChemAxon. Marvin. http://www.chemaxon.com (2024).
Acknowledgements
This work was supported by NSFC projects (Nos.22333003, 22361132528 and 21933011) and the Opening Project of Key Laboratory of Optoelectronic Chemical Materials and Devices of Ministry of Education, Jianghan University (JDGD202216).
Author information
Authors and Affiliations
Contributions
Conceptualization: Z.L. Data Curation: Y.Z. and M.L. Formal Analyses: Y.Z. and Z.L. Funding Acquisition: Z.L. Investigation: Y.Z., M.L. and Z.L. Methodology: Y.Z. Project Administration: Z.L. Resources: C.X. and Z.L. Software: Y.Z. and C.X. Supervision: C.X. and Z.L. Validation: Y.Z. and M.L. Visualization: Y.Z. and M.L. Writing - Original Draft Preparation: Y.Z. Writing - Review & Editing: Y.Z., M.L., C.X. and Z.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, Y., Li, M., Xu, C. et al. Quantum Chemistry Dataset with Ground- and Excited-state Properties of 450 Kilo Molecules. Sci Data 11, 948 (2024). https://doi.org/10.1038/s41597-024-03788-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03788-x
- Springer Nature Limited